Abstract
The electroencephalography (EEG) technique has the capability of identifying individual traits. Previous work has already used functional connectivity (FC) features obtained from EEG data for biometric purposes. In this work, we explored two FC methods not yet used in this context: motifs synchronization and space-time recurrences. Fifty subjects with two resting-state EEG acquisitions (one with eyes open and another with eyes closed) were included in the study. FC matrices for 1 s and 5 s epochs were computed for each acquisition. Subject’s identification was sought by comparing the FC matrices from both acquisitions using the Pearson correlation coefficient. The motifs method achieved 48% accuracy for both epoch sizes, and the space-time recurrences achieved 36% and 38% accuracies for 1 s and 1 s epochs respectively. Although the accuracies were low, they were well above the 2% chance level. Also, unlike other similar studies, the comparison was made between signals acquired in different conditions. In general, the obtained low accuracies illustrate the challenging problem of performing biometry from EEG and the need for further adjustments in the feature extraction and classification stages.
M. V. A. Davanço and M. C. de Paulo—These authors contributed equally to this work.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
The Electroencephalography (EEG) technique has been used for several clinical applications, particularly for diagnosis of some neurological diseases such as epilepsy [1] and sleep disorders [2]. Former studies have also shown that this technique provides information about differences between individuals related to anatomical and functional brain traits [3, 4]. More recently, the idea of using EEG signals to distinguish between different individuals aiming to implement a biometrical system [5, 6] has been explored in greater depth in several works [7,8,9], usually considering resting state acquisitions [10,11,12]. Indeed, the resting state paradigm has the benefit of enabling the use of data ac-quired from any individual, including those with restricted mobility, as well as diminishing the problem of movement artifacts, since the person must be still during the EEG evaluation.
Among the studies that aim to obtain biometric information from EEG signals are those that have used characteristics extracted from specific electrodes, and those that have explored the relationships between EEG signals obtained by different electrodes, a method known as brain connectivity. Connectivity-based approaches assume that many brain functions are executed under a specific engagement of different brain regions [13] or networks. Thus, the understanding of how these interactions take place may play a key role in providing additional information regarding the individual.
Functional connectivity (FC) is a data-driven, exploratory method, which seeks to find similarities between the dynamics of different brain regions. This method establishes relationships between the regions, which can be analyzed through graph theory [14]. For that, the graph nodes and a measure of similarity to provide the links between the nodes must be defined. For EEG data, the nodes are usually chosen to be the electrodes. In the context of using FC obtained from EEG to identify individuals, some similarity measures have already been explored, such as spectral coherence [9, 15, 16], the Spearman correlation applied to the Hilbert transform of the time series [17], the phase-locking value [17, 18], the imaginary part of the phase-locking value [19], the phase lag index or phase lag index [15], and mutual information applied to ordinal patterns [20], among others.
This work aims to analyze resting-state EEG under the perspective of graph-based measures to identify individuals. Two different FC similarity measures are used here: spatial-temporal recurrences [21] and motifs synchronization [22] based on ordinal patterns [23, 24]. It is important to stress that, to the best of our knowledge, neither type of similarity measure has yet been used in this context.
This article is divided as follows: Sect. 2 presents the EEG database used, the preprocessing steps and the two similarity methods used to evaluate the FC; Sect. 3 presents the identification results; and Sect. 4 presents a discussion about these and a brief conclusion of the work.
2 Materials and Methods
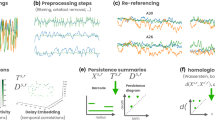
Figure 1 shows a flowchart summarizing the signal processing pipeline adopted in this work, including the chosen database, the preprocessing steps, the feature extraction approach, and the methodological analysis. All these steps are described in more detail in the following sections.
Signal processing pipeline. The database consisted of 50 subjects with two EEG acquisitions each (R1 and R2). These were preprocessed and 1 s or 5 s epochs were extracted from the time series (four epochs from R1 and one epoch from R2). FC matrices were calculated from these epochs, using the motifs synchronization and space-time recurrences methods. A mean FC matrix from R1 was used as reference and the FC matrix from R2 was used as test. These were compared among all subjects using Pearson’s correlation coefficient.
2.1 Database and Preprocessing
The EEG data used was from the Physionet database [25, 26], in which data from 109 subjects were recorded using a 64-channel EEG BCI2000 system, with electrodes positioned following the 10–10 system (Fig. 2). The subjects performed 14 experimental runs. The first two runs were acquired in resting state, during one minute each, with eyes open (R1 – condition) and closed (R2 - condition), respectively. These runs were used in the analysis performed here.
After downloading the data in EDF format, the preprocessing was performed using EEGLAB [27], and consisted of four steps: first, removal of artifacts by simple inspection; second, decomposition of the data using Independent Component Analysis (ICA) and removal of undesired components; third, removal of alpha and power grid frequencies; fourth, Common Average Referencing (CAR) of the data [28].
In order to remove more blatant artifacts, the “Inspect/Reject data by eye” tool was used, in which sections of the signals could be marked for removal. Sections that had greater (at least five-fold) amplitude than the rest of the signal were removed.
Then, the signal was decomposed into independent components (ICA), using the tool “Decompose Data by ICA”. This tool displays the obtained independent components through scalp map projections of the EEG activity. Components related to muscle movements, eye blinks and other eye movements can be easily recognized, and were thus removed.
Next, the alpha band was removed, using a stop band filter (with the “Basic FIR Filter” tool of EEGlab, considering an interval of 7 Hz to 13 Hz, and selecting the option “Notch filter the data instead of pass band”). This was done because we wanted to compare signals obtained from eyes closed and eyes open paradigms, and this band is known to be strikingly different between opened and closed eyes signals. Finally, the signal was bandpass filtered (again with the same tool, but without selecting the option “Notch filter the data instead of pass band”) between 4 and 50 Hz, to eliminate low-frequency artifacts and high-frequency noise.
10–10 electrode positioning system. Obtained from https://upload.wikimedia.org/wikipedia/commons/3/38/International_10-20_system_for_EEG-MCN.png. Author: Brylie Christopher Oxley.
The final preprocessing step implied in a spatial filter for re-referencing the signals using CAR [28]. This method consists in calculating the mean of the signals over electrodes and then subtracting this value from each electrode signal.
2.2 Functional Connectivity Matrices
All the database was preprocessed using the four steps aforementioned, however, only data from 50 subjects were used in this work. These subjects were selected considering the duration of the acquisitions after preprocessing. Sub-jects with acquisitions with less than 45 s were discarded.
From the R1 acquisition, four epochs were extracted, starting at seconds 10, 20, 30 and 40. From the R2 acquisition, only one epoch was extracted, starting at second 30. Lengths of 1 s and 5 s were tested for these epochs. Then, FC matrices were computed for both R1 and R2 epochs for all subjects, to be used as features in the identification problem. A template matching approach was used, in which the R1 matrices were further averaged to give one reference FC matrix per subject, while the R2 matrix was used as a test sample.
Two different similarity methods were used to compute the FC matrices: motifs synchronization [22] and space-time recurrences [21]. Both methods were implemented in MATLAB (2018, Natick, Massachusetts: The MathWorks Inc). These methods are detailed in the following.
Motifs Synchronization.
A motif series is basically a series of behavior patterns in the EEG signal. In this work, motifs with three points were used, as in Fig. 3. Thus, a temporal series of an EEG electrode can be “translated” into a motif series, according to the types of motifs in the signal.
The motif series of two electrodes can be used to evaluate the similarity between signals considering different lag values. In this work, a lag \(t=0\) was used. Mathematically, the similarity between the motif series of electrodes \(i\) and \(j\) can be calculated using the coefficient \({c}_{ij}\), defined as follows [22]:
where \({L}_{M}\) is the motif series length, \({J}_{k}=1\) if the motif at position \(k\) is the same in both series, and \({J}_{k}=0\) otherwise.
Then, the degree of synchronization between electrodes \(i\) and \(j\) is calculated:
which varies from 0 to 1.
With that, an \(N\times N\) connectivity matrix is obtained, where \(N\) is the number of electrodes used for the acquisition (in this work, \(N=64\)), and each element of the matrix is the degree of synchronization between the row electrode and the column electrode.
Space-Time Recurrences.
Space-time recurrences is a method used to identify whether a system has returned to a previous configuration during a given time period [29].
The space-time recurrence between two time series \({x}_{i}\) and \({x}_{j}\) is defined as:
The structure \(STR\) is called the space time recurrence matrix: a tridimensional data structure of \(N\times N\times {N}_{s}\), with \(N\) being the number of channels (or electrodes; in this case, \(N=64\)); and \({N}_{s}\) the total number of samples in the chosen time frame (e.g. \({N}_{s}=160\) for 1 s frames or \({N}_{s}=800\) for 5 s frames, since the sampling rate was \(160\) Hz). \(\varTheta \) is the Heaviside function, therefore: \(\varTheta \left(x\right)= 0\) if \(x<0\) and \(\varTheta (x)= 1\) if \(x\ge 0\). Finally, \(\varepsilon \) is an arbitrary distance threshold. In the present work, we chose \(\varepsilon = 50\%\) of the maximum distance (\(|{x}_{i}(n) -{x}_{j}(n)|\)) between electrode time series.
From the \(STR\) it is possible to calculate the connectivity matrix, which consists in normalizing the sum of the values of each electrode pair in the \({STR}_{i,j}\) structure:
Thus, it is possible to reduce the dimension of the problem, since \({A}_{i,j}\) is a two dimensional \(N\times N\) matrix. Each element of the matrix describes the similarity between two temporal series of EEG.
2.3 Comparison by Pearson Correlation Coefficient
To evaluate the similarity among the signals, and thus identify a given subject, the Pearson correlation coefficient was calculated between the mean R1 (eyes open) matrix and the R2 (eyes closed) matrix of all subjects.
If the highest correlation value was for R1 and R2 of the same individual, it was possible to identify the person, because it indicated greater similarity between different acquisitions of the same person. If not, it was not possible to identify the person.
Finally, the methods were compared in terms of their hit rate, or accuracy (i.e., percentage of correctly classified individuals).
3 Results
Table 1 shows the accuracy values obtained for subject identification, for each FC method and epoch length.
Using motif synchronization, for both epoch lengths (1 s and 5 s), 24 individuals were correctly identified among 50 analyzed, which corresponds to 48% of accuracy in both cases. Interestingly, the epoch length did not seem to influence the performance of this method.
The space-time recurrences method was able to correctly identify 18 out of 50 individuals for FC matrices computed using 1 s data, which corresponds to 36% accuracy, and for 5 s matrices it could identify 19 individuals among 50, which corresponds to a 38% accuracy. Therefore, the results using 5 s epochs to compute the FC matrices were slightly better.
4 Discussion
Regarding a comparison between methods, the motifs method achieved better accuracy than the space-time recurrences method, for all epoch lengths. This indicates that the motifs method was more capable than the recurrences method of extracting relevant information from the EEG signals regarding individual traits. The motifs method has been shown to be more efficient than other usual EEG FC methods, such as mean squared coherence and imaginary coherence, for extracting relevant information regarding interictal epileptiform discharges [30].
Nevertheless, the accuracies obtained with both methods used here, for all epoch lengths, were too low for practical purposes. Indeed, in [9], La Rocca and colleagues achieved up to 100% recognition rates with this same database, using features obtained both from power spectral density (PSD) and from coherence-based connectivity. They looked at individual electrode PSD features and individual channel (electrode pair) coherence features, and then combined the features from a given region (e.g., central, parietal or frontal) and fed them to a classifier based on the Mahalanobis distance. However, it is important to note that they only compared epochs within a given acquisition (eyes open or eyes closed); they did not attempt to use one acquisition to predict the other, as done here.
This work has several limitations. The number of subjects was low for the type of application (biometry). Notwithstanding, it is important to note that when the number of subjects is increased, the rate of accuracy decreases, since more comparisons are being made and the chance that there will be a correlation coefficient smaller than that of the right person increases. We previously tested the method with a sample of 11 subjects and the accuracies were indeed much better (64% for both methods).
The number of acquisitions per subject was also low, and additionally, the two acquisitions used did not follow exactly the same conditions, since despite both being in resting-state, one was acquired with eyes open and the other with eyes closed. Closing one’s eyes is known to increase the amplitude of alpha band oscillations in EEG signals. In a first analysis (not reported here) we attempted to use these different signals without subtracting the alpha band, but the results were worse than the ones reported here.
Also, the first preprocessing step (artifact removal by visual inspection) is somewhat subjective and may not have been exactly the same for all signals. Additionally, the STR requires adjusting the recurrence threshold for optimum FC evaluation [22] and a further detailed analysis considering a specific dataset for hyperparameter tuning outlines a natural perspective.
Nevertheless, it is important to highlight that the method presented may be taken further by exploring different options in each step of the methodology. Preprocessing could benefit from an automatic artifact removal algorithm such as SOUND [31], which would take away the subjectivity of removing signal stretches and ICA components by simple inspection. Also, other types of referencing methods, such as REST [32], could be tried instead of CAR. In the feature extraction step, graph parameters computed from the FC matrices could be explored. The motifs method could be improved by looking at delays other than zero, as in [22], while STR can be improved by means of threshold adaptations. Finally, in the classification step, a very simple classification method was used, namely, the Pearson correlation coefficient, but comparatively more sophisticated classification approaches could be investigated, such as Linear Discriminant Analysis (LDA), Support Vector Machine (SVM) or even deep neural networks.
In conclusion, both methods of FC calculation, motif synchronization and space-time recurrences, produced results that remained below what would be considered an accurate pattern of subject identification. That said, these results were highly above the chance level (which, for 50 subjects, would have been 2%), showing that the methods have potential for this application. Also, our results were obtained attempting to match two signals acquired in different moments, while other works in the literature using similar approaches have compared only signal epochs within the same acquisition (and condition). Finally, this was a pilot study, which aimed to explore the use of two FC measures that, to the best of our knowledge, had not yet been applied to biometry studies based on EEG data.
References
Koutroumanidis, M., et al.: The role of EEG in the diagnosis and classification of the epilepsy syndromes: a tool for clinical practice by the ILAE Neurophysiology Task Force (Part 1). Epileptic Disord. 19(3), 233–298 (2017)
Campbell, I.G.: EEG recording and analysis for sleep research. Curr. Protoc. Neurosci. 49(1), 10–2 (2009)
Berkhout, J., Walter, D.O.: Temporal stability and individual differences in the human EEG: an analysis of variance of spectral values. IEEE Trans. Biomed. Eng. BME 15(3), 165–168 (1968)
Van Dis, H., Corner, M., Dapper, R., Hanewald, G., Kok, H.: Individual differences in the human electroencephalogram during quiet wakefulness. Electroencephalogr. Clin. Neurophysiol. 47(1), 87–94 (1979)
Chan, H.L., Kuo, P.C., Cheng, C.Y., Chen, Y.S.: Challenges and future perspectives on electroencephalogram-based biometrics in person recognition. Front. Neuroinf. 12, 66 (2018)
Fraschini, M., Meli, M., Demuru, M., Didaci, L., Barberini, L.: EEG fingerprints under naturalistic viewing using a portable device. Sensors 20(22), 6565 (2020)
Campisi, P., La Rocca, D., Scarano, G.: EEG for automatic person recognition. Computer 45(7), 87–89 (2012)
Marcel, S., Millan, J.R.: Person authentication using brainwaves (EEG) and maximum a posteriori model adaptation. IEEE Trans. Pattern Anal. Mach. Intell. 29(4), 743–752 (2007)
La Rocca, D., et al.: Human brain distinctiveness based on EEG spectral coherence connectivity. IEEE Trans. Biomed. Eng. 61(9), 2406–2412 (2014)
Mantini, D., Perrucci, M.G., Del Gratta, C., Romani, G.L., Corbetta, M.: Electrophysiological signatures of resting state networks in the human brain. Proc. Natl. Acad. Sci. 104(32), 13170–13175 (2007)
Campisi, P., La Rocca, D.: Brain waves for automatic biometric-based user recognition. IEEE Trans. Inf. Forensics Secur. 9(5), 782–800 (2014)
Moctezuma, L.A., Molinas, M.: Towards a minimal EEG channel array for a biometric system using resting-state and a genetic algorithm for channel selection. Sci. Rep. 10(1), 1–14 (2020)
Pessoa, L.: Understanding brain networks and brain organization. Phys. Life Rev. 11(3), 400–435 (2014)
de Vico Fallani, F., Richiardi, J., Chavez, M., Achard, S.: Graph analysis of functional brain networks: practical issues in translational neuroscience. Phil. Trans. Royal Soc. B: Biol. Sci. 369(1653), 20130521 (2014)
Garau, M., Fraschini, M., Didaci, L., Marcialis, G.L.: Experimental results on multi-modal fusion of EEG-based personal verification algorithms. In: 2016 International Conference on Biometrics (ICB), pp. 1–6 (2016)
Boutorabi, S., Sheikhani, A.: Evaluation of electroencephalogram signals of the professional pianists during iconic memory and working memory tests using spectral coherence. J. Med. Signals Sensors 8(2), 87 (2018)
Cox, R., Schapiro, A.C., Stickgold, R.: Variability and stability of large-scale cortical oscillation patterns. Netw. Neurosci. 2(4), 481–512 (2018)
Pereda, E., García-Torres, M., Melián-Batista, B., Mañas, S., Méndez, L., González, J.J.: The blessing of dimensionality: feature selection outperforms functional connectivity-based feature transformation to classify ADHD subjects from EEG patterns of phase synchronisation. PLoS ONE 13(8), e0201660 (2018)
Dimitriadis, S.I., Salis, C., Tarnanas, I., Linden, D.E.: Topological filtering of dynamic functional brain networks unfolds informative chronnectomics: a novel data-driven thresholding scheme based on orthogonal minimal spanning trees (OMSTs). Front. Neuroinf. 11, 28 (2017)
Kottlarz, I., et al.: Extracting robust biomarkers from multichannel EEG time series using nonlinear dimensionality reduction applied to ordinal pattern statistics and spectral quantities. Front. Physiol. 11, 614565 (2021)
Rodrigues, P.G., Filho, C.A.S., Attux, R., Castellano, G., Soriano, D.C.: Space-time recurrences for functional connectivity evaluation and feature extraction in motor imagery brain-computer interfaces. Med. Biol. Eng. Comput. 57(8), 1709–1725 (2019)
Rosário, R.S., Cardoso, P.T., Muñoz, M.A., Montoya, P., Miranda, J.G.V.: Motif-synchronization: a new method for analysis of dynamic brain networks with EEG. Phys. A Stat. Mech. its Appl. 439, 7–19 (2015)
Olofsen, E., Sleigh, J.W., Dahan, A.: Permutation entropy of the electroencephalogram: a measure of anaesthetic drug effect. Br. J. Anaesth. 101(6), 810–821 (2008)
Quintero-Quiroz, C., Montesano, L., Pons, A.J., Torrent, M.C., García-Ojalvo, J., Masoller, C.: Differentiating resting brain states using ordinal symbolic analysis. Chaos Interdiscip. J. Nonlinear Sci. 28(10), 106307 (2018)
Schalk, G., McFarland, D.J., Hinterberger, T., Birbaumer, N., Wolpaw, J.R.: BCI2000: a general-purpose brain-computer interface (BCI) system. IEEE Trans. Biomed. Eng. 51(6), 1034–1043 (2004)
Goldberger, A.L., et al.: PhysioBank, PhysioToolkit, and PhysioNet: components of a new research resource for complex physiologic signals. Circulation 101(23), e215–e220 (2000)
Delorme, A., Makeig, S.: EEGLAB: an open source toolbox for analysis of single-trial EEG dynamics including independent component analysis. J. Neurosci. Methods 134(1), 9–21 (2004)
Ludwig, K.A., Miriani, R.M., Langhals, N.B., Joseph, M.D., Anderson, D.J., Kipke, D.R.: Using a common average reference to improve cortical neuron recordings from microelectrode arrays. J. Neurophysiol. 101(3), 1679–1689 (2009)
Eckmann, J.-P., Kamphorst, S.O., Ruelle, D.: Recurrence plots of dynamical systems. Europhys. Lett. 4(9), 973–977 (1987)
Costa, L.R.D., Campos, B.M.D., Alvim, M.K., Castellano, G.: EEG signal connectivity for characterizing interictal activity in patients with mesial temporal lobe epilepsy. Front. Neurol. 12, 673559 (2021)
Mutanen, T.P., Metsomaa, J., Liljander, S., Ilmoniemi, R.J.: Automatic and robust noise suppression in EEG and MEG: the SOUND algorithm. Neuroimage 166, 135–151 (2018)
Dong, L., et al.: MATLAB toolboxes for reference electrode standardization technique (REST) of scalp EEG. Front. Neurosci. 11, 601 (2017)
Acknowledgement
We thank PIBIC/SAE-UNICAMP, CNPq (grant 304008/2021–4) and FAPESP (grant 2013/07759–3) for financial support.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Davanço, M.V.A., de Paulo, M.C., Rodrigues, P.G., Soriano, D.C., Castellano, G. (2024). Motif Synchronization and Space-Time Recurrences for Biometry from Electroencephalography Data: A Proof-of-Concept. In: Marques, J.L.B., Rodrigues, C.R., Suzuki, D.O.H., Marino Neto, J., García Ojeda, R. (eds) IX Latin American Congress on Biomedical Engineering and XXVIII Brazilian Congress on Biomedical Engineering. CLAIB CBEB 2022 2022. IFMBE Proceedings, vol 99. Springer, Cham. https://doi.org/10.1007/978-3-031-49404-8_4
Download citation
DOI: https://doi.org/10.1007/978-3-031-49404-8_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-49403-1
Online ISBN: 978-3-031-49404-8
eBook Packages: EngineeringEngineering (R0)