Abstract
Antibody production in response to infection or vaccination is a crucial process to combat and prevent infectious diseases. Regarding SARS-CoV-2, different features have been observed and literature still lacks consensus about it. Here, we discuss the production of antibodies during SARS-CoV-2 infection and its therapeutic relevance.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
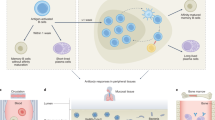
The humoral immune response is an arm of adaptive response, also known as antibody-mediated immune response. It is responsible to protect the extracellular fluids, such as blood and lymph, through the production of effector and memory B cells, which B cells generate antibodies, leading to neutralization, opsonization, complement activation, and modulation of inflammation. Also, it promotes an immunological memory, capable of protecting against future reinfection to the pathogen [1,2,3]. Upon a reexposure to the pathogen, or, more specifically, to an antigen, the humoral memory response has three typical characteristics: (1) it is more robust and faster than the primary antibody response; (2) it is dominated by high affinity, isotype-switched antibodies; and (3) it is long-lived and self-sustaining, allowing for a rapid complement cascade activation and antibody production [4, 5]. Given all that, antibody production following natural infection or vaccination is essential to combat and prevent infectious diseases.
In 2020, the World Health Organization (WHO) declared COVID-19 as a pandemic disease, which challenged the researchers to understand how the virus stimulates the immune system as soon as possible, in order to discover a way to treat the disease and stop the viral transmission. This chapter intends to discuss the humoral immune response against SARS-CoV-2 infection.
Pathogens and Antibodies Evolve Together
Antibodies are glycoprotein molecules composed of four chains: two light chains (L) and two heavy chains (H). The domains are linked by disulfide bonds. The intersection of the L and H chains forms the hypervariable region, or complementarity-determining region (CDR). The CDR regions comprise the paratope, which is responsible for the interaction with the antigen’s epitope. The analysis of the evolution of CDRs in response to an infection is important: the changes in CDR results from natural selection, aiming to increase the affinity for the target molecule, the epitope, from 1000 to 10,000 times [6,7,8].
On the other hand, some pathogens present high mutation rates. The exchange of amino acids allows for better adaptation to the environment and improves the performance of the pathogens in front of challenges, like the host’s immune response [9].
Given that, through the accumulation of genetic mutations, the host’s antibodies and pathogen’s antigens coevolve, acting as forces of selection for each other. The coevolution between environment, pathogens, and host was originally proposed by the “Red Queen Theory”: it proposes that a successful evolution from one species produces a negative effect on the other species, and vice versa [10, 11].
Antibody Kinetics After SARS-CoV-2 Infection
SARS-CoV-2 presents two main proteins that are highly immunogenic and able to trigger humoral response: the Spike (S) and the Nucleocapsid (N) proteins. S is divided into S1 and S2 subunits: the first mediates the binding with angiotensin-converting enzyme (ACE)-2 by the receptor binding domain (RBD) and the second mediates the fusion of the virus and the cellular membrane. N is the most abundant viral protein, it binds with the RNA and mediates virion assembly. Membrane (M) and Envelope (E), the other two structural proteins, induce a poor humoral response, probably because of their small molecular size; however, such proteins are studied in cellular response [12, 13].
The immune response to pathogens usually presents initial IgM seroconversion, a result of T-independent humoral response, which is a mark of acute disease that decreases within a few weeks. The following IgG seroconversion, after T cell activation and class switch, is a mark of maturation of immune response and immunologic memory. IgG is the main class of antibody found systemically, and it is often desired for an adequate immune response [2].
Antibody response to acute viral infection is found in patients with COVID-19. As expected, the first antibody detected is IgM, followed by IgG, once the seroconversion rate and antibody levels increase fast during the first 2 weeks following infection. The cumulative seropositive rate reaches 50% on the 11th day and 100% on the 39th day [14]. The IgG titers increase until 2 months after diagnostic, then it reaches a plateau [15]. One study demonstrated that, after 6 months, the positive rate for IgG was maintained, ranging from 92.3% to 95.5%, while the positive IgM rate decreased from 90.4% to 22.7% [16]. Another study demonstrated a durable B cell response, until 8 months after infection [17].
It was demonstrated that 6 months after the infection, the patients continue having an anti-SARS-CoV-2 B cells response, being observed an accumulation of somatic mutations in these cells, and production of antibodies with increased neutralizing breadth and potency [18].
However, subsequent studies described a concomitant IgM/IgG seroconversion. It has also been suggested the value of IgA seroconversion as the first mark of the humoral response against SARS-CoV-2. Thus, the combined serology of IgA/IgG presents higher sensitivity and specificity than IgM/IgG to detect past exposure to the virus [14, 19, 20].
In summary, there are a great number of studies about antibody kinetics after SARS-CoV-2 infection. As expected, not all the results agree with each other, but, in general, it has been proposed that IgA seroconversion happens within 4–6 days post symptoms onset, peaking around 16–20 days and declining after 31–41 days. For IgM, seroconversion starts 4 to 6 days after symptoms onset, the peak happens on days 11 to 15, and then it decreases [20].
The Role of Antibodies in COVID-19
The description of the immune response in COVID-19 has been an issue and several studies have focused on it. Despite the differences between the investigations conducted, it can be stated that the sole presence of antibodies cannot be used to infer protection against SARS-CoV-2. Studies support that the ideal response is probably a synergetic one, that comprises the innate, humoral, and cellular mechanisms [21].
Antibody Titers
Studies point that severe infection patients present higher antibody titers than mild infection patients—which could lead one to suggest that antibodies would not bring benefits to the patients. A work demonstrated that this high antibody secretion in severe infection patients could mediate pathogenesis by multiple mechanisms, including tissue damage by activation of inflammatory macrophages [22]. The possible explanation for this is the lack of viral replication control, which induces a persistent viremia and causes an intense or prolonged B cell activation, resulting in a pathogenic B cell production [23].
These high loads of IgG in the alveoli form immune complexes with viral particles, capable of activating the complement system and inducing inflammation in the lungs, a serious issue in COVID-19 [24]. The worry about IgG response was also related to antibody-dependent enhancement (ADE). It happens when antibodies produced by a previous, poor immune response, which are not capable of neutralizing activity or present lower affinity by the pathogen, intensify the current infection, allowing for internalization mediated by the Fcγ receptor, thus favoring the release of pro-inflammatory cytokines and immunopathology. Several studies about SARS-CoV-2 do not corroborate this hypothesis, but it was important to state how the quality of antibodies, rather than the quantity, should be assessed [25].
The expressive presence of anti-N antibodies in severe patients also points to higher viremia, since large amounts of N protein are incorporated into the virion. It is also supported by children showing high anti-S but low anti-N titers: there is a decrease of ACE-2 expression in this age group, which has been related to their reduced risk of suffering from COVID-19; thus, their viremia is expected to be lower when compared with that of adults. Of note, the induction of antibodies through vaccination, training the immune system before the exposure, should not be directly compared with natural infection response [12, 26, 27].
It is still controversial how antibodies and the severity of the disease may affect each other. A study demonstrated that antibody levels were significantly higher in severe than in nonsevere patients, between the second and fifth week after disease onset; but there is no observation for IgG or IgM alone [14]. Another study shows that 3 weeks after the disease, the levels of IgM and IgG to S and N proteins were higher in non-severe and RNA-negative patients than in severe and RNA-positive patients [28]. The same controversial results were observed when the antibody titer is correlated with age and symptoms. In some studies, the age was positively correlated with IgG, IgM, and IgA titers; and especially IgG was correlated with specific COVID-19 symptoms, like fever, sore throat, shortness of breath, and nausea [28, 29]. On the other hand, a study shows that antibody response was independent of patient age, sex, and most preexistent comorbidities [30]. It was demonstrated that male sex, older age, and hospitalization for COVID-19 were associated with increasing antibody response [31].
Antibodies Functionality
Generally, antibody avidity increases during the infection and remains elevated. The same was observed to SARS-CoV-2: low antibody avidity was reported during early infection, until 3 weeks after symptom onset [32]. However, other studies report that the avidity of naturally induced antibodies did not improve with time [33]. It was also observed that the avidity is higher in hospitalized than in nonhospitalized patients. As an indicator of functionality, anti-spike avidity was correlated with higher neutralizing antibodies (nAbs) titers [34, 35].
It is described that nAbs are needed for virus clearance and it has been considered a key for the protection or treatment of COVID-19. Diverse studies found that nAb levels in asymptomatic or mild cases were lower than moderate or severe cases [14, 26, 36]. Such results have led previous reports to question the efficacy of nAb-mediated protection in COVID-19 severe cases and have suggested that the enhancement of nAbs is associated with a worst clinical condition [17, 37]. Similarly to antibody titers, which are usually higher in severe cases, the neutralizing activity of the plasma of most symptomatic COVID-19 patients persists up to 6 months [28], whereas in asymptomatic patients, it gradually disappears in 2 months [38]. The interplay between viral load and antibody titers discussed above could also affect the nAb titers.
The study of IgA against SARS-CoV-2 has been encouraged, given the involvement of mucosa in COVID-19 [24]. The seric IgA could reflect the mucosa implication of COVID-19. It was described as the main antibody responsible for early neutralization of SARS-CoV-2, even in less quantity than IgG; thus, it would be capable of penetrating epithelial cells, neutralizing intracellular virus [20, 39]. The secretory-IgA (sIgA), locally produced in the mucosa, was considered as a potential biomarker of SARS-CoV-2 early infection, which could be tested in saliva. With better elucidation of duration and functionality of the immune response, sIgA could also be a correlate of protection, given its ability to control the infection when the virus first enters the host [40]. Moreover, patients with nAbs and anti-spike IgA demonstrated a faster viral control [41].
When the production of antibodies against the structural proteins was analyzed, it was verified that SARS-CoV-2 specific IgM recognition of S and N proteins was transient and disappeared around the 12th week; thus, the IgM response would not contribute to sustained immunity against the virus. Also, there was no correlation between IgM response and the ability of plasma to neutralize the virus in cell culture. Differently, IgG antibodies that recognize the S and N proteins maintain high positive rates for up to 6 months, and particularly RBD-specific IgG were correlated with neutralizing activity, being associated with early virus control [3, 28]. Thus, titers of IgG was not correlated with severe acute respiratory distress syndrome [30].
It was postulated that anti-S or, more specifically, anti-RBD IgG, would be ideal for protection, since it could neutralize the virus by impairing the RBD-ACE-2 binding. Serology studies also suggest that anti-S protein antibodies are maintained over longer periods when compared with anti-N antibodies. Indeed, the S protein or its subunits has been used as vaccine antigens [12, 20, 42, 43].
Because of the lack of drugs capable of inhibiting SARS-CoV-2, convalescent plasma (plasma obtained from recovered patients that present high levels of nAbs) was indicated as a therapeutic option for COVID-19 severe infection patients. The studies that followed this type of intervention varied a lot regarding the number of patients and how the plasma was obtained, which limits the comparisons, but good results were described overall [44].
Serology presents a limited role in diagnosing SARS-CoV-2 infection and assessing the protective status of a person. However, determining the humoral response is an interesting tool for public health, to verify the prevalence of COVID-19 [43]. Despite that, the study of neutralizing activity is useful for immune-based therapy trials [45].
Antibodies and Reinfection
Until 2020, sporadic cases of reinfection by SARS-CoV-2 were described around the world. In some cases, it was more severe, but in others, an increasing severity was observed. However, with the emergence of variants and as time passed, more cases were documented [46,47,48]. It was suggested that some people would fail to develop a protective immunity, which would explain the reinfections [17].
However, with the pandemic ongoing, four hypotheses have been developed to explain the cases of more severe reinfection: (1) a very high dose of virus might have led to this second infection and induced more severe disease; (2) the reinfection was caused by another, more virulent, strain; (3) the mechanism of antibody-dependent enhancement might be the cause; and (4) the incomplete avidity maturation after COVID-19 infection did not confer a protective immunity [33, 48,49,50].
The current knowledge leads to suggest that the most accurate hypothesis is that natural infection is prone to failure in developing an efficient immunologic memory, because of impaired affinity maturation, and high-coverage vaccination is needed to control the pandemic, since it provides a more adequate immune response [33].
Antibody Production Modulated by Cytokines
As described before, COVID-19 hospitalized patients usually present a stronger IgG avidity and higher nAbs titers than nonhospitalized patients [51]. A study showed that nAb longevity was associated with sustained levels of inflammatory cytokines, up to 180 days after symptoms onset in COVID-19 [52]; furthermore, pro-inflammatory cytokine milieu was correlated with antibody levels against the virus [53].
Some cytokines play an important role in B cell development, as interleukin (IL)-7, which aids in survival and proliferation; IL-4 and IL-6, which influence isotype switching; IL-10, which is important for regulation of the immune response; and Interferon-gamma (INF-γ), IL-12, and IL-17, which participate in B cell development [54,55,56,57].
Some studies show that patients with severe COVID-19 exhibit higher levels of IL-2, IL-6, IL-7, IL-10, INF-γ, tumor necrosis factor-alpha (TNF-α), inducible protein (IP)-10, monocyte chemoattractant protein (MCP)-1, macrophage inflammatory protein 1α, and granulocyte-colony stimulating factor than patients with mild and moderate infections. Such cytokine environment stimulates the antibody production and functionality [19, 58, 59].
Even though the cytokine storm induced by SARS-CoV-2 infection contributes to the humoral response, it can also be correlated with increased severity of the disease and favoring uncontrolled inflammation [60]. Considering that, cytokine production becomes a double-edged sword in the case of COVID-19.
Maternal Antibodies
Maternal antibodies are transferred from mother to child to protect them during their immune system maturation in the first year of life. The majority of maternal antibodies are of IgG isotype, which are preferentially transferred before the birth in utero across placenta; these passively-transferred antibodies enter the bloodstream of offspring and act as a protective shield in the same way as active antibodies [61]. Different from IgG isotype, secretory IgA is transferred to breast milk from mother and protects the gastrointestinal tract against pathogens [62, 63].
It is well described that vaccination of pregnant women can increase neonatal antibodies against influenza, tetanus, diphtheria, and pertussis [64, 65]. Moreover, the WHO reports a 96% reduction of death by neonatal tetanus through the recommendation of certain good practices from 1988 to 2015, including the vaccination of pregnant women [66].
In a study that analyzed the seroconversion of newborns from pregnant women infected with SARS-CoV-2, it was demonstrated that SARS-CoV-2 IgG positive rate among parturients was 80.8%, and half of their infants obtained maternal IgG.
If the mothers were infected earlier and later than 2 weeks before delivery, the IgG rates were, respectively, 18.8% and 81.8% in their infants; after that, they presented a reduction of IgG in the first 2 months of life [67]. In this way, the study demonstrated that the passage of naturally induced maternal antibodies against SARS-CoV-2 is low. On the other side, when prenatal BNT16b2 mRNA vaccination was analyzed, it was observed a robust maternal humoral response, which was effectively transferred to the fetus [68], showing the importance of vaccination against SARS-CoV-2 during pregnancy.
Conclusion
In this chapter, we have reviewed the humoral response after COVID-19. Our knowledge regarding SARS-CoV-2 has increased dramatically with the pandemic and we still have a long way to go to completely understand the virus and the response it triggers in the human immune system.
It should be noted that the substitution of amino acids in the variable portions of the immunoglobulins brings the advantage of high repertoire variability and greater chances of expression of a highly effective antibody, but this mechanism suffers an important restriction caused by the stability of the resulting protein. The stability of protein folding is constant when analyzing the evolution of proteins [69]; however, the mutations that generate the most specific paratope do not necessarily result in the most stable CDR region—it is a dynamic process. Affinity maturation is one of the easily observable examples of Darwinian evolution: genetic mutations are continuously happening in the coding regions of CDRs and selected immediately, since B cells which mutations are neutral or beneficial rapidly expand. Emerging pathogens present an excellent opportunity to learn about the coevolution of pathogens and the immune system: evolution is happening right in front of our eyes.
References
Dunkelberger JR, Song WC. Complement and its role in innate and adaptive immune responses. Cell Res. 2010;20:34–50. https://doi.org/10.1038/cr.2009.139.
Forthal DN. Functions of antibodies. In: Antibodies for infectious diseases. New York: American Society of Microbiology, Wiley; 2014. p. 25–48.
Assadiasl S, Fatahi Y, Zavvar M, Nicknam MH. COVID-19: significance of antibodies. Hum Antibodies. 2020;28:287–97. https://doi.org/10.3233/HAB-200429.
Inoue T, Moran I, Shinnakasu R, et al. Generation of memory B cells and their reactivation. Immunol Rev. 2018;283:138–49. https://doi.org/10.1111/imr.12640.
Shishido SN, Varahan S, Yuan K, et al. Humoral innate immune response and disease. Clin Immunol. 2012;144:142–58. https://doi.org/10.1016/j.clim.2012.06.002.
Eisen HN, Siskind GW. Variations in affinities of antibodies during the immune response. Biochemistry. 1964;3:996–1008. https://doi.org/10.1021/bi00895a027.
Berek C, Milstein C. Mutation drift and repertoire shift in the maturation of the immune response. Immunol Rev. 1987;96:23–41. https://doi.org/10.1111/j.1600-065X.1987.tb00507.x.
Wang S, Mata-Fink J, Kriegsman B, et al. Manipulating the selection forces during affinity maturation to generate cross-reactive HIV antibodies. Cell. 2015;160:785–97. https://doi.org/10.1016/j.cell.2015.01.027.
Van Valen L. Molecular evolution as predicted by natural selection. J Mol Evol. 1974;3:89–101. https://doi.org/10.1007/BF01796554.
Liow LH, Van Valen L, Stenseth NC. Red queen: from populations to taxa and communities. Trends Ecol Evol. 2011;26:349–58. https://doi.org/10.1016/j.tree.2011.03.016.
Brockhurst MA, Chapman T, King KC, et al. Running with the red queen: the role of biotic conflicts in evolution. Proc R Soc B Biol Sci. 2014;281:20141382. https://doi.org/10.1098/rspb.2014.1382.
Dai L, Gao GF. Viral targets for vaccines against COVID-19. Nat Rev Immunol. 2021;21:73–82. https://doi.org/10.1038/s41577-020-00480-0.
Paces J, Strizova Z, Smrz D, Cerny J. COVID-19 and the immune system. Physiol Res. 2020;69:379–88. https://doi.org/10.33549/PHYSIOLRES.934492.
Zhao J, Yuan Q, Wang H, et al. Antibody responses to SARS-CoV-2 in patients with novel coronavirus disease 2019. Clin Infect Dis. 2020;71:2027–34. https://doi.org/10.1093/cid/ciaa344.
Gudbjartsson DF, Norddahl GL, Melsted P, et al. Humoral immune response to SARS-CoV-2 in Iceland. N Engl J Med. 2020;383:1724–34. https://doi.org/10.1056/nejmoa2026116.
Liu C, Yu X, Gao C, et al. Characterization of antibody responses to SARS-CoV-2 in convalescent COVID-19 patients. J Med Virol. 2021;93:2227–33. https://doi.org/10.1002/jmv.26646.
Dan JM, Mateus J, Kato Y, et al. Immunological memory to SARS-CoV-2 assessed for up to 8 months after infection. Science. 2021;371:eabf4063. https://doi.org/10.1126/science.abf4063.
Gaebler C, Wang Z, Lorenzi JCC, et al. Evolution of antibody immunity to SARS-CoV-2. Nature. 2021;591:639–44. https://doi.org/10.1038/s41586-021-03207-w.
Huang C, Wang Y, Li X, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395:497–506. https://doi.org/10.1016/S0140-6736(20)30183-5.
Ma H, Zeng W, He H, et al. Serum IgA, IgM, and IgG responses in COVID-19. Cell Mol Immunol. 2020;17:773–5. https://doi.org/10.1038/s41423-020-0474-z.
Vardhana SA, Wolchok JD. The many faces of the anti-COVID immune response. J Exp Med. 2020;217:1–10. https://doi.org/10.1084/JEM.20200678.
Hoepel W, Chen H-J, Geyer CE, et al. High titers and low fucosylation of early human anti-SARS-CoV-2 IgG promote inflammation by alveolar macrophages. Sci Transl Med. 2021;13(596):eabf8654. https://doi.org/10.1126/scitranslmed.abf8654.
Woodruff MC, Ramonell RP, Nguyen DC, et al. Extrafollicular B cell responses correlate with neutralizing antibodies and morbidity in COVID-19. Nat Immunol. 2020;21:1506–16. https://doi.org/10.1038/s41590-020-00814-z.
Béné MC, de Carvalho BM, Eveillard M, Le Bris Y. Good IgA bad IgG in SARS-CoV-2 infection? Clin Infect Dis. 2020;71:897–8. https://doi.org/10.1093/cid/ciaa426.
Halstead SB, Katzelnick L. COVID-19 vaccines: should we fear ADE? J Infect Dis. 2020;222:1946–50. https://doi.org/10.1093/infdis/jiaa518.
Atyeo C, Fischinger S, Zohar T, et al. Distinct early serological signatures track with SARS-CoV-2 survival. Immunity. 2020;53:524–532.e4. https://doi.org/10.1016/j.immuni.2020.07.020.
Gallo O, Locatello LG, Mazzoni A, et al. The central role of the nasal microenvironment in the transmission, modulation, and clinical progression of SARS-CoV-2 infection. Mucosal Immunol. 2021;14:305–16. https://doi.org/10.1038/s41385-020-00359-2.
Wu J, Liang B, Chen C, et al. SARS-CoV-2 infection induces sustained humoral immune responses in convalescent patients following symptomatic COVID-19. Nat Commun. 2021;12:1–9. https://doi.org/10.1038/s41467-021-22034-1.
Weisberg SP, Connors TJ, Zhu Y, et al. Distinct antibody responses to SARS-CoV-2 in children and adults across the COVID-19 clinical spectrum. Nat Immunol. 2021;22:25–31. https://doi.org/10.1038/s41590-020-00826-9.
Cervia C, Nilsson J, Zurbuchen Y, et al. Systemic and mucosal antibody responses specific to SARS-CoV-2 during mild versus severe COVID-19. J Allergy Clin Immunol. 2021;147:545–557.e9. https://doi.org/10.1016/j.jaci.2020.10.040.
Klein SL, Pekosz A, Park HS, et al. Sex, age, and hospitalization drive antibody responses in a COVID-19 convalescent plasma donor population. J Clin Invest. 2020;130:6141–50. https://doi.org/10.1172/JCI142004.
Valdivia A, Torres I, Huntley D, et al. Qualitative assessment of SARS-CoV-2-specific antibody avidity by lateral flow immunochromatographic IgG/IgM antibody assay. J Med Virol. 2021;93:1141–4. https://doi.org/10.1002/jmv.26344.
Bauer G. The potential significance of high avidity immunoglobulin G (IgG) for protective immunity towards SARS-CoV-2. Int J Infect Dis. 2021;106:61–4. https://doi.org/10.1016/j.ijid.2021.01.061.
Benner SE, Patel EU, Laeyendecker O, et al. SARS-CoV-2 antibody avidity responses in COVID-19 patients and convalescent plasma donors. J Infect Dis. 2020;222:1974–84. https://doi.org/10.1093/infdis/jiaa581.
Gaspar EB, De Gaspari E. Avidity assay to test functionality of anti-SARS-Cov-2 antibodies. Vaccine. 2021;39:1473–5. https://doi.org/10.1016/j.vaccine.2021.02.003.
Zhang B, Zhou X, Zhu C, et al. Immune phenotyping based on the neutrophil-to-lymphocyte ratio and IgG level predicts disease severity and outcome for patients with COVID-19. Front Mol Biosci. 2020;7:157. https://doi.org/10.3389/fmolb.2020.00157.
Wang K, Long Q-X, Deng H-J, et al. Longitudinal dynamics of the neutralizing antibody response to severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection. Clin Infect Dis. 2020;73:e531–9. https://doi.org/10.1093/cid/ciaa1143.
Lei Q, Li Y, Yan HH, et al. Antibody dynamics to SARS-CoV-2 in asymptomatic COVID-19 infections. Allergy. 2021;76:551–61. https://doi.org/10.1111/all.14622.
Sterlin D, Mathian A, Miyara M, et al. IgA dominates the early neutralizing antibody response to SARS-CoV-2. Sci Transl Med. 2021;13(577):eabd2223. https://doi.org/10.1126/scitranslmed.abd2223.
Chao YX, Rötzschke O, Tan EK. The role of IgA in COVID-19. Brain Behav Immun. 2020;87:182–3. https://doi.org/10.1016/j.bbi.2020.05.057.
Dispinseri S, Secchi M, Pirillo MF, et al. Neutralizing antibody responses to SARS-CoV-2 in symptomatic COVID-19 is persistent and critical for survival. Nat Commun. 2021;12:2670. https://doi.org/10.1038/s41467-021-22958-8.
Karthik K, Senthilkumar TMA, Udhayavel S, Raj GD. Role of antibody-dependent enhancement (ADE) in the virulence of SARS-CoV-2 and its mitigation strategies for the development of vaccines and immunotherapies to counter COVID-19. Hum Vaccin Immunother. 2020;16:3055–60. https://doi.org/10.1080/21645515.2020.1796425.
West R, Kobokovich A, Connell N, Gronvall GK. COVID-19 antibody tests: a valuable public health tool with limited relevance to individuals. Trends Microbiol. 2021;29:214–23. https://doi.org/10.1016/j.tim.2020.11.002.
Lu L, Zhang H, Zhan M, et al. Antibody response and therapy in COVID-19 patients: what can be learned for vaccine development? Sci China Life Sci. 2020;63:1833–49. https://doi.org/10.1007/s11427-020-1859-y.
Sewell HF, Agius RM, Kendrick D, Stewart M. Vaccines, convalescent plasma, and monoclonal antibodies for COVID-19. BMJ. 2020;370:m2722. https://doi.org/10.1136/bmj.m2722.
To KKW, Hung IF-N, Ip JD, et al. Coronavirus disease 2019 (COVID-19) re-infection by a phylogenetically distinct severe acute respiratory syndrome coronavirus 2 strain confirmed by whole genome sequencing. Clin Infect Dis. 2020;73(9):e2946–51. https://doi.org/10.1093/cid/ciaa1275.
Van Elslande J, Vermeersch P, Vandervoort K, et al. Symptomatic severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) reinfection by a phylogenetically distinct strain. Clin Infect Dis. 2020;73:354–6. https://doi.org/10.1093/cid/ciaa1330.
Tillett RL, Sevinsky JR, Hartley PD, et al. Genomic evidence for reinfection with SARS-CoV-2: a case study. Lancet Infect Dis. 2021;21:52–8. https://doi.org/10.1016/S1473-3099(20)30764-7.
Guallar MP, Meiriño R, Donat-Vargas C, et al. Inoculum at the time of SARS-CoV-2 exposure and risk of disease severity. Int J Infect Dis. 2020;97:290–2. https://doi.org/10.1016/j.ijid.2020.06.035.
Yip MS, Leung NHL, Cheung CY, et al. Antibody-dependent infection of human macrophages by severe acute respiratory syndrome coronavirus. Virol J. 2014;11:1–11. https://doi.org/10.1186/1743-422X-11-82.
Moura A, Costa HH, Correa V, et al. S assessment of avidity related to IgG subclasses in SARS-CoV-2 Brazilian infected patients. Sci Rep. 2021;11:17642. https://doi.org/10.1038/s41598-021-95045-z.
Chia WN, Zhu F, Ong SWX, et al. Dynamics of SARS-CoV-2 neutralising antibody responses and duration of immunity: a longitudinal study. Lancet Microbe. 2021;6:e240–9. https://doi.org/10.1016/S2666-5247(21)00025-2.
Wu F, Liu M, Wang A, et al. Evaluating the Association of Clinical Characteristics with neutralizing antibody levels in patients who have recovered from mild COVID-19 in Shanghai, China. JAMA Intern Med. 2020;180:1356–62. https://doi.org/10.1001/jamainternmed.2020.4616.
Abed NS, Chace JH, Fleming AL, Cowdery JS. Interferon-γ regulation of B lymphocyte differentiation: activation of B cells is a prerequisite for IFN-γ-mediated inhibition of B cell differentiation. Cell Immunol. 1994;153:356–66. https://doi.org/10.1006/cimm.1994.1034.
Metzger DW. Interleukin-12 as an adjuvant for induction of protective antibody responses. Cytokine. 2010;52:102–7. https://doi.org/10.1016/j.cyto.2010.06.011.
Vazquez MI, Catalan-Dibene J, Zlotnik A. B cells responses and cytokine production are regulated by their immune microenvironment. Cytokine. 2015;74:318–26. https://doi.org/10.1016/j.cyto.2015.02.007.
Shibui A, Shimura E, Nambu A, et al. Th17 cell-derived IL-17 is dispensable for B cell antibody production. Cytokine. 2012;59:108–14. https://doi.org/10.1016/j.cyto.2012.03.018.
Chen G, Wu D, Guo W, et al. Clinical and immunological features of severe and moderate coronavirus disease 2019. J Clin Invest. 2020;130:2620–9. https://doi.org/10.1172/JCI137244.
Liu J, Li S, Liu J, et al. Longitudinal characteristics of lymphocyte responses and cytokine profiles in the peripheral blood of SARS-CoV-2 infected patients. EBioMedicine. 2020;55:102763. https://doi.org/10.1016/j.ebiom.2020.102763.
Hojyo S, Uchida M, Tanaka K, et al. How COVID-19 induces cytokine storm with high mortality. Inflamm Regen. 2020;40:37. https://doi.org/10.1186/s41232-020-00146-3.
Fouda GG, Martinez DR, Swamy GK, Permar SR. The impact of IgG transplacental transfer on early life immunity. Immuno Horizons. 2018;2:14–25. https://doi.org/10.4049/immunohorizons.1700057.
Langel SN, Otero CE, Martinez DR, Permar SR. Maternal gatekeepers: how maternal antibody fc characteristics influence passive transfer and infant protection. PLoS Pathog. 2020;16:e1008303. https://doi.org/10.1371/journal.ppat.1008303.
Niewiesk S. Maternal antibodies: clinical significance, mechanism of interference with immune responses, and possible vaccination strategies. Front Immunol. 2014;5:446. https://doi.org/10.3389/fimmu.2014.00446.
Albrecht M, Pagenkemper M, Wiessner C, et al. Infant immunity against viral infections is advanced by the placenta-dependent vertical transfer of maternal antibodies. Vaccine. 2021;40(11):1563–71. https://doi.org/10.1016/j.vaccine.2020.12.049.
Albrecht M, Arck PC. Vertically transferred immunity in neonates: mothers, mechanisms and mediators. Front Immunol. 2020;11:555. https://doi.org/10.3389/fimmu.2020.00555.
World Health Organization. Protecting all against tetanus guide to sustaining maternal and neonatal tetanus elimination (MNTE) and broadening tetanus protection for all populations. 2019. https://apps.who.int/iris/bitstream/handle/10665/329882/9789241515610-eng.pdf?sequence=1&isAllowed=y
Wang X, Yang P, Zheng J, et al. Dynamic changes of acquired maternal SARS-CoV-2 IgG in infants. Sci Rep. 2021;11:8021. https://doi.org/10.1038/s41598-021-87535-x.
Beharier O, Plitman Mayo R, Raz T, et al. Efficient maternal to neonatal transfer of SARS-CoV-2 and BNT162b2 antibodies. J Clin Invest. 2021;131(13):e15031. https://doi.org/10.1172/JCI150319.
Lässig M, Mustonen V, Walczak AM. Predicting evolution. Nat Ecol Evol. 2017;1:77. https://doi.org/10.1038/s41559-017-0077.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Correa, V.A., Portilho, A.I., Gaspar, E.B., De Gaspari, E. (2022). Humoral Immune Response in SARS-CoV-2 Infection and Its Therapeutic Relevance. In: Adibi, S., Griffin, P., Sanicas, M., Rashidi, M., Lanfranchi, F. (eds) Frontiers of COVID-19. Springer, Cham. https://doi.org/10.1007/978-3-031-08045-6_2
Download citation
DOI: https://doi.org/10.1007/978-3-031-08045-6_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-08044-9
Online ISBN: 978-3-031-08045-6
eBook Packages: MedicineMedicine (R0)