Abstract
The diatoms evolved within the stramenopiles, an ecologically important and diverse assemblage of eukaryotes that includes both photosynthetic macrophytes and microalgae, as well as non-photosynthetic heterotrophs and parasites. The evolutionary history of the stramenopiles, which stretches back to the Palaeozoic, has been marked by the acquisition of chloroplasts in a recent common ancestor of their photosynthetic members, the ochrophytes; and progressive gains of genes in the nuclear genome by horizontal and endosymbiotic gene transfer. Here, we place diatoms in their actual evolutionary context within the stramenopiles; identify gene transfers that have shaped the coding content of the diatom nucleus; and profile sources of differences in chloroplast and mitochondrial genome content between different stramenopiles including diatoms. We underline the importance of considering diatoms as evolutionary mosaics, supported by genes of bacterial, red, green and other eukaryotic algal origins, as illustrated by multiple phylogenomic studies realised over the last two decades; and the relatively limited changes to organelle genome content in diatoms compared to other stramenopile lineages. We further identify a previously undocumented transfer of a novel open reading frame of the chloroplasts of green algae into the ochrophytes, underlining the importance of changes in organelle and nuclear gene content, in defining the current biology of diatoms.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 The Evolutionary Context of Diatoms within the Stramenopiles
1.1 Diversity of Photosynthetic and Non-photosynthetic Stramenopile Groups
Diatoms belong to the stramenopile phylum which sits within the “SAR clade” (Stramenopiles, Alveolates and Rhizaria) of the tree of life (Dorrell and Bowler 2017; Ichinomiya et al. 2016; Adl et al. 2012) (Fig. 1). Besides stramenopiles, the SAR clade contains the alveolates (including dinoflagellates which are the principal photosynthetic component of corals, and apicomplexans such as the malaria parasite Plasmodium), and rhizarians (including the important fossil markers, forams). The group is distantly related both to plants and to animals, sharing a last common ancestor with each no more recently than one billion years ago (Parfrey et al. 2011). The stramenopiles themselves are evolutionarily ancient, radiating at least six hundred million years before the present (Brown and Sorhannus 2010), placing them on a common level of antiquity to plants and animals (Dorrell and Smith 2011).
Evolutionary positions of diatoms within the eukaryotes and the stramenopiles. Top: a schematic tree of eukaryotic diversity, adapted from Dorrell and Smith (2011); bottom, a close-up of stramenopile lineages, adapted from Dorrell and Smith (2017), showing the global and local evolutionary context of diatoms. Cell images are reprinted from Encyclopaedia of Life (www.eol.org), Prof. Connie Lovejoy (Université Laval), and Dr. Zhanru Shao (Institute of Oceanology, Chinese Academy of Sciences), with permissions
The stramenopiles are an ecologically diverse, and environmentally important group of eukaryotes. They are characterised, in particular, by their flagellar organisation which typically includes one short flagellum and one long flagellum covered in tubular « hair »-like structures, with the uneven length of their flagella giving rise to an alternate name of « heterokonts » (Adl et al. 2012; Cavalier-Smith 1998). Beyond this, stramenopiles contain few conserved features, although their monophyly is well supported by molecular data (Burki et al. 2007; Elias et al. 2009).
Alongside the diatoms, the stramenopiles contain numerous other photosynthetic taxa, referred to collectively as “ochrophytes” (Fig. 1). These include, but are not limited to, giant macrophytic kelps (found within the phaeophytes) which can grow to nearly one hundred meters in length (Fork et al. 1991), and unicellular and colonial algae (e.g. the chrysophytes, or “golden algae,” dictyochophytes, or “silicoflagellates” and pelagophytes) which can have cell diameters of only a couple of micrometers. Several other stramenopile groups (particularly phaeophytes, pelagophytes, dictyochophytes and chrysophytes) are important contributors to marine and freshwater photosynthesis (Dorrell and Smith 2011) Some of the ochrophyte groups are silicifying, like diatoms (e.g. silicoflagellates within the dictyochophytes; synurophytes within the chrysophytes), while others are not (Dorrell and Bowler 2017; Ichinomiya et al. 2016; Hendry et al. 2018). Some of these photosynthetic groups are also known to engage in mixotrophic strategies, either through the phagocytotic consumption of bacteria (e.g. in chrysophytes and dictyochophytes (Graupner et al. 2018; Walker et al. 2011)), or through the osmotrophic uptake of extracellular organic nutrients (e.g. in diatoms and pelagophytes) (Villanova et al. 2017), while in others (e.g. kelps), neither strategy has been reported (Dorrell and Bowler 2017).
Moreover, there is a substantial non-photosynthetic component to stramenopile diversity (Fig. 1). These include some lineages that resolve within otherwise photosynthetic groups, for example, non-photosynthetic diatoms (Kamikawa et al. 2015a; Pendergrass et al. 2020), dictyochophytes (Sekiguchi et al. 2002; Kayama et al. 2020), and chrysophytes (Graupner et al. 2018; Dorrell et al. 2019), and accordingly retain leucoplasts (non-photosynthetic plastids). Other non-photosynthetic stramenopile groups contain no photosynthetic members and no trace of plastids. These “aplastidic stramenopiles” include important pathogens: oomycetes, parasites of algae and plants, such as the causative agents of potato blight and sudden oak death (Stiller et al. 2009; Levesque et al. 2010); labyrinthulomycetes which are important pathogens of marine algae and invertebrates (Tsui et al. 2009); and Blastocystis, a human gut commensal which may have harmful effects in immuno-deficient hosts (Eme et al. 2017). In addition, the aplastidic stramenopiles include important marine saproptrophs (e.g. hypochytriomycetes, related to oomycetes (Leonard et al. 2018)) and predator groups (e.g. bicosoecids, related to Blastocystis (Jirsová et al. 2019)).
1.2 Phylogenetic Arrangement and Endosymbiotic Histories of the Stramenopiles
The phylogenetic relationships within the stramenopiles have been revealed progressively by morphology, electron microscopy and biochemistry (Adl et al. 2012; Cavalier-Smith 1998), alongside single-gene, and subsequently multigene phylogenies (Parks et al. 2018; Ichinomiya et al. 2016; Derelle et al. 2016) (Fig. 1). The exclusively non-photosynthetic groups form the most basally divergent stramenopile nodes, with oomycetes forming the closest major sister-group to the photosynthetic ochrophytes (Ichinomiya et al. 2016; Derelle et al. 2016) (Fig. 1). Within the ochrophytes, three major groups are apparent (Fig. 1): the “chrysista” which include phaeophytes; chrysophytes, and the oil-producing alga Nannochloropsis (Ichinomiya et al. 2016; Rodolfi et al. 2009); the « hypogyristea», containing pelagophytes and dictyochophytes (Ichinomiya et al. 2016; Dorrell et al. 2017a); and the “khakista” or diatoms and their immediate sister group, a single-genus group alternatively labelled Bolidomonas or Triparma (Ichinomiya et al. 2016; Tajima et al. 2016). The hypogyristea and khakista are clearly sister-groups, whereas the monophyly of chrysista is uncertain, with some phylogenomic studies resolving the constituent lineages as a single group (Derelle et al. 2016), and others indicating a basally divergent group of pinguiophytes, synchromophytes, synurophytes and chrysophytes which precede the divergence of phaeophytes, raphidophytes and eustigmatophytes from the remaining ochrophytes (Dorrell et al. 2021; Burki et al. 2016).
Chloroplasts originate via the endosymbiotic uptake of free-living photosynthetic bacteria (in the case of primary endosymbiosis), of single-celled red or green eukaryotic algae containing chloroplasts (in the case of secondary endosymbiosis), or even of eukaryotic algae containing secondary chloroplasts (tertiary or higher endosymbioses; Fig. 1). The engulfed cells are converted into stable, intracellular organelles; and this process has been observed to occur in multiple algal groups across the tree of life (Fig. 1) (Walker et al. 2011). The chloroplasts found in ochrophytes are typically characterised by the presence of four surrounding membranes, the outermost of which is contiguous to the endoplasmic reticulum (Ishida et al. 2000) and possess chlorophyll c as a light-harvesting pigment (Kowallik et al. 1995), although exceptions to these paradigms are known (Dorrell and Bowler 2017; Wetherbee et al. 2019) (Fig. 2). Both of these features are characteristic of chloroplasts acquired through the secondary or higher endosymbioses of red algae (i.e. those found in cryptomonads, haptophytes, dinoflagellates and photosynthetic apicomplexans; Fig. 1), although are not known in red algae themselves (Dorrell and Bowler 2017). In contrast, ochrophytes do not retain phycobiliprotein subunits which are found in red algae and in cryptomonad chloroplasts (Bhattacharya et al. 2013; Sturm et al. 2013)). The ochrophyte chloroplast additionally is defined by a « girdle lamella », a ring-like structure consisting of three apressed thylakoids that cover the exterior of the chloroplast stroma (Andersen et al. 1993), and additional synapomorphies (the accessory pigments diatoxanthin and diadinoxanthin) unifies the Khakista (diatoms and Bolidomonas) and hypogyristea (pelagophytes and dictyochophytes) (Dorrell and Bowler 2017; Buck et al. 2019; Kuczynska et al. 2015). (Figs. 1 and 2).
Consistent with the classical paradigm based on biochemical and ultrastructural data, multigene phylogenies of chloroplast genomes robustly resolve ochrophytes as a monophyletic group, with the chloroplast originating within red algae, and closely related to other chloroplast groups acquired through the secondary endosymbiosis of red algae (Dorrell et al. 2017a; Muñoz-Gómez et al. 2017; Stiller et al. 2014). Thus, the most parsimonious scenario for the origin of the ochrophyte chloroplast is through a single endosymbiosis event in a recent common ancestor (Dorrell et al. 2017a), prior to the divergence of diatoms (Fig. 1).
2 Phylogenomic Insights into Diatom Evolution
Prior to the genomics era, the deep evolutionary history of the eukaryotic tree was resolvable only through ultrastructural similarities, and single-gene trees realised with individual markers (e.g. 18S and chloroplast 16S ribosomal DNA (Cavalier-Smith 1998, 1999)). Based on the conserved ultrastructural features of their chloroplasts, and single-gene phylogenetic data, it was posited that the major groups of algae with chloroplasts of secondary red origin, that is, stramenopiles, cryptomonads, haptophytes and dinoflagellates, descended from a common ancestor which acquired its chloroplast through a single secondary endosymbiosis involving a red alga (Cavalier-Smith 1998, 1999). These lineages were therefore proposed to form a monophyletic group, termed the “chromalveolates”; non-photosynthetic members of these lineages, such as oomycetes, were posited to have therefore once possessed chloroplasts, and subsequently to have lost them (Cavalier-Smith 1998).
Whole-genome sequencing in diatoms commenced with the centric species Thalassiosira pseudonana and the pennate Phaeodactylum tricornutum (Armbrust et al. 2004; Bowler et al. 2008). This has also opened new windows of insight into the phylogenetic composition of the diatom nucleus (Armbrust et al. 2004; Bowler et al. 2008; Moustafa et al. 2009) which has undergone substantial elaborations over its evolutionary history. The advent of high-throughput genomic sequencing, and multigene phylogenomics, revolutionised our understanding of diatom evolution, for example, through the incorporation of rhizaria (which do not contain red algal chloroplasts) with stramenopiles and alveolates, as part of the “SAR clade” (Burki et al. 2007; Walker et al. 2011). Further studies have positioned the nuclear lineages of cryptomonads as being more closely related to that of plants and red algae than that of diatoms, directly questioning the occurrence of a single, secondary endosymbiosis event (Burki et al. 2016; Lax et al. 2018). Moreover, genomic investigations of major non-photosynthetic stramenopile groups, such as oomycetes (Stiller et al. 2009) and hypochytriomycetes (Leonard et al. 2018), have revealed an absence of genes of red algal origin that are shared with photosynthetic lineages (Stiller et al. 2014; Wang et al. 2017), placing the acquisition of the ochrophyte chloroplast after their divergence from non-photosynthetic relatives (Fig. 1).
The acquisition of a chloroplast by the common ancestor of the ochrophytes dramatically altered the composition of their genomes, adding a new organelle, with its own complex biochemical and physiological needs, to the host genome. Some of these functions are encoded in the chloroplast genome itself, and others in the genome of the stramenopile nucleus (Fig. 2). These nuclear genes, encoding chloroplast-targeted proteins, might have originated within the nucleus itself, and adapted to support the biology of the chloroplast (Novák Vanclová et al. 2020; Larkum et al. 2007; Morozov and Galachyants 2019), or might have been derived from either the nucleus or the chloroplast of the red algal ancestor of the ochrophyte chloroplast, and relocated to the host via endosymbiotic gene transfer (Dorrell and Bowler 2017; Dorrell et al. 2017a) (Fig. 2). At least, some of the proteins in the chloroplast proteome may additionally have bifunctional roles in other organelles, for example, through dual-targeting to the mitochondria (Dorrell et al. 2017a, 2019; Gile et al. 2015) or to the plasma membrane (Kazamia et al. 2018) (Fig. 2).
Alongside this, both the nuclear and mitochondrial genomes of ochrophytes and aplastidic stramenopiles have undergone their own, independent gene transfer events: for example the relocation of mitochondria-encoded functions to the nucleus and the horizontal acquisition of non-chloroplast related genes of non-stramenopile origin by the stramenopile nuclear genome (Bowler et al. 2008; Keeling and Palmer 2008; Rastogi et al. 2018). Here, we summarise current knowledge (as of 2022) regarding different evolutionary sources of novelty in diatom nuclear genomes that have arisen through the course of stramenopile evolution, focusing on the genome of the model diatom P. tricornutum, which remains the best-studied system for phylogenomic reconstructions of gene transfer in the diatom lineage (Bowler et al. 2008; Rastogi et al. 2018) (Table 1). Subsequently, we explore the dynamic evolution of diatom mitochondria and chloroplasts, and identify features underpinning the progressive loss, and occasional gain, of novel coding functions in the organelle genomes of individual diatom species.
2.1 Methods and Problems when Exploring Origins of Diatom Nuclear Genes
Dynamic transitions in the evolutionary history of the diatom nuclear, mitochondrial and chloroplast genomes can be identified, and quantified, by comparative genomic and phylogenomic approaches (Bowler et al. 2008; Moustafa et al. 2009; Rastogi et al. 2018) (Table 1). Understanding these transitions is important as it may reveal adaptations of diatoms to their current environments and allow us to pinpoint genomic novelties that may explain their success over red or green algae, or even other stramenopile groups, in the contemporary ocean (Dorrell and Smith 2011). However, the exact size, timing, reason and even the veracity of individual gene transfer events can be controversial (Deschamps and Moreira 2012; Dagan and Martin 2009), depending particularly on the methodology used.
Firstly, different results may be obtained depending on the reference library used which may be constrained both by taxonomic undersampling and oversampling of key lineages. For example, early estimates of the green algal signal in diatom genomes (Armbrust et al. 2004; Bowler et al. 2008; Ponce-Toledo et al. 2019) were likely to be overinflated by the paucity of red algal gene models at the time of analysis, being largely dependent on the highly reduced genome of the extremophilic alga Cyanidioschyzon merolae as the only publicly available red algal gene reference (Matsuzaki et al. 2004; Collén et al. 2013).
In contrast, the indiscriminate use of transcriptome data and poorly curated reference genomes can lead to the misidentification of contaminants as horizontally transferred genes (Dorrell et al. 2017a; Vancaester et al. 2020; Burki et al. 2012), or the preferential identification of genes with unresolvable or ambiguous origins as being related to oversampled taxa (Deschamps and Moreira 2012). These strategies may be avoided by judicious taxonomic sampling when constructing reference datasets (Dorrell et al. 2017a; Morozov and Galachyants 2019): alongside the use of new long-read sequencing technologies (e.g.; PacBio (Rhoads and Au 2015)), assembly approaches (e.g. Dovetail (Moll et al. 2017)), gene annotation pipelines (incorporative of transcriptome or proteomic data (Rastogi et al. 2018; Yang et al. 2018)), and in silico cleaning of reference genome and transcriptome libraries (e.g. through considering nucleotide composition, (Dorrell et al. 2021; Hehenberger et al. 2016; Sato et al. 2020)) to improve curation of gene models, and remove probable contaminant sequences.
Different results can also be obtained because of the methodology used, for example, BLAST-based analyses such as ranking of top hits (Stiller et al. 2014; Armbrust et al. 2004; Rastogi et al. 2018; Méheust et al. 2016) versus whole-genome phylogenomic analysis (Dorrell et al. 2021; Morozov and Galachyants 2019; Deschamps and Moreira 2012; Vancaester et al. 2020; Fan et al. 2020). BLAST top hit analysis is less computationally intensive than phylogenetic techniques and provides several technical advantages, for example, the ability to infer possible evolutionary histories for genes that are too short or divergently evolved (e.g. chimeric gene fusions) to be resolved through classical phylogenetic approaches (Dorrell et al. 2017a, 2021; Méheust et al. 2016), but its resolution is necessarily poorer: for example not being able to determine the directions of gene transfers that can be inferred from the topology (e.g. monophyly versus paraphyly of stramenopile lineages) within a phylogenetic output (Dorrell et al. 2017a; Morozov and Galachyants 2019).
The threshold stringencies applied when identifying evolutionary origins may also bias the evolutionary relationships reconstructed. Too permissive criteria can admit poorly resolved or artefactual relationships, whereas too stringent parameters can conversely eliminate usable phylogenetic signals. For example using phylogenomic approaches and automated tree sorting, Moustafa et al. identified 418 red and 1757 green genes in the P. tricornutum genome (Moustafa et al. 2009) (Table 1). Following manual reinspection of the green gene dataset identified by Moustafa et al. (Moustafa et al. 2009), Deschamps and Moreira found that 91 of the putative green gene trees had topologies consistent with a vertical transfer of genes from the green lineage into the diatoms, whereas 89 of the putative green genes had histories more consistent with a transfer from the red lineage into the diatoms (Deschamps and Moreira 2012), although they did not perform a comparable analysis of the red genes identified by Moustafa et al. (Table 1).
Finally, even within well-sampled datasets, the exact questions that are posed by a study of horizontal gene transfer will determine the results obtained. Early studies on diatoms necessarily focused on gene transfers from animals, fungi, bacteria, red algae and green algae, as these were the reference genomic models available at the time (Armbrust et al. 2004; Bowler et al. 2008; Moustafa et al. 2009). However, diatoms may also have exchanged genes with other algae with secondary chloroplasts (e.g. cryptomonads, haptophytes and dinoflagellates): either as part of chloroplast endosymbiosis events (Dorrell et al. 2017a, 2021; Stiller et al. 2014) or through independent horizontal gene transfers (Nonoyama et al. 2019; Kazamia et al. 2018), and some of the “bacterial,” “red” or “green” genes identified in early studies may have passed through one or more of these algal groups as intermediates prior to arriving in diatom genomes (Dorrell et al. 2021). Genes acquired horizontally into diatom genomes may have variously been received in early ancestors of the SAR clade, stramenopiles or ochrophytes, in the diatom ancestor, or even specifically within the diatom species considered; and recent studies have profited from the expanded number of diatom and stramenopile genomes and transcriptomes available to infer the probable timing of individual gene transfers (Dorrell et al. 2021; Vancaester et al. 2020; Fan et al. 2020). These more densely sampled datasets may even be able to identify occasions in which diatoms or their ancestors have acted as donors, rather than recipients, in gene transfers with other lineages (Dorrell et al. 2017a, 2021).
In Table 1, we profile the key results of different studies of horizontal gene transfers involving the diatom nucleus, focussing on the T. pseudonana and P. tricornutum genomes (Dorrell et al. 2017a, 2021; Armbrust et al. 2004; Bowler et al. 2008; Moustafa et al. 2009; Rastogi et al. 2018; Deschamps and Moreira 2012; Vancaester et al. 2020; Fan et al. 2020). The most recent of these analyses, involving a densely sampled reference dataset of all currently available eukaryotic and prokaryotic genomes and transcriptomes, and manual sorting of trees by partner lineage, timing and direction of horizontal transfer (Dorrell et al. 2021), identified 1347/12177 (11.0%) genes in the P. tricornutum genome to have been horizontally acquired since the origin of the ochrophytes, and a further 1771 examples, distributed over 1184 (9.7%) genes, of gene transfers from the ochrophytes into the other branches of the tree of life (Dorrell et al. 2021). Below, we discuss the likely origin points and functions of different horizontally transferred genes, including genes acquired from bacteria, red algae and their endosymbiotic descendants, green algae, as well as gene transfers from diatoms into other eukaryotic lineages.
2.2 Prokaryotic Signals
Phylogenomic annotations of the P. tricornutum have identified large numbers of genes of prokaryotic (i.e., bacterial and archaeal) origin, inferred to constitute 7.4% (784 genes) of the original annotation (Bowler et al. 2008) and reduced to a still substantial 285 genes (2.3%) (Dorrell et al. 2021) in the most recent estimates from the third genome annotation with more densely sampled taxonomic references (Table 1) (Dorrell et al. 2021). Most of these genes pertain to bacteria, with limited or zero estimated archaeal and viral contributions into diatom genomes (Dorrell et al. 2021; Rastogi et al. 2018). These genes have been acquired progressively through diatom evolution, with studies focused on Phaeodactylum (Dorrell et al. 2021; Rastogi et al. 2018); and indeed on wider diatom pan-genomes (Dorrell et al. 2021; Vancaester et al. 2020; Fan et al. 2020), detecting large numbers of bacterially derived genes into early ancestors of the SAR clade, stramenopiles or ochrophytes; into the diatom common ancestor; and even into individual diatom groups or species (Table 1).
Two recent studies from Vancaester et al. (2020) and from our group (Dorrell et al. 2021), which have reconciled automatically resolved bacterial transfers from densely sampled datasets with multigene reference tree topologies for SAR clade members, have attributed large numbers of bacterial gene transfer to early ancestors of the diatoms, following their divergence from other stramenopiles (Dorrell et al. 2021; Vancaester et al. 2020) (Table 1). This may reflect a greater flux of bacterial DNA into diatoms than related algae which may contribute to their comparative ecological success (Dorrell et al. 2021). However, it is possible that this asymmetry reflects the greater number of diatom reference sequence libraries in which gene transfers may be identified, and a parallel study by Fan et al. (2020) using a smaller number of diatom genomes, but balanced taxonomic sampling of diatom and non-diatom species, did not appear to yield greater numbers of bacterial genes in diatoms than other lineages (Fan et al. 2020) (Table 1). An alternative explanation is a high rate of transfer, but also secondary loss of bacterial-derived genes into diatoms which may enable specific diatom species to explore new evolutionary niches and fluctuating environments (Dorrell et al. 2021).
Regardless of the frequencies of their appearance, bacterial genes contribute a wide range of different functions to diatoms. For example, within the dataset of 770 genes encoding chloroplast-targeted proteins that are shared across (and presumably ancestral to) all ochrophytes, we identified 49 of probable prokaryotic origin, including many implicated in the expression of the chloroplast genome (Dorrell et al. 2017a). More recently, considering both in silico targeting predictions and GFP labelling, we have shown that bacterial genes acquired early during ochrophyte evolution are enriched in chloroplast-targeted proteins, consistent with origins tied to the ochrophyte chloroplast, whereas more recently acquired proteins predominantly function in the diatom secretome (Dorrell et al. 2021). Many of these more recently acquired bacterial genes have functions pertaining to environmental stress: ice-binding proteins in polar-adapted diatoms (Sorhannus 2011; Raymond and Kim 2012), and proteins involved in the synthesis of Vitamin B12 which may be a limiting micronutrient in many of the Southern Ocean habitats where diatoms are abundant (Vancaester et al. 2020; Browning et al. 2017).
2.3 Red-Algal-Related Signals
Early estimates posited approximately 400 genes (4% of the coding content) in the P. tricornutum and T. pseudonana genomes were of red algal origin (Table 1) (Bowler et al. 2008; Moustafa et al. 2009; Rastogi et al. 2018). These genes principally encode proteins of chloroplast function, and are shared across ochrophytes, for example, constituting one-half (255 of the 502 genes) of the chloroplast-targeted proteins shared across all ochrophytes identified by Dorrell et al. 2017a, b for which a tractable phylogenetic signal could be found (Table 1) (Dorrell et al. 2017a). In contrast, only four of these genes were found to be shared with non-photosynthetic stramenopile groups such as oomycetes and labyrinthulomycetes (Dorrell et al. 2017a) (Table 1). This, alongside the previously discussed lack of red algal signal in the genomes of non-photosynthetic stramenopiles (Stiller et al. 2009, 2014; Leonard et al. 2018; Wang et al. 2017), is consistent with a late acquisition of the ochrophyte chloroplast, after their divergence from oomycetes (Dorrell and Bowler 2017).
Despite the massive expansion in the availability of red algal genome sequences and transcriptomes in the last decade (Dorrell et al. 2017a; Collén et al. 2013; Brawley et al. 2017), the number of red algal-derived genes identified in stramenopile genomes has remained relatively constant. For example, whereas Moustafa et al. (2009) identified 418 genes of putative red origin in the P. tricornutum genome using the extremophilic red alga C. merolae as a principal reference (Moustafa et al. 2009), only 459 red genes were identified by Rastogi et al. (2018) using a much more exhaustive dataset including five complete red algal genomes and thirteen different red algal transcriptomes (Rastogi et al. 2018) (Table 1). One possible reason for this quite limited red contribution is the relatively high level of reduction observed in red algal nuclear genomes, which lack many of the accessory functions identified in diatoms and in other eukaryotic groups (Rastogi et al. 2018; Qiu et al. 2017).
An alternative scenario is that the red algal signal in stramenopile lineages does not itself directly come from red algae, but from a secondary, red-chloroplast containing lineage that was acquired by the stramenopiles through a tertiary or more complex endosymbiosis event. Using an innovative methodology based on the similarity of BLAST hit signals, Stiller et al. (2014) concluded a probable cryptomonad origin for the ochrophyte chloroplast (Stiller et al. 2014). In our most recent analyses, considering algae with secondary chloroplasts as possible donors into diatom genomes, we could only manually identify 193 P. tricornutum genes that resolved phylogenetically as a direct sister to red algae, but found a further 514 that resolved with cryptomonads, haptophytes or dinoflagellates, with haptophytes contributing the greatest number (Table 1) (Dorrell et al. 2021). In this case, whatever functions that might have been inherited by the stramenopile ancestor from red algae would have been winnowed by not only one, but potentially two or more successive endosymbiotic events. Ultimately, identifying why stramenopiles contain the red algal genes that they do will depend on tracing the permeation of the red signal across all major groups of algae with secondary, red-derived chloroplasts.
2.4 Green-Algal-Derived Signals
As discussed above, diatoms contain a sizeable number of genes of green algal origin, although the number identified varies based on the methodology employed (Moustafa et al. 2009; Morozov and Galachyants 2019; Deschamps and Moreira 2012) (Table 1). In our most recent estimates, we identified 260 (2%) genes in the P. tricornutum genome showing relationships consistent with a direct gene transfer from the green algae into the ochrophytes, most of which resolve to deep nodes in ochrophyte evolution (Table 1) (Dorrell et al. 2021), and 105 out of 502 (20%) of conserved ochrophyte chloroplast-targeted proteins for which a tractable origin was identified for green algae (Dorrell et al. 2017a) (Table 1). The exact numbers and identities of the green genes in diatom genomes are likely to undergo further revision with deeper sequencing of red algae revealing previously unidentified homologues of diatom genes (e.g., xanthophyll cycle genes (Dautermann and Lohr 2017)). Other green genes may be reassigned as having more direct cryptomonad, haptophyte or dinoflagellate origins, although even the most stringent pipelines and sampling thresholds fail to completely eliminate the green signal from diatom genomes (Dorrell et al. 2021; Deschamps and Moreira 2012).
Beyond the presence and size of the green signal in diatoms, other questions remain, namely: where do these genes come from, and how were they acquired? Phylogenomic analyses typically pinpoint green genes as arising within chlorophytes (i.e. single-celled green algae) rather than streptophytes (plants and their closest relatives) which helps argue against their origin as being artefactual because of phylogenetic misannotation (Dorrell et al. 2017a; Moustafa et al. 2009), although it remains to be determined where within the chlorophytes this signal preferentially falls (Table 1). The green genes that are present in diatom genomes are enriched in genes encoding chloroplast-targeted proteins (Dorrell et al. 2017a) which might be consistent with a chloroplast endosymbiotic origin. This could be due to a cryptic green endosymbiosis during the evolutionary history of the stramenopile host, or the acquisition of a tertiary or higher chloroplast containing mixed signals of red and green origin (Dorrell and Bowler 2017; Dorrell and Smith 2011). Answering precisely how green genes were transferred into diatoms will depend both on deeper sequencing of the green algal tree (particularly in the context of early-diverging members, e.g. Prasinoderma (Li et al. 2020)), and ideally the verification of the exact evolutionary origin of the ochrophyte chloroplast (from red algae, cryptomonads or another group entirely) (Stiller et al. 2014).
Perhaps most interesting is to consider what functional advantages green genes might contribute to diatoms. Many of the green genes encoding chloroplast-targeted proteins have functions in biosynthetic pathways (e.g. chloroplast and carotenoid synthesis (Dorrell and Bowler 2017; Coesel et al. 2008; Frommolt et al. 2008)), and their presence may change the metabolic functions observed in the diatom chloroplast. Other diatom green genes have functions in both chloroplast- and non-chloroplast-related environmental stress responses: Lhcx light-harvesting complex genes implicated in photoadaptation to aberrant light conditions (Buck et al. 2019; Büchel 2015), and the iron-stress-related protein ISIP2a which mediates non-reductive iron uptake across the plasma membrane (McQuaid et al. 2018; Allen et al. 2008). Understanding the functional significance of the green footprint in diatom genomes will depend on large-scale characterisation of their encoded properties: using environmental sequence datasets and targeted mutagenesis in model taxa (Dorrell and Smith 2011).
2.5 Have Diatoms Donated Genes to Other Organisms?
Alongside the mosaic origin of diatom genomes, it is possible that diatoms, or ochrophytes in general, have donated genes to other organisms. For example we previously identified 243 conserved chloroplast-targeted proteins that have been shared between the ochrophytes and the haptophytes (Dorrell et al. 2017a) which we have subsequently expanded to 817 genes (6.7%) of the Phaeodactylum genome supporting horizontal gene transfer from ochrophytes to haptophytes (Dorrell et al. 2021) (Table 1). Phylogenetic analysis of these genes indicated a mass transfer event from the hypogyristea (pelagophytes and dictyochophytes) into a common haptophyte ancestor (Dorrell and Bowler 2017; Dorrell et al. 2017a, 2021). These genes predominantly have chloroplast-targeted functions and may be footprints of an ancient endosymbiotic transfer of the ochrophyte chloroplast into the haptophytes (Dorrell et al. 2017a; Stiller et al. 2014). We have subsequently inferred the same enrichment in pelagophyte/dictyochophyte affinities for genes transferred from the ochrophytes to the dinoflagellates, which may relate to direct or indirect (e.g. via haptophyte) gene transfers (Dorrell et al. 2021) (Table 1). Further phylogenetic analysis of the chloroplast genomes of the photosynthetic alveolates Chromera and Vitrella has revealed a possible relationship with the ochrophytes, potentially involving a chrysophyte endosymbiotic transfer (Ševčíková et al. 2015; Dorrell et al. 2021; Kim et al. 2017). These serial transfer events may help resolve the still uncertain origins, dynamics and functions of the red and green algal signals associated with marine algal genomes.
Despite the gene transfers associated with other ochrophytes, there is relatively little evidence that diatoms themselves have transferred large numbers of genes into other algal groups—with one exception. The « dinotoms », a closely related group of dinoflagellate algae within the order Peridiniales, possess whole cell endosymbionts of diatom origin (Imanian et al. 2010; Yamada et al. 2019). These endosymbionts, which retain a complete chloroplast, mitochondria and nucleus, appear to have been acquired, lost and replaced on multiple occasions, from both pennate and centric diatom lineages (Kretschmann et al. 2018; Yamada et al. 2017, 2020). Phylogenomic analysis of dinotom nuclear transcriptomes reveals very little evidence for the loss of functions from the symbionts, or transfer of genes to the host, consistent with their relatively transient evolutionary associations (Hehenberger et al. 2016; Burki et al. 2014). It remains to be determined what genetic integration events, if any, are required for the stable domestication of the dinotom endosymbionts.
Despite their relatively limited evolutionary interactions with other algal groups, diatoms may exchange genes with each other which may have adaptive functions to certain environmental stresses. For example the iron-stress-associated protein ISIP1, which facilitates the non-reductive uptake of extracellular siderophores by endocytosis, shows a complex phylogeny, consistent with either multiple transfer events between diatoms, or independent paralogy and gene loss events (Kazamia et al. 2018). Similarly, phylogenies of ice-binding proteins, which confer cold stress tolerance in polar native algae, reveal probable horizontal transfer events between centric and pennate species (e.g. between the centric diatom Chaetoceros and the pennate Navicula sp. or between the centric Attheya and the pennate Amphora sp. (Sorhannus 2011; Raymond and Kim 2012)). Resolving these more recent gene transfer events is more challenging, as they are more likely to be biased by limitations in taxonomic sampling and limited resolution of short protein sequences (Swenson 2009). However, explorations of adaptive and within-genus diatom gene transfers is becoming increasingly possible due to dense transcriptomic sampling of specific genera (e.g. Thalassiosira, Chaetoceros, Fig. 1 (Ichinomiya et al. 2016)) or genomes (Thalassiosira, Pseudo-nitzschia; (Basu et al. 2017)).
3 Evolution of the Diatom Organelle Genomes
3.1 Diatom Mitochondria
Diatom mitochondrial genomes display a largely well-conserved organisation, forming a single, circular chromosome with a single repeat region (Kamikawa et al. 2018; Crowell et al. 2019). These genomes typically contain 33 core-protein-coding genes, alongside genes for tRNAs and rRNAs (Fig. 3a) (Crowell et al. 2019; Ravin et al. 2010; Oudot-Le Secq and Green 2011). This coding content is less than in some microbial eukaryotes (e.g. 47 protein-coding genes in the free-living stramenopile relative Ancoracystis), but somewhat greater than the three protein-coding genes retained in dinoflagellate mitochondria (Janouškovec et al. 2017; Nash et al. 2007). The coding content of diatom mitochondria is similar to that of both ochrophytes and non-photosynthetic stramenopiles (Fig. 4a), except for the probable ancestral loss of atp1 in diatoms which is mitochondria-encoded in both photosynthetic eustigmatophytes (Ševčíková et al. 2016) and non-photosynthetic oomycetes and bicosoecids (Jirsová et al. 2019). A further seven genes have more sporadic distributions across diatom mitochondria, including tatA, which is only mitochondria-encoded in a few select genera (Cattolico et al. 2008; Crowell et al. 2019; Guillory et al. 2018) (Fig. 3b).
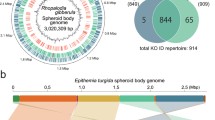
Organisation of the diatom mitogenome. Top: Venn Diagram of mitochondrial-genome-encoded proteins assignable to the common ancestors of diatoms, other ochrophytes (chrysista), and aplastidic stramenopiles (oomycetes, labyrinthulomycetes, biocosiceds). Bottom: heatmap of the occurrence of seven genes with sporadic distribution in published diatom mitochondrial genomes. Red cells indicate presence and blank cells indicate absence
Plastid genomes of diatoms and their relatives. Venn Diagram of chloroplast–encoded genes commonly found in photosynthetic diatoms (Yu et al. 2018), non-photosynthetic diatoms: Nitzschia (Kamikawa et al. 2018), ochrophytes assigned to the hypogyristea (pelagophytes, dictyochophytes, Triparma) (Han et al. 2019; Ong et al. 2010; Tajima et al. 2016), and chrysista (chrysophytes, Nannochloropsis, phaeophytes) (Ševčíková et al. 2015; Cattolico et al. 2008; Le Corguillé et al. 2009; Ševcíková et al. 2019). Genes identified in fewer than 25% of published genomes within a particular lineage are not shown
Alongside this relatively conserved genomic content, more unusual organisations have evolved in individual mitochondrial genomes. These include structural changes, for example, a possible linear organisation of the mitochondrial genome in the secondarily non-photosynthetic diatom Nitzschia (Kamikawa et al. 2018), and expansions (Oudot-Le Secq and Green 2011) or losses (An et al. 2016) of the repeat regions in different diatom species. These also include changes likely to impact on the expression pathways of individual genes: the dynamic transfer and independent inheritance of introns in diatom mitochondrial coxI and rnL genes (Crowell et al. 2019; Ravin et al. 2010; Guillory et al. 2018), independent origins of translationally fused gene pairs in the pennate species Halamphora and Phaeodactylum (Crowell et al. 2019; Oudot-Le Secq and Green 2011), and the possible use of UGA-stop codons to encode tryptophan in the centric diatoms Thalassiosira and Skeletonema (Ehara et al. 2000).
Most dramatically, a change in the post-translational mitochondrial biology of raphid pennate diatoms (including Fragilariopsis, Phaeodactylum and the diatom endosymbionts of dinotom algae) has been noted, in which the nad11 gene is divided into two separately located and independently transcribed ORFs, corresponding to the iron-sulphur-binding and the molybdopterin-binding domains (Oudot-Le Secq and Green 2011; An et al. 2016; Imanian et al. 2012). This configuration is not known in araphid pennate or centric diatom mitochondria, and its functional consequences remain unknown (Imanian et al. 2012).
3.2 Diatom Chloroplast Genomes
The chloroplast genomes associated with diatoms are typically arranged as a single, circular chromosome, with 134–180 protein-coding genes, alongside ribosomal and transfer RNAs (Fig. 4) (Dorrell and Bowler 2017; Yu et al. 2018; Hamsher et al. 2019; Prasetiya et al. 2019). Phylogenomic analysis, and the presence of discrete molecular synapomorphies (e.g. the presence of a form ID type rubisco), robustly places this chloroplast genome within red algae and closely related to the chloroplasts of cryptomonads and haptophytes (Muñoz-Gómez et al. 2017; Janouskovec et al. 2010; Tabita et al. 2008). However, the coding content of the diatom chloroplast genome is somewhat less than the ca. 250 genes associated with red algal chloroplasts (Muñoz-Gómez et al. 2017; Qiu et al. 2017), and indeed is less than the chloroplast gene contents of other ochrophyte groups (Dorrell and Bowler 2017): raphidophytes (Heterosigma) (Cattolico et al. 2008), phaeophytes (Ectocarpus) (Le Corguillé et al. 2009) and chrysophytes (Ochromonas) (Ševčíková et al. 2015), pointing to both endosymbiotic and post-endosymbiotic reductions in diatom chloroplast genome content (Fig. 3). This notwithstanding, the diatom chloroplast genome has undergone less dramatic reductions than some other ochrophyte groups (e.g. pelagophytes, dictyochophytes), in which even the loss of the chloroplast inverted repeat is known (Han et al. 2019; Ong et al. 2010).
Alongside these general trends, different diatom species have retained and lost different patterns of genes (Fig. 5). For example the basally divergent genus Leptocylindrus (Parks et al. 2018) retains a chloroplast petJ gene encoding cytochrome c§ which is also retained in other ochrophytes (e.g. Ectocarpus) (Dorrell and Bowler 2017; Le Corguillé et al. 2009), suggesting it was present in the diatom common ancestor and lost from other species. Similarly, the ilvB and ilvH genes, encoding the small and large subunits of acetolactate synthase, have been retained in the chloroplast genomes of the diatom genera Leptocylindrus, Coscinodiscus, Cerataulina, Acanthoceros and Eunotia (Yu et al. 2018; Sabir et al. 2014). Conversely, the light-independent protochlorophyllide oxidoreductase complex, encoded by the chlB, chlL and chlN genes, is uniquely chloroplast-encoded in the diatom Toxarium (Ruck et al. 2017). These diatoms are distantly positioned to one another and (except for Leptocylindrus) to the base of the diatom tree (Parks et al. 2018), but single-gene phylogenies reveal likely vertical origins of each gene (Ruck et al. 2017), suggesting that they have been independently lost in a wide range of other diatoms.
Chloroplast-encoded genes with sporadic distributions across diatom species. Diatom species are sorted reflecting their underlying phylogenetic relationships with most basally divergent members at the left, following (Yu et al. 2018; Parks et al. 2018). Blue cells indicate presence and blank cells indicate absence
Other diatoms have lost genes typically retained in the chloroplast genomes of other species (Figs. 5, and 6). Substantial losses of chloroplast-encoded genes have been noted in the centric species Proboscia and the pennate species Astrosyne radiata which form extremely long branches in diatom chloroplast gene trees (Yu et al. 2018; Ren et al. 2020). Astrosyne was isolated from a shallow-water coral reef habitat in Guam (Ashworth et al. 2012), whereas Proboscia was isolated from the Red Sea, and it remains to be determined if this reductive evolution is correlated to high-light or temperature adaptations in either species (Yu et al. 2018).
Dynamic of evolutionary rates in diatom chloroplast genomes. Scatterplots showing the mean pairwise identities observed by BLAST searches between complete (>90% coverage) protein-coding sequences with universal presence (green), identified to be infrequently lost (red), or to have sporadic distributions (serC, tsf, ilvB, ilvH) in diatom chloroplast genomes, plotted against similar pairwise identity scores for pairs of non-diatom stramenopile sequences (left), and pairs of diatom and non-diatom sequences (right). The regression gradients for the chloroplast-encoded proteins with incomplete presences in diatom chloroplasts are much steeper than those with universal presence: a small decrease in non-diatom chloroplast sequence identity correlates to a much larger decrease in diatom chloroplast sequence identity. This indicates that the chloroplast-encoded proteins that are variably present in diatom chloroplasts are more rapidly diverging, even within their chloroplast milieu, than genes that are never lost
3.3 Diatom Leucoplast Genomes
An even more dramatic degree of reduction is known in some diatoms within the genus Nitzschia, which have secondarily lost the capacity for photosynthesis, and may have originated through one (Onyshchenko et al. 2019) or independent evolutionary events (Kamikawa et al. 2015a). These species retain vestigial, non-photosynthetic plastids known as « leucoplasts » which perform essential biosynthetic functions and still retain genomes, but lack plastid-encoded genes associated with photosynthetic functions: genes for photosystem I, II, cytochrome b6f, the Calvin Cycle and chlorophyll biosynthesis (Fig. 4) (Kamikawa et al. 2017, 2018). Notably, the Nitzschia leucoplast genome retains genes encoding subunits of plastid ATP synthase (Fig. 4) which has been proposed to function in this organelle in the catabolic consumption of ATP, allowing the maintenance of a thylakoid proton gradient that would permit function of the thylakoid Tat protein import complex (Kamikawa et al. 2015b, 2017). Comparative analysis of different non-photosynthetic Nitzschia species reveals largely conserved leucoplast genome content, consistent either with convergent trajectories following independent losses of photosynthesis, or limited divergence in genome content following a single loss of photosynthesis (Kamikawa et al. 2018).
The Nitzschia leucoplast genome is notably less reduced in coding content compared to that of other secondarily non-photosynthetic plastids of similar evolutionary origin, for example those identified in non-photosynthetic chrysophytes or in apicomplexans (Dorrell et al. 2019; Hadariová et al. 2018) which have lost a wider range of leucoplast-encoded functions, including ATP synthase. It has been proposed that the osmotrophic feeding strategies of non-photosynthetic Nitzschia spp. may limit their ability to supplement plastidial functions with heterotrophically acquired metabolites, explaining the greater relative functional autonomy of their leucoplasts compared to phagotrophic (chrysophyte) or parasitic (apicomplexan) lineages (Dorrell et al. 2019; Kamikawa et al. 2017). Ultimately, further functional characterisation is required to understand the metabolic contributions of the Nitzschia leucoplast in the context of other, secondarily non-photosynthetic algal species. which will be aided by the recent completion of a nuclear transcriptome (Kamikawa et al. 2017) and genome (Pendergrass et al. 2020).
3.4 Why Are Certain Genes Lost from Diatom Chloroplasts?
It remains to be determined what physiological processes underpin the reductive evolution of diatom organelle genomes. This is the case both for genes that have been lost from diatom chloroplasts compared to other algal groups, and genes that have been lost from individual diatom lineages. In other groups of eukaryotes, for example, plants and dinoflagellates, the loss of genes from the chloroplast genome is correlated to mutation rate and a loss in constraining selective pressure (Dorrell et al. 2017b; Magee et al. 2010), and it is notable that Astrosyne and Proboscia, which have undergone substantial amounts of chloroplast gene loss, are also two of the fastest-evolving diatom species, considering chloroplast-encoded substitution rates (Ren et al. 2020).
As a test of this principle, we have compared the degree of conservation between proteins encoded with universal, sporadic and occasional presence in diatom chloroplast genomes (Fig. 6). As controls, we have considered the conservation observed between orthologues from non-diatom stramenopiles, and between diatom and non-diatom species (Fig. 6). We note that the proteins with sporadic distributions in diatom species are much more divergent than would be expected compared to both proteins with conserved distributions and orthologues from non-diatom stramenopiles (Fig. 6). Thus, the genes that are most frequently lost from diatom chloroplast genomes are likely to be those already under relaxed selection in the diatom chloroplast.
3.5 Loss Versus Transfer of Diatom Organelle Genes
Another related question is under what circumstances genes are lost completely from diatom organelles, versus being relocated to the nucleus. Typically, genes are lost from chloroplast and mitochondrial genomes if they no longer perform necessary functions in these organisms.
In contrast, genes may be relocated to the nucleus to permit differential regulation, to protect from elevated chloroplast mutation rates or to change the stoichiometry of their expression relative to chloroplast-encoded copies (Magee et al. 2010; Dorrell and Howe 2012; Noordally et al. 2013). In the context of diatoms, the open ocean species Thalassiosira oceanica, which is adapted to tolerate chronic iron limitation (Gao et al. 2021), has been proposed to have relocated the gene petF, encoding the iron-sulphur protein ferredoxin, to the nucleus, to allow its regulation in response to environmental iron availability (Lommer et al. 2010). In another case, the psb28 gene of the centric diatom T. pseudonana has been shown to be present as both chloroplast- and nucleus-encoded copies, suggesting it is midway through a functional chloroplast to nuclear transfer event (Jiroutova et al. 2010). Psb28 has a non-essential role in photosynthetic complex assembly, but its absence from the cyanobacteria Synechococcus and Synechocystis leads to retarded growth under high light conditions (Bečková et al. 2017). It remains to be determined whether the nucleus-encoded copies of these proteins permit photo-adaptation in T. pseudonana.
3.6 Gains and Transfers of Novel Diatom Organelle ORFs
Finally, alongside the loss and reduction of organelle-encoded genes, diatom chloroplasts and mitochondria possess novel ORFs not identified in other lineages. For example some pennate diatom chloroplasts are known to contain serC and tsf genes, encoding serine and tyrosine recombinases (Hamsher et al. 2019). These genes are frequently located on plasmids and may represent the traces of mobile genetic elements integrated into diatom chloroplasts (Hildebrand et al. 1991; Jacobs et al. 1992).
Other ORFs are either conserved across all diatoms, or indeed all stramenopiles, hence might form ancestral components of stramenopile organelle genomes. Among these is ycf66, a gene found in plant and green algal chloroplast genomes, and conserved in cyanobacteria (Stoebe et al. 1998), although it is frequently lost in some lineages (Leliaert et al. 2016; Gao et al. 2011), and is not known in the chloroplasts of red algae. A phylogeny of ycf66 positions the ochrophytes as a monophyletic group, within the green lineage, as a sister-group to the streptophyte lineage (i.e. land plants and their close relatives), except for the early-branching species Klebsormidium (Fig. 7).
Evolution of diatom chloroplast ycf66. Consensus MrBayes and RAxML tree, realised under three substitution matrices (GTR, Jones/JTT, WAG), for a 31 taxa x 93 aa alignment of ycf66 from cyanobacteria and chloroplast genomes. The MrBayes topology supports placement of the ochrophyte sequences within the green algae, as a sister-group to all streptophyte species except Klebsormidium sp.
The most parsimonious explanation for this topology is a transfer of the ycf66 gene from the green chloroplast lineage into a common ancestor of the ochrophyte chloroplast, concomitant with its endosymbiotic uptake, which would represent the first known example of a horizontal gene transfer event between two distantly related chloroplast genomes. Understanding the functional consequences of this transfer will depend on characterisation of the physiological role performed by ycf66 which remains unknown.
4 Concluding Remarks
In this chapter, we have explored the phylogenetic context of the diatoms within their broader constituent lineage, the ochrophytes, within the stramenopiles (Fig. 1). Diatoms are evolutionary mosaics, containing complex chloroplasts with complicated evolutionary histories (Fig. 2), and nuclear genomes that have been enriched via by the horizontal and endosymbiotic acquisition of genes with chloroplast and non-chloroplast functions (Table 1). We show that the diatom nucleus is supported by genes of prokaryotic, red, green and other eukaryotic algal origin (Table 1). While the exact numbers, and even the individual evolutionary events that have given rise to these genes remain uncertain, their presence is indelible and may have contributed unique biological functions to diatom cells.
Alongside this, we have compared reductive trajectories in mitochondrial (Fig. 3) and chloroplast genome content (Figs. 4, and 5) across the stramenopile tree of life, focusing on sources of evolutionary difference within the diatom lineage. Diatom organelles are marked by reductive evolution, although greater extremes exist in both non-diatom and diatom branches of the stramenopile tree (e.g. non-photosynthetic members of the genus Nitzschia) than the organelle genome reductions associated with the common ancestor of the diatom lineage following its divergence from other ochrophyte groups. We show that the genes are frequently lost from diatom chloroplast genomes and tend to exhibit lower global similarity than well-conserved genes (Fig. 6). It remains to be determined whether this is driven by chloroplast mutation rate, which appears to be somewhat lower in diatoms than that of the mitochondria, despite having a more conserved coding content (Krasovec et al. 2019) Figs. 4, and 6), or via relaxed selection pressure which has been shown to substantially vary across different chloroplast genes in other algal lineages (e.g. dinoflagellates (Klinger et al. 2018)).
Finally, a possible gain of a chloroplast-encoded function has been identified, the uncharacterised open reading frame ycf66 which appears to have been acquired by ochrophytes via horizontal gene transfer of an equivalent open reading frame from green algal chloroplasts (Fig. 7). The role of this transfer event awaits functional and environmental characterisation. We stress the importance of considering function alongside the number, tempo and mode of acquisition of novel diatom genes, when exploring the extraordinary environmental success of this lineage. In this context, emergent transformable systems (Mock et al. 2017; Brunson et al. 2018; Sharma et al. 2018), and environmental sequence datasets (Caputi et al. 2019; Carradec et al. 2018), may cast new functional insights into the consequences of dynamic evolutionary events in diatom nuclei and organellular genomes.
Abbreviations
- ATP:
-
adenosine triphosphate
- BLAST:
-
Basic Local Alignment Search Tool
- DNA:
-
deoxyribonucleic acid
- ER:
-
endoplasmic reticulum
- GFP:
-
green fluorescent protein
- GTR:
-
generalised time reversible substitution model
- HGT:
-
horizontal gene transfer
- ISIP:
-
iron-stress-induced protein
- JTT:
-
Jones-Taylor-Thornton substitution model
- Lhc:
-
light harvesting complex
- ORF:
-
open reading frame
- Pt:
-
Phaeodactylum tricornutum
- rRNA:
-
ribosomal ribonucleic acid
- SAR:
-
stramenopiles, alveolates and rhizaria
- Tat:
-
twin arginine transporter
- Tp:
-
Thalassiosira pseudonana
- tRNA:
-
transfer ribonucleic acid
- WAG:
-
Whelan and Goldman substitution model
References
Adl SM et al (2012) The revised classification of eukaryotes. J Euk Microbiol 59:429–493
Allen AE et al (2008) Whole-cell response of the pennate diatom Phaeodactylum tricornutum to iron starvation. Proc Natl Acad Sci U S A 105:10438–10443
An SM, Noh JH, Choi DH, Lee JH, Yang EC (2016) Repeat region absent in mitochondrial genome of tube-dwelling diatom Berkeleya fennica (Naviculales, Bacillariophyceae). Mitochondrial DNAl 27:2137–2138
Andersen RA, Saunders GW, Paskind MP, Sexton JP (1993) Ultrastructure and 18S ribosomal RNA gene sequence for Pelagomonas calceolata gen. Et sp. nov. and the description of a new algal class, the pelagophyceae classis nov. J Phycol 29:701–715
Armbrust EV et al (2004) The genome of the diatom Thalassiosira pseudonana: ecology, evolution, and metabolism. Science 306:79–86
Ashworth MP, Ruck EC, Lobban CS, Romanovicz DK, Theriot EC (2012) A revision of the genus Cyclophora and description of Astrosyne gen. Nov (Bacillariophyta), two genera with the pyrenoids contained within pseudosepta. Phycologia 51:684–699
Basu S et al (2017) Finding a partner in the ocean: molecular and evolutionary bases of the response to sexual cues in a planktonic diatom. New Phytol 215:140–156
Bečková M et al (2017) Association of Psb28 and Psb27 proteins with PSII-PSI supercomplexes upon exposure of Synechocystis sp. PCC 6803 to high light. Mol Plant 10:62–72
Bhattacharya D et al (2013) Genome of the red alga Porphyridium purpureum. Nat Commun 4:1941
Bowler C et al (2008) The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 456:239–244
Brawley SH et al (2017) Insights into the red algae and eukaryotic evolution from the genome of Porphyra umbilicalis (Bangiophyceae, Rhodophyta). Proc Natl Acad Sci U S A 114:6361–6370
Brown JW, Sorhannus U (2010) A molecular genetic timescale for the diversification of autotrophic stramenopiles (Ochrophyta): substantive underestimation of putative fossil ages. PLoS One 6:12759
Browning TJ et al (2017) Nutrient co-limitation at the boundary of an oceanic gyre. Nature 551:242–246
Brunson JK et al (2018) Biosynthesis of the neurotoxin domoic acid in a bloom-forming diatom. Science 361:1356–1358
Büchel C (2015) Evolution and function of light harvesting proteins. J Plant Physiol 172:62–75
Buck JM et al (2019) Lhcx proteins provide photoprotection via thermal dissipation of absorbed light in the diatom Phaeodactylum tricornutum. Nat Commun 10:4167
Burki F et al (2007) Phylogenomics reshuffles the eukaryotic supergroups. PLoS One 2:000790
Burki F et al (2012) Re-evaluating the green versus red signal in eukaryotes with secondary plastid of red algal origin. Genom Biol Evol 4:626–635
Burki F et al (2014) Endosymbiotic gene transfer in tertiary plastid-containing dinoflagellates. Eukaryot Cell 13:00299–00213
Burki F et al (2016) Untangling the early diversification of eukaryotes: a phylogenomic study of the evolutionary origins of Centrohelida, Haptophyta and Cryptista. Proc Biol Sci 283:20152802
Caputi L et al (2019) Community-level responses to iron availability in open ocean planktonic ecosystems. Glob Biogeochem Cycles 10:1029
Carradec Q et al (2018) A global ocean atlas of eukaryotic genes. Nat Commun 9:373
Cattolico RA et al (2008) Chloroplast genome sequencing analysis of Heterosigma akashiwo CCMP452 (West Atlantic) and NIES293 (West Pacific) strains. BMC Genomics 9:211
Cavalier-Smith T (1998) A revised six-kingdom system of life. Biol Rev 73:203–266
Cavalier-Smith T (1999) Principles of protein and lipid targeting in secondary symbiogenesis: euglenoid, dinoflagellate, and sporozoan plastid origins and the eukaryote family tree. J Euk Microbiol 46:347–366
Coesel S, Obornik M, Varela J, Falciatore A, Bowler C (2008) Evolutionary origins and functions of the carotenoid biosynthetic pathway in marine diatoms. PLoS One 3:2896
Collén J et al (2013) Genome structure and metabolic features in the red seaweed Chondrus crispus shed light on evolution of the Archaeplastida. Proc Natl Acad Sci U S A 110:5247–5252
Crowell RM, Nienow JA, Cahoon AB (2019) The complete chloroplast and mitochondrial genomes of the diatom Nitzschia palea (Bacillariophyceae) demonstrate high sequence similarity to the endosymbiont organelles of the dinotom Durinskia baltica. J Phycol 55:352–364
Dagan T, Martin W (2009) Seeing green and red in diatom genomes. Science 324:1651–1652
Dautermann O, Lohr M (2017) A functional zeaxanthin epoxidase from red algae shedding light on the evolution of light-harvesting carotenoids and the xanthophyll cycle in photosynthetic eukaryotes. Plant J 92:879–891
Derelle R, López-García P, Timpano H, Moreira D (2016) A phylogenomic framework to study the diversity and evolution of stramenopiles (heterokonts). Mol Biol Evol 33:2890–2898
Deschamps P, Moreira D (2012) Re-evaluating the green contribution to diatom genomes. Genome Biol Evol 4:683–688
Dorrell RG, Bowler C (2017) Secondary plastids of stramenopiles. In: Hirakawa Y (ed) Adv. Bot. Res.: secondary endosymbiosis, vol 84. Elsevier, pp 57–103
Dorrell RG, Howe CJ (2012) What makes a chloroplast? Reconstructing the establishment of photosynthetic symbioses. J Cell Sci 125:1865–1875
Dorrell RG, Smith AG (2011) Do red and green make brown?: perspectives on plastid acquisitions within chromalveolates. Eukaryot Cell 10:856–868
Dorrell RG et al (2017a) Chimeric origins of ochrophytes and haptophytes revealed through an ancient plastid proteome. elife 6:23717
Dorrell RG et al (2017b) Progressive and biased divergent evolution underpins the origin and diversification of peridinin dinoflagellate plastids. Mol Biol Evol 34:361–379
Dorrell RG et al (2019) Principles of plastid reductive evolution illuminated by nonphotosynthetic chrysophytes. Proc Natl Acad Sci U S A 116:6914–6923
Dorrell RG et al (2021) Phylogenomic fingerprinting of tempo and functions of horizontal gene transfer within ochrophytes. Proc Natl Acad Sci U S A 118:2009974118
Ehara M, Inagaki Y, Watanabe KI, Ohama T (2000) Phylogenetic analysis of diatom coxI genes and implications of a fluctuating GC content on mitochondrial genetic code evolution. Curr Genet 37:29–33
Elias M, Patron NJ, Keeling PJ (2009) The RAB family GTPase Rab1A from plasmodium falciparum defines a unique paralog shared by chromalveolates and Rhizaria. J Eukaryot Microbiol 56:348–356
Eme L, Gentekaki E, Curtis B, Archibald JM, Roger AJ (2017) Lateral gene transfer in the adaptation of the anaerobic parasite Blastocystis to the gut. Curr Biol 27:807–820
Fan X et al (2020) Phytoplankton pangenome reveals extensive prokaryotic horizontal gene transfer of diverse functions. Sci Adv 6:0011
Fork DC, Herbert SK, Malkin S (1991) Light energy distribution in the brown alga Macrocystis pyrifera (giant kelp). Plant Physiol 95:731–739
Frommolt R et al (2008) Ancient recruitment by chromists of green algal genes encoding enzymes for carotenoid biosynthesis. Mol Biol Evol 25:2653–2667
Gao X, Bowler C, Kazamia E (2021) Iron metabolism strategies in diatoms. J Exp Bot 72:2165–2180
Gao L, Zhou Y, Wang ZW, Su YJ, Wang T (2011) Evolution of the rpoB-psbZ region in fern plastid genomes: notable structural rearrangements and highly variable intergenic spacers. BMC Plant Biol 11:64
Gile GH, Moog D, Slamovits CH, Maier UG, Archibald JM (2015) Dual organellar targeting of aminoacyl-tRNA synthetases in diatoms and cryptophytes. Genome Biol Evol 7:1728–1742
Graupner N et al (2018) Evolution of heterotrophy in chrysophytes as reflected by comparative transcriptomics. FEMS Microbiol Ecol 94:fiy039
Guillory WX et al (2018) Recurrent loss, horizontal transfer, and the obscure origins of mitochondrial introns in diatoms (Bacillariophyta). Genom Biol Evol 10:1504–1515
Hadariová L, Vesteg M, Hampl V, Krajčovič J (2018) Reductive evolution of chloroplasts in non-photosynthetic plants, algae and protists. Curr Genet 64:365–387
Hamsher SE et al (2019) Extensive chloroplast genome rearrangement amongst three closely related Halamphora spp. (Bacillariophyceae), and evidence for rapid evolution as compared to land plants. PLoS One 14:e0217824
Han KY et al (2019) Dictyochophyceae plastid genomes reveal unusual variability in their organization. J Phycol 55:1166–1180
Hehenberger E, Burki F, Kolisko M, Keeling PJ (2016) Functional relationship between a dinoflagellate host and its diatom endosymbiont. Mol Biol Evol 33:2376–2390
Hendry KR et al (2018) Competition between silicifiers and non-silicifiers in the past and present ocean and its evolutionary impacts. Front Mar Sci 5:22
Hildebrand M et al (1991) Plasmids in diatom species. J Bacteriol 173:5924–5927
Ichinomiya M, Lopes dos Santos A, Gourvil P, Yoshikawa S, Kamiya M, Ohki K, Audic S, de Vargas C, Noël MH, Vaulot D, Kuwata A (2016) Diversity and oceanic distribution of the Parmales (Bolidophyceae), a picoplanktonic group closely related to diatoms. ISME J 10:2419–2436
Imanian B, Pombert JF, Dorrell RG, Burki F, Keeling PJ (2012) Tertiary endosymbiosis in two dinotoms has generated little change in the mitochondrial genomes of their dinoflagellate hosts and diatom endosymbionts. PLoS One 7:e43763
Imanian B, Pombert JF, Keeling PJ (2010) The complete plastid genomes of the two 'dinotoms' Durinskia baltica and Kryptoperidinium foliaceum. PLoS One 5:107111
Ishida K, Cavalier-Smith T, Green BR (2000) Endomembrane structure and the chloroplast protein targeting pathway in Heterosigma akashiwo (Raphidophyceae, Chromista). J Phycol 36:1135–1144
Jacobs JD et al (1992) Characterization of two circular plasmids from the marine diatom Cylindrotheca fusiformis: plasmids hybridize to chloroplast and nuclear DNA. Mol Gen Genet 233:302–310
Janouskovec J, Horák A, Oborník M, Lukes J, Keeling PJ (2010) A common red algal origin of the apicomplexan, dinoflagellate, and heterokont plastids. Proc Natl Acad Sci U S A 107:10949–10954
Janouškovec J et al (2017) A new lineage of eukaryotes illuminates early mitochondrial genome reduction. Curr Biol 27:3717–3724.e3715
Jiroutova K, Koreny L, Bowler C, Obornik M (2010) A gene in the process of endosymbiotic transfer. PLoS One 5:013234
Jirsová D, Füssy Z, Richtová J, Gruber A, Oborník M (2019) Morphology, ultrastructure, and mitochondrial genome of the marine non-photosynthetic bicosoecid Cafileria marina gen. Et sp. nov. Microorganisms 7:240
Kamikawa R, Azuma T, Ishii KI, Matsuno Y, Miyashita H (2018) Diversity of organellar genomes in non-photosynthetic diatoms. Protist 169:351–361
Kamikawa R et al (2015a) Multiple losses of photosynthesis in Nitzschia (Bacillariophyceae). Phycol Res 63:19–28
Kamikawa R et al (2015b) Proposal of a twin arginine translocator system-mediated constraint against loss of ATP synthase genes from nonphotosynthetic plastid genomes. Mol Biol Evol 32:2598–2604
Kamikawa R et al (2017) A non-photosynthetic diatom reveals early steps of reductive evolution in plastids. Mol Biol Evol 34:2355–2366
Kayama M et al (2020) Highly reduced plastid genomes of the non-photosynthetic dictyochophyceans Pteridomonas spp. (Ochrophyta, SAR) are retained for tRNA-Glu-based organellar heme biosynthesis. Front Plant Sci 11:602455
Kazamia E et al (2018) Endocytosis-mediated siderophore uptake as a strategy for Fe acquisition in diatoms. Sci Adv 4:4536
Keeling PJ, Palmer JD (2008) Horizontal gene transfer in eukaryotic evolution. Nat Rev Genet 9:605–618
Kim JI et al (2017) Evolutionary dynamics of cryptophyte plastid genomes. Genome Biol Evol 9:1859–1872
Klinger CM et al (2018) Plastid transcript editing across dinoflagellate lineages shows lineage-specific application but conserved trends. Genom Biol Evol 10:1019–1038
Kowallik KV, Stoebe B, Schaffran I, Kroth-Pancic P, Freier U (1995) The chloroplast genome of a chlorophyll a+c-containing alga, Odontella sinensis. Plant Mol Biol Reporter 13:336–342
Krasovec M, Sanchez-Brosseau S, Piganeau G (2019) First estimation of the spontaneous mutation rate in diatoms. Genom Biol Evol 11:1829–1837
Kretschmann J, Žerdoner Čalasan A, Gottschling M (2018) Molecular phylogenetics of dinophytes harboring diatoms as endosymbionts (Kryptoperidiniaceae, Peridiniales), with evolutionary interpretations and a focus on the identity of Durinskia oculata from Prague. Mol Phylogenet Evol 118:392–402
Kuczynska P, Jemiola-Rzeminska M, Strzalka K (2015) Photosynthetic pigments in diatoms. Mar Drugs 13:5847–5881
Larkum AWD, Lockhart PJ, Howe CJ (2007) Shopping for plastids. Trends Plant Sci 12:189–195
Lax G et al (2018) Hemimastigophora is a novel supra-kingdom-level lineage of eukaryotes. Nature 564:410–414
Le Corguillé G et al (2009) Plastid genomes of two brown algae, Ectocarpus siliculosus and Fucus vesiculosus: further insights on the evolution of red-algal derived plastids. BMC Evol Biol 9:253
Leliaert F et al (2016) Chloroplast phylogenomic analyses reveal the deepest-branching lineage of the chlorophyta, Palmophyllophyceae class nov. Sci Rep 6:25367
Leonard G et al (2018) Comparative genomic analysis of the 'pseudofungus' Hyphochytrium catenoides. Open Biol 8:170184
Levesque CA et al (2010) Genome sequence of the necrotrophic plant pathogen Pythium ultimum reveals original pathogenicity mechanisms and effector repertoire. Genome Biol 11:73
Li L et al (2020) The genome of Prasinoderma coloniale unveils the existence of a third phylum within green plants. Nat Ecol Evol 4:1220–1231
Lommer M et al (2010) Recent transfer of an iron-regulated gene from the plastid to the nuclear genome in an oceanic diatom adapted to chronic iron limitation. BMC Genomics 11:718
Magee AM et al (2010) Localized hypermutation and associated gene losses in legume chloroplast genomes. Genome Res 20:1700–1710
Matsuzaki M et al (2004) Genome sequence of the ultrasmall unicellular red alga Cyanidioschyzon merolae 10D. Nature 428:653–657
McQuaid JB et al (2018) Carbonate-sensitive phytotransferrin controls high-affinity iron uptake in diatoms. Nature 555:534–537
Méheust R, Zelzion E, Bhattacharya D, Lopez P, Bapteste E (2016) Protein networks identify novel symbiogenetic genes resulting from plastid endosymbiosis. Proc Natl Acad Sci U S A 113:3579–3584
Mock T et al (2017) Evolutionary genomics of the cold-adapted diatom Fragilariopsis cylindrus. Nature 541:536–540
Moll KM et al (2017) Strategies for optimizing BioNano and dovetail explored through a second reference quality assembly for the legume model, Medicago trunculata. BMC Genom 18:578
Morozov AA, Galachyants YP (2019) Diatom genes originating from red and green algae: implications for the secondary endosymbiosis models. Mar Genomics 45:72–78
Moustafa A et al (2009) Genomic footprints of a cryptic plastid endosymbiosis in diatoms. Science 324:1724–1726
Muñoz-Gómez SAM-F, Durnin FG, Colp K, Grisdale M, Archibald CJ, Slamovits JM (2017) The new red algal subphylum proteorhodophytina comprises the largest and most divergent plastid genomes known. Curr Biol 27:1677–1684
Nash EA et al (2007) Organization of the mitochondrial genome in the dinoflagellate Amphidinium carterae. Mol Biol Evol 24:1528–1536
Nonoyama T et al (2019) Metabolic innovations underpinning the origin and diversification of the diatom chloroplast. Biomol Ther 9:322
Noordally ZB et al (2013) Circadian control of chloroplast transcription by a nuclear-encoded timing signal. Science 339:1316–1319
Novák Vanclová AMG et al (2020) Metabolic quirks and the colourful history of the Euglena gracilis secondary plastid. New Phytol 225:1578–1592
Ong HC et al (2010) Analyses of the commplete chloroplast genome sequences of two members of the Pelagophyceae: Aureococcus anophageferrens CCMP1984 and Aureooumbra lagunensis CCMP 1507. J Phycol 46:602–615
Onyshchenko A, Ruck EC, Nakov T, Alverson AJ (2019) A single loss of photosynthesis in the diatom order Bacillariales (Bacillariophyta). Am J Bot 106:560–572
Oudot-Le Secq MP, Green BR (2011) Complex repeat structures and novel features in the mitochondrial genomes of the diatoms Phaeodactylum tricornutum and Thalassiosira pseudonana. Gene 476:20–26
Parfrey LW, Lahr DJ, Knoll AH, Katz LA (2011) Estimating the timing of early eukaryotic diversification with multigene molecular clocks. Proc Natl Acad Sci U S A 108:13624–13629
Parks MB, Wickett NJ, Alverson AJ (2018) Signal, uncertainty, and conflict in phylogenomic data for a diverse lineage of microbial eukaryotes (diatoms, Bacillariophyta). Mol Biol Evol 35:80–93
Pendergrass A, Roberts WR, Ruck EC, Lewis JA, Alverson AJ (2020) The genome of the nonphotosynthetic diatom, Nitzschia sp.: insights into the metabolic shift to heterotrophy and the rarity of loss of photosynthesis in diatoms. bioRXiv 05.28.115543
Ponce-Toledo RI, López-García P, Moreira D (2019) Horizontal and endosymbiotic gene transfer in early plastid evolution. New Phytol 224:618–624
Prasetiya F et al (2019) Haslea nusantara (Bacillariophyceae), a new blue diatom from the Java Sea, Indonesia: morphology, biometry and molecular characterization. Plant Ecol Evol 152(15)
Qiu H, Lee JM, Yoon HS, Bhattacharya D (2017) Hypothesis: gene-rich plastid genomes in red algae may be an outcome of nuclear genome reduction. J Phycol 53:715–719
Rastogi A et al (2018) Integrative analysis of large scale transcriptome data draws a comprehensive landscape of Phaeodactylum tricornutum genome and evolutionary origin of diatoms. Sci Rep 8:4834
Ravin NV et al (2010) Complete sequence of the mitochondrial genome of a diatom alga Synedra acus and comparative analysis of diatom mitochondrial genomes. Curr Genet 56:215–223
Raymond JA, Kim HJ (2012) Possible role of horizontal gene transfer in the colonization of sea ice by algae. PLoS One 7:e35968
Ren Y et al (2020) Nucleotide substitution rates of diatom plastid encoded protein genes are correlated with genome architecture. Sci Rep 10:14358
Rhoads A, Au KF (2015) PacBio sequencing and its applications. Genom Proteom Bioinformat 13:278–289
Rodolfi L et al (2009) Microalgae for oil: strain selection, induction of lipid synthesis and outdoor mass cultivation in a low-cost photobioreactor. Biotechnol Bioeng 102:100–112
Ruck EC, Linard SR, Nakov T, Theriot EC, Alverson AJ (2017) Hoarding and horizontal transfer led to an expanded gene and intron repertoire in the plastid genome of the diatom, Toxarium undulatum (Bacillariophyta). Curr Genet 63:499–507
Sabir JSM et al (2014) Conserved gene order and expanded inverted repeats characterize plastid genomes of Thalassiosirales. PLoS One 9:107854
Sato S et al (2020) Genome-enabled phylogenetic and functional reconstruction of an araphid pennate diatom Plagiostriata sp. CCMP470, previously assigned as a radial centric diatom, and its bacterial commensal. Sci Rep 10:9449
Sekiguchi H, Moriya M, Nakayama T, Inouye I (2002) Vestigial chloroplasts in heterotrophic stramenopiles Pteridomonas danica and Ciliophrys infusionum (Dictyochophyceae). Protist 153:157–167
Ševčíková T et al (2015) Updating algal evolutionary relationships through plastid genome sequencing: did alveolate plastids emerge through endosymbiosis of an ochrophyte? Sci Rep 5:10134
Ševčíková T et al (2016) A comparative analysis of mitochondrial genomes in eustigmatophyte algae. Genom Biol Evol 8:705–722
Ševcíková T et al (2019) Plastid genomes and proteins illuminate the evolution of eustigmatophyte algae and their bacterial endosymbionts. Genom Biol Evol 11:362–379
Sharma AK, Nymark M, Sparstad T, Bones AM, Winge P (2018) Transgene-free genome editing in marine algae by bacterial conjugation - comparison with biolistic CRISPR/Cas9 transformation. Sci Rep 8:14401
Sorhannus U (2011) Evolution of antifreeze protein genes in the diatom genus fragilariopsis: evidence for horizontal gene transfer, gene duplication and episodic diversifying selection. Evol Bioinformatics Online 7:279–289
Stiller JW, Huang JL, Ding Q, Tian J, Goodwillie C (2009) Are algal genes in nonphotosynthetic protists evidence of historical plastid endosymbioses? BMC Genomics 10:484
Stiller JW et al (2014) The evolution of photosynthesis in chromist algae through serial endosymbioses. Nat Commun 5:5764
Stoebe B, Martin W, Kowallik K (1998) Distribution and nomenclature of protein-coding genes in 12 sequenced chloroplast genomes. Plant Mol Biol Rep 16:13
Sturm S et al (2013) A novel type of light-harvesting antenna protein of red algal origin in algae with secondary plastids. BMC Evol Biol 13:159
Swenson NG (2009) Phylogenetic resolution and quantifying the phylogenetic diversity and dispersion of communities. PLoS One 4:e4390
Tabita FR, Satagopan S, Hanson TE, Kreel NE, Scott SS (2008) Distinct form I, II, III, and IV rubisco proteins from the three kingdoms of life provide clues about rubisco evolution and structure/function relationships. J Exp Bot 59:1515–1524
Tajima N et al (2016) Sequencing and analysis of the complete organellar genomes of Parmales, a closely related group to Bacillariophyta (diatoms). Curr Genet 62:887–896
Tsui CKM et al (2009) Labyrinthulomycetes phylogeny and its implications for the evolutionary loss of chloroplasts and gain of ectoplasmic gliding. Mol Phylogenet Evol 50:129–140
Vancaester E, Depuydt T, Osuna-Cruz CM, Vandepoele K (2020) Comprehensive and functional analysis of horizontal gene transfer events in diatoms. Mol Biol Evol 37:3243–3257
Villanova V et al (2017) Investigating mixotrophic metabolism in the model diatom Phaeodactylum tricornutum. Philos Trans R Soc B 372:20160404
Walker G, Dorrell RG, Schlacht A, Dacks JB (2011) Eukaryotic systematics: a user's guide for cell biologists and parasitologists. Parasitology 138:1638–1663
Wang Q, Sun H, Huang J (2017) Re-analyses of “algal” genes suggest a complex evolutionary history of oomycetes. Front Plant Sci 8:1540
Wetherbee R et al (2019) The golden paradox - a new heterokont lineage with chloroplasts surrounded by two membranes. J Phycol 55:257–278
Yamada N, Sakai H, Onuma R, Kroth PG, Horiguchi T (2020) Five non-motile dinotom dinoflagellates of the genus Dinothrix. Front Plant Sci 11:591050
Yamada N, Sym SD, Horiguchi T (2017) Identification of highly divergent diatom-derived chloroplasts in dinoflagellates, including a description of Durinskia kwazulunatalensis sp. nov. (Peridiniales, Dinophyceae). Mol Biol Evol 34:1335–1351
Yamada N et al (2019) Discovery of a kleptoplastic 'dinotom' dinoflagellate and the unique nuclear dynamics of converting kleptoplastids to permanent plastids. Sci Rep 9:10474
Yang M, Lin X, Liu X, Zhang J, Ge F (2018) Genome annotation of a model diatom Phaeodactylum tricornutum using an integrated proteogenomic pipeline. Mol Plant 11:1292–1307
Yu M et al (2018) Evolution of the plastid genomes in diatoms. In: Chaw S, Jansen R (eds) Adv. Bot. Res. Plastid genome evolution, vol 85. Elsevier, pp 129–155
Acknowledgements
RGD acknowledges a CNRS Momentum Fellowship (awarded 2019-2021) and an ANR JCJC (“PanArctica”, grant number ANR-21-CE02-0014-01, awarded 2022-2025).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Dorrell, R.G., Liu, F., Bowler, C. (2022). Reconstructing Dynamic Evolutionary Events in Diatom Nuclear and Organelle Genomes. In: Falciatore, A., Mock, T. (eds) The Molecular Life of Diatoms. Springer, Cham. https://doi.org/10.1007/978-3-030-92499-7_6
Download citation
DOI: https://doi.org/10.1007/978-3-030-92499-7_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-92498-0
Online ISBN: 978-3-030-92499-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)