Abstract
Mitochondria control a myriad of intracellular processes including ATP synthesis, redox balance, ion homeostasis and metabolism of amino acids and lipids. Maintaining a healthy and demand-matched pool of mitochondria is critical for supporting the immune system. Changes in mitochondrial mass, size, number, morphology, connectiveness and distribution occur in a dynamic process mainly driven by oscillations in energy demand and supply in health and disease. Therefore, disruption of mitochondrial biogenesis and dynamics likely results in mitochondrial dysfunction-associated diseases. This chapter reviews the molecular mechanisms that regulate mitochondrial content, number and morphology. It also highlights the clinical implications of defective mitochondrial biogenesis and dynamics.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Introduction
Mitochondria are double-membrane-bound organelles exclusively found in eukaryotic cells and best known for its role in the generation of adenosine triphosphate (ATP) [1]. The endosymbiotic hypothesis proposes that mitochondria arise from the integration of a free-living aerobic bacterium into a host cell over a billion of years ago. In this relationship, the host cell provided a safe and nutrient-rich environment for the aerobic bacterium. It also acquired a new source of oxygen dependent-energy [2]. More recently, it has been suggested that the phagocytosed bacterium may have provided defense molecules for the host cell, also connecting the advantages of this endosymbiotic relationship to immunity [3]. Throughout evolution, a massive transfer of genes to the host cell allowed the evolvement of the endosymbiotic bacterium as a permanent organelle—the mitochondrion (mitochondria for plural) [4].
Derived from two Greek words: “mitos”—thread and “chondros”—granule, the organelle displays two lipid bilayer-membranes enclosing two aqueous compartments. The outer mitochondrial membrane surrounds the intermembrane space, while the inner mitochondrial membrane, which contains invaginations denominated cristae, encloses the matrix compartment [5, 6]. The inner membrane accommodates the oxidative phosphorylation system (OXPHOS)—a five multimeric protein complexes (Complex I–V) that uses redox reactions to generates ATP. At the expense of oxygen as a final electron acceptor, a sequential transfer of electrons from Complex I to Complex IV generates a proton electrochemical gradient across the inner membrane, also known as membrane potential, that is used by Complex V to drive ATP synthesis [1]. The mitochondrial membrane potential also helps metabolites transport and ion homeostasis.
Often referred as the powerhouse of the eukaryotic cells, the role of mitochondria goes beyond ATP production. The organelle orchestrates a myriad of other processes including reactive oxygen species (ROS) formation [7, 8], aldehyde metabolism [9], heat production [10], ion homeostasis [11] and programmed cell death [12] that ultimately dictate cell fate. This functional versatility is intimately linked to the content, size and number of mitochondria. Their morphological complexity is controlled by the processes termed mitochondrial biogenesis and dynamics. While mitochondrial biogenesis increases the number and content of the organelles in a coordinated effort with the nucleus [13], mitochondrial dynamics drives the formation of larger or smaller organelles through the antagonist activities of fusion and fission [14].
Mitochondria are recently recognized by their dynamic nature. In order to meet the cellular requirements for ATP, the mitochondrial network are under constantly remodeling. Indeed, metabolic cues (i.e. starvation, exercise) trigger not only fusion and fission machineries in order to create elongated or fragmented mitochondria [15], but also drive transcription factors activation to increase mitochondrial mass and boost oxidative metabolism [16]. Moreover, this dynamism helps impaired mitochondria to be rescued or eliminated. In the first case, fusion events allow damaged components to be diluted throughout the network, thereby avoiding the propagation of stress that might cause mitochondrial dysfunction or collapse [17]. On the opposite way, the fission process segregates part of dysfunctional mitochondria that now can be addressed for degradation in the lysosome, a process termed mitophagy [18].
Exciting new findings have revealed mitochondria as a major intracellular signaling platform regulating immune cell function. Indeed, the cellular metabolic plasticity provided by mitochondria not only allows immune cells to grow, but it is also required during transition from a metabolically quiescent stage to a highly active state [19]. Moreover, proteins located on the outer mitochondrial membrane, as well as mitochondrial DNA can dictate immune cell activation [20, 21]. Finally, due to the reciprocal crosstalk between mitochondrial metabolism and morphology, fluctuations in shape, size and position of the organelle within the cell likey affect both phenotype and activity of immune cells [22].
Considering the extensive knowledge highlighting mitochondria as the powerhouse of the cells as well as emerging evidence placing mitochondria at the heart of immunity, this chapter reviews the general processes regulating mitochondrial biogenesis and dynamics, and discuss the critical role of these processes in health and disease.
2 Mitochondrial Biogenesis
Mitochondrial biogenesis is a simplified term used to describe a complex process involving the increase in mass of pre-existing mitochondria. Due to their bacterial origin, mitochondria possess their own genetic material, which includes DNA and the translational/transcriptional system. The mitochondrial DNA (mtDNA) is a circular double strand DNA molecule containing ~16.5 kb that encodes only 37 genes: 22 transfer RNA and 2 ribosomal RNA (12S and 16S) required for translating 13 messenger RNA. Moreover, maternal inheritance, lack of introns (non-coding sections of a gene) and several copies per cell (1–10 copies per mitochondrion) are among the unique features that differ mtDNA from the nuclear DNA [23].
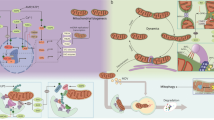
The entire protein-coding capacity of mtDNA relies on 13 essential subunits of the electron transport chain (ETC) that are replicated and transcribed within the mitochondrial matrix: 7 subunits of NADH: Ubiquinone oxidoreductase (Complex I), 1 subunit of Ubiquinone: Cytochrome c oxidoreductase (Complex III), 3 subunits of Cytochrome c Oxidase (Complex IV) and 2 subunits of ATP synthase (Complex V) [24]. The ~1100 remaining mitochondrial proteins [25, 26] have to be transcribed in the nucleus, translated in cytosolic ribosomes and imported into the organelle (Fig. 1). Therefore, mitochondrial biogenesis faces several challenges before promoting an increase in the mitochondrial content.
Summary of the transcriptional regulation of mitochondrial biogenesis. The expression of mitochondrial genes encoded by both nDNA (nuclear DNA) and mtDNA (mitochondrial DNA) is mainly regulated by a family of transcriptional coactivators named PGC-1 [peroxisome proliferator-activated receptor (PPAR) gamma coactivator 1]. PGC-1 members bind to and coactivate NRFs (nuclear respiratory factors) to induce the expression of multiple components of the OXPHOS (oxidative phosphorylation system), ETC (electron transport chain) and mtDNA replication. NRFs also regulate the levels of TFAM and TFB (mitochondrial transcription factors A and B, respectively) involved in the expression of genes encoded by the mtDNA. Interaction between PGC-1 and specific transcription factors such as PPARs and EERs (estrogen-related receptors) control the expression of many genes involved in FAO (fatty acid oxidation), TCA cycle (tricarboxylic acid cycle), glucose and lipid metabolism, and detoxifying enzymes. Nuclear-encoded mitochondrial proteins are translated in cytosolic ribosomes and imported into the organelle
The first challenge relies on coordinating the gene expression between two genomes located into distinct subcellular compartments. Indeed, to ensure a proper OXPHOS, the number of ETC subunits must be stoichiometrically balanced [27]. mtDNA occurs in the ratio of ~1000:1 copies relative to nuclear DNA [23]. Second, the majority of mitochondrial proteins are translated in the cytosol; thus, demanding a synchronized cellular machinery to properly target, import and assemble these nuclear-encoded proteins [28, 29]. Failure in addressing these proteins to mitochondria not only impairs ETC subunitse stoichiometry, but also compromises mtDNA replication, which is orchestrated by the nuclear-encoded protein DNA polymerase gamma (POLG) [30]. For a complete description about how mitochondrial genome is replicated, transcribed and translated, please see reviews [28, 31, 32]. Finally, mitochondrial dynamics, which will be discussed above, must also be coordinated.
2.1 Transcription Factors Regulating Mitochondrial Biogenesis
The transcription of both nuclear and mitochondrial genomes is coordinated by specific proteins termed transcription factors. Transcription factors are able to modulate the rate of gene expression by binding to specific regulatory regions of DNA. These proteins contain effector domains that allow the interaction not only with other proteins essential for transcription, including the RNA polymerase, but also with other transcription factors; thereby regulating the amount of messenger RNA produced per gene [33].
Nuclear Respiratory Factors 1 (NRF-1) and 2 (NRF-2) are considered critical players in mitochondrial biogenesis. Together, these transcription factors display DNA-binding sites for most of the genes encoding respiratory subunits. First identified in 1989 as a transcriptional activator of the cytochrome c gene [34], NRF-1 targeted genes are now branded for encoding subunits of all five respiratory complexes of the ETC [35]. A couple of years later, NRF-2 was discovered by its specific binding to the cytochrome oxidase subunit IV promoter [36]. Although often recognized by their power of binding to antioxidant response element (ARE) and promoting gene expression of detoxifying enzymes [37], functional NRF-2 sites have been implicated in the expression of subunits of Complex II, IV and V of the OXPHOS [38].
The regulatory network of NRFs also targets other nuclear genes whose products function in the mitochondria, including components for assembling and importing mitochondrial proteins [39], and constituents of the mtDNA transcription and replication machinery [40]. Indeed, NRF-1 is able to stimulate the expression of mitochondrial transcription factors A (TFAM) [40] and B (TFB) [41]—two nuclear‐encoded transcription factors essential for replication, maintenance, and transcription of mtDNA [42, 43]. Moreover, not only TFAM and TFB have been recently added to the list of genes controlled by NRF-2 [38], but NRF-2 indirectly regulates mitochondrial biogenesis by driving the gene expression of NRF-1 [44]. Due to this essential role in coordinating bi-genomic respiratory subunits, deficiency of NRF can lead to a severe impairment of mitochondrial biogenesis [45, 46].
Members of the nuclear receptor superfamily also control the transcription of respiratory apparatus. The peroxisome proliferator-activated receptor (PPAR) family is composed by three isoforms: PPARα [47], PPAR β/δ [48], PPARγ [49]. The expression of PPAR isoforms differs among tissues and these transcription factors regulate metabolic pathways at different levels. While PPARγ is involved in glucose metabolism and regulation of fatty acid storage, PPARα and PPAR β/δ promote changes in cellular lipid metabolism by upregulating genes involved in mitochondrial fatty acid oxidation [50]. Estrogen-related receptors α (ERRα) and γ (ERRγ) represent another class of nuclear receptors targeting ~700 nuclear-encoded mitochondrial genes. The controlling of these transcription factors, expressed in mitochondrion-enriched tissues such as skeletal muscle and heart [51], is attached to all aspects of energy homeostasis, including mtDNA replication, OXPHOS, ion homeostasis and mitochondrial detoxifying mechanisms (reviewed in [52]). Moreover, ERRα can regulate the levels of PPARα transcripts [53], therefore magnifying the control over mitochondrial fatty acid oxidation pathway.
Finally, a relative small number of other transcription factors have been shown to activate or repress nuclear genes encoding mitochondrial proteins, including stimulatory protein 1 (Sp1), ying yang 1 transcription factor (YY1), cAMP-responsive element-binding protein (CREB) and myocyte enhancer factor 2 (MEF-2), and a detailed consideration of those is covered elsewhere [54].
2.2 The Role of Transcriptional Coactivators in Mitochondrial Biogenesis: PGC-1 Family
As described above, mitochondrial biogenesis requires the coordination of several transcription factors to proper ensure the expression of both nuclear and mitochondrial genes. Adding complexity to this process, mitochondrial metabolism and content differs widely among cells, tissues and organs; thereby demanding an extra layer of regulation. While transcription factors bind to DNA in a sequence-dependent manner, transcriptional coactivators interact with them and amplify the activity of the transcriptional machinery by recruiting multi-protein complexes to modify chromatin folding, interact with the RNA polymerase II complex and process messenger RNA [38]. Although the fundamental mechanisms of how mitochondrial biogenesis is orchestrated are still elusive, a major breakthrough came with the discovery of a family of transcriptional coactivators termed PPARγ coactivator 1 (PGC-1) [55]. PGC-1 proteins have emerged as major players in the transcriptional regulatory circuits controlling mitochondrial biogenesis and function.
Conserved across many species, PGC-1 family is formed by three members that share similar domain structures to interact with nuclear receptors [56]. The first and most studied member of this family is PGC-1α. First identified in brown adipose tissue during adaptive thermogenesis—a process that regulates heat production in response to cold and diet, PGC-1α is considered the master regulator of mitochondrial biogenesis in mammals [55]. Similar to PGC-1α, PGC-1β is predominantly expressed in tissues with abundant mitochondria (e.g. heart and skeletal muscle [57]). However, it is not upregulated upon cold exposure [56]. The third member of this family is the PGC-1 related coactivator (PRC). Despite the relatively low homology with the other two isoforms, PRC is ubiquitously expressed and supports mitochondrial biogenesis during early embryogenesis [58, 59]. Together, they bind to and coactivate most of the transcription factors regulating expression of mitochondrial proteins encoded by the nucleus.
Several studies have shown that PGC-1α is capable of regulating virtually every aspect of mitochondrial content [60]. Indeed, by binding to and coactivating NRF-1 and NRF-2, PGC-1α promotes not only a powerful induction of nuclear-encoded mitochondrial respiratory chain subunits, but also leads to the transcription of the mitochondrial genome through the induction of TFAM [61]. Moreover, the interaction between PGC-1α and transcription factors such as PPARα [62], PPARδ [63], ERRα [64], EERγ [64, 65], thyroid hormone receptor [66] and estrogen receptor [57, 67], controls fat and glucose metabolism. And since PGC-1α and PGC-1β share similar molecular structures and functions, it is not surprising that the mitochondrial gene expression driven by these two coactivators overlaps [68, 69]. Interestingly, their work results in mitochondria with different metabolic features [70]; thereby suggesting that distinct upstream pathways modulate PGC-1α and PGC-1β.
The pioneering work of Puigserver and coworkers first showed in 1998 that PGC-1α is dramatically induced (up to 50-fold) upon cold exposure in brown fat and skeletal muscle [55]. Since then, many studies have determined that the expression of PGC-1 family members are controlled by a variety of external stimuli, such as exercise, cold and nutrient deprivation, in a tissue-dependent manner (reviewed in [71]). Among the transcription factors regulating PGC1-α levels, CREB is responsible for integrating multiple signaling pathways in different cell types to boost mitochondrial function. For example, CREB-dependent induction of PGC1-α occurs in fasted liver [72], in exercised skeletal muscle [73], as well as in brown adipose tissue during cold [55]. Moreover, in a positive autoregulatory loop, PGC-1α regulates its own expression when binding to some of its transcription factors targets such as MEF2 [73] and ERRγ [74]. With equal importance of transcriptional levels, posttranslational modifications of PGC-1α also control mitochondrial biogenesis.
Posttranslational modifications refer to biochemical modifications of a protein (e.g. phosphorylation, acetylation, methylation) capable of influencing not only its structure, but also its activity [75]. The fine-tuning of PGC-1α activity occurs via posttranslational mechanisms. First, phosphorylation of PGC-1α protein is able to triple its half-life, which is relatively short (~2.3 h) [76]. Second, posttranslational modifications interfere with PGC-1α signal transduction by either increasing or inhibiting its activity. In response to bioenergetics imbalance, PGC-1α displays increased activity when phosphorylated by AMP-activated protein kinase (AMPK) [77] and deacetylated by Sirtuin 1 (SIRT1) [78]. On the contrary, PGC-1α can be phosphorylated by glycogen synthase kinase 3β (GSK3β) leading to its inhibition and degradation [79]. Third, most of the signaling pathways conducting these protein modifications have their gene expression regulated by PGC-1 family; thus, reinforcing the feed forward loop [80].
Finally, although the molecular mechanisms are not fully elucidated, it has been demonstrated that posttranslational modifications of PGC-1α result in a preferential induction of biogenesis in a time-, tissue- and subset of mitochondrial genes-dependent manner [78, 81]. This can be explained, at least in part, by the discovery of different splicing variants of PGC-1α: novel truncated PGC-1α (NT-PGC-1α), PGC-1α-β and PGC-1α4. While NT-PGC-1α [82] and PGC-1α-β [83] specifically affect energy metabolism by promoting mitochondrial biogenesis in brown adipose tissue and skeletal muscle, respectively, PGC-1α4 leads to skeletal muscle hypertrophy by regulating a non-mitochondrial gene program [84]. Interestingly, exercise is able to induce and activate all these variants [84, 85].
3 Mitochondrial Dynamics
The high-resolution electron microscopy images of mitochondria, published by Palade [5] and Sjostrand [6] in the 1950s, revealed for the first time the unique ultrastructure of these organelles. Those images also showed a lack of physical connection between mitochondria; thus, suggesting that the organelle was stationary and working independently. Two decades later, descriptions of mega-mitochondria formation in tissues such as liver [86] and skeletal muscle [87] started to question this independency. In the 1990s, advances in electron microscopy along with the development of mitochondrial-targeted fluorescent proteins allowed the observations that mitochondrial can dynamically rearrange their structure over time [88, 89]. Since then, a complete set of genes driving these morphological changes was discovered (reviewed in [90]) and mitochondrial dynamics has been consolidated as a new area of study in mitochondrial biology.
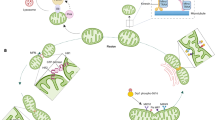
Mitochondrial dynamics refers to a set of processes including the regulation of mitochondrial morphology and connectivity, as well as their position inside the cells. The mitochondrial ability to reshape, rebuild and redistribute itself is orchestrated by the opposite role of fusion and fission processes [90]. Members of a large family of dynamin guanosine triphosphatases (GTPases) use the hydrolysis of guanosine triphosphate (GTP) to create conformational changes in the mitochondrial membrane that will lead to either the union between two organelles or the division of one mitochondrion in two organelles [91] (Fig. 2).
Simplified model for mitochondrial fusion and fission. The OMM (outer mitochondrial membrane) fuses through interaction of homo- or hereto-oligomers Mfn1 (Mitofusin 1) and Mfn2 (Mitofusin 2) of two opposing mitochondria. Following OMM fusion, OPA1 (Optic atrophy 1) drives IMM (inner mitochondrial membrane) fusion. Please note that, as membrane-bound proteins, Mitofusins and OPA1 are still present in the new fused membranes, but are now disassembled. Mitochondrial fragmentation requires activation of cytosolic Drp1 (Dynamin-related protein 1) and recruitment to the organelle via OMM-bound receptors (R). At these sites, the Drp1 oligomerizes in a ring-like structure and constricts the mitochondria into 2 daughters. Of interest, asymmetrical fission of a damaged or senescent mitochondrion produces 1 dysfunctional organelle that can either be eliminated by mitophagy or re-enter the mitochondrial network and regenerate by fusing with other healthy organelles
Despite often viewed as a separate phenomenon, the recycling of mitochondria through mitophagy—a specific form of autophagy, is influenced by mitochondrial fission and therefore directly interferes with the dynamic nature of the organelle. To a detailed description of mitophagy, readers are referred to excellent reviews on this topic [92, 93]. Together, mitochondrial fusion-fission machinery, mitochondrial biogenesis and mitophagy comprise a well-conserved quality control axis capable of controlling the function of the organelle, and as consequence, interfering with cellular physiology [94].
3.1 Mitochondrial Fusion
Mitochondrial fusion is an evolutionary conserved process that merges two neighboring mitochondria. By allowing the exchange of mitochondrial proteins, metabolites and mtDNA, mitochondrial fusion maximizes cellular respiration [95]. It is also required for the maintenance of mtDNA integrity [96]. Moreover, the newly elongated fused organelle prevents erroneous degradation of mitochondria [17]. Considering that mitochondria have outer and inner membranes, the fusion process requires bringing together four membranes in separated events. First, mitofusin 1 (Mfn1) and mitofusin 2 (Mfn2) proteins are responsible for fusing the outer mitochondrial membrane. Later, optic atrophy factor 1 (OPA1) governs the union of the inner mitochondrial membrane [97]. As nuclear-encoded mitochondrial proteins, they need to be synthesized in the cytosol and imported into mitochondria, therefore reinforcing the connection between mitochondrial biogenesis and dynamics. Additionally, these GTPases contain a transmembrane domain that anchors part of them to the lipid bilayer, whereas their free part can physically interact with other GTPases to promote the tethering [98].
Mfn1 and Mfn2 are the major players in promoting outer membrane shape-changes. These isoforms display high homology (~80%) and initiate the fusion of the outer mitochondrial membrane by the formation of homo- (Mfn1-Mfn1 or Mfn2-Mfn2) or hetero-oligomers (Mfn1-Mfn2) between adjacent organelles [99]. Despite widely expressed and essential for embryonic development [100], Mfn1 is more abundant in heart and liver, while Mfn2 predominates in skeletal muscle, brain and adipose tissue [101]. Interestingly, each mitofusin not only differently affects mitochondrial morphology, but also plays distinct roles in cellular physiology. The absence of Mfn1 leads to highly fragmented mitochondria when compared to Mfn2 downregulation [100]. Moreover, Mfn2 also participates in calcium regulation by tethering the mitochondria to the endoplasmic reticulum [102, 103].
In order to complete the fusion process, OPA1 drives the unification of the two inner mitochondrial membranes. This intermembrane space-localized GTPase not only suffers alternative splicing—a mechanism by which different messenger RNA are generated from the same gene, but its activity is also regulated by proteolytic processing [104]. Because of that, there are at least eight variants of OPA1 in humans containing one or two proteolytic sites [105]. The mitochondrial proteases OMA1 and YME1L1 are responsible for generating long and short OPA1 isoforms, in a membrane potential dependent manner [106, 107]. Despite the fact that both isoforms are required for full fusion events, an excessive processing of short OPA1 limits fusion; therefore, triggering mitochondrial fragmentation [108,109,110]. Finally, regardless governing the delicate balance between fusion and fission, OPA1 variants are able to control apoptosis by regulating the cristae morphology and consequent release of cytochrome c—a component of ETC that triggers programed cell death [109, 111].
3.2 Mitochondrial Fission
Mitochondrial fission process is responsible for the asymmetrical segregation of portions of the organelle. Whereas this new spherical and smaller organelle facilitates motility throughout the cell, it is also involved in mtDNA replication and inheritance during cellular proliferation [112]. Fragmentation of the mitochondrial network also permits the selective removal of damaged organelles by mitophagy [18]. Unlike mitochondrial fusion, the division of the outer and inner membranes of the organelle is catalyzed by a single GTPase effector—dynamin-related protein 1 (Drp1) [113]. Unlike mitochondrial fusion-related proteins, Drp1 is a nuclear-encoded protein that resides in the cytosol as a small oligomer, thereby demanding recruitment to the mitochondrial surface. The assembly of several Drp1 oligomers forms a ring-like structure around the outer mitochondrial membrane and cut mitochondria into two separate entities in a GTP hydrolysis-dependent manner [114].
A multi-step process is required before completing mitochondrial membrane remodeling. First, Drp1 needs to be activated in order to translocate from the cytosol to mitochondria. Among the posttranslational modifications regulating Drp1 activity, phosphorylation has been extensively studied and serves as an efficient way to synchronize intracellular signaling pathways and mitochondrial metabolism. For example, protein kinase A (PKA) phosphorylation of Drp1 at serine-637 blocks fission and protects mitochondrial from degradation during starvation [115]. Dephosphorylation of the same residue by the phosphatase calcineurin triggers fission in a calcium-induced mitochondrial dysfunction environment [116, 117]. Ubiquitination of Drp1 by E3 ligases can either induce mitochondrial fragmentation or inhibit fission by promoting Drp1 degradation [118, 119]. Additional posttranslational modifications of Drp1 (e.g. SUMOylation and S-nitrosylation) also dictate mitochondrial dynamics. These regulatory mechanisms are reviewed elsewhere [120, 121].
Once activated, the second step involves the recruitment of Drp1 to specific regions of the outer mitochondrial membrane. Four specific adaptor proteins, also termed Drp1 receptors, facilitate this anchoring process: mitochondrial fission 1 protein (Fis1), mitochondrial fission factor (Mff) and mitochondrial dynamics proteins of 49 and 51 kDa (MiD49 and MiD51, respectively) [113, 122, 123]. This receptor-mediated recruitment of Drp1 assists mitochondrial fragmentation by allowing the self-assemble of Drp1 into oligomeric complexes at specific sites of the outer membrane pre-constricted by the endoplasmic reticulum [124, 125]. Similar to Drp1, these receptors can be activated by posttranslational modifications. In particular, MFF can be phosphorylated by AMPK in response to nutrient excess, favoring mitochondrial fission [126, 127]. Finally, recent evidence place another GTPase—dynamin 2 (Dyn2), as a mechanoenzyme involved in terminating membrane scission. It has been proposed that Drp1-mediated constriction allows Dyn2 assembly to complete the fission event [128].
4 Mitochondrial Dynamics in Health and Disease
Given the fact that mitochondrial biogenesis and dynamics interfere with a variety of intracellular processes including ATP production, ROS release and apoptosis, it is not surprising that they are critical in the context of both physiological and pathological events [129]. Disruption of mitochondrial homeostasis follows the clinical progression of a variety of chronic degenerative diseases (e.g. Heart Failure and Diabetes) [130]. Moreover, mitochondrial dysfunction is a common feature of rare inherited mitochondrial diseases, which are driven by mutations in either nuclear or mitochondrial DNA (e.g. Leigh syndrome and Friedreich’s ataxia) [131]. Either way, the inability of maintaining a healthy mitochondrial population has been placed as a central determinant of several diseases. Here, we discuss mitochondrial biogenesis and dynamics in the context of cardiac, metabolic and neurodegenerative diseases, as well as in mitochondrial diseases.
Since heart contractility requires elevated and sustained levels of ATP, an overall failure of mitochondrial function has been placed as a hallmark of cardiac diseases [132, 133]. Disrupted mitochondrial morphology—characterized by increase number of smaller organelles [134], has been detected in cardiac patients suggesting imbalance between fusion and fission as critical factor for heart pathophysiology. Indeed, while absence of Mfn1 or Mfn2 [135,136,137], or excessive OPA1 cleavage [138] are sufficient to disrupt mitochondrial fusion leading to cardiomyopathy in mice, inactivation of Drp1 blunts excessive fission of the organelle, thus counteracting cardiac dysfunction [139]. Likewise, small molecules capable of blocking fission (i.e. Mdivi-1 and P110) [139, 140] or improving fusion (SAMβA) [141], as well as exercise [142], reestablish mitochondrial dynamics and improve clinical outcome in preclinical models of cardiac diseases. Of interest, failing hearts display loss of mtDNA along with reduced expression of mitochondrial biogenesis markers [143]. Moreover, cardiac specific ablation of PGC-1α leads to cardiac dysfunction in mice [144]. Because of that, activators of AMPK (i.e. Metformin, AICAR) capable of stimulating mitochondrial biogenesis [145], are emerging as promising therapies to treat cardiovascular diseases [146].
Metabolic disorders including type 2 Diabetes and obesity not only arises from a complex combination of genetic and environmental factors such as insulin resistance, dyslipidemia, erroneous food intake and physical inactivity [147], but also display mitochondrial dysfunction as a common feature [148]. Part of this phenotype is due to impaired mitochondrial biogenesis and dynamics in a wide spectrum of tissues. Reduced expression of PGC-1 members along with defective translation of genes encoding subunits of respiratory chain have been observed in skeletal muscle from diabetic patients [149] and adipose tissue from obese subjects [150]. Strengthening these results, mice lacking PGC-1α in adipose tissue develop insulin resistance and abnormal thermogenic response [151]. Moreover, mitochondrial biogenesis have been linked to the beneficial effects of agonists of AMPK [152, 153] and PPAR [154, 155]—widely used drugs for the treatment of type 2 Diabetes. The excessive nutrient environment observed in metabolic disorders also promotes disruption of mitochondrial dynamics. Consistent with reduction of Mfn2 levels [156, 157], mitochondrial fragmentation associated with insulin sensitivity and altered metabolism has been observed in obesity and type 2 Diabetes [15, 158].
Along with progressive loss of neuronal systems, disruption of mitochondrial homeostasis plays a role in the pathogenesis of neurodegenerative disorders such as Parkinson´s, Alzheimer´s and Huntington´s diseases [159, 160]. Analysis of human brains from Alzheimer´s patients revealed not only structurally abnormal mitochondria [161, 162], but also indicated a strong link between Drp1-mediated mitochondrial fission and neurodegeneration [163]. Indeed, blocking mitochondrial fragmentation exhibits beneficial effects in preclinical models of Huntington´s [164] and Parkinson’s [165] disease, and Amyotrophic lateral sclerosis [166]. On the contrary, loss-of-function Drp1 mutations leading to giant and aberrant mitochondria are often associated with lethal neurological disorders including microcephaly [167] and refractory epilepsy [168]; therefore, reinforcing the role of an exquisite balance of mitochondrial fusion and fission events in cellular physiology. In the context of mitochondrial biogenesis, deficiencies in the ETC are related to mtDNA mutations in Alzheimer’s patients, which suppress mitochondrial transcription and replication [169]. Similarly, studies in animals have shown that whereas impaired mitochondrial biogenesis leads to loss of neurons [170], PGC-1α upregulation protects neural cells against oxidative stress-induced death [171].
Mostly driven by loss-of-function mutations in mtDNA or nuclear DNA, mitochondrial diseases refer to a heterogeneous group of disorders triggered by mitochondrial dysfunction [172]. Regardless the disease etiology, there is an overall decrease in content and function of respiratory chain subunits [173,174,175]. In this context, PGC-1α overexpression can boost ATP production by increasing the amount of the organelle in Leigh syndrome [174]. Inducers of mitochondrial biogenesis (i.e. AICAR) also delay the progression of mitochondrial myopathies in mice [174, 176]. Despite the fact that most of these disorders arise from defects in OXPHOS components, progressive neuronal degeneration along with aberrant mitochondrial morphology are observed in preclinical models of Leigh syndrome [175]. Progressive loss of vision observed in autosomal dominant optic atrophy disease is associated with OPA1 mutations [177]. Moreover, impaired mitochondrial fusion or fission by mutations in Mfn2 [178] and Dyn2 genes [179] cause the inherited Charcot Marie Tooth disease. Of interest, due to its involvement in mtDNA replication [112, 180], disruption of mitochondrial dynamics may increase the susceptibility to these inborn errors. Finally, highlighting the dynamic nature of mitochondria, gene therapy is the latest and attractive strategy to restore mitochondrial function and counteract clinical progression of primary mitochondrial diseases [181,182,183,184].
5 Concluding Remarks
The dynamic behavior of mitochondria morphology, controlled by mitochondrial biogenesis and fission-fusion machineries, are determinant for the whole-body homeostasis. Fluctuations in mitochondrial quantity, size and cellular position occur in response to numerous stress and metabolic conditions, which will lead to divergent outcomes. If transient, perturbations of mitochondrial mass and morphology enable metabolic adaptations to meet energetic requirements. On the contrary, sustained stress-induced mitochondrial dysfunction often triggers mitochondrial fragmentation and induces cell death. Moreover, due to the dynamic nature of mitochondria, studying the physiological and pathological significance of mitochondrial network in a time-, tissue- and stress-dependent manner is a challenging task. The development of advanced techniques capable of tracking fusion and fission events in vivo, as well as the identification of new players controlling biogenesis and dynamics will be crucial not only to overcome these obstacles, but also to open up new avenues for pharmacological interventions.
References
Mitchell P, Moyle J (1967) Chemiosmotic hypothesis of oxidative phosphorylation. Nature 213(5072):137–139
Sagan L (1967) On the origin of mitosing cells. J Theor Biol 14(3):255–274
Mills EL, Kelly B, O’Neill LAJ (2017) Mitochondria are the powerhouses of immunity. Nat Immunol 18(5):488–498
Roger AJ, Munoz-Gomez SA, Kamikawa R (2017) The origin and diversification of mitochondria. Curr Biol 27(21):R1177–R1192
Palade GE (1953) An electron microscope study of the mitochondrial structure. J Histochem Cytochem 1(4):188–211
Sjostrand FS (1953) Electron microscopy of mitochondria and cytoplasmic double membranes. Nature 171(4340):30–32
Boveris A, Chance B (1973) The mitochondrial generation of hydrogen peroxide. General properties and effect of hyperbaric oxygen. Biochem J 134(3):707–716
Boveris A, Oshino N, Chance B (1972) The cellular production of hydrogen peroxide. Biochem J 128(3):617–630
Walkenstein SS, Weinhouse S (1953) Oxidation of aldehydes by mitochondria of rat tissues. J Biol Chem 200(2):515–523
Rafael J, Klaas D, Hohorst HJ (1968) Mitochondria from brown fat: enzymes and respiratory chain phosphorylation during the pre- and postnatal development of the interscapular fat body of the guinea pig. Hoppe Seylers Z Physiol Chem 349(12):1711–1724
Rossi CS, Lehninger AL (1964) Stoichiometry of respiratory stimulation, accumulation of Ca++ and phosphate, and oxidative phosphorylation in rat liver mitochondria. J Biol Chem 239:3971–3980
Kerr JF, Wyllie AH, Currie AR (1972) Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br J Cancer 26(4):239–257
Rabinowitz M, Swift H (1970) Mitochondrial nucleic acids and their relation to the biogenesis of mitochondria. Physiol Rev 50(3):376–427
Bereiter-Hahn J (1990) Behavior of mitochondria in the living cell. Int Rev Cytol 122:1–63
Liesa M, Shirihai OS (2013) Mitochondrial dynamics in the regulation of nutrient utilization and energy expenditure. Cell Metab 17(4):491–506
Lin J, Handschin C, Spiegelman BM (2005) Metabolic control through the PGC-1 family of transcription coactivators. Cell Metab 1(6):361–370
Chen H, Chomyn A, Chan DC (2005) Disruption of fusion results in mitochondrial heterogeneity and dysfunction. J Biol Chem 280(28):26185–26192
Twig G, Elorza A, Molina AJ, Mohamed H, Wikstrom JD, Walzer G et al (2008) Fission and selective fusion govern mitochondrial segregation and elimination by autophagy. EMBO J 27(2):433–446
Pearce EL, Pearce EJ (2013) Metabolic pathways in immune cell activation and quiescence. Immunity 38(4):633–643
Mohanty A, Tiwari-Pandey R, Pandey NR (2019) Mitochondria: the indispensable players in innate immunity and guardians of the inflammatory response. J Cell Commun Signal. 13(3):303–318
Zhang Q, Raoof M, Chen Y, Sumi Y, Sursal T, Junger W et al (2010) Circulating mitochondrial DAMPs cause inflammatory responses to injury. Nature 464(7285):104–107
Rambold AS, Pearce EL (2018) Mitochondrial dynamics at the interface of immune cell metabolism and function. Trends Immunol 39(1):6–18
Robin ED, Wong R (1988) Mitochondrial DNA molecules and virtual number of mitochondria per cell in mammalian cells. J Cell Physiol 136(3):507–513
Andersson SG, Zomorodipour A, Andersson JO, Sicheritz-Ponten T, Alsmark UC, Podowski RM et al (1998) The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature 396(6707):133–140
Calvo SE, Clauser KR, Mootha VK (2016) MitoCarta2.0: an updated inventory of mammalian mitochondrial proteins. Nucleic Acids Res 44(D1):D1251–D1257
Pagliarini DJ, Calvo SE, Chang B, Sheth SA, Vafai SB, Ong SE et al (2008) A mitochondrial protein compendium elucidates complex I disease biology. Cell 134(1):112–123
Letts JA, Sazanov LA (2017) Clarifying the supercomplex: the higher-order organization of the mitochondrial electron transport chain. Nat Struct Mol Biol 24(10):800–808
Garesse R, Vallejo CG (2001) Animal mitochondrial biogenesis and function: a regulatory cross-talk between two genomes. Gene 263(1–2):1–16
Jovaisaite V, Auwerx J (2015) The mitochondrial unfolded protein response-synchronizing genomes. Curr Opin Cell Biol 33:74–81
Weissbach A, Baltimore D, Bollum F, Gallo R, Korn D (1975) Nomenclature of eukaryotic DNA polymerases. Science 190(4212):401–402
Yasukawa T, Kang D (2018) An overview of mammalian mitochondrial DNA replication mechanisms. J Biochem 164(3):183–193
D’Souza AR, Minczuk M (2018) Mitochondrial transcription and translation: overview. Essays Biochem 62(3):309–320
Vaquerizas JM, Kummerfeld SK, Teichmann SA, Luscombe NM (2009) A census of human transcription factors: function, expression and evolution. Nat Rev Genet 10(4):252–263
Evans MJ, Scarpulla RC (1989) Interaction of nuclear factors with multiple sites in the somatic cytochrome c promoter. Characterization of upstream NRF-1, ATF, and intron Sp1 recognition sequences. J Biol Chem 264(24):14361–8.
Evans MJ, Scarpulla RC (1990) NRF-1: a trans-activator of nuclear-encoded respiratory genes in animal cells. Genes Dev 4(6):1023–1034
Virbasius JV, Virbasius CA, Scarpulla RC (1993) Identity of GABP with NRF-2, a multisubunit activator of cytochrome oxidase expression, reveals a cellular role for an ETS domain activator of viral promoters. Genes Dev 7(3):380–392
Itoh K, Chiba T, Takahashi S, Ishii T, Igarashi K, Katoh Y et al (1997) An Nrf2/small Maf heterodimer mediates the induction of phase II detoxifying enzyme genes through antioxidant response elements. Biochem Biophys Res Commun 236(2):313–322
Kelly DP, Scarpulla RC (2004) Transcriptional regulatory circuits controlling mitochondrial biogenesis and function. Genes Dev 18(4):357–368
Blesa JR, Hernandez JM, Hernandez-Yago J (2004) NRF-2 transcription factor is essential in promoting human Tomm70 gene expression. Mitochondrion 3(5):251–259
Virbasius JV, Scarpulla RC (1994) Activation of the human mitochondrial transcription factor A gene by nuclear respiratory factors: a potential regulatory link between nuclear and mitochondrial gene expression in organelle biogenesis. Proc Natl Acad Sci U S A 91(4):1309–1313
Gleyzer N, Vercauteren K, Scarpulla RC (2005) Control of mitochondrial transcription specificity factors (TFB1M and TFB2M) by nuclear respiratory factors (NRF-1 and NRF-2) and PGC-1 family coactivators. Mol Cell Biol 25(4):1354–1366
Larsson NG, Wang J, Wilhelmsson H, Oldfors A, Rustin P, Lewandoski M et al (1998) Mitochondrial transcription factor A is necessary for mtDNA maintenance and embryogenesis in mice. Nat Genet 18(3):231–236
Falkenberg M, Gaspari M, Rantanen A, Trifunovic A, Larsson NG, Gustafsson CM (2002) Mitochondrial transcription factors B1 and B2 activate transcription of human mtDNA. Nat Genet 31(3):289–294
Piantadosi CA, Carraway MS, Babiker A, Suliman HB (2008) Heme oxygenase-1 regulates cardiac mitochondrial biogenesis via Nrf2-mediated transcriptional control of nuclear respiratory factor-1. Circ Res 103(11):1232–1240
Huo L, Scarpulla RC (2001) Mitochondrial DNA instability and peri-implantation lethality associated with targeted disruption of nuclear respiratory factor 1 in mice. Mol Cell Biol 21(2):644–654
Chen H, Hu Y, Fang Y, Djukic Z, Yamamoto M, Shaheen NJ et al (2014) Nrf2 deficiency impairs the barrier function of mouse oesophageal epithelium. Gut 63(5):711–719
Issemann I, Green S (1990) Activation of a member of the steroid hormone receptor superfamily by peroxisome proliferators. Nature 347(6294):645–650
Schmidt A, Endo N, Rutledge SJ, Vogel R, Shinar D, Rodan GA (1992) Identification of a new member of the steroid hormone receptor superfamily that is activated by a peroxisome proliferator and fatty acids. Mol Endocrinol 6(10):1634–1641
Greene ME, Blumberg B, McBride OW, Yi HF, Kronquist K, Kwan K et al (1995) Isolation of the human peroxisome proliferator activated receptor gamma cDNA: expression in hematopoietic cells and chromosomal mapping. Gene Expr 4(4–5):281–299
Madrazo JA, Kelly DP (2008) The PPAR trio: regulators of myocardial energy metabolism in health and disease. J Mol Cell Cardiol 44(6):968–975
Bookout AL, Jeong Y, Downes M, Yu RT, Evans RM, Mangelsdorf DJ (2006) Anatomical profiling of nuclear receptor expression reveals a hierarchical transcriptional network. Cell 126(4):789–799
Eichner LJ, Giguere V (2011) Estrogen related receptors (ERRs): a new dawn in transcriptional control of mitochondrial gene networks. Mitochondrion 11(4):544–552
Huss JM, Torra IP, Staels B, Giguere V, Kelly DP (2004) Estrogen-related receptor alpha directs peroxisome proliferator-activated receptor alpha signaling in the transcriptional control of energy metabolism in cardiac and skeletal muscle. Mol Cell Biol 24(20):9079–9091
Scarpulla RC, Vega RB, Kelly DP (2012) Transcriptional integration of mitochondrial biogenesis. Trends Endocrinol Metab 23(9):459–466
Puigserver P, Wu Z, Park CW, Graves R, Wright M, Spiegelman BM (1998) A cold-inducible coactivator of nuclear receptors linked to adaptive thermogenesis. Cell 92(6):829–839
Lin J, Puigserver P, Donovan J, Tarr P, Spiegelman BM (2002) Peroxisome proliferator-activated receptor gamma coactivator 1beta (PGC-1beta ), a novel PGC-1-related transcription coactivator associated with host cell factor. J Biol Chem 277(3):1645–1648
Knutti D, Kaul A, Kralli A (2000) A tissue-specific coactivator of steroid receptors, identified in a functional genetic screen. Mol Cell Biol 20(7):2411–2422
Andersson U, Scarpulla RC (2001) Pgc-1-related coactivator, a novel, serum-inducible coactivator of nuclear respiratory factor 1-dependent transcription in mammalian cells. Mol Cell Biol 21(11):3738–3749
He X, Sun C, Wang F, Shan A, Guo T, Gu W et al (2012) Peri-implantation lethality in mice lacking the PGC-1-related coactivator protein. Dev Dyn 241(5):975–983
Finck BN, Kelly DP (2006) PGC-1 coactivators: inducible regulators of energy metabolism in health and disease. J Clin Invest 116(3):615–622
Wu Z, Puigserver P, Andersson U, Zhang C, Adelmant G, Mootha V et al (1999) Mechanisms controlling mitochondrial biogenesis and respiration through the thermogenic coactivator PGC-1. Cell 98(1):115–124
Vega RB, Huss JM, Kelly DP (2000) The coactivator PGC-1 cooperates with peroxisome proliferator-activated receptor alpha in transcriptional control of nuclear genes encoding mitochondrial fatty acid oxidation enzymes. Mol Cell Biol 20(5):1868–1876
Wang YX, Lee CH, Tiep S, Yu RT, Ham J, Kang H et al (2003) Peroxisome-proliferator-activated receptor delta activates fat metabolism to prevent obesity. Cell 113(2):159–170
Schreiber SN, Knutti D, Brogli K, Uhlmann T, Kralli A (2003) The transcriptional coactivator PGC-1 regulates the expression and activity of the orphan nuclear receptor estrogen-related receptor alpha (ERRalpha). J Biol Chem 278(11):9013–9018
Huss JM, Kopp RP, Kelly DP (2002) Peroxisome proliferator-activated receptor coactivator-1alpha (PGC-1alpha) coactivates the cardiac-enriched nuclear receptors estrogen-related receptor-alpha and -gamma. Identification of novel leucine-rich interaction motif within PGC-1alpha. J Biol Chem 277(43):40265–40274
Zhang Y, Ma K, Song S, Elam MB, Cook GA, Park EA (2004) Peroxisomal proliferator-activated receptor-gamma coactivator-1 alpha (PGC-1 alpha) enhances the thyroid hormone induction of carnitine palmitoyltransferase I (CPT-I alpha). J Biol Chem 279(52):53963–53971
Tcherepanova I, Puigserver P, Norris JD, Spiegelman BM, McDonnell DP (2000) Modulation of estrogen receptor-alpha transcriptional activity by the coactivator PGC-1. J Biol Chem 275(21):16302–16308
Lai L, Leone TC, Zechner C, Schaeffer PJ, Kelly SM, Flanagan DP et al (2008) Transcriptional coactivators PGC-1alpha and PGC-lbeta control overlapping programs required for perinatal maturation of the heart. Genes Dev 22(14):1948–1961
Espinoza DO, Boros LG, Crunkhorn S, Gami H, Patti ME (2010) Dual modulation of both lipid oxidation and synthesis by peroxisome proliferator-activated receptor-gamma coactivator-1alpha and -1beta in cultured myotubes. FASEB J 24(4):1003–1014
St-Pierre J, Lin J, Krauss S, Tarr PT, Yang R, Newgard CB et al (2003) Bioenergetic analysis of peroxisome proliferator-activated receptor gamma coactivators 1alpha and 1beta (PGC-1alpha and PGC-1beta) in muscle cells. J Biol Chem 278(29):26597–26603
Handschin C, Spiegelman BM (2006) Peroxisome proliferator-activated receptor gamma coactivator 1 coactivators, energy homeostasis, and metabolism. Endocr Rev 27(7):728–735
Herzig S, Long F, Jhala US, Hedrick S, Quinn R, Bauer A et al (2001) CREB regulates hepatic gluconeogenesis through the coactivator PGC-1. Nature 413(6852):179–183
Handschin C, Rhee J, Lin J, Tarr PT, Spiegelman BM (2003) An autoregulatory loop controls peroxisome proliferator-activated receptor gamma coactivator 1alpha expression in muscle. Proc Natl Acad Sci U S A 100(12):7111–7116
Wang L, Liu J, Saha P, Huang J, Chan L, Spiegelman B et al (2005) The orphan nuclear receptor SHP regulates PGC-1alpha expression and energy production in brown adipocytes. Cell Metab 2(4):227–238
Santos AL, Lindner AB (2017) Protein posttranslational modifications: roles in aging and age-related disease. Oxid Med Cell Longev 2017:5716409
Knutti D, Kressler D, Kralli A (2001) Regulation of the transcriptional coactivator PGC-1 via MAPK-sensitive interaction with a repressor. Proc Natl Acad Sci U S A 98(17):9713–9718
Zong H, Ren JM, Young LH, Pypaert M, Mu J, Birnbaum MJ et al (2002) AMP kinase is required for mitochondrial biogenesis in skeletal muscle in response to chronic energy deprivation. Proc Natl Acad Sci U S A 99(25):15983–15987
Rodgers JT, Lerin C, Haas W, Gygi SP, Spiegelman BM, Puigserver P (2005) Nutrient control of glucose homeostasis through a complex of PGC-1alpha and SIRT1. Nature 434(7029):113–118
Anderson RM, Barger JL, Edwards MG, Braun KH, O’Connor CE, Prolla TA et al (2008) Dynamic regulation of PGC-1alpha localization and turnover implicates mitochondrial adaptation in calorie restriction and the stress response. Aging Cell 7(1):101–111
Fernandez-Marcos PJ, Auwerx J (2011) Regulation of PGC-1alpha, a nodal regulator of mitochondrial biogenesis. Am J Clin Nutr 93(4):884S-S890
Winder WW, Holmes BF, Rubink DS, Jensen EB, Chen M, Holloszy JO (1985) Activation of AMP-activated protein kinase increases mitochondrial enzymes in skeletal muscle. J Appl Physiol 88(6):2219–2226
Zhang Y, Huypens P, Adamson AW, Chang JS, Henagan TM, Boudreau A et al (2009) Alternative mRNA splicing produces a novel biologically active short isoform of PGC-1alpha. J Biol Chem 284(47):32813–32826
Tadaishi M, Miura S, Kai Y, Kano Y, Oishi Y, Ezaki O (2011) Skeletal muscle-specific expression of PGC-1alpha-b, an exercise-responsive isoform, increases exercise capacity and peak oxygen uptake. PLoS One 6(12):e28290
Ruas JL, White JP, Rao RR, Kleiner S, Brannan KT, Harrison BC et al (2012) A PGC-1alpha isoform induced by resistance training regulates skeletal muscle hypertrophy. Cell 151(6):1319–1331
Ydfors M, Fischer H, Mascher H, Blomstrand E, Norrbom J, Gustafsson T (2013) The truncated splice variants, NT-PGC-1alpha and PGC-1alpha4, increase with both endurance and resistance exercise in human skeletal muscle. Physiol Rep 1(6):e00140
Asano M, Wakabayashi T, Ishikawa K, Kishimoto H (1977) Induction of megamitochondria in mouse hepatocytes by nialamide. J Electron Microsc (Tokyo) 26(2):141–144
Bakeeva LE, Chentsov YuS, Skulachev VP (1978) Mitochondrial framework (reticulum mitochondriale) in rat diaphragm muscle. Biochim Biophys Acta 501(3):349–369
Bereiter-Hahn J, Voth M (1994) Dynamics of mitochondria in living cells: shape changes, dislocations, fusion, and fission of mitochondria. Microsc Res Tech 27(3):198–219
Cortese JD (1998) Stimulation of rat liver mitochondrial fusion by an outer membrane-derived aluminum fluoride-sensitive protein fraction. Exp Cell Res 240(1):122–133
Liesa M, Palacin M, Zorzano A (2009) Mitochondrial dynamics in mammalian health and disease. Physiol Rev 89(3):799–845
Ishihara N, Otera H, Oka T, Mihara K (2013) Regulation and physiologic functions of GTPases in mitochondrial fusion and fission in mammals. Antioxid Redox Signal 19(4):389–399
Youle RJ, Narendra DP (2011) Mechanisms of mitophagy. Nat Rev Mol Cell Biol 12(1):9–14
Hamacher-Brady A, Brady NR (2016) Mitophagy programs: mechanisms and physiological implications of mitochondrial targeting by autophagy. Cell Mol Life Sci 73(4):775–795
Eisner V, Picard M, Hajnoczky G (2018) Mitochondrial dynamics in adaptive and maladaptive cellular stress responses. Nat Cell Biol 20(7):755–765
Rambold AS, Kostelecky B, Elia N, Lippincott-Schwartz J (2011) Tubular network formation protects mitochondria from autophagosomal degradation during nutrient starvation. Proc Natl Acad Sci U S A 108(25):10190–10195
Chen H, Vermulst M, Wang YE, Chomyn A, Prolla TA, McCaffery JM et al (2010) Mitochondrial fusion is required for mtDNA stability in skeletal muscle and tolerance of mtDNA mutations. Cell 141(2):280–289
Malka F, Guillery O, Cifuentes-Diaz C, Guillou E, Belenguer P, Lombes A et al (2005) Separate fusion of outer and inner mitochondrial membranes. EMBO Rep 6(9):853–859
Koshiba T, Detmer SA, Kaiser JT, Chen H, McCaffery JM, Chan DC (2004) Structural basis of mitochondrial tethering by mitofusin complexes. Science 305(5685):858–862
Ishihara N, Eura Y, Mihara K (2004) Mitofusin 1 and 2 play distinct roles in mitochondrial fusion reactions via GTPase activity. J Cell Sci 117(Pt 26):6535–6546
Chen H, Detmer SA, Ewald AJ, Griffin EE, Fraser SE, Chan DC (2003) Mitofusins Mfn1 and Mfn2 coordinately regulate mitochondrial fusion and are essential for embryonic development. J Cell Biol 160(2):189–200
Eura Y, Ishihara N, Yokota S, Mihara K (2003) Two mitofusin proteins, mammalian homologues of FZO, with distinct functions are both required for mitochondrial fusion. J Biochem 134(3):333–344
de Brito OM, Scorrano L (2008) Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature 456(7222):605–610
Naon D, Zaninello M, Giacomello M, Varanita T, Grespi F, Lakshminaranayan S et al (2016) Critical reappraisal confirms that Mitofusin 2 is an endoplasmic reticulum-mitochondria tether. Proc Natl Acad Sci U S A 113(40):11249–11254
Del Dotto V, Fogazza M, Carelli V, Rugolo M, Zanna C (2018) Eight human OPA1 isoforms, long and short: what are they for? Biochim Biophys Acta Bioenerg 1859(4):263–269
Delettre C, Griffoin JM, Kaplan J, Dollfus H, Lorenz B, Faivre L et al (2001) Mutation spectrum and splicing variants in the OPA1 gene. Hum Genet 109(6):584–591
Song Z, Chen H, Fiket M, Alexander C, Chan DC (2007) OPA1 processing controls mitochondrial fusion and is regulated by mRNA splicing, membrane potential, and Yme1L. J Cell Biol 178(5):749–755
Ehses S, Raschke I, Mancuso G, Bernacchia A, Geimer S, Tondera D et al (2009) Regulation of OPA1 processing and mitochondrial fusion by m-AAA protease isoenzymes and OMA1. J Cell Biol 187(7):1023–1036
Anand R, Wai T, Baker MJ, Kladt N, Schauss AC, Rugarli E et al (2014) The i-AAA protease YME1L and OMA1 cleave OPA1 to balance mitochondrial fusion and fission. J Cell Biol 204(6):919–929
Ishihara N, Fujita Y, Oka T, Mihara K (2006) Regulation of mitochondrial morphology through proteolytic cleavage of OPA1. EMBO J 25(13):2966–2977
Head B, Griparic L, Amiri M, Gandre-Babbe S, van der Bliek AM (2009) Inducible proteolytic inactivation of OPA1 mediated by the OMA1 protease in mammalian cells. J Cell Biol 187(7):959–966
Olichon A, Baricault L, Gas N, Guillou E, Valette A, Belenguer P et al (2003) Loss of OPA1 perturbates the mitochondrial inner membrane structure and integrity, leading to cytochrome c release and apoptosis. J Biol Chem 278(10):7743–7746
Mishra P, Chan DC (2014) Mitochondrial dynamics and inheritance during cell division, development and disease. Nat Rev Mol Cell Biol 15(10):634–646
Gandre-Babbe S, van der Bliek AM (2008) The novel tail-anchored membrane protein Mff controls mitochondrial and peroxisomal fission in mammalian cells. Mol Biol Cell 19(6):2402–2412
Smirnova E, Griparic L, Shurland DL, van der Bliek AM (2001) Dynamin-related protein Drp1 is required for mitochondrial division in mammalian cells. Mol Biol Cell 12(8):2245–2256
Gomes LC, Di Benedetto G, Scorrano L (2011) During autophagy mitochondria elongate, are spared from degradation and sustain cell viability. Nat Cell Biol 13(5):589–598
Cereghetti GM, Stangherlin A, Martins de Brito O, Chang CR, Blackstone C, Bernardi P, et al (2008) Dephosphorylation by calcineurin regulates translocation of Drp1 to mitochondria. Proc Natl Acad Sci U S A 105(41):15803–15808
Cribbs JT, Strack S (2007) Reversible phosphorylation of Drp1 by cyclic AMP-dependent protein kinase and calcineurin regulates mitochondrial fission and cell death. EMBO Rep 8(10):939–944
Yonashiro R, Ishido S, Kyo S, Fukuda T, Goto E, Matsuki Y et al (2006) A novel mitochondrial ubiquitin ligase plays a critical role in mitochondrial dynamics. EMBO J 25(15):3618–3626
Poole AC, Thomas RE, Andrews LA, McBride HM, Whitworth AJ, Pallanck LJ (2008) The PINK1/Parkin pathway regulates mitochondrial morphology. Proc Natl Acad Sci U S A 105(5):1638–1643
Chang CR, Blackstone C (2010) Dynamic regulation of mitochondrial fission through modification of the dynamin-related protein Drp1. Ann N Y Acad Sci 1201:34–39
Benard G, Karbowski M (2009) Mitochondrial fusion and division: regulation and role in cell viability. Semin Cell Dev Biol 20(3):365–374
Loson OC, Song Z, Chen H, Chan DC (2013) Fis1, Mff, MiD49, and MiD51 mediate Drp1 recruitment in mitochondrial fission. Mol Biol Cell 24(5):659–667
Palmer CS, Osellame LD, Laine D, Koutsopoulos OS, Frazier AE, Ryan MT (2011) MiD49 and MiD51, new components of the mitochondrial fission machinery. EMBO Rep 12(6):565–573
Friedman JR, Lackner LL, West M, DiBenedetto JR, Nunnari J, Voeltz GK (2011) ER tubules mark sites of mitochondrial division. Science 334(6054):358–362
Ji WK, Chakrabarti R, Fan X, Schoenfeld L, Strack S, Higgs HN (2017) Receptor-mediated Drp1 oligomerization on endoplasmic reticulum. J Cell Biol 216(12):4123–4139
Toyama EQ, Herzig S, Courchet J, Lewis TL, Jr., Loson OC, Hellberg K, et al (2016) Metabolism. AMP-activated protein kinase mediates mitochondrial fission in response to energy stress. Science 351(6270):275–281
Ducommun S, Deak M, Sumpton D, Ford RJ, Nunez Galindo A, Kussmann M et al (2015) Motif affinity and mass spectrometry proteomic approach for the discovery of cellular AMPK targets: identification of mitochondrial fission factor as a new AMPK substrate. Cell Signal 27(5):978–988
Lee JE, Westrate LM, Wu H, Page C, Voeltz GK (2016) Multiple dynamin family members collaborate to drive mitochondrial division. Nature 540(7631):139–143
Nunnari J, Suomalainen A (2012) Mitochondria: in sickness and in health. Cell 148(6):1145–1159
Sorrentino V, Romani M, Mouchiroud L, Beck JS, Zhang H, D’Amico D et al (2017) Enhancing mitochondrial proteostasis reduces amyloid-beta proteotoxicity. Nature 552(7684):187–193
Pfeffer G, Majamaa K, Turnbull DM, Thorburn D, Chinnery PF (2012) Treatment for mitochondrial disorders. Cochrane Database Syst Rev (4):CD004426
Campos JC, Bozi LH, Bechara LR, Lima VM, Ferreira JC (2016) Mitochondrial Quality Control in Cardiac Diseases. Front Physiol 7:479
Campos JC, Gomes KM, Ferreira JC (2013) Impact of exercise training on redox signaling in cardiovascular diseases. Food Chem Toxicol 62:107–119
Nan J, Zhu W, Rahman MS, Liu M, Li D, Su S et al (2017) Molecular regulation of mitochondrial dynamics in cardiac disease. Biochim Biophys Acta Mol Cell Res 1864(7):1260–1273
Song M, Franco A, Fleischer JA, Zhang L, Dorn GW (2017) Abrogating mitochondrial dynamics in mouse hearts accelerates mitochondrial senescence (2nd edn). Cell Metab 26(6):872–883, e5
Chen Y, Dorn GW 2nd (2013) PINK1-phosphorylated mitofusin 2 is a Parkin receptor for culling damaged mitochondria. Science 340(6131):471–475
Chen Y, Liu Y, Dorn GW 2nd (2011) Mitochondrial fusion is essential for organelle function and cardiac homeostasis. Circ Res 109(12):1327–1331
Wai T, Garcia-Prieto J, Baker MJ, Merkwirth C, Benit P, Rustin P, et al (2015) Imbalanced OPA1 processing and mitochondrial fragmentation cause heart failure in mice. Science 350(6265):aad0116
Disatnik MH, Ferreira JC, Campos JC, Gomes KS, Dourado PM, Qi X, et al (2013) Acute inhibition of excessive mitochondrial fission after myocardial infarction prevents long-term cardiac dysfunction. J Am Heart Assoc 2(5):e000461
Givvimani S, Munjal C, Tyagi N, Sen U, Metreveli N, Tyagi SC (2012) Mitochondrial division/mitophagy inhibitor (Mdivi) ameliorates pressure overload induced heart failure. PLoS One 7(3):e32388
Ferreira JCB, Campos JC, Qvit N, Qi X, Bozi LHM, Bechara LRG et al (2019) A selective inhibitor of mitofusin 1-betaIIPKC association improves heart failure outcome in rats. Nat Commun 10(1):329
Campos JC, Queliconi BB, Bozi LHM, Bechara LRG, Dourado PMM, Andres AM et al (2017) Exercise reestablishes autophagic flux and mitochondrial quality control in heart failure. Autophagy 13(8):1304–1317
Pisano A, Cerbelli B, Perli E, Pelullo M, Bargelli V, Preziuso C et al (2016) Impaired mitochondrial biogenesis is a common feature to myocardial hypertrophy and end-stage ischemic heart failure. Cardiovasc Pathol 25(2):103–112
Arany Z, He H, Lin J, Hoyer K, Handschin C, Toka O et al (2005) Transcriptional coactivator PGC-1 alpha controls the energy state and contractile function of cardiac muscle. Cell Metab 1(4):259–271
Herzig S, Shaw RJ (2018) AMPK: guardian of metabolism and mitochondrial homeostasis. Nat Rev Mol Cell Biol 19(2):121–135
Beauloye C, Bertrand L, Horman S, Hue L (2011) AMPK activation, a preventive therapeutic target in the transition from cardiac injury to heart failure. Cardiovasc Res 90(2):224–233
Han TS, Lean ME (2016) A clinical perspective of obesity, metabolic syndrome and cardiovascular disease. JRSM Cardiovasc Dis 5:2048004016633371
Bhatti JS, Bhatti GK, Reddy PH (2017) Mitochondrial dysfunction and oxidative stress in metabolic disorders—A step towards mitochondria based therapeutic strategies. Biochim Biophys Acta Mol Basis Dis 1863(5):1066–1077
Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J et al (2003) PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34(3):267–273
Heinonen S, Buzkova J, Muniandy M, Kaksonen R, Ollikainen M, Ismail K et al (2015) Impaired mitochondrial biogenesis in adipose tissue in acquired obesity. Diabetes 64(9):3135–3145
Kleiner S, Mepani RJ, Laznik D, Ye L, Jurczak MJ, Jornayvaz FR et al (2012) Development of insulin resistance in mice lacking PGC-1alpha in adipose tissues. Proc Natl Acad Sci U S A 109(24):9635–9640
Foretz M, Guigas B, Bertrand L, Pollak M, Viollet B (2014) Metformin: from mechanisms of action to therapies. Cell Metab 20(6):953–966
Zhou G, Myers R, Li Y, Chen Y, Shen X, Fenyk-Melody J et al (2001) Role of AMP-activated protein kinase in mechanism of metformin action. J Clin Invest 108(8):1167–1174
Bogacka I, Xie H, Bray GA, Smith SR (2005) Pioglitazone induces mitochondrial biogenesis in human subcutaneous adipose tissue in vivo. Diabetes 54(5):1392–1399
Diamant M, Heine RJ (2003) Thiazolidinediones in type 2 diabetes mellitus: current clinical evidence. Drugs 63(13):1373–1405
Bach D, Naon D, Pich S, Soriano FX, Vega N, Rieusset J et al (2005) Expression of Mfn2, the charcot-marie-tooth neuropathy type 2A gene, in human skeletal muscle: effects of type 2 diabetes, obesity, weight loss, and the regulatory role of tumor necrosis factor alpha and interleukin-6. Diabetes 54(9):2685–2693
Bach D, Pich S, Soriano FX, Vega N, Baumgartner B, Oriola J, et al (2003) Mitofusin-2 determines mitochondrial network architecture and mitochondrial metabolism. A novel regulatory mechanism altered in obesity. J Biol Chem 278(19):17190–171907
Wai T, Langer T (2016) Mitochondrial dynamics and metabolic regulation. Trends Endocrinol Metab 27(2):105–117
Lin MT, Beal MF (2006) Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature 443(7113):787–795
Johri A, Beal MF (2012) Mitochondrial dysfunction in neurodegenerative diseases. J Pharmacol Exp Ther 342(3):619–630
Wang X, Su B, Lee HG, Li X, Perry G, Smith MA et al (2009) Impaired balance of mitochondrial fission and fusion in Alzheimer’s disease. J Neurosci 29(28):9090–9103
Burte F, Carelli V, Chinnery PF, Yu-Wai-Man P (2015) Disturbed mitochondrial dynamics and neurodegenerative disorders. Nat Rev Neurol 11(1):11–24
Cho DH, Nakamura T, Fang J, Cieplak P, Godzik A, Gu Z et al (2009) S-nitrosylation of Drp1 mediates beta-amyloid-related mitochondrial fission and neuronal injury. Science 324(5923):102–105
Guo X, Disatnik MH, Monbureau M, Shamloo M, Mochly-Rosen D, Qi X (2013) Inhibition of mitochondrial fragmentation diminishes Huntington’s disease-associated neurodegeneration. J Clin Invest 123(12):5371–5388
Rappold PM, Cui M, Grima JC, Fan RZ, de Mesy-Bentley KL, Chen L et al (2014) Drp1 inhibition attenuates neurotoxicity and dopamine release deficits in vivo. Nat Commun 5:5244
Joshi AU, Saw NL, Vogel H, Cunnigham AD, Shamloo M, Mochly-Rosen D (2018) Inhibition of Drp1/Fis1 interaction slows progression of amyotrophic lateral sclerosis. EMBO Molecul Med 10(3)
Waterham HR, Koster J, van Roermund CW, Mooyer PA, Wanders RJ, Leonard JV (2007) A lethal defect of mitochondrial and peroxisomal fission. N Engl J Med 356(17):1736–1741
Vanstone JR, Smith AM, McBride S, Naas T, Holcik M, Antoun G et al (2016) DNM1L-related mitochondrial fission defect presenting as refractory epilepsy. Eur J Hum Genet 24(7):1084–1088
Coskun PE, Beal MF, Wallace DC (2004) Alzheimer’s brains harbor somatic mtDNA control-region mutations that suppress mitochondrial transcription and replication. Proc Natl Acad Sci U S A 101(29):10726–10731
Jiang H, Kang SU, Zhang S, Karuppagounder S, Xu J, Lee YK, et al (2016) Adult conditional knockout of PGC-1alpha leads to loss of dopamine neurons. eNeuro 3(4)
St-Pierre J, Drori S, Uldry M, Silvaggi JM, Rhee J, Jager S et al (2006) Suppression of reactive oxygen species and neurodegeneration by the PGC-1 transcriptional coactivators. Cell 127(2):397–408
Craven L, Alston CL, Taylor RW, Turnbull DM (2017) Recent advances in mitochondrial disease. Annu Rev Genomics Hum Genet 18:257–275
Ast T, Meisel JD, Patra S, Wang H, Grange RMH, Kim SH, et al (2019) hypoxia rescues frataxin loss by restoring iron sulfur cluster biogenesis. Cell 177(6):1507–1521 e16
Viscomi C, Bottani E, Civiletto G, Cerutti R, Moggio M, Fagiolari G et al (2011) In vivo correction of COX deficiency by activation of the AMPK/PGC-1alpha axis. Cell Metab 14(1):80–90
Quintana A, Kruse SE, Kapur RP, Sanz E, Palmiter RD (2010) Complex I deficiency due to loss of Ndufs4 in the brain results in progressive encephalopathy resembling Leigh syndrome. Proc Natl Acad Sci U S A 107(24):10996–11001
Khan NA, Auranen M, Paetau I, Pirinen E, Euro L, Forsstrom S et al (2014) Effective treatment of mitochondrial myopathy by nicotinamide riboside, a vitamin B3. EMBO Mol Med 6(6):721–731
Delettre C, Lenaers G, Griffoin JM, Gigarel N, Lorenzo C, Belenguer P et al (2000) Nuclear gene OPA1, encoding a mitochondrial dynamin-related protein, is mutated in dominant optic atrophy. Nat Genet 26(2):207–210
Zuchner S, Mersiyanova IV, Muglia M, Bissar-Tadmouri N, Rochelle J, Dadali EL et al (2004) Mutations in the mitochondrial GTPase mitofusin 2 cause charcot-marie-tooth neuropathy type 2A. Nat Genet 36(5):449–451
Zuchner S, Noureddine M, Kennerson M, Verhoeven K, Claeys K, De Jonghe P et al (2005) Mutations in the pleckstrin homology domain of dynamin 2 cause dominant intermediate charcot-marie-tooth disease. Nat Genet 37(3):289–294
Sato M, Sato K (2011) Degradation of paternal mitochondria by fertilization-triggered autophagy in C. elegans embryos. Science 334(6059):1141–1144
Perdomini M, Belbellaa B, Monassier L, Reutenauer L, Messaddeq N, Cartier N et al (2014) Prevention and reversal of severe mitochondrial cardiomyopathy by gene therapy in a mouse model of Friedreich’s ataxia. Nat Med 20(5):542–547
Jang YH, Lim KI (2018) Recent advances in mitochondria-targeted gene delivery. Molecules 23(9)
Gerard C, Xiao X, Filali M, Coulombe Z, Arsenault M, Couet J et al (2014) An AAV9 coding for frataxin clearly improved the symptoms and prolonged the life of Friedreich ataxia mouse models. Mol Ther Methods Clin Dev. 1:14044
Yu H, Koilkonda RD, Chou TH, Porciatti V, Ozdemir SS, Chiodo V et al (2012) Gene delivery to mitochondria by targeting modified adenoassociated virus suppresses Leber’s hereditary optic neuropathy in a mouse model. Proc Natl Acad Sci U S A 109(20):E1238–E1247
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Campos, J.C., Bozi, L.H.M., Ferreira, J.C.B. (2022). Mitochondrial Biogenesis and Dynamics in Health and Disease. In: Camara, N.O.S., Alves-Filho, J.C., Moraes-Vieira, P.M.M.d., Andrade-Oliveira, V. (eds) Essential Aspects of Immunometabolism in Health and Disease. Springer, Cham. https://doi.org/10.1007/978-3-030-86684-6_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-86684-6_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-86683-9
Online ISBN: 978-3-030-86684-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)