Abstract
Healthcare is one of the most important concerns for living beings. Prediction of the toxicity of a drug is a great challenge over the years. It is quite an expensive and complex process. Traditional approaches are laborious as well as time-consuming. The era of computational intelligence has started and gives new insights into drug toxicity prediction. The quantitative structure-activity relationship has accomplished significant advancements in the field of toxicity prediction. Nine machine learning algorithms are considered such as Gaussian Process, Linear Regression, Artificial Neural Network, SMO, Kstar, Bagging, Decision Tree, Random Forest, and Random Tree to predict the toxicity of a drug. In the study, we developed an optimized regression model (Optimized KRF) by ensembling Kstar and Random Forest algorithm. For the mentioned machine learning models, evaluation parameters are assessed. The 10-fold cross-validation is applied to validate the model. The optimized model gave a coefficient of correlation, coefficient of determination, mean absolute error, root mean squared error, and accuracy of 0.9, 0.81, 0.23, 0.3, and 77% respectively. Further, the Saw score is calculated in two aspects as W-Saw score and the L-Saw score. The W-Saw score for the optimized ensembled model is 0.83 which is the maximum and L-Saw score is 0.27 which is the lowest in comparison to other classifiers. Saw score provides the strength to an ensemble model. These parameters indicate that the optimized ensembled model is more reliable and made predictions that were more accurate than earlier models. As a result, this model could be efficiently utilized to forecast toxicity.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction and Background
Toxicity means the extent to which a drug compound is toxic to living beings. Prediction of toxicity is a great challenge [1]. Toxicity can cause death, allergies, or adverse effects on a living organism, and it is associated with the number of chemical substances inhaled, applied, or injected [2]. There is a narrow gap between the effective quality of a drug and the toxic quality of the drug. A drug is required to help in illness, diagnosis of a disease, or prevention of disease [3]. The development of a new drug or chemical compound is quite an expensive and complex process.
A subset of artificial intelligence is machine learning [4]. It is a study of computer algorithm that is automatically improved through experience. Machine learning algorithm creates models based on training data to make predictions without explicit programming [5]. It can learn and enhance the ability to decision-making when introduced to new data. So with the help of these algorithms, models can gain knowledge from experience and enhance their capacity for acting, planning, and thinking [6]. The field of health care has made substantial use of machine learning techniques [7].
Feature selection is a method for choosing pertinent features from a dataset and removing irrelevant features [8]. Feature selection is employed to demonstrate the ranking of each feature with the variances. The input variables used in machine learning models are called features. Essential and non-essential features are part of the input variables [9]. The irrelevant and non-essential features can make the optimal model weaker and slower. Two main feature selection techniques are supervised and unsupervised. Algorithms are essential to anticipate toxicity in the age of artificial intelligence [10]. These techniques make it easier for models to infer intended outcomes from historical data and incidents. Every machine learning technique must ensure an optimal model that will predict the desired outcome best [11].
The ensemble method is a technique that combines multiple base classifiers to generate the best prediction model [12]. The ensembling technique focuses on considering a number of the base model into account and optimizing/averaging these models to provide one final model instead of constructing an individual model and expecting it to predict the paramount outcome [13].
2 Literature Review
In this section, the related work based on various techniques used in machine learning models is deliberated. Ai utilized SVM and the Recursive Feature Elimination (RFE) approach, he created a regression model [14]. Hooda et al. introduced a better feature selection ensemble framework for classifying hazardous compounds, using imbalanced and complex pharmacological data of high dimensions to create an improved model [15]. The Real Coded Genetic Algorithm was used by Pathak et al. to assess the significance of each feature, and k cross-validation was employed to assess the resilience of the best prediction model [16]. Collado et al. worked on a class balancing problem and provided an effective solution for class imbalance datasets to predict toxicity [17]. Cai et al. discussed the challenge in the analysis of high dimensional data in ML and provided effective feature selection methods to improve the learning model [18]. Austin et al. assessed the impact of missing members on the accuracy of the forecast and looked at the impacts missing members had on a voting-based ensemble and a stacking-based ensemble [19]. Invasive ductal carcinoma (IDC) stage identification is very time-consuming and difficult for doctors, as Roy et al. explained, thus they created a computer-assisted breast cancer detection model employing ensembling [20]. Takci et al. discussed the problem of the prediction of heart attack is necessary, especially in low-income countries, and determined the ML model to predict heart attacks [21]. Gambella et al. presented mathematical optimized models for advanced learning. The strengths and weaknesses of the models are discussed and a few open obstacles are highlighted [22]. Tharwat et al. proposed a new version of Grey Wolf optimization to adopt prominent features and to reduce the computational time for the process. These encouraging findings mark a significant step forward in the development of a completely automated toxicity test using photos of zebrafish embryos employing machine learning techniques and the next iteration of GWO [23].
The rest of the paper is organized as Sect. 3 explains the research methodology and the results with discussions are explained in Sect. 4. Finally, the last Sect. 5 concludes the work performed.
3 Proposed Methodology
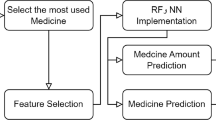
Computer-aided models are examined in this research. Nine machine learning algorithms are considered such as Gaussian Process, Linear Regression, Artificial Neural Network, SMO, Kstar, Bagging, Decision Tree, Random Forest, and Random Tree to predict toxicity. We developed an optimized regression model (Optimized KRF) by ensembling Kstar and Random Forest algorithm. For the mentioned machine learning models, evaluation parameters are assessed. The results in terms of accuracy are compared and assessed. Ten folds of cross-validation are used to create a robust model. The proposed methodology’s workflow procedure is depicted in Fig. 1.
Figure 2 represents the methodology for an ensembled model. Classifier -1 and classifier- 2 are applying a lazy and eager algorithm for prediction. Further ensembling is performed using different algorithms.
4 Results and Discussion
In this paper, the toxicity dataset is acquired from UCI machine learning datasets “UCI Machine Learning Repository: QSAR aquatic toxicity Data Set” and is used to assess how well learning models perform. The dataset consists of 546 occurrences and 9 attributes (one class attribute and eight predictive attributes). Table 1 lists the specifics of the ranking-related attributes. The ranking of important features is done using the correlation attribute evaluator method.
The coefficient of correlation in Table 2 is calculated with the help of Eq. (1) and mentioned as:
where \({\rho }_{PQ}\) is correlation coefficients, n represents the size, P, and Q are selected features and Ʃ is the summation symbol.
We have considered 9 machine learning algorithms such as Gaussian Process, Linear Regression, Artificial Neural Network, SMO, Kstar, Bagging, Decision Tree, Random Forest, and Random Tree to predict toxicity. Parameters are evaluated for the mentioned machine learning models. We calculated and compared how accurate each model is to select the best predictive model. The model is validated using the tenfold cross-validation method.
In the study, we developed an optimized regression model (Optimized KRF) by ensembling Kstar and Random Forest algorithm. Further Saw score is calculated in two aspects as W-Saw score and the L-Saw score. Gaussian Process, Linear Regression, Artificial neural Network, SMO, Kstar, Bagging, Decision Tree, Random Forest, and Random Tree achieved 53%, 58%, 50%, 57%, 64%, 60%, 54%, 63%, 60% accuracy respectively. The optimized model gave a coefficient of correlation, coefficient of determination, mean absolute error, root mean square error, and accuracy of 0.9, 0.81, 0.23, 0.3, and 77% respectively.
The W-Saw score for the optimized ensembled model is 0.83 which is the maximum and the L-Saw score is 0.27 which is the lowest in comparison to other classifiers. Saw score provides the strength to an ensemble model. Table 3 shows the state of art parameters evaluated and Fig. 3 represents the Comparison of the coefficient of correlation and determination graphically.

The prediction and ensembling algorithm is presented above in terms of lazy and eager classifiers. Table 4 represents the accuracy of several models.
Figure 4 depicts a comparison of accuracy for several models graphically. The saw score is a multi-attribute score based on the concept of weighted summation. This will seek weighted averages of rating the performance of each alternative. W-Saw score in Table 5 will be the highest score among all alternatives and is recommended as shown in Eq. (2). Figure 5 represents W-Saw scores for different models.
Highest Score Recommender
L-Saw Score in Table 6 is the score evaluated among alternatives and the lowest score is recommended as shown in Eq. (3). Figure 6 represents L-Saw scores for different models.
Lowest Score Recommender.
5 Concluding Remarks and Scope
To reduce the period and complexity of toxicity prediction, we have to develop intelligent systems for living beings so that they can reveal the possibilities of toxicity. Machine learning has significance in toxicity prediction. In this study, nine machine learning algorithms were taken into account such as Gaussian Process, Linear Regression, Artificial neural Network, SMO, Kstar, Bagging, Decision Tree, Random Forest, and Random Tree to predict the toxicity of a drug. In the study, we developed an optimized regression model (Optimized KRF) by ensembling Kstar and Random Forest algorithm. Parameters are evaluated for the mentioned machine learning models. The technique of tenfold cross-validation is used to validate the model. The optimized ensembled model gave a correlation coefficient, coefficient of determination, mean absolute error, root mean square error, and accuracy of 0.9, 0.81, 0.23, 0.3, and 77% respectively. Further Saw score is calculated in two aspects as W-Saw score and the L-Saw score. The W-Saw value in the ensembled model is 0.83 which is the maximum and the L-Saw value for the ensembled model is 0.27 which is the lowest in comparison to other classifiers. Saw score provides the strength to an ensemble model. These parameters indicate that the optimized ensembled model is more reliable and made predictions more accurately than earlier methods. The study can be extended to the ensembling of other classifiers to get higher accuracy and fewer errors. Other techniques can be applied for feature selection, class balancing, and optimization.
References
Karim, A., et al.: Quantitative toxicity prediction via meta ensembling of multitask deep learning models. ACS Omega 6(18), 12306–12317 (2021)
Borrero, L.A., Guette, L.S., Lopez, E., Pineda, O.B., Castro, E.B.: Predicting toxicity properties through machine learning. Procedia Comput. Sci. 170, 1011–1016 (2020)
Zhang, L., et al.: Applications of machine learning methods in drug toxicity prediction. Curr. Top. Med. Chem. 18(12), 987–997 (2018)
Ali, M.M., Paul, B.K., Ahmed, K., Bui, F.M., Quinn, J.M., Moni, M.A.: Heart disease prediction using supervised machine learning algorithms: performance analysis and comparison. Comput. Biol. Med. 136, 1–10 (2021)
Alyasseri, Z.A.A., et al.: Review on COVID-19 diagnosis models based on machine learning and deep learning approaches. Expert. Syst. 39(3), 1–32 (2022)
Tuladhar, A., Gill, S., Ismail, Z., Forkert, N.D.: Building machine learning models without sharing patient data: a simulation-based analysis of distributed learning by ensembling. J. Biomed. Inform. 106, 1–9 (2020)
Ramana, B.V., Boddu, R.S.K.: Performance comparison of classification algorithms on medical datasets. In: 9th Annual Computing and Communication Workshop and Conference, CCWC 2019, pp. 140–145. IEEE (2019)
Latief, M.A., Siswantining, T., Bustamam, A., Sarwinda, D.: A comparative performance evaluation of random forest feature selection on classification of hepatocellular carcinoma gene expression data. In: 3rd International Conference on Informatics and Computational Sciences, ICICOS 2019, pp. 1–6. IEEE (2019)
Babaagba, K.O., Adesanya, S.O.: A study on the effect of feature selection on malware analysis using machine learning. In: 8th International Conference on Educational and Information Technology, ICEIT 2019, pp. 51-55. ACM, Cambridge (2019)
Feng, L., Wang, H., Jin, B., Li, H., Xue, M., Wang, L.: Learning a distance metric by balancing kl-divergence for imbalanced datasets. IEEE Trans. Syst. Man Cybern. Syst. 49(12), 2384–2395 (2018)
Rawat, D., Pawar, L., Bathla, G., Kant, R.: Optimized deep learning model for lung cancer prediction using ANN algorithm. In: 3rd International Conference on Electronics and Sustainable Communication Systems, ICESC 2022, pp. 889–894. IEEE (2022)
Sarwar, A., Ali, M., Manhas, J., Sharma, V.: Diagnosis of diabetes type-II using hybrid machine learning based ensemble model. Int. J. Inf. Technol. 12, 419–428 (2020)
Uçar, M.K.: Classification performance-based feature selection algorithm for machine learning: P-score. IRBM 41(4), 229–239 (2020)
Ai, H., et al.: QSAR modelling study of the bioconcentration factor and toxicity of organic compounds to aquatic organisms using machine learning and ensemble methods. Ecotoxicol. Environ. Saf. 179, 71–78 (2019)
Hooda, N., Bawa, S., Rana, P.S.: B2FSE framework for high dimensional imbalanced data: a case study for drug toxicity prediction. Neurocomputing 276, 31–41 (2018)
Pathak, Y., Rana, P.S., Singh, P.K., Saraswat, M.: Protein structure prediction (RMSD ≤ 5 Å) using machine learning models. Int. J. Data Min. Bioinform. 14(1), 71–85 (2016)
Antelo-Collado, A., Carrasco-Velar, R., García-Pedrajas, N., Cerruela-García, G.: Effective feature selection method for class-imbalance datasets applied to chemical toxicity prediction. J. Chem. Inf. Model. 61(1), 76–94 (2020)
Cai, J., Luo, J., Wang, S., Yang, S.: Feature selection in machine learning: a new perspective. Neurocomputing 300, 70–79 (2018)
Austin, A., Benton, R.: Effects of missing members on classifier ensemble accuracy. In: IEEE International Conference on Big Data, ICBD 2020, pp. 4998–5006. IEEE (2020)
Roy, S.D., Das, S., Kar, D., Schwenker, F., Sarkar, R.: Computer aided breast cancer detection using ensembling of texture and statistical image features. Sensors 21(11), 1–17 (2021)
Takci, H.: Improvement of heart attack prediction by the feature selection methods. Turk. J. Electr. Eng. Comput. Sci. 26(1), 1–10 (2018)
Gambella, C., Ghaddar, B., Naoum-Sawaya, J.: Optimization problems for machine learning: a survey. Eur. J. Oper. Res. 290(3), 807–828 (2021)
Tharwat, A., Gaber, T., Hassanien, A.E., Elhoseny, M.: Automated toxicity test model based on a bio-inspired technique and adaBoost classifier. Comput. Electr. Eng. 71, 346–358 (2018)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Rawat, D., Meenakshi, Bajaj, R. (2024). Optimized Ensembled Predictive Model for Drug Toxicity. In: Challa, R.K., et al. Artificial Intelligence of Things. ICAIoT 2023. Communications in Computer and Information Science, vol 1929. Springer, Cham. https://doi.org/10.1007/978-3-031-48774-3_23
Download citation
DOI: https://doi.org/10.1007/978-3-031-48774-3_23
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-48773-6
Online ISBN: 978-3-031-48774-3
eBook Packages: Computer ScienceComputer Science (R0)