Abstract
The COVID-19 pandemic caused by SARS-CoV-2 has shown a rapid increase in the number of infected patients with a remarkable mortality rate, making it a global public health concern. Because there is currently no specific anti-viral drug for the treatment of COVID-19, repurposing of already approved drugs for other diseases may be explored. Drug repurposing has become a promising approach due to the opportunity to reduce development timelines and overall costs. In this chapter, we will discuss various computational drug repositioning strategies, the current COVID-19 treatment scenario, and challenges to the correct interpretation of existing preclinical/clinical evidence, as well as the generation of new evidence related to drug repurposing.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
The process of discovering new uses for previously approved, discontinued, delayed, or drugs under investigation is referred to as drug repurposing. It is also referred to as repositioning, drug reprofiling, indication expansion, or indication shift [1]. Although drug reuse is not a new concept, it has gained popularity over the past few decades. More than a third of the authorized pharmaceutical products have been reconstituted, resulting in a 25% increase in overall annual revenue for the pharmaceutical operation [2].
According to CMR International’s recent Pharmaceutical Research and Development Handbook, more than 55 drugs were phased out during Phase III clinical development between 2008 and 2010 [3]. Although some drugs have failed in preclinical and early human studies, they are safe. This aspect of drug safety is very attractive for drug repositioning. In general, approved drugs are more likely to be safe in new indications and patient populations. The growing body of drug knowledge will shorten the development cycle and reduce the risk of development costs and costs associated with new molecular entities [4].
While drug repurpose is possible at any stage, it is most promising for drugs that are already approved [5]. Currently, advanced computer technology is being utilized to forecast novel drug targets or drug reuse. In comparison to high-throughput screening, which requires the assessment of hundreds of compounds, computer technology is rapid and affordable and may be used as a preliminary filtering approach. They are also beneficial for high-priority therapies that require more investigation and testing. The rationale for drug recycling is that numerous diseases may use the same metabolic processes [6]. Drug repurposing has significant regulatory and scientific consequences, notably in situations like the COVID-19. This chapter addresses drug repurposing techniques and methodologies.
Drug Repurposing
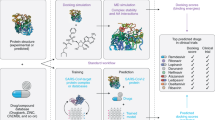
Traditional de novo drug discovery is an expensive and risky approach. Computational techniques for drug repurposing can significantly speed up the traditional drug discovery process (Fig. 24.1). Developing drugs traditionally is costly and time-consuming. It takes an average of 14 years and costs more than U.S. $2 billion to bring a drug to market. Approximately 90% of drugs fail throughout the drug development process owing to safety concerns or a lack of efficacy. The computational drug repurposing technique, which combines and analyses huge data sets on tens of thousands of drugs and diseases automatically, has the potential to significantly speed the traditional drug development process. As the COVID-19 epidemic expanded, two more impediments emerged: the speed with which therapeutic approval could be obtained and the urgency with which clinical requirements could be met. Appropriate regulatory measures should take the risk–benefit ratio into account. Drugs must be developed and authorized quickly to halt the spread of diseases [7]. Even if the proposed drug shows early efficacy in animal and clinical studies, it will take at least 2 years to reach the market. Because the manufacturer will conduct safety studies before the start of the clinical trial, and the clinical study can be delayed for up to 2 years [8,9,10].
While rapid development and decision-making can result in a more rapid release of drugs, complete safety, and effectiveness data are compromised. Although drug development takes an average of 12 to 15 years, it can be completed in as little as 12 to 18 months if the process is accelerated. To accelerate the completion of phase III clinical trials, shorter, fewer, or no phase III trials are required [11]. As a result, drug approval will require less information than usual in the event of a pandemic. This is possible when the period of low incidence coincides with the pandemic’s recruitment phase [12, 13]. Although COVID-19 lacks a broad legal framework because it is based on disease incidence, a small number of reusable drugs have been approved, owing in part to the evaluation of COVID-19’s benefit–risk profile and biomarker evidence [14]. Drugs that have been repurposed for COVID-19 have been approved for an emergency, conditional marketing, or early access to drugs with limited clinical data [15].
Although postmarketing studies are commonly used to expand Phase II studies, approval decisions are typically based on data from Phase III clinical trials [16,17,18,19,20]. It is generally recognized that the safety information provided for all pharmaceutical drugs at the time of approval is insufficient. The goal of efficacy-focused clinical development is to improve efficacy. As a result, the postauthorization risk management plan is data-driven to address any safety concerns that occur following authorization. These problems are exacerbated by the need for rapid development and approval. As a result, this is strongly advised, although, in some circumstances, authorities will need to undertake a complete postapproval examination and report to the Food and Drug Administration (FDA) regularly [21].
In Silico Drug Repurposing Methods
According to the Pharmaceutical Research and Manufacturers of America (PhRMA), only one out of every 5000–10,000 compounds discovered in drug research is likely to be commercialized [22]. The information age has transformed the process of drug development, providing massive amounts of data that have assisted in our knowledge of the molecular pathways behind human disease. While significant advances have been achieved in the postgenomic era, researchers continue to encounter problems in finding, collecting, and analyzing all relevant data on any human disease effectively and completely. This problem necessitates the efficient use of bioinformatics and computational methods. The primary obstacles include the following: (i) collecting relevant information from terabytes of data from various sources. (ii) data mining and knowledge management techniques are combined using coherent and manual search methods. (iii) effective data analysis to generate clinically relevant test hypotheses [23,24,25].
The complete sequencing of the human genome at the turn of the twentieth century reshaped the landscape of drug development. Currently, a large amount of data, such as the genome, transcriptome, proteome, metabolome, and pharmacological data, is dispersed among many databases that are publicly available through the Internet [26]. In addition, with the creation of complex protein and signaling pathway databases, the number of these databases has increased, and these databases collectively reflect the current understanding of disease mechanisms [27,28,29,30]. Likewise, the number of publications in scientific journals has increased along with the enormous amount of data. Due to these diverse resources, the development of bioinformatics and computational methods for collecting, analyzing, and interpreting data is essential [31]. This strategy has also led to the development of new theories linking disease with experimental or commercial drugs in conjunction with predictive algorithms [32,33,34]. These computational approaches to drug repositioning can be divided into the five strategies: knowledge mining and integration strategies, genetic analysis methods, connectivity map strategy, network analysis methods, and computer-aided drug design (Fig. 24.2).
Knowledge Mining and Integration Strategies
Drug development is often divided into five stages: preclinical and discovery, safety evaluation, clinical research, FDA review, and FDA postmarketing surveillance. This is a lengthy and costly process with a high rate of failure. On the other side, drug repositioning is a four-stage process: The FDA is responsible for the identification, acquisition, development, and postmarket safety monitoring of drugs [35]. Recent research has produced a large amount of experimental data on COVID-19. To discover new and hidden knowledge, a computer evaluation of the use of data science techniques must be carried out. To fully understand the biology and mechanism of SARS-CoV-2 and its process, the data must be fully comprehended.
There has been a wealth of solutions because of the challenge computer biologists confront in gathering and understanding the most relevant data from many sources for the goal of hypothesis creation. At a high level, the main differences between these approaches relate to identifying and evaluating key sources. Consequently, the establishment of an opportunity to reposition a drug requires both a major computer effort and a biological evaluation of the feasibility of action for the new drug suggestion. A computer strategy for new drug suggestions in its systemic form is typically an automated approach. This oversees the evaluation of proposed diseases as well as the design of experimental study concepts to test with the assistance of biologists or disease experts [36].
To incorporate knowledgeable disease-related data, a target-based information technology method can be combined with high-throughput data mining strategies. The challenge is not only determining which data to extract or which methodologies to employ but also which high-quality rich-content databases to use for hypothesis creation [37]. A critical component of any new approach for disease indicators is the final prioritization or classification of the different types of data necessary to conduct a thorough evaluation of the evidence for each disease assumption. In addition, one disease hypothesis is compared to another via an absolute score system. The efficacy data from Phase II/III clinical trials are the most significant evidence for combining a biological target and a new indication for premarket drug candidates. Ongoing trial with no published data from clinical trials suggests preclinical data with slightly lower evidence [38]. The third level of evidence for target participation in a novel indication would be genetic evidence from functional polymorphic or associated human disease research, as well as mouse knockout data. Finally, evidence gathered from a variety of bibliographic databases can be used to generate new hypotheses for indications that can then be included in the disease classification system using advanced text mining techniques [39,40,41,42].
Genetic Analysis Methods
Using genetic studies to identify “druggable” targets is one method to improve the chances of successful drug repositioning. The advent of large-scale genetic research, mostly in the form of genome-wide association studies (GWAS), has significantly expanded our understanding of the genetic basis of a variety of diseases and enabled researchers to use this information to develop therapeutic targets. Many GWAS conducted in recent years represents a large number of potential new drug retargeting libraries. Disease–gene associations are identified by combining functional genomic data with advanced computational approaches. Genetic analysis is an effective strategy for coming up with new therapeutic indication hypotheses. This information is especially useful when it is backed up by other sources of information, such as clinical expression analysis or preclinical research [43,44,45]. The approach can provide several high-end repositioning hypotheses depending on the nature and source of genomic data. Because of the success of this technology, many sequencing, expression, and phenotype genetic analytical methodologies have been developed [46, 47]. One of the limitations of the GWAS study is that some of the top genes identified may not be druggable. Second, relying exclusively on the impact of the most significant SNPs may lead to the omission of physiologically important target genes. Third, focusing upon only candidate genes may result in the omission of multi-target drugs, which may be more effective in some cases than single-target drugs. Fourth, due to the complexity of the human genome, there is no one-size-fits-all approach to accurate annotation.
Connectivity Map Strategy
The Connectivity Map concept may be the most unique strategy for discovering new indications in the process of drug repositioning. This is a computational method that uses microarray-based transcription profile data to generate new ideas for disease indications and is easy to apply to other “omics” platforms [48]. This technique can theoretically link small molecule drugs to their mode of action as well as novel diseases by using their corresponding gene expression. The stronger the relationship between the disease signature and the inverse drug profile, the more likely the drug is to reverse the disease genotype/phenotype [49].
Connectivity mapping is a method for assembling reference transcriptional profiles obtained from microarrays by assessing the differential expression of a cell line treated with a series of drugs in comparison to untreated controls. The differential expressions’ rank order for each compound is then generated. Disease signatures can be derived from a variety of sources, including disease microarrays and previously published data. By ranking all connection levels in descending order and setting the relevance threshold, the drug with the greatest score is chosen [50, 51]. As a result, the ideal treatment will have an exact inverse signature of the disease state, restoring the normal phenotype. This technique has the advantage of being platform-independent for compound reference profiles and disease signature sources. Because it is a computational approach, it has the potential to quickly identify many probable direct and inverse correlations between a wide range of disease conditions. The Connectivity Map method is based on the Kolmogorov-Smirnov statistic and employs Gene Set Enrichment Analysis, a nonparametric, rank-based pattern-matching tool. The expression profiles of the compound reference genes are also nonparametrically represented, but each gene is rank ordered depending on its degree of differential expression relative to untreated controls [52]. This method has a limitation in terms of experimental replicates, which can be challenging given that most small compounds have only one copy per cell line per experiment. Another potential limitation is the occurrence of batch processing effects.
Network Analysis Methods
Network analysis tools greatly simplify the investigation of the complexity of biological systems and the diversity of different types of data defining a disease state. These techniques can be used to mathematically and graphically represent the various protein interactions that occur in higher species. In general, these strategies focus on concepts that are overrepresented in the pattern of the nodes and edges of a biological network [53, 54]. This approach could be extended to network hubs that appear to be key proteins in protein interactions. These hubs can be useful as important intervention sites in a specific disease condition, making them potential drug targets [55]. The current state of knowledge about molecular interaction networks is incomplete, and the associated configuration profiles are also very noisy, which are the limitations. Interactomes, on the other hand, only provide static snapshots of biological systems, whereas it contains dynamic systems. There is a paucity of information on precise interaction kinetics, and there is no obvious link between the molecular origin and the organism’s reaction.
Transcriptome data in the gene regulatory network may capture the dynamic features of cells and give an in-depth knowledge of drug action mechanisms. A network analysis approach for differential expression data has been developed to identify genes from specific disease linkage and associated regions. Most gene prioritization methods necessitate prior knowledge of disease processes to identify potential drug targets. Differential gene expression studies show that gene expression patterns change systemically as diseases develop. Consequently, diverse gene expression patterns might be used as input to prioritize prospective therapeutic targets. However, this technique has certain drawbacks, such as the difficulty in defining robust gene features owing to noise in the expression of some genes. Furthermore, drug targets and genes controlled by targets may not always have significant expression changes [56].
Compounds and metabolites are depicted as nodes in the metabolic network. Excessive chemical concentrations (mass flow) produced by specific enzymes can result in disease. These enzymes may be therapeutic targets for this disease due to their ability to manipulate the concentration of disease-causing compounds via drug manipulation. Flux balance analysis is an example of an approach for identifying pharmacological targets.
The protein–protein interaction network (PPIN) is a form of a molecular interaction network that displays the interaction between a known drug target and other proteins, as well as proteins that interact indirectly with the target. Due to the difficulty of scanning the entire PPIN subnetwork space, advanced mathematical techniques will be required to detect out-of-tune subnetworks effectively. While PPIN’s drug repositioning strategy has been extremely successful, it does have certain limitations. PPINs include links to undefined potential functionalities derived from several experimental sources. Additionally, the required data is noisy and incomplete, resulting in a bias in the generated network.
The network-based method is an effective way to link the molecular and phenotypic levels to determine drug targets. Determining the interaction between the drug and the target protein is an important step in drug discovery. Drug discovery and design are mainly based on the interaction between the drug and the target. Many drugs are nonspecific and respond to other targets in addition to the main target. In this case, a drug interaction is used to clarify the relationship between two drugs based on their similarity. The association with drug diseases includes various association modes, such as drug indications and drug side effects. In the disease–disease interaction, it has been proposed that two drugs with similar molecular pathophysiology can be interchanged. When effective drug repositioning is required, each network approach has limitations that can be overcome by combining data from several sources, such as molecular interaction networks and gene expression profiles.
Computer-Aided Drug Design
Drug design may be divided into two categories: structural drug design and ligand-based drug design. A structural drug design is predicated on knowledge of the three-dimensional structure of the protein target as determined by X-ray crystallography or nuclear magnetic resonance spectroscopy. To carry out the structural drug design paradigm, an atomic-resolution three-dimensional protein structure of the receptor is required. A well-resolved crystal structure is preferred, with a resolution of at least 2.5 is often regarded as sufficient. If the target’s three-dimensional structure is unknown, a protein model can be built by homology modeling to the nearest target-related protein having a known and accessible three-dimensional structure. Molecular docking can anticipate the intermolecular framework formed by a protein and a small molecule or another protein, as well as the binding modes that inhibit the protein. The virtual screening (VS) method compares the target protein to databases that contain millions of drug-like or lead-like compounds with well-defined three-dimensional structures [57]. To conduct the computational screening, the ligands’ binding affinities are compared using a docking method. Ligand-based drug design is a method for finding compounds that bind to a protein target without knowledge of the three-dimensional receptor. The quantitative structure–activity relationship (QSAR) and pharmacophore modeling techniques are essential in ligand-oriented drug discovery because they provide statistical models for finding and optimizing leads [58]. However, using molecular docking for drug repositioning has several drawbacks. Docking applications are severely limited by the requirement for known chemical ligands and three-dimensional (3D) structures of protein targets because the structures of many physiologically significant proteins are still unknown. Furthermore, the molecular docking method is computationally intensive, which could lead to longer processing times. Furthermore, molecular docking studies have a high rate of false positives due to structural flaws in some proteins and inadequate modeling of atomic and molecular interactions. Machine learning approaches appear to be less expensive than docking simulations, since they can test more potential candidates for subsequent experimental screening.
In Silico Drug Repurposing for COVID-19
Previously approved FDA drugs for other diseases were repositioned for COVID-19 treatment using various computational methods. Network-based algorithms, expression-based algorithms, and docking simulations were used to identify the drug that was predicted for drug repurposing in COVID-19 therapies. However, the accuracy of the predictions can be determined by comparing the reported computer studies’ final candidate drug lists to the drugs currently undergoing clinical trials on clinicaltrials.gov (https://clinicaltrials.gov/). Numerous computational studies predicted and repurposed drugs for COVID-19 treatment, include chloroquine, hydroxychloroquine, remdesivir, lopinavir–ritonavir, ivermectin, favipiravir, oseltamivir, ribavirin, corticosteroids, and tocilizumab. This section will examine the clinical success of computationally predicted repurposed drugs [6]. This section contains comprehensive information on the drugs that have been largely repurposed and are currently being used to control SARS-CoV-2 infections as determined through computational methods. It is attempted to compile and review research on selected drugs using data from the WHO COVID guidelines, clinicaltrials.org, and a variety of other sources (at the time of writing this manuscript, September 2021).
Chloroquine
Chloroquine has been repurposed as a COVID-19 drug due to its potent antiviral activity. Chloroquine is used to make chloroquine, a malarial drug that was traditionally found in the bark of Peruvian Kinchon trees. According to in-vitro research, chloroquine is a powerful bioactive agent against RNA viruses. Chloroquine has been found to have potential therapeutic effects against the coronavirus of SARS-CoV-1. The extensive antiviral activity against COVID-19 evaluated for chloroquine. While these results are preliminary, they are widely welcomed by the media, and some well-known personalities, including certain health regulatory authorities, have encouraged the use of chloroquine and hydroxychloroquine to treat COVID-19 [59]. However, little is known about the effect of chloroquine and hydroxychloroquine on the frequency and severity of adverse drug reactions (ADRs). Significant clinical studies have demonstrated that hydroxychloroquine and chloroquine are unlikely to be effective in treating or preventing COVID-19, prompting the U.S. Food and Drug Administration (FDA) to revoke the drug’s emergency use authorization on June 15, 2020 [60]. According to the current meta-analysis of randomized studies, the use of chloroquine in COVID-19 patients has no benefit [61].
Hydroxychloroquine
Hydroxychloroquine (a chloroquine derivative) is an antimalarial drug with anti-inflammatory characteristics. It has been used successfully as a safe anti-inflammatory medicine in auto-immune diseases, and preliminary research suggests that it may decrease pro-inflammatory cytokine expression in COVID-19. However, whether it is effective in eliminating the SARS-CoV-2 virus in ICU patients with overactive immune systems, particularly when the lungs are severely inflamed, has yet to be scientifically demonstrated [62, 63]. The available clinical evidence appears to be insufficient to prove the efficacy of HCQ in severely ill COVID-19 patients [61]. However, hydroxychloroquine, like chloroquine, has been demonstrated to be unsuccessful in the treatment or prevention of COVID-19, forcing the U.S. Food and Drug Administration (FDA) to revoke the drug’s emergency use authorization on June 15, 2020. The recent solidarity trial, the UK’s recovery trial, and a Cochrane review of other evidence on hydroxychloroquine conclusively showed that hydroxychloroquine did not reduce deaths among hospitalized COVID-19 patients.
Lopinavir–Ritonavir
In vitro and animal studies have shown that lopinavir is effective against MERS-CoV. Additionally, lopinavir inhibits SARS-CoV by inhibiting a critical reproductive protease that appears to be highly conserved in SARS-CoV-2. In several countries, lopinavir–ritonavir therapy is recommended as first- or second-line treatment for COVID-19. Although several observational studies have demonstrated that lopinavir–ritonavir is associated with decreased viral shedding and fever in patients with COVID-19. Recent research on COVID-19, however, has found that regular lopinavir–ritonavir supplementation provides no benefit [64]. The lopinavir/ritonavir did not show efficacy in two large randomized controlled trials in hospitalized patients with COVID-19. The COVID-19 treatment guidelines panel recommends against the use of lopinavir/ritonavir for the treatment of COVID-19 in hospitalized patients [65].
Remdesivir
Remdesivir is a novel and effective intravenous antiviral medication. It is effective against COVID-19 as well as other beta-coronaviruses in the same family. The laboratory investigation has shown that human cells can be protected from becoming infected with COVID-19. Furthermore, the findings suggest that in vitro remediation is highly successful in eradicating COVID-19 infection. Remdesivir was approved for emergency use in over 50 countries during the COVID-19 pandemic. The FDA has approved Remdesivir for the treatment of COVID-19 infection in adults and hospitalized children (over the age of 12 and weighing less than 45 kg). In adult COVID 19 patients, remdesivir, on the other hand, was not associated with statistically significant therapeutic benefits. Additional international research is necessary to establish the drug’s efficacy and safety in COVID-19 patients [66]. An interesting study revealed that RDV’s parent nucleotide, GS-441524, is more effective and less hazardous than its prodrug form and has been proven to be efficacious in vivo veterinary settings. As a result, future research into the parent nucleotide’s usage against COVID-19 should proceed at a quicker speed.
Favipiravir
Favipiravir is a novel RNA-dependent RNA polymerase (RdRp) inhibitor that is successful in the treatment of Ebola. Favipiravir is considered as a feasible treatment for COVID-19 due to its efficacy against a variety of viral diseases. Some research has demonstrated that favipiravir can accelerate virological clearance and clinical improvement, but most investigations are hampered by potential confounding factors (e.g., concurrent use of immunomodulators and other therapies). The study found no improvement in terms of mortality. A systematic review found that when given during the first seven days of hospitalization, there is a possibility of clinical improvement, but no statistically significant decrease in mortality for any of the groups studied, including hospitalized patients and those with mild or moderate symptoms. Other well-designed studies on dosage and duration of treatment, are essential to draw clear conclusions [67]. Any approval for the use of favipiravir, on the other hand, will require more clinical research, followed by approval for public use by the country’s competent regulatory body [68].
Oseltamivir
Oseltamivir inhibits the neuraminidase enzyme, which is expressed on the viral surface against the influenza virus and is also effective for various avian influenza virus strains. The enzyme promotes the release of virus from infected cells and facilitates viral movement within the respiratory tract. In the presence of neuraminidase inhibitors, virions stay attached to the membrane of infected cells and are also entrapped in respiratory secretions [69]. Clinical trials are also being conducted using oseltamivir in combination with various chloroquine and favipiravir regimens. A study showed that the drug exhibited no positive result on COVID-19. Additional clinical trials and larger, randomized controlled trials are required to demonstrate Oseltamivir’s efficacy in patients with COVID-19 [70].
Ribavirin
Ribavirin is an antiviral drug that prevents viruses from replicating and spreading [71]. Ribavirin has been approved for COVID-19 therapy in combination with interferon alfa or lopinavir–ritonavir. Ribavirin is effective against the Middle East respiratory syndrome corona virus (MERS-CoV) in vitro and in vivo, and case reports demonstrate that ribavirin paired with interferon alfa resulted in virologic clearance and survival. Clinical trial evidence demonstrating ribavirin’s efficacy in treating COVID-19 is still insufficient [72].
Ivermectin
Ivermectin is a medication used to treat parasite infestations and it was repositioned for COVID-19 treatment. Throughout the COVID-19 epidemic, misinformation suggesting that ivermectin helps treat and prevent COVID-19 has been extensively propagated. These claims are unsupported by reliable scientific evidence. Multiple major health organizations, including the Food and Drug Administration, the United States Centres for Disease Control, the European Medicines Agency, and the World Health Organization, have declared that ivermectin is not authorized or approved to treat COVID-19 [73].
Corticosteroid
Corticosteroids are anti-inflammatory drugs that suppress the immune system. Dexamethasone is a glucocorticoid drug used in the treatment of rheumatoid arthritis, a range of skin diseases, severe allergies, asthma, and chronic obstructive pulmonary disease. According to WHO guidelines, corticosteroids should be used only in patients with severe or critical COVID-19 infection and not in individuals with nonsevere COVID-19 infection (absence of criteria for severe or critical infection). Dexamethasone was authorized by the European Medicines Agency (EMA) in September 2020 for use in adults and adolescents (12 years of age and a weight of at least 40 kg) who need supplemental oxygen therapy. Prolonged corticosteroid treatment may lead to the development of the so-called long COVID syndrome, which is characterized by tiredness and psychological problems and may be exacerbated by steroid-related adverse medication responses such as myopathy, neuromuscular weakness, and mental symptoms. Thus, corticosteroids seem to be a double-edged sword in the battle against COVID-19 and should be used with caution, considering the risk–benefit ratio, as a short-course treatment drug in a limited group of patients with COVID-19 who have been documented to benefit from survival. The safety and efficacy of corticosteroids in combination with antiviral medication for the treatment of COVID-19 have not been extensively studied in clinical trials [74].
Tocilizumab
Tocilizumab (TCZ) is a recombinant humanized anti-interleukin-6 receptor monoclonal antibody used to treat systemic juvenile idiopathic arthritis (SJIA), polyarticular juvenile idiopathic arthritis (pJIA), and rheumatoid arthritis. In June 2021, the FDA issued tocilizumab an emergency use authorization for the treatment of COVID-19 in hospitalized patients aged two years and older who need supplementary oxygen, noninvasive or invasive mechanical ventilation. The COVID-19 Treatment Guidelines panel recommends tocilizumab in combination with dexamethasone in certain hospitalized patients who are having rapid respiratory decompensation due to COVID-19 [75]. The European Medicines Agency (EMA) is now assessing tocilizumab for the treatment of hospitalized individuals with severe COVID-19 who are already undergoing corticosteroid therapy and need additional oxygen or mechanical ventilation as of August 2021.
Opportunities and Challenges
The traditional approach to pharmaceutical drug development is costly, time-consuming, and prone to failure. Repositioning, on the other hand, is a low-risk strategy that saves time and money. Experiments like cell-based assays, protein-based assays, animal models, and clinical trials provide a direct, evidence-based, and accurate understanding of the drug–disease connection. As a result of the availability of experimental data, computational techniques for drug repositioning have gained popularity in recent years, and they are frequently merged with them to provide accurate results. While there are many excellent computer models for drug repositioning, creating a reliable model is a difficult and time-consuming process. One of the major challenges is that theoretical calculation techniques are difficult to put into practice due to the difficulty of mapping such theoretical approaches to replicate biological activity, as well as other barriers such as missing, distorted, or erroneous data. For example, creating an accurate gene expression profile may be difficult due to a variety of factors, such as changes in experimental conditions throughout multiple trials, resulting in data mismatches in gene expression features and hence data bias. Furthermore, when these genes are employed as pharmacological targets, significant changes in gene expression may not occur consistently, resulting in incorrect findings. Furthermore, chemical structure and molecular information techniques are challenging to find probable drug–target interactions due to a lack of high-resolution structural data on drug targets. Another issue with computational drug repositioning models is the lack of a reliable gold standard data set for assessing their efficacy. Furthermore, the model’s recommended potential repositioning drugs cannot be evaluated without clinical validation of safety criteria and proof of their effectiveness against anticipated diseases.
Clinical trials are typically preceded by preclinical investigations, both in vitro and in vivo. In the event of drug repurposing, preclinical research on the impact of a drug on disease should be prioritized. However, it requires a thorough knowledge of the disease process, which in the case of COVID-19 may be difficult. In contrast to clinical trials, preclinical research may rapidly assess whether a certain pharmacological approach is likely to be worth pursuing. Preclinical studies, by definition, offer an inadequate picture of disease biology and may produce inconsistent findings, and the knowledge they can provide regarding drug safety and efficacy is severely restricted. Randomized controlled trials are the gold standard for gathering evidence on drug efficacy in clinical trials because they allocate possible confounders to treated and untreated patients at random. The clinical proof is needed to demonstrate a drug’s effectiveness and safety, even if preclinical research indicates biological plausibility. During a pandemic, conducting methodologically sound research may be difficult since the rapid spread of low-quality findings may have severe implications. Several considerations highlight the necessity of interpreting current clinical data with great care.
As a result, effective drug repositioning necessitates a combination of computer prediction and in-vitro validation or retrospective clinical history analysis. The comprehensive methodology, which includes a combination of computational and experimental methodologies, enables a comprehensive assessment of all repositioning possibilities. Effectiveness and timeliness of repositioned drugs are greatly improved using a multimodal approach to pharmacological reconstitution [76]. Certain legal concerns may make it difficult to patent a novel therapeutic application and/or enforce patent rights, reducing the incentives for drug repurposing. Certain national regulations make obtaining a patent for a second or subsequent medical use more challenging, although repurposed medicinal uses are protected in most of the major pharmaceutical markets. While patents can be awarded for off-patent drugs, enforcement may be an issue if the new indication makes use of currently available strengths and dose formulas.
Combining and integrating all the approaches will open up a plethora of new possibilities for drug development, most notably through the construction and access to massive databases of drug and disease omics data. Researchers now have access to the most up-to-date, reliable tools and data to investigate unknown mechanisms of action/pathways based on the target protein and/or biomarkers associated with disease progression. Thanks to advancements in techniques such as genomics, proteomics, transcriptomics, and metabolomics, numerous datasets and tools and pathway analysis are publicly available [77]. Several algorithms have been developed to improve the speed and convenience of the recalculation methods. Pharmaceutical repositioning is similarly fraught with difficulties. Repositioning an existing drug by adding a new therapeutic indication is a difficult and complex task because it involves numerous factors, like technology, commercial strategy, patents, investment, and market demand. Selection of an appropriate medicinal product sector, clinical trial issues such as outdated or inadequate clinical or preclinical data on the original pharmaceutical or drug product are few prominent challenges.
Conclusion
There are currently no drugs available that are effective in treating COVID-19 patients. While research continues, some countries are experimenting with various combinations to treat their patients. While computing can aid in repurposing, data from in vitro drug screening, in vivo research, including animal models, ongoing clinical trials, electronic health records, literature mining, or expert knowledge must be used to evaluate computational drug repurposing methods. Numerous candidate drugs continue to exhibit experimental flaws, necessitating comprehensive validation of candidate therapeutics to establish a baseline for technique accuracy. Because this is impractical, it has become even more critical to combine projections with expert knowledge. Researchers and scientists should avoid duplicating studies and organize studies so that the outcomes can be compared. It is also necessary to conduct a critical analysis of existing data to determine the efficacy and safety of a drug for possible repurposing. As a result, additional clinical trials and large randomized control studies are required through international collaboration to improve the treatment options and safety of COVID-19 patients.
References
Saini M, Parihar N, Soni SL, Sharma V. Drug repurposing: an overview. Asian J Pharm Res Dev. 2020;8(4):194–212.
Talevi A, Bellera CL. Challenges and opportunities with drug repurposing: finding strategies to find alternative uses of therapeutics. Expert Opin Drug Discovery. 2020;15(4):397–401.
Poduri R. Pharmaceutical industry, academia, regulatory authorities and end user collaboration in successful drug discovery and development. In: Drug Discovery and Development: From Targets and Molecules to Medicines. Singapore: Springer; 2021. p. 465–71.
Sonaye HV, Sheikh RY, Doifode CA. Drug repurposing: iron in the fire for older drugs. Biomed Pharmacother. 2021;1(141):111638.
Shyr ZA, Gorshkov K, Chen CZ, Zheng W. Drug discovery strategies for SARS-CoV-2. J Pharmacol Exp Ther. 2020;375(1):127–38.
Galindez G, Matschinske J, Rose TD, Sadegh S, Salgado-Albarrán M, Späth J, Baumbach J, Pauling JK. Lessons from the COVID-19 pandemic for advancing computational drug repurposing strategies. Nat Comput Sci. 2021;1(1):33–41.
Garrison LP Jr, Towse A, Bresnahan BW. Assessing a structured, quantitative health outcomes approach to drug risk-benefit analysis. Health Aff. 2007;26(3):684–95.
Ross JS, Mulvey GK, Hines EM, Nissen SE, Krumholz HM. Trial publication after registration in ClinicalTrials. Gov: a cross-sectional analysis. PLoS Med. 2009;6(9):e1000144.
Paul SM, Mytelka DS, Dunwiddie CT, Persinger CC, Munos BH, Lindborg SR, Schacht AL. How to improve R&D productivity: the pharmaceutical industry’s grand challenge. Nat Rev Drug Discov. 2010;9(3):203–14.
Sultana J, Crisafulli S, Gabbay F, Lynn E, Shakir S, Trifirò G. Challenges for drug repurposing in the COVID-19 pandemic era. Front Pharmacol. 2020;6(11):1657.
Chong CR, Sullivan DJ. New uses for old drugs. Nature. 2007;448(7154):645–6.
Eichler HG, Pignatti F, Flamion B, Leufkens H, Breckenridge A. Balancing early market access to new drugs with the need for benefit/risk data: a mounting dilemma. Nat Rev Drug Discov. 2008;7(10):818–26.
Eichler HG, Oye K, Baird LG, Abadie E, Brown J, Drum CL, Ferguson J, Garner S, Honig P, Hukkelhoven M, Lim JC. Adaptive licensing: taking the next step in the evolution of drug approval. Clinical Pharmacology & Therapeutics. 2012;91(3):426–37.
Eichler HG, Bloechl-Daum B, Brasseur D, Breckenridge A, Leufkens H, Raine J, Salmonson T, Schneider CK, Rasi G. The risks of risk aversion in drug regulation. Nat Rev Drug Discov. 2013;12:907–16.
Sarkar C, Mondal M, Torequl Islam M, Martorell M, Docea AO, Maroyi A, Sharifi-Rad J, Calina D. Potential therapeutic options for COVID-19: current status, challenges, and future perspectives. Front Pharmacol. 2020;15(11):1428.
Baumfeld Andre E, Reynolds R, Caubel P, Azoulay L, Dreyer NA. Trial designs using real-world data: the changing landscape of the regulatory approval process. Pharmacoepidemiol Drug Saf. 2020;29(10):1201–12.
Schneeweiss S, Gagne JJ, Glynn RJ, Ruhl M, Rassen JA. Assessing the comparative effectiveness of newly marketed medications: methodological challenges and implications for drug development. Clin Pharmacol Therapeut. 2011;90(6):777–90.
Sharma MR, Schilsky RL. Role of randomized phase III trials in an era of effective targeted therapies. Nat Rev Clin Oncol. 2012;9(4):208.
Raphael MJ, Gyawali B, Booth CM. Real-world evidence and regulatory drug approval. Nat Rev Clin Oncol. 2020;17(5):271–2.
Swift B, Jain L, White C, Chandrasekaran V, Bhandari A, Hughes DA, Jadhav PR. Innovation at the intersection of clinical trials and real-world data science to advance patient care. Clin Transl Sci. 2018;11(5):450–60.
Angus DC. Optimizing the trade-off between learning and doing in a pandemic. JAMA. 2020;323(19):1895–6.
Prasad S, McOsker C, Foley M. Filling the drug discovery abyss. Drug Discovery. 2016;17:25.
Leelananda SP, Lindert S. Computational methods in drug discovery. Beilstein J Org Chem. 2016;12(1):2694–718.
Ou-Yang SS, Lu JY, Kong XQ, Liang ZJ, Luo C, Jiang H. Computational drug discovery. Acta Pharmacol Sin. 2012 Sep;33(9):1131–40.
Kumar N, Hendriks BS, Janes KA, de Graaf D, Lauffenburger DA. Applying computational modeling to drug discovery and development. Drug Discov Today. 2006;11(17–18):806–11.
Kumar S. Online resource and tools for the development of drugs against novel coronavirus. In: Silico Modeling of Drugs Against Coronaviruses. Methods in Pharmacology and Toxicology. New York, NY: Humana; 2021. https://doi.org/10.1007/7653_2020_53.
Kumar S. Protein–Protein Interaction Network for the Identification of New Targets Against Novel Coronavirus. In: Silico Modeling of Drugs Against Coronaviruses. Methods in Pharmacology and Toxicology. New York, NY: Humana; 2021. https://doi.org/10.1007/7653_2020_62.
Kumar S. COVID-19: A drug repurposing and biomarker identification by using comprehensive gene-disease associations through protein-protein interaction network analysis. Preprints. 2020:2020030440. https://doi.org/10.20944/preprints202003.0440.v1.
Nana KS, Karuppanan K, Kumar S. Identification of common key genes and pathways between Covid-19 and lung cancer by using protein-protein interaction network analysis. bioRxiv. 2021; https://doi.org/10.1101/2021.02.16.431364
Kumar S, Mathavan S, Jin WJ, Azman NA, Subramanaiam D, Zainalabidin NA, Lingadaran D, Sattar ZB, Manickam DL, Anbananthan PS, Taqiyuddin JA. COVID-19 Vaccine Candidates by Identification of B and T Cell Multi-Epitopes Against SARS-CoV-2. Preprints 2020. https://doi.org/10.20944/preprints202008.0092.v1.
Mathavan S, Kumar S. Evaluation of the Effect of D614G, N501Y and S477N Mutation in SARS-CoV-2 through Computational Approach. Preprints 2020. https://doi.org/10.20944/preprints202012.0710.v1.
Kumar S, Ramanujam M. Computational prediction of novel broad-spectrum drug targets against vibrio cholerae by integrated genomics and proteomics approach. Malaysian J Med Health Sci. 2020;16(2):99–104.
Kumar S. In Silico Identification of Novel Tuberculosis Drug Targets in Mycobacterium tuberculosisP450 Enzymes by Interaction Study with Azole Drugs. Preprints 2020.
Kumar S. Computational identification and binding analysis of orphan human cytochrome P450 4X1 enzyme with substrates. BMC Res Notes. 2015;8(1):1.
Karaman B, Sippl W. Computational drug repurposing: current trends. Curr Med Chem. 2019;26(28):5389–409.
Oprea TI, Nielsen SK, Ursu O, Yang JJ, Taboureau O, Mathias SL, Kouskoumvekaki I, Sklar LA, Bologa CG. Associating drugs, targets and clinical outcomes into an integrated network affords a new platform for computer-aided drug repurposing. Mol inform. 2011;30(2–3):100–11.
Xue H, Li J, Xie H, Wang Y. Review of drug repositioning approaches and resources. Int J Biol Sci. 2018;14(10):1232.
Mazzarella R, Webb C. Computational and bioinformatic strategies for drug repositioning. Drug Repositioning. 2012;9:91.
Deftereos SN, Andronis C, Friedla EJ, Persidis A, Persidis A. Drug repurposing and adverse event prediction using high-throughput literature analysis. Wiley Interdiscip Rev Syst Biol Med. 2011;3(3):323–34.
Rebholz-Schuhmann D, Oellrich A, Hoehndorf R. Text-mining solutions for biomedical research: enabling integrative biology. Nat Rev Genet. 2012;13(12):829–39.
Andronis C, Sharma A, Virvilis V, Deftereos S, Persidis A. Literature mining, ontologies and information visualization for drug repurposing. Brief Bioinform. 2011;12(4):357–68.
Loging W, Rodriguez-Esteban R, Hill J, Freeman T, Miglietta J. Cheminformatic/bioinformatic analysis of large corporate databases: application to drug repurposing. Drug Discov Today Ther Strateg. 2011;8(3–4):109–16.
Nabirotchkin S, Peluffo AE, Rinaudo P, Yu J, Hajj R, Cohen D. Next-generation drug repurposing using human genetics and network biology. Curr Opin Pharmacol. 2020;51:78–92.
Vilar S, Hripcsak G. The role of drug profiles as similarity metrics: applications to repurposing, adverse effects detection and drug–drug interactions. Brief Bioinform. 2017;18(4):670–81.
Grenier L, Hu P. Computational drug repurposing for inflammatory bowel disease using genetic information. Comput Struct Biotechnol J. 2019;17:127–35.
Hamid JS, Hu P, Roslin NM, Ling V, Greenwood CM, Beyene J. Data integration in genetics and genomics: methods and challenges. Hum Genomics Proteomics. 2009;2009:869093.
Hasin Y, Seldin M, Lusis A. Multi-omics approaches to disease. Genome Biol. 2017;18(1):1–5.
Michnick SW. The connectivity map. Nat Chem Biol. 2006;2(12):663–4.
Reka AK, Kuick R, Kurapati H, Standiford TJ, Omenn GS, Keshamouni VG. Identifying inhibitors of epithelial-mesenchymal transition by connectivity map–based systems approach. J Thorac Oncol. 2011;6(11):1784–92.
De Bastiani MA, Pfaffenseller B, Klamt F. Master regulators connectivity map: a transcription factors-centered approach to drug repositioning. Front Pharmacol. 2018;9:697.
Koehler AN. Microarrays in chemical biology. Wiley Encyclopedia Chem Biol. 2007;14:1.
Lamb J, Crawford ED, Peck D, Modell JW, Blat IC, Wrobel MJ, Lerner J, Brunet JP, Subramanian A, Ross KN, Reich M. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science. 2006;313(5795):1929–35.
Eckhardt M, Hultquist JF, Kaake RM, Hüttenhain R, Krogan NJ. A systems approach to infectious disease. Nat Rev Genet. 2020;21(6):339–54.
Ideker T, Galitski T, Hood L. A new approach to decoding life: systems biology. Annu Rev Genomics Hum Genet. 2001;2(1):343–72.
Cheng F, Desai RJ, Handy DE, Wang R, Schneeweiss S, Barabási AL, Loscalzo J. Network-based approach to prediction and population-based validation of in silico drug repurposing. Nat Commun. 2018;9(1):1–2.
Sadegh S, Matschinske J, Blumenthal DB, Galindez G, Kacprowski T, List M, Nasirigerdeh R, Oubounyt M, Pichlmair A, Rose TD, Salgado-Albarrán M. Exploring the SARS-CoV-2 virus-host-drug interactome for drug repurposing. Nat Commun. 2020;11(1):1–9.
Surabhi S, Singh BK. Computer aided drug design: an overview. J Drug Deliv Ther. 2018;8(5):504–9.
Abdolmaleki A, Ghasemi B, J, Ghasemi F. Computer aided drug design for multi-target drug design: SAR/QSAR, molecular docking and pharmacophore methods. Curr Drug Targets. 2017;18(5):556–75.
Cortegiani A, Ingoglia G, Ippolito M, Giarratano A, Einav S. A systematic review on the efficacy and safety of chloroquine for the treatment of COVID-19. J Crit Care. 2020;57:279–83.
Manivannan E, Karthikeyan C, Moorthy NS, Chaturvedi SC. The rise and fall of chloroquine/hydroxychloroquine as compassionate therapy of COVID-19. Front Pharmacol. 2021;12:1057.
Axfors C, Schmitt AM, Janiaud P, van’t Hooft J, Abd-Elsalam S, Abdo EF, Abella BS, Akram J, Amaravadi RK, Angus DC, Arabi YM. Mortality outcomes with hydroxychloroquine and chloroquine in COVID-19 from an international collaborative meta-analysis of randomized trials. Nat Commun. 2021;12(1):1–3.
Gasmi A, Peana M, Noor S, Lysiuk R, Menzel A, Benahmed AG, Bjørklund G. Chloroquine and hydroxychloroquine in the treatment of COVID-19: the never-ending story. Appl Microbiol Biotechnol. 2021;30:1–1.
Chen Y, Li MX, Lu GD, Shen HM, Zhou J. Hydroxychloroquine/chloroquine as therapeutics for COVID-19: truth under the mystery. Int J Biol Sci. 2021;17(6):1538.
Patel TK, Patel PB, Barvaliya M, Saurabh MK, Bhalla HL, Khosla PP. Efficacy and safety of lopinavir-ritonavir in COVID-19: a systematic review of randomized controlled trials. J Infect Public Health. 2021;14:740–8.
Young B, Tan TT, Leo YS. The place for remdesivir in COVID-19 treatment. Lancet Infect Dis. 2021;21(1):20.
Horby PW, Mafham M, Bell JL, Linsell L, Staplin N, Emberson J, Palfreeman A, Raw J, Elmahi E, Prudon B, Green C. Lopinavir–ritonavir in patients admitted to hospital with COVID-19 (RECOVERY): a randomised, controlled, open-label, platform trial. Lancet. 2020;396(10259):1345–52.
Dabbous HM, Abd-Elsalam S, El-Sayed MH, Sherief AF, Ebeid FF, Abd El Ghafar MS, Soliman S, Elbahnasawy M, Badawi R, Tageldin MA. Efficacy of favipiravir in COVID-19 treatment: a multi-center randomized study. Arch Virol. 2021;166(3):949–54.
Butler CC, van der Velden AW, Bongard E, Saville BR, Holmes J, Coenen S, Cook J, Francis NA, Lewis RJ, Godycki-Cwirko M, Llor C. Oseltamivir plus usual care versus usual care for influenza-like illness in primary care: an open-label, pragmatic, randomised controlled trial. Lancet. 2020;395(10217):42–52.
Hassanipour S, Arab-Zozani M, Amani B, Heidarzad F, Fathalipour M, Martinez-de-Hoyo R. The efficacy and safety of favipiravir in treatment of COVID-19: a systematic review and meta-analysis of clinical trials. Sci Rep. 2021;11(1):1.
Rosa SG, Santos WC. Clinical trials on drug repositioning for COVID-19 treatment. Rev Panam Salud Publica. 2020;8(44):e40.
Thomas E, Ghany MG, Liang TJ. The application and mechanism of action of ribavirin in therapy of hepatitis C. Antivir Chem Chemother. 2012;23(1):1–2.
Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y, Yu T. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395(10223):507–13.
Ali MJ, Hanif M, Haider MA, Ahmed MU, Sundas FN, Hirani A, Khan IA, Anis K, Karim AH. Treatment options for COVID-19: a review. Front Med. 2020;31(7):480.
Singh AK, Majumdar S, Singh R, Misra A. Role of corticosteroid in the management of COVID-19: a systemic review and a Clinician’s perspective. Diabetes Metab Syndr Clin Res Rev. 2020;14(5):971–8.
Salama C, Han J, Yau L, Reiss WG, Kramer B, Neidhart JD, Criner GJ, Kaplan-Lewis E, Baden R, Pandit L, Cameron ML. Tocilizumab in patients hospitalized with Covid-19 pneumonia. N Engl J Med. 2021;384(1):20–30.
Hurle MR, Yang L, Xie Q, Rajpal DK, Sanseau P, Agarwal P. Computational drug repositioning: from data to therapeutics. Clin Pharmacol Ther. 2013;93(4):335–41.
Jin G, Wong ST. Toward better drug repositioning: prioritizing and integrating existing methods into efficient pipelines. Drug Discov Today. 2014;19(5):637–44.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Kumar, S. (2022). In Silico Drug Repositioning for COVID-19: Progress and Challenges. In: Adibi, S., Griffin, P., Sanicas, M., Rashidi, M., Lanfranchi, F. (eds) Frontiers of COVID-19. Springer, Cham. https://doi.org/10.1007/978-3-031-08045-6_24
Download citation
DOI: https://doi.org/10.1007/978-3-031-08045-6_24
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-08044-9
Online ISBN: 978-3-031-08045-6
eBook Packages: MedicineMedicine (R0)