Abstract
Corynebacterium diphtheriae is the leading causing agent of respiratory and cutaneous diphtheria, an acute disease with local and systemic manifestations, which remains as an important cause of morbidity and mortality in different continents. Diphtheria vaccination programs implemented in industrialized and developing countries led to an increasing number of atypical cases of diphtheria in addition to localized and systemic infections, including fully immunized adults. Changes in the clinical epidemiology and virulence features of diphtheria pathogens have been investigated. Cases of infections due to diphtheria toxin (DT)-producing and non-DT-producing C. diphtheriae and Corynebacterium ulcerans, a zoonotic pathogen, have been increasingly reported. The timely and precise diagnosis of DT-producing Corynebacterium strains is indispensable for the patient management and for establishment of surveillance and control strategy of disease. Different molecular methods, such as real-time PCR (polymerase chain reaction) and multiplex PCR, have been used for the characterization of C. diphtheriae, C. ulcerans, and C. pseudotuberculosis and the detection of the gene for DT (tox). Recent investigations by genomic sequencing and chemotaxonomic analyses reported DT-producing C. diphtheriae subsp. lausannense and Corynebacterium belfantii sp. nov. in addition to Corynebacterium rouxii sp. nov. Genotyping methods have been used as essential epidemiological tools for C. diphtheriae and C. ulcerans infection prevention and control, including pulsed-field gel electrophoresis (PFGE), random amplification of polymorphic DNA (RAPD), and multilocus sequence typing (MLST) assays. Molecular typing methods are required in studies involving characterization, virulence potential, and susceptibility to antimicrobial agents of C. diphtheriae and C. ulcerans clinical isolates; origin, routes, and transmission of diphtheria and atypical invasive infections; and endemicity, outbreaks, recurrent infections, and trace cross-transmission caused by non-DT-producing and DT-producing Corynebacterium spp.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Corynebacterium
- Corynebacterium diphtheriae
- Corynebacterium ulcerans
- Corynebacterium pseudotuberculosis
- Diphtheria toxin
- Diphtheria
- Zoonosis
- Endocarditis
- Corynebacterium infections
- Molecular typing
- DNA sequence analysis
- Pulsed-field gel electrophoresis
- Multilocus sequence typing
- Random amplified of polymorphic DNA
- Virulence
1 Introduction
Molecular typing techniques have been successfully applied for determining virulence potential, origin, and routes of diphtheria and atypical invasive infections, confirming endemicity, outbreaks, and trace cross-transmission caused by Corynebacterium diphtheriae and zoonotic toxin-producing Corynebacterium species.
2 Clinical Significance and Epidemiology of Diphtheria Toxin-Producing Corynebacterium spp.
Corynebacterium diphtheriae is a major etiologic agent of classic respiratory diphtheria, including local pharyngeal symptoms and systemic manifestations, mainly caused by the action of diphtheria toxin (DT). Symptoms typically begin 2 to 5 days after infection. C. diphtheriae usually localizes in the upper respiratory tract, ulcerates the mucosa, and induces the formation of an inflammatory pseudomembrane. Systemic toxicity increases as the pseudomembrane spreads from the tonsillopharyngeal area. A form of malignant diphtheria is associated with extensive “membranous pharyngitis” plus massive swelling of the tonsils, uvula, cervical lymph nodes, submandibular region, and anterior neck (the so-called bull neck of toxic diphtheria). Acute disease of the respiratory tract usually involves one or more of the following: tonsillar zones, larynx, soft palate, uvula, and nasal cavities or, less commonly, in the stomach or lungs. The exceedingly potent DT is absorbed into the circulation, and lesions may also occur in vital organs, including the heart (myocarditis), nervous system, and kidneys, potentially resulting in death [1].
Diphtheria toxin is an extracellular protein that inhibits protein synthesis and ultimately exerts death of susceptible eukaryotic cells. DT was the first member to be identified of a group of bacterial protein toxins that act by ADP ribosylation of a target protein. DT contains a toxic A subunit (active toxin with enzymatic activity) and the receptor binding B subunit. The B subunit (fragment) facilitates translocation of the A subunit from the phagosome to the cytosol, followed by separation, allowing full activity of the A subunit on its target protein elongation factor-2 (EF-2). EF-2 transfers polypeptidyl-transfer RNA from acceptor to donor sites on the ribosome of the host cell. The A subunit catalyzes the transfer of adenine, ribose, and phosphate from NAD to EF-2 (ADP ribosylation), inactivating EF-2 and turning off inhibiting protein synthesis. DT causes local destruction at the site of membrane formation and may be also absorbed into the bloodstream and distributed, resulting in systemic complications including demyelinating neuritis and myocarditis. The tox gene that encodes DT is present in β and ω corynephages, and DT is only produced by C. diphtheriae isolates that harbor tox+ corynephages. Although the tox gene is part of the phage genome, the regulation of DT expression is under bacterial control, as the corresponding iron-sensing regulator DtxR is encoded by a gene on the C. diphtheriae chromosome [2,3,4].
Cutaneous diphtheria is the most common nonrespiratory clinical manifestation of infection due to C. diphtheriae strains. The disease is characterized by the presence of shallow skin ulcers, usually chronic, which can occur anywhere on the body, mostly on the legs, feet, and hands. This type of diphtheria may cause pain, redness, and swelling similar to other bacterial cutaneous infections. Cutaneous diphtheria is likely to be diagnosed less quickly than respiratory infection due to the nonspecific clinical appearance and often coinfections with pathogens, mostly Staphylococcus aureus and Streptococcus pyogenes. Cutaneous diphtheria frequently occurs in warm tropical climates and is normally associated with colonization of preexisting skin lesions, including surgical wounds, burns, impetigo, psoriasis, leishmaniotic ulcers, and insect bites [5, 6].
In the United Kingdom, cases of travel-related cutaneous diphtheria were reported, including patients with high diphtheria vaccination coverage and from tropical countries. The authors emphasized that with increasing travel to and from diphtheria-endemic countries, more cases may occur. These lesions are an important reservoir of infection and can cause respiratory and cutaneous infections in contacts as well as outbreaks. In several outbreaks, secondary transmission has been higher in contacts of patients with cutaneous infection than in those with respiratory tract infection. Cutaneous diphtheria may also cause greater environmental contamination, through dust and fomites. The potential for secondary transmission leads to a large number of contacts requiring follow-up, especially children at school [6]. Therefore, awareness of clinicians and microbiologists of the importance of obtaining swab specimens from any chronic nonhealing skin lesions in patients who have traveled to or a disease-endemic area is necessary, especially in tropical countries. Wound swab samples from these patients should be examined for diphtheria toxin-producing Corynebacterium species. Early diagnoses and reporting are crucial to trigger effective public health control measures (skin ulcers, which can occur anywhere on the body and are usually chronic) [5,6,7,8].
In many countries, diphtheria is considered an infrequent disease once there are treatment and diphtheria toxoid-containing vaccines to prevent it. Since 1990, diphtheria reemerged in the Russian Federation and spread to all Newly Independent States (NIS) and Baltic states. Awareness of characteristics of the largest diphtheria epidemic in the last decades that seized several European countries should be used to help predict the spread of future epidemics. The epidemic demonstrated conclusively the potential susceptibility of adults to diphtheria in the vaccine era. Important characteristics included, among several other factors, the emergence of distinct epidemic clonal group, a progressive spread of disease from urban centers to rural areas. However, epidemic diphtheria outbreaks remain poorly understood and continues to challenge industrialized and developing countries. Higher risk of acquiring C. diphtheriae infections and potentially life-threatening complications may be possible related to inadequately immunized or unimmunized conditions of persons and travelling from/to countries with endemic diphtheria [5, 9,10,11,12].
Although immunization is one of the most successful and cost-effective health interventions known, there are still many regions of the world with low vaccine coverage. Diphtheria caused by C. diphtheriae is still endemic worldwide, mostly among developing countries, including Nigeria, Venezuela, India, and Brazil. Diphtheria resurgence or epidemic outbreaks remain an important cause of morbidity and mortality that may occur in places where vaccination programs are not maintained or there is a proportion of adults susceptible to disease due to decline in antibody levels provided by vaccine, especially in low socioeconomic and health conditions areas (Fig. 1.1) [14,15,16,17,18,19,20,21]. Previous investigations reported an epidemic outbreak in Dhule, a predominantly tribal and rural district in Northern Maharashtra, India, with diphtheria cases mostly observed among adolescents (10–15 years), despite poor immunization coverage (below 50%) [17]. A diphtheria outbreak was also verified in villages of three municipalities of Maranhão, a northern state from Brazil. Most cases occurred in partially or completely immunized patients, including pharyngitis without pseudomembrane formation [22].
Diphtheria global immunization 1980–2019. Global coverage from three doses of a diphtheria toxoid-containing vaccine (DTP) at 85% in 2019 [13]
Clinical features of diphtheria in partially vaccinated patients may still be similar to those that were observed in the pre-vaccine era. However, mass vaccination has also altered clinical features of some diphtheria cases, independent of immunization status and age of individuals. Therefore, health professionals should be aware of the possibility of atypical cases of DT-producing C. diphtheriae infections, including pharyngitis without pseudomembrane formation. Cases of coincidental respiratory diphtheria with infectious mononucleosis were also reported [5, 17, 22, 23].
C. diphtheriae has been increasingly reported not only as the etiological agent of diphtheria but also as the causative agent of atypical invasive infections. C. diphtheriae was originally characterized as an extracellular pathogen with local growth pharynx mucosa. During the last decades, cases of atypical and/or invasive infections caused by both non-DT-producing and DT-producing C. diphtheriae strains have been reported, such as pneumonia, arthritis, endocarditis, bacteremia, and catheter-related infections, including cancer patients, leading patients to death in varied opportunities independent of age and sex. Septicemia, renal failure, and/or arthritis are frequently reported in patients with C. diphtheriae endocarditis [24,25,26,27,28,29,30].
C. diphtheriae is usually transmitted by respiratory droplets, direct contact, and fomites when individuals are at home or during occupational activities, especially in laboratory and hospital environments. In 1941, during the pre-vaccine era, the occurrence and persistence of diphtheria bacilli in floor dust of hospital wards and the resultant contamination of the air were verified. DT-producing C. diphtheriae strains were found capable to survive in dust and clothing for an extended period of time [31]. Once considered a strictly human pathogen, C. diphtheriae strains have been found to be able to infect animals, including cows, horses, and cats [32,33,34,35].
Changes of varied aspects in the epidemiology of diphtheria pathogens have been occurring worldwide. Starting in the middle of the 1980s, DT-producing and non-DT-producing Corynebacterium ulcerans have been increasingly identified as the etiologic agent of diphtheria of zoonotic nature and extrapharyngeal infections in varied industrialized and developing countries, including immunized individuals in the American continent [8, 12, 36,37,38,39,40]. Similar to C. diphtheriae strains, C. ulcerans strains were found to produce clinical syndromes of the lower respiratory tract, such as pneumonia and pulmonary granulomatous nodules, independent of DT production [36, 41,42,43,44]. At first, zoonotic diphtheria cases were mainly restricted to rural populations and associated with contact with dairy cattle and consumption of unpasteurized dairy products. Lately, C. ulcerans has been increasingly isolated as emerging zoonotic agent from companion animals such as cats and dogs. Therefore, there is a potentially large reservoir of infection with little knowledge about the risks of zoonotic transmission, since C. ulcerans strains were already found among animals from farms, domestic and natural settings [8, 37,38,39, 45].
Since the epidemic in European countries during the 1990s, the number of diphtheria cases due to C. ulcerans was found to exceed the number of reported cases related to C. diphtheriae [45]. Although C. ulcerans have been increasingly recognized in several industrialized countries as an emerging zoonotic pathogen, its capacity to cause disease in humans, including among the inhabitants of urban centers, is still often neglected [38]. Detection of C. ulcerans strains in Canada, during the period of 2006–2019, showed that 77% of the isolates were from humans and mostly obtained from cutaneous sites and 23% were from animals – mink (lung), dog (ear), and cat and horse (abscess and skin) – comprising 45% DT-producing strains [39]. In Brazil, a case of concurrent zoonotic diphtheria by C. ulcerans and infectious mononucleosis (IM) was first reported in the literature [46]. Moreover, a case of fatal pulmonary disease caused by a unusual penicillin and clindamycin resistant non-DT-producing C. ulcerans disseminated from primary nonhealing lesions on lower legs was documented. Both legs of the elderly patient with C. ulcerans-invasive infection presented skin ulcers covered by yellowish membranes [36]. Previous studies also reported the isolation of C. ulcerans strains from nares and/or skin wounds of asymptomatic dogs (companion dogs or kept in animal shelter), from Duque de Caxias and Niterói cities, located at the metropolitan area of Rio de Janeiro [37, 47].
Similar to C. ulcerans, Corynebacterium pseudotuberculosis is also a zoonotic etiologic agent and also considered a public health concern. C. pseudotuberculosis is a diphtheria toxin-producing pathogen of medical, veterinary, and biotechnological interest that mainly affects small ruminants, causing caseous lymphadenitis (CLA), throughout the world and generates significant economic losses. Sheep and goats are the most common animals infected within the broad spectrum of hosts in which C. pseudotuberculosis causes clinical disease. This zoonotic pathogen may also infect bovines, pigs, and equines. Therefore, contamination of meat and milk by C. pseudotuberculosis may possibly occur, putting children and adult consumers at risk. However, human infections due to C. pseudotuberculosis remain apparently rare and have been mostly reported among those with close contact to animals, including farm workers and travelers to rural areas [48,49,50].
In many countries, routine procedures for identification of DT-producing Corynebacterium spp. are uncommonly undertaken by diagnostic laboratories due to a low prevalence of diphtheria cases alongside the difficulties of diagnosis through conventional biochemical tests and the ever-increasing need for cost-effectiveness. This fact contributes to justify the low number of reported cases of human infections by zoonotic C. pseudotuberculosis strains over the years [49].
In a previous investigation of zoonotic potential of C. pseudotuberculosis, a summary data from all 33 cases of human infections reported over a period of 42 years (from 1966 to 2008) showed that a main group of non-DT-producing C. pseudotuberculosis strain-infected patients presented a characteristic of lymphadenopathy. Profiles of most of these patients included adult males, 21–40 years old, and previously exposed to raw milk or meat, farm animals (mostly sheep), and/or rural areas. Only two cases involved clinical presentations other than the characteristic lymphadenopathy: from the United States, a 28-year-old (male) veterinary student who worked with equines, diagnosed with eosinophilic pneumonia, and from China, a 63-year-old (male) with ocular infection post-retinal reattachment intervention. Until the present moment, only one case of human infection due to DT-producing C. pseudotuberculosis was reported. The zoonotic pathogen was isolated in the United Kingdom from the aortic root vegetation of an intravenous drug user with endocarditis [50,51,52,53].
3 Treatment and Prevention of DT-Producing Corynebacterium spp. Infections
Antibiotics are needed to kill DT-producing Corynebacterium spp., eliminate diphtheria toxin production, limit carriage that may persist even after clinical recovery, and prevent further transmission from asymptomatic carriers and colonization of close contacts. Penicillin and erythromycin have long been the drugs of choice for the eradication of DT-producing strains of Corynebacterium spp. The World Health Organization (WHO) has recently added azithromycin as part of the standard antibiotics for these pathogens. However, the increasing problems of resistance to penicillin, oxacillin, erythromycin, and other drugs including rifampicin, tetracycline, and clindamycin are examples of challenges confronting both industrialized and developing countries. Resistance to ß-lactams should also be considered in invasive infections, since failure to eliminate C. diphtheriae in cases of endocarditis treated with penicillin has been reported. Data emphasize the need for a continuous survey of antibiotic susceptibility for these pathogens, especially in tropical and developing countries where diphtheria is endemic and invasive infections may occur [20, 22, 28, 46, 54,55,56,57].
In cases of classic and zoonotic diphtheria, patients with severe infections should be immediately admitted to a hospital intensive care unit and given diphtheria antitoxin (DAT), consisting of antibodies isolated from the serum of horses that have been challenged with diphtheria toxin. Since antitoxin does not neutralize toxin that is already bound to tissues, delaying its administration increases risk of death. Therefore, the decision to administer diphtheria antitoxin is based on clinical diagnosis, and should not await laboratory diagnosis [20]. Administration of diphtheria vaccine is recommended during convalescence because diphtheria infection does not always confer immunity [53].
In the early 1880s, C. diphtheriae was first visualized in stained specimens from pseudomembranes and shown to be the cause of diphtheria isolated by bacteriologists Edwin Klebs and Friedrich Loeffler. In 1890, Emil von Behring isolated the first diphtheria antitoxin from blood samples of an infected horse. A few years later, William H. Park and Anna W. Williams isolated a C. diphtheriae strain that produced an unusually large amount of diphtheria toxin, later named the Park-Williams no. 8 (PW8) strain. Since the 1920s, a diphtheria toxoid vaccine has been produced from diphtheria toxin treated with formalin to inactivate the toxicity and to maintain the immunogenicity of the protein. C. diphtheriae PW8 is lysogenized by two copies of corynephage ωtox+, suggesting that the enhanced DT synthesis is due to a gene dosage effect of the tox gene [3].
Protection against diphtheria is mainly due to the development of neutralizing toxin antibodies. Diphtheria antitoxin production, primarily of IgG type, can be induced by absorption of native toxin during clinical infection or in the carrier state or by immunization with diphtheria toxoid. It is believed that a circulating diphtheria antitoxin level of 0.01 IU/ml, as determined by the neutralization test in animals or in cell culture, provides clinical immunity against disease. The outcome of revaccination of adults depends on several factors, including the immunization schedule, potency, and time since the last dose of toxoid [58, 59]. In developing countries where diphtheria is endemic, the process of maintaining immunity usually operates through natural mechanisms, including frequent skin infections caused by C. diphtheriae. Nowadays, adults might become susceptible to diphtheria due to reduced opportunities of subclinical infections. Since diphtheria infection may also occur among previously vaccinated persons, the immunity gap observed among adults should be closed by regular diphtheria boosters [9, 60,61,62].
4 Identification of DT-Producing Corynebacterium spp. in Diagnostic Laboratories
Accurate and fast diphtheria laboratory diagnosis is not only a matter of acute patient management but also an important issue in public health due to international notification and management requirements, and there is an urgent need for a reliable, robust, and fast laboratory method for diagnosing DT-producing Corynebacterium spp., especially in the light of the continuing loss of laboratory expertise even in national reference laboratories for diphtheria [5, 7, 8, 22, 63].
Phenotypic characterization of C. diphtheriae, C. ulcerans, and C. pseudotuberculosis may be performed by conventional biochemical assays and semiautomatized systems, including API Coryne System (bioMérieux). However, it takes at least 16 hours after isolation of suspicious colonies from screening plates of catalase-positive irregular Gram-positive bacilli (IGPB) [7, 22, 29, 36, 64].
In many countries, routine procedures for identification of C. diphtheriae and C. ulcerans are not commonly undertaken by diagnostic laboratories due to a low prevalence of diphtheria cases in recent years alongside the ever-increasing need for cost-effectiveness. Consequently, many diagnostic laboratories have suspended screening for diphtheria etiologic agents, further increasing the potential for missed and delayed diagnoses. Therefore, screening tests remain currently essential for the presumptive identification of these pathogens in clinical microbiology laboratories [5, 8, 22, 28, 65].
The use of DNase screening test provided a substantial improvement in the existing standard identification algorithm of DT-producing Corynebacterium spp. of routine diagnostic laboratories. DNase assays have been useful for differentiating DNAse-positive C. diphtheriae and C. ulcerans from DNAse-negative C. pseudotuberculosis and other suspected pathogenic corynebacteria, particularly in the surveillance of cases of diphtheria, asymptomatic carriers, and invasive infections in endemic or epidemic areas with unfavorable economic conditions [64]. The reverse CAMP test is particularly a screening assay effectively used as part of the identification of C. ulcerans and C. pseudotuberculosis zoonotic pathogens. A reverse CAMP test is based on the inhibition of hemolytic activity of beta-hemolysin from S. aureus through the production of phospholipase D by C. ulcerans and C. pseudotuberculosis [36].
The application of molecular techniques for the identification of bacterial pathogens has been expanded for use in clinical microbiology laboratories. Molecular procedures have been also proposed for the identification of Corynebacterium species. Improvements should become widely available for the rapid and precise detection of DT-producing Corynebacterium spp., including direct analysis of swabs and other clinical samples, as already done with C. diphtheriae and C. ulcerans in some laboratories [66,67,68].
5 MALDI-TOF Assays
Matrix-assisted laser desorption ionization-time-of-flight mass spectrometry (MALDI-TOF MS) is one of the most recently established technologies used for species identification based on the protein composition of microbial cells. MALDI-TOF MS has been increasingly applied worldwide in routine analysis of clinical microbiology laboratories due to easy procedure, rapid results (15 minutes), and accurate identification of several bacterial species, including misidentified pathogens in specific clinical specimens. MALDI-TOF MS became a powerful tool that has initiated a revolution in the clinical microbiology laboratory for identification of nosocomial pathogens, including Corynebacterium spp. Misidentification of some human-pathogenic clinical isolates commonly occurs when using conventional and commercially available methods in microbiology laboratories. The ability to rapidly identify bacterial species, including rarely described as pathogens in specific clinical specimens, may help to study the clinical burden resulting from the emergence of these species as human pathogens and MALDI-TOF MS may be considered an alternative to DNA-based methods in clinical laboratories. Due to the difficulties of diagnosis in the laboratory routine through conventional biochemical tests, MALDI-TOF MS also represents an important alternative method for the identification of C. diphtheriae, C. ulcerans, and C pseudotuberculosis strains [39, 46, 69,70,71].
6 DNA-Based Methods
The timely and precise diagnosis of corynebacterial infections, especially those involving DT-producing strains, is indispensable for the patient management and for establishment of surveillance and control strategy of the disease. Consequently, different molecular methods, such as end-point and real-time PCR (polymerase chain reaction), have been used since the 1990s for the characterization of C. diphtheriae, C. ulcerans, and C. pseudotuberculosis [7, 49, 66, 68, 72,73,74,75,76,77,78]. Since the investigation of the toxigenic potential of clinical isolates is one of the most critical aspects of diphtheria diagnosis, conventional end-point PCR assays targeting the tox gene were the first to be developed. Subsequently, the tox gene detection was combined with species identification PCR targets, as dtxR (diphtheria toxin repressor gene) from C. diphtheriae, in multiplex assays [7, 49, 66, 68, 72, 73]. Since detection of the tox gene only provides presumption of toxigenicity, additional phenotypic investigations such as Elek test and Vero cell cytotoxicity assays have been currently used to demonstrate DT production by Corynebacterium strains [7, 66, 74].
Multiplex PCR represents a fast, simple, and reliable methodology for identification and differentiation between DT-producing and non-DT-producing strains of C. diphtheriae, C. ulcerans, and C. pseudotuberculosis. In Brazil, an mPCR protocol was developed, and it has been used for clinical diagnosis and epidemiological and virulence research, during the last decade. Direct analyses of swabs and other clinical samples have also been done. Brazilian mPCR allows the detection of tox gene from potentially DT-producing Corynebacterium spp., in addition to 16S rRNA from both C. pseudotuberculosis and C. ulcerans, pld from C. pseudotuberculosis, dtxR from C. diphtheriae, and rpoB from Corynebacterium spp. [22, 27, 37, 46, 49].
Real-time PCR instruments are increasingly common in public health laboratories, mostly in industrialized countries. Real-time PCR (qPCR for quantitative PCR) presents some advantages than classical PCR, including faster data collection, low contamination risks, and high sensitivity, especially for pathogen detection in host carriers and clinical samples which often contain components that inhibit PCR [69, 79].
During the last decades, different qPCR assays have been developed for detection of the tox gene and/or identification of C. diphtheriae, C. ulcerans, and C. pseudotuberculosis directly from clinical samples. However, similarly to end-point PCR, currently available qPCR assays allow only the detection of tox gene from Corynebacterium spp. strains. However, the confirmation of DT expression still requires phenotypic investigations [66, 69, 75, 80, 81].
7 rpoB Gene Sequencing Technique
The genus Corynebacterium is a heterogeneous group of species comprising human and animal pathogens and environmental bacteria. It is defined on the basis of several phenotypic characters and the results of DNA-DNA relatedness and, more recently, 16S rRNA gene sequencing. The rpoB gene, encoding the beta-subunit of RNA polymerase, has emerged as a core gene candidate for phylogenetic analyses and identification of corynebacteria, especially when studying closely related isolates. However, the 16S rRNA gene is not polymorphic enough to ensure reliable phylogenetic studies and needs to be completely sequenced for accurate identification. Previous studies verified that higher proportions (91%) of corynebacterial isolates were positively identified by partial rpoB gene determination than by that based on 16S rRNA gene sequences [82,83,84].
8 Diphtheria Toxin-Producing Group Becoming Diverse: A Novel C. diphtheriae Complex
C. diphtheriae was historically classified into four biovars – gravis, mitis, intermedius, and belfanti – based on biochemical phenotypic testing [4, 85]. Recent investigations documented C. diphtheriae to be genetically heterogeneous and that genomics does not support the use of biovars to reliably classify diphtheria bacilli isolates, since C. diphtheriae strains within a certain biovar were found to be genetically more distant than between biovars [3, 86, 87].
A switch in populations causing endemic infections from DT-producing to non-DT-producing isolates in the 1990s and the 2000s and other countries with vaccination coverage has been documented as a direct consequence of the large-scale use of diphtheria toxoid. In Brazil, several C. diphtheriae isolates were found capable of degrading sucrose, a phenotypic characteristic rarely described in other parts of the world. Results of rpoB sequence analysis confirmed all sucrose-fermenting isolates as C. diphtheriae species. Sucrose-fermenting and DT-producing C. diphtheriae strains were predominantly isolated from human respiratory tract of diphtheria patients. However, cases of endocarditis due to sucrose-fermenting C. diphtheriae phenotypes were also observed in South American countries [28, 88, 89].
During the 1980s and 1990s, studies dealing with C. diphtheriae biovar belfanti were occasionally reported in literature. C. diphtheriae biovar belfanti were mostly isolated from human respiratory samples. In Brazil, the case of pulmonary infection in a cancer patient was reported [90]. The tox gene, which codes for diphtheria toxin, was infrequently reported in isolates of biovar belfanti [7, 90,91,92,93].
Taxonomic status of C. diphtheriae biovars has been increasingly investigated [91, 93, 94]. Phylogenic analysis described two lineages of non-DT-producing C. diphtheriae biovar belfanti obtained from 18 countries, covering a time period from 1957 to 2006 [91]. In France, the number of non-DT-producing C. diphtheriae biovar belfanti increased between 1977 and 2011, and it is the most frequent biovar recovered in recent years. Non-DT-producing belfanti isolates were mostly isolated from human respiratory samples, including from a woman presenting with rhinitis. However, there were also found belfanti phenotypes isolated from blood, skin, and bone lesions. Phylogenic analyses of French non-DT-producing C. diphtheriae biovar belfanti human isolates were distributed among three distinct lineages. Almost all belfanti isolates belonged to a single clonal complex. A third new lineage was composed of a single clinical isolate (rhinitis) and phylogenetically distant from other two partially studied belfanti lineages from France [91, 94].
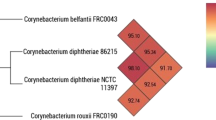
Recent investigations by genomic sequencing, biochemical, and chemotaxonomic analyses indicated that C. diphtheriae biovar belfanti represents a branch that is clearly demarcated from C. diphtheriae biovar mitis and gravis. Data, including the inability to reduce nitrate, allowed to differentiate biovar belfanti from other C. diphtheriae strains. Consequently, the name Corynebacterium belfantii sp. nov. for the group of nitrate-negative strains, previously considered as C. diphtheriae biovar belfanti, was recently reported [95].
In a further study, it was proposed that C. diphtheriae taxon should be subdivided into two subspecies, C. diphtheriae subsp. diphtheriae and nitrate-negative C. diphtheriae subsp. lausannense [93]. However, given that C. belfantii was validly published a few months before the taxonomic proposal C. diphtheriae subsp. lausannense was validated, the latter subspecies was suggested to be a heterotypic synonym of C. belfantii [92].
Most recently, a group of clinical isolates previously identified as C. diphtheriae biovar belfanti strains isolated from human cutaneous or peritoneum infections and from one dog were characterized by genomic sequencing, biochemical analysis, and MALDI-TOF assays as Corynebacterium rouxii sp. nov. for the novel group. Phenotyping data revealed an atypical negative or heterogeneous intermediate maltose fermentation reaction for both human and animal isolates. Atypical, maltose-negative, and DT-negative C. diphtheriae biovar belfanti isolated from domestic cats, including with severe otitis, was also previously described in the United States [33, 92].
C. pseudotuberculosis are classified into biovars equi and ovis based on the ability to convert nitrate to nitrite, due to genetic characteristic that includes the presence of the nitrate reduction operon: equi, nitrate-positive strains, and ovis, nitrate-negative strains. Disease caused by C. pseudotuberculosis biovars has different clinical manifestations in the susceptible hosts, and biovar identification is important for understanding the epidemiology of infection and consequently for disease control. C. pseudotuberculosis biovar equi strains are etiologic agents of ulcerative lymphangitis in horses, cows, camels, buffaloes, and occasionally humans. C. pseudotuberculosis biovar ovis strains are the causative agents of caseous lymphadenitis (CLA) in small ruminants, mostly ovine and caprine herds. CLA causes important economic losses in ovine and caprine herds by reducing wool, meat, and milk production. Cases of human infections due to non-DT-producing C. pseudotuberculosis biovar ovis were more frequently described worldwide than cases by biovar equi. Lymphadenitis in human hosts due to C. pseudotuberculosis was also reported in literature, mostly occurring in those who were visitors or were occupationally exposed to animals in rural areas, especially sheep farms [78, 96, 97].
Only one case of DT-producing C. pseudotuberculosis was described in literature. The zoonotic pathogen was isolated from the aortic root vegetation of an intravenous drug user with endocarditis; this patient had no history of animal contact, and no possible source of infection was identified. This isolation occurred in the United Kingdom and biovar was not reported [53].
9 Pathogenomics of Potentially DT-Producing Corynebacterium spp.
Diphtheria toxin is one of the best investigated bacterial toxins and a leading virulence factor of toxigenic C. diphtheriae and C. ulcerans strains. Different investigations demonstrated that tox genes of C. diphtheriae strains showed similar nucleotide sequence identity. Phylogenetic analyses of C. ulcerans revealed diverse diphtheria toxin suggesting that C. ulcerans tends to acquire mutations more frequently than C. diphtheriae. Two possible explanations for this phenomenon are that C. ulcerans strains are maintained by various animals and have a phage-independent pathway to acquire the DT-encoding gene, increasing its diversity compared with C. diphtheriae [2, 3, 98, 99].
The diphtheria toxin repressor DtxR is known as an iron-dependent regulator that controls the transcription of the diphtheria toxin gene tox and a complex gene regulatory network involved in iron homeostasis. Variations of dtxR genes and in the regulatory network of DtxR might lead to differences in iron supply of the bacterial cell, thereby influencing the expression of the tox gene and the virulence of DT-producing Corynebacterium spp. [3, 100, 101]. Therefore, naturally occurring diversity of tox genes and variations on the expression of diphtheria toxin due to dtxR regulatory activities may exert influence on efficacy of diphtheria toxoid vaccine and diphtheria antitoxin for preventing and treating infections caused by DT-producing Corynebacterium spp. pathogens [3, 5, 99, 100].
The occurrence of diphtheria among immunized persons, as well as the increasing frequency of cases of atypical and invasive diseases, caused by non-DT-invasive clones also points the relevance of multiple virulence factors of the potentially DT-producing Corynebacterium spp. [26, 30, 37, 102,103,104,105]. In previously reported cases of invasive infections, non-DT-producing C. diphtheriae and C. ulcerans strains were found capable of expressing additional proteins with cytotoxic effects similar to Shiga-like toxins, characterized as ribosome-binding proteins (Rbps). Experimental evidence for the cytotoxic function of Rbps toxins were provided by the interaction of C. diphtheriae and C. ulcerans wild-type, mutant, and complementation as well as overexpression strains with invertebrate model systems, Caenorhabditis elegans and Galleria mellonella, and on various animal and human macrophage and epithelial cell lines, including Vero cells [28, 36, 105, 106].
The dermonecrotic phospholipase D (PLD) exotoxin may be also produced by both zoonotic pathogens that have been investigated as a prominent virulence factor, especially for C. pseudotuberculosis. The pld gene encoding the phospholipase D is included among the subset of genes homologous for C. pseudotuberculosis and C. ulcerans species. Studies with C. pseudotuberculosis strains with inactivated PLD have convincingly demonstrated the necessity of PLD for establishment of diseases in animals, including caseous lymphadenitis [8, 46, 98, 107, 108]. However, a case of diphtheria due to C. ulcerans strain that is unable to express both PLD and DT activities was recently reported. These data emphasize that virulence mechanisms and pathogenic potential of C. ulcerans species may arise independent of PLD and DT production. C. ulcerans virulence potential and zoonotic pathogenicity traits need further investigation [36,37,38, 46, 98, 104, 105, 109].
Basic mechanisms and specific virulence determinants, other than DT, involved in the pathogenic potential of C. diphtheriae have been investigated for almost half a century [110]. Since 2003, data from whole-genome sequencing (WGS) of C. diphtheriae , including a pangenomic study with Brazilian clinical isolates from cases of classical diphtheria, endocarditis, and pneumonia, involving sucrose-fermenting C. diphtheriae strains, exposed horizontal gene transfer of virulence factors, such as adhesins, fimbrial proteins, and iron uptake systems [3, 111].
In an attempt to further investigate mechanisms that promote C. diphtheriae survival within different environmental conditions, infection and dissemination through host tissues, several features have been of concern [112,113,114]. A putative determinant (CDCE8392_813 gene), coding for tellurite (TeO32−) –resistance (TeR) was detected, and the influence on virulence attributes of C. diphtheriae strains was verified. Tellurium (Te) is a metalloid that exists as a trace component in natural environments. Although TeO32− is toxic to most microorganisms, TeR bacteria, including C. diphtheriae, exist in nature. The presence of TeR determinants in pathogenic bacteria might provide selective advantages in the natural environment. The C. diphtheriae TeR-disrupted mutant strain expressed increased susceptibility to TeO32− and reactive oxygen species (hydrogen peroxide) but not to other antimicrobial agents. Moreover, TeR determinants contributed to the survival of C. diphtheriae strains by using in vivo and in vitro models of infection [115].
The ability of biofilm formation on varied biotic and abiotic surfaces was also investigated. Low-dose antibiotics was reported to favor biofilm formation by C. diphtheriae, similar to observations for other human pathogens. C. diphtheriae strains expressed higher cell-surface hydrophobicity and biofilm formation on different abiotic surfaces in the presence of penicillin and erythromycin. Moreover, C. diphtheriae was also recognized as a potential cause of catheter-related infections, independent of DT production [29, 89, 102, 115,116,117,118].
C. diphtheriae was also found to express the ability to invade and survive within different types of human cells and the capacity to cause invasive bloodstream infections. Systemic complications of C. diphtheriae bacteremia are not unusual and include endocarditis, joint infections, and peripheral embolic disease. A strain-dependent ability to induce osteomyelitis by DT-negative C. diphtheriae strains was probed by an in vivo assay using Swiss Webster mice, as first reported for C. ulcerans [27, 104].
Mechanisms of interaction with different cell types have been also investigated, such as erythrocytes, macrophages, and endothelial and epithelial cells [26, 27, 30, 102, 115, 119]. The pathogenic role of aggregative-adhering properties in C. diphtheriae-invasive disease was investigated. C. diphtheriae biovar mitis and gravis, isolated from cases of endocarditis, expressed aggregative adherence (AA) patterns to human epithelial cells. The predominance of localized (LA) and diffuse adherence (DA) patterns have been reported for C. diphtheriae strains mostly isolated from throat and skin lesions [26, 64]. C. elegans nematodes have been also applied as an infection model system for C. diphtheriae and C. ulcerans with invasive phenotypes [102, 115, 120,121,122].
During years of research, the adhesive properties of C. diphtheriae strains have been already defined as multifactorial, relying on specific and general mechanisms. Functions and mechanisms of action of fimbriae; non-fimbrial adhesins – 67-72p (DIP0733) and DIP2093 – trans-sialidase; hydrophobins; and sugar residues are already recognized at different levels, especially how they jointly participate in the adherence to the host cells and in the colonization of these cells during bacterial infection [113, 117, 123,124,125].
Genomic analysis of C. diphtheriae revealed the identification of three distinct pili clusters (spaABC, spaDEF, spaGHI) together with five sortase-encoding genes (srtA–E), which are essential for pilus assembly. Adherence rates are not strictly correlated with pili formation, and the pili repertoire of C. diphtheriae strains is highly variable. spaA-type is the pilus mostly detected among C. diphtheriae strains. As shown by genome comparisons, it is necessary to investigate various isolates on a molecular level to understand and to predict the colonization process of different C. diphtheriae strains [3, 123, 124, 126].
The DIP0733 was initially described as a non-fimbrial 67-72p protein responsible for the adherence of C. diphtheriae strains to human erythrocytes. Further studies demonstrated DIP0733 protein as a microbial surface component recognizing adhesive matrix molecule (MSCRAMM). The influence of DIP0733 in C diphtheriae interaction with human epithelial cells and macrophages in addition to the ability to induce host cell death, giving a signal for apoptosis in the early stages of infection, was also reported. These findings support the idea that DIP0733 is a multifunctional virulence factor of C. diphtheriae that enhances the ability to spread throughout the whole human body via the bloodstream [125, 127, 128].
10 Molecular Typing Methods for DT-Producing Corynebacterium spp.
Molecular typing methods are expected to be reproducible with high discriminatory power, stable and cost-effective, and easy to perform and interpret. Several typing methods have been developed to investigate epidemiological relationship of strains in disease outbreaks. Investigation of outbreaks is possible due to phylogenetic analysis of origin of strains and patterns of local and global dissemination over the years, among other features [23, 129, 130].
In the past, epidemiological surveillance of diphtheria was limited, depended largely on phenotypic characterization of strains, first by differentiation into biotypes and subsequently by serotyping and phage and bacteriocin typing. At the request of the World Health Organization Regional Office for Europe, the European Laboratory Working Group on Diphtheria (ELWGD) was formed in July 1993 because of the re-emergence of diphtheria to epidemic levels in the Russian Federation and Newly Independent States. The main objectives were to form a network of laboratories for microbiological surveillance, to standardize laboratory diagnostic methods, and to understand the molecular epidemiology and characteristics of C. diphtheriae strains at that time. In 2001, the network was expanded to become a Diphtheria Surveillance Network (DIPNET) (http://www.dipnet.org) concerned with epidemiological and microbiological aspects of diphtheria and other infections caused by other DT-producing Corynebacterium species, including Brazilian scientists. The main purpose of DIPNET is to establish a Pan-European network of expertise for the prevention of diphtheria and other related infections across the EU Member States and beyond. Among the specific objectives of DIPNET are to (i) determine the disease prevalence and characteristics of toxigenic and nontoxigenic C. diphtheriae and C. ulcerans in a variety of populations with emphasis upon higher-risk countries, (ii) expand the DIPNET external quality assurance schemes for laboratory diagnosis to include epidemiological typing and serological immunity, and (iii) develop novel tools for integrated molecular epidemiological characterization so as to gain a clearer understanding of the spread of epidemic clones throughout the WHO European Region [3, 23, 108, 131,132,133,134,135,136].
Over the years, several molecular typing methods have been applied for C. diphtheriae, including ribotyping, pulsed-field gel electrophoresis (PFGE), random amplification of polymorphic DNA (RAPD), multilocus enzyme electrophoresis (MEE), multilocus sequence typing (MLST), and spoligotyping. Some of these genotyping approaches have been also used for epidemiologic investigations of both C. ulcerans and C. pseudotuberculosis zoonotic pathogens [92, 137,138,139,140,141].
10.1 Ribotyping Assays
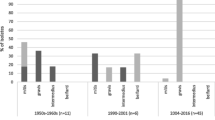
Ribotyping methods , based on restriction patterns of ribosomal RNA genes, had been previously considered the gold standard procedure for C. diphtheriae epidemiological surveillance, due to its high discriminatory power, reproducibility, and optimal typeability [142,143,144]. At the beginning, two ribotypes were identified by the restriction enzyme BstEII and were named G and M, since they seemed to be related to biovars gravis and mitis, respectively. However, G ribotypes were found in C. diphtheriae biovar mitis strains, and M ribotypes were also found in C. diphtheriae biovar gravis strains. The ten most frequent ribotypes of C. diphtheriae strains from Russia during the period of 1984 to 1996 were shown in Fig. 1.2. During an accurate interlaboratory comparative analysis, a revised nomenclature for the designation of ribotypes was proposed. Prefixes “G” and “M” were both replaced by prefix “D” (diphtheria). In 2004, an international ribotyping database was established at the Pasteur Institute and several collaborating laboratories supported by ELWGD/WHO. The ribotype nomenclature was revised and named using the geographical origin to reflect the location where one of the strains was isolated or studied. Eighty-six ribotypes were identified by the restriction patterns using BstEII digestion of the DNA [142, 145].
Ribotyping assay based on restriction patterns of ribosomal RNA genes. Predominant riboprofiles identified in Corynebacterium diphtheriae strains (n = 156) from Russia during the period of 1984 to 1996. Tox, diphtheria toxin-producing. G, gravis. M, mitis. (Reprinted from Popovic et al. [142])
Ribotyping methods have been also used in studies of C. ulcerans zoonotic pathogen. In Japan, DT-producing C. ulcerans strains isolated from pharyngeal swabs of two patients attended at a hospital unit were indistinguishable by PFGE analysis and distinguished by ribotyping methods [146]. In 2016, a case of asphyxia death due to pseudomembrane caused by a DT-producing C. ulcerans strain, also recovered from the patient’s domestic cat, was reported in a Japanese woman. Ribotyping analysis during this case in 2016 detected identical ribotype observed for C. ulcerans 0102 strain isolated from the first case in Japan during the year 2001 [57]. In a previous study, 9 different ribotypes, designated U1 to U9, were identified when evaluating 81 C. ulcerans strains, 50 of which were clinical isolates from the United Kingdom (90% toxigenic), 7 isolates from domestic cat, and the remaining from different places and sources of origin. The U1 ribotype was the predominant pattern found among human clinical isolates from the United Kingdom (20 isolates) and four isolates from other countries (Germany, France, Ukraine, and Italy). The seven domestic cat isolates also generated ribotypes found among human clinical isolates [147].
Limitations of ribotyping as a genotyping method for C. ulcerans were also reported. Genome sequence of C. ulcerans FH2016-1 isolated from the first fatal case described above was sequenced and compared with genomes of C. ulcerans strains of the first and second cases from Japan, 0102 and 0211, respectively. Although the analyses demonstrated a low variability between genomes, the isolate FH2016-1 was genetically distinct from 0102 and 0211, indicating that conventional ribotyping did not accurately reflect the strain with consequent inaccurate classification [148].
10.2 Pulsed-Field Gel Electrophoresis (PFGE) Technique
PFGE technique consists of the separation of DNA fragments with high molecularweight of bacterial genoma, obtained by using restriction enzymes [149]. PFGE technique consists of the separation of DNA fragments with high molecular weight of bacterial genoma, obtained by using restriction enzymes were used, and the SfiI restriction endonuclease was chosen, producing 18 to 25 DNA fragments ranging in size from 24 to 290 kb. In comparison with ribotyping, PFGE was not able to distinguish three ribotypes [137]. Thereafter, minor changes in the PFGE protocol were done, and both PFGE and ribotyping showed identical discriminatory ability. In that opportunity, all isolates grouped in one PFGE type were grouped in a ribotype, and vice versa. In addition to the PFGE protocol changes, some explanations seemed plausible for the results found [150]. A protocol using difference in three or more bands was used to distinguish C. diphtheriae PFGE types. In a later study, a difference in only two bands to distinguish the PFGE types of a limited number of C. diphtheriae strains was used [137, 150].
Nowadays, PFGE typing method has been scarcely used to investigate epidemiological relationship of C. diphtheriae strains in disease outbreaks. During the diphtheria outbreak in Northeastern Brazil in 2010, most of the confirmed cases occurred in partially or completely immunized children, including three fatal cases. Molecular analysis demonstrated the spread of predominant PFGE type related strains (Fig. 1.3) [22].
Pulsed-field gel electrophoresis (PFGE) types of Corynebacterium diphtheriae biovar intermedius strains isolated from children with diphtheria living in the state of Maranhão, Brazil. Lane 1, λ DNA ladder PFGE marker; lanes 2–5, PFGE type Ia (MA19, MA23, MA52, MA131 strains, respectively); lane 6, PFGE type II (MA136 strain); lane 7, PFGE type Ib (MA150 strain). Other Brazilian DT-producing C. diphtheriae strains: lane 8, profile III (sucrose-positive TR241 biovar mitis strain); lane 9, profile IV (sucrose-negative VA01 biovar gravis strain). Toxoid vaccine producer strain: lane 10, profile V (PW8 strain). (Reprinted from Santos et al. [22])
PFGE typing assays have been also used in investigations of origin, transmission, and dissemination of zoonotic C. ulcerans strains, especially in Japan. DT-producing C. ulcerans strains were isolated from pharyngeal swabs of two patients from the same hospital unit during 2001 and 2002 and were characterized by PFGE and ribotyping. The isolates could not be distinguished by PFGE ; however, ribotyping showed discriminatory results [146]. In contrast, a study of a fatal case due to a DT-producing C. ulcerans strain, PFGE analyses, and ribotyping of C. ulcerans strains from the patient and his cat belonged to the same molecular type [57].
10.3 Multilocus Enzyme Electrophoresis (MEE)
This electrophoresis technique detects amino acid substitutions capable of altering charge and structural conformation of cellular housekeeping enzymes. Each electromorph or mobility variants of the same enzyme are visualized in a starch gel matrix as bands with different migration rates. Twenty-seven enzymes are tested, and each electromorph is considered a different allele of the specific enzyme. An electromorph profile defines the electrophoretic type (ET) of each bacterial strain. The genetic distance of ET is calculated as a dendrogram generated by the average linkage method of clustering the ETs [151]. Several studies had used MEE methods to estimate C. diphtheriae genetic diversity and epidemiological features of endemic and epidemic diseases. Diphtheria epidemic in the 1990s, which initiated in the Russian Federation and dispersed to several European countries, was characterized by the simultaneous presence of different ET that were also detected in C. diphtheriae strains isolated during the pre-epidemic period. The majority of C. diphtheriae strains with D1 and D4 ribotype patterns (previously named G1 and G4) belonged to the clonal group called ET8 complex [139].
10.4 Random Amplification of Polymorphic DNA (RAPD) Assays
The RAPD technique aims to amplify random segments of DNA with single primers of arbitrary nucleotide sequence that may be used to construct genetic maps in a variety of species [152]. The RAPD assay for C. diphtheriae was recently standardized by using C. diphtheriae strains isolated worldwide. Initially, C. diphtheriae strains from Russia (1985–1994) were evaluated, and primers 3 and 4 classified the isolates previously ribotyped into 19 and 24 genotypic profiles, respectively. However, epidemic ribotypes D1 and D4 could not be differentiated by primers 3 and 4. Subsequently, 120 C. diphtheriae strains isolated from Russia (1994–1995), Kazakhstan (1996), and the Republic of Georgia (1995–1996) were included and presented RAPD profiles typical of the epidemic ribotypes D1 and D4 [153]. Difficulties in the standardization of RAPD assays for C. diphtheriae such as the use of crude DNA and different thermocyclers resulted in poor amplifications, and nonreproducible patterns were also verified [138]. RAPD assays have advantages for being simple, rapid, and inexpensive. However, RAPD assays for C. diphtheriae demonstrated low reproducibility in some opportunities [138, 150, 153].
In a previous study, DT-producing and non-DT-producing C. diphtheriae strains isolated from cases of infective endocarditis showed different RAPD profiles , demonstrating the invasive properties and circulation of these different clones in Brazil [28]. Analysis performed with purified DNA of several C. diphtheriae strains from 26 countries resulted in the differentiation of Eastern European epidemic ribotypes D1 and D4 corresponding to the RAPD profiles Rp1 and Rp4, respectively [143] (Fig. 1.4).
Thirteen RAPD profiles of C. diphtheriae isolates illustrating the differentiation of epidemic ribotypes D1 e D4 as Rp1 and Rp4, respectively. (Reprinted from De Zoysa et al. [143])
10.5 Amplified Fragment Length Polymorphism (AFLP)
AFLP technique is based on PCR amplification of restriction fragments from total genomic DNA digestion, by using generic primers that do not require prior information of the target DNA sequence. DNA restriction and ligation of oligonucleotide adapters are made to form the binding sites. Subsequently, the selective amplification of the restriction fragments occurs, and finally the fragments are visualized in gel [154]. Evaluation of the AFLP technique was verified during a study conducted with C. diphtheriae strains (n = 57) presenting nine different ribotypes. A total of ten AFLP profiles were assigned to C. diphtheriae tested strains; however, it was not able to discriminate the predominant ribotype during the Eastern European epidemic. AFLP is a PCR fingerprint method easy to perform, rapid, inexpensive, and suitable for most laboratories. Moreover, the AFLP standards are representative of the complete genome [155]. However, AFLP method was less discriminatory than ribotyping in some studies [143, 156].
10.6 Multilocus Sequence Typing (MLST) Method
MLST assays have been widely used in different countries for molecular typing of circulating C. diphtheriae strains and investigations of epidemic outbreaks. The method aims to group strains related to cloned complex after sequencing and analyzing fragments of seven constitutive genes that encode essential functions for microbial metabolism. C. diphtheriae MLST scheme uses fragments of the following housekeeping genes: ATP synthase alpha chain (atpA), DNA polymerase III alpha subunit (dnaE), chaperone protein (dnaK), elongation factor G (fusA), 2-isopropylmalate synthase (leuA), 2-oxoglutarate dehydrogenase E1 and E2 components (odhA), and DNA-directed RNA polymerase beta chain (rpoB) [91, 144].
The PubMLST website hosts a collection of open-access, curated databases that integrate sequence data with phenotype information for many microbial species, including C. diphtheriae and C. ulcerans. Currently, more than 700 categorized types are deposited (October 2020) in the PubMLST database (http://pubmlst.org/cdiphtheriae/). In 2010, the MLST scheme was proposed for C. diphtheriae by using the sequences of the 7 constitutive genes described above, with a total of 150 tested strains (toxigenic isolates n = 96) from 18 different countries, during the period of 1957–2006. The results were consistent with previous ribotyping data, which was considered the “gold standard” typing method of C. diphtheriae for many years [91].
Although MLST is recognized as a valuable fast and simple PCR-based method used for the tracking the spread of important clones and evolutionary investigation of bacteria, this methodology has some limitations, such as the identification of hypervirulent clones, since the MLST data are based on changes in the core genome, while changes in the accessory genome are responsible for C. diphtheriae virulent variants. Each ST can be represented by toxigenic noninvasive, nontoxigenic invasive, and nontoxigenic noninvasive strains [157, 158]. Furthermore, some C. diphtheriae strains with identical ST may differ in up to 290 genes; noncorrelated results between MLST and biotype tests may also occur [158, 159].
The ST-8 clone was responsible for the beginning of the Eastern European epidemic and spread of more than 157,000 registered cases of diphtheria, which resulted in approximately 5,000 deaths. Non-DT-producing profiles of ST-8 strains, previously isolated as toxigenic in Russia, were recently described in Poland. This change was attributed to the environmental pressure exerted by the increase in the number of vaccinated individuals. ST-8 continued to circulate after the epidemic period, as reported realized in Germany, which points to the persistence of ST-8 until today [158, 160, 161]. From 2016 to 2017 in Germany, there was an increase in the circulation of nontoxigenic C. diphtheriae characterized mostly by the ST-8. This ST is the most abundant found in the MLST database for C. diphtheriae and probably in Europe [160].
Since invasive infections caused by C. diphtheriae in vaccinated and non-vaccinated individuals have been reported in Brazil, a genetic relationship of C. diphtheriae strains isolated from classic diphtheria and invasive infections in Rio de Janeiro metropolitan area was investigated by using MLST. Four strains presented an atypical sucrose-fermenting ability and corresponded to new STs. Interestingly, a sucrose-fermenting C. diphtheriae biovar mitis strain, isolated in 1999 patient with endocarditis, formed a clonal complex with a DT-positive C. diphtheriae biovar mitis strain isolated in Argentina (1995) causing classic diphtheria, suggesting a C. diphtheriae biovar mitis clonal complex circulation in South America. Moreover, a sucrose-negative strain isolated from a case of endocarditis in 2003 generated an MLST profile that had been previously deposited in the database, ST128, a single locus variant (SLV) of ST-80, the clonal complex that comprises strains currently isolated in different countries including France (ST128) and Canada. These data indicated that C. diphtheriae clonal complexes comprising clinical strains related to diphtheria disease may also include invasive phenotypes [9, 28, 91, 94, 111, 162].
An MLST protocol C. ulcerans was based on protocol described for C. diphtheriae. The website PUBMLST comprises data for both species [91, 163, 164]. MLST methods have been also used in phylogenetic analyses, epidemiology, and zoonotic transmission investigations of C. pseudotuberculosis [55, 97, 163, 165].
11 In Silico-Based Approaches
Bacterial in silico typing based on repetitive DNA sequences has been also established with the objective of genomic characterization and epidemiological surveillance of diphtheria. Two approaches named CRISPR (clustered regularly interspaced short palindromic repeats) loci and VNTR (variable number tandem repeats) were investigated. In Poland, a study used the complete genome sequence of the NCTC 13129 C. diphtheriae strain to identify 75 VNTR loci, of which 14 were selected. Primers were designed, and PCR conditions were optimized to amplify the selected VNTR markers. Fourteen markers were tested and eight were considered potentially useful. This approach showed discriminatory genotyping ability for the C. diphtheriae tested strains (n = 28), but the preliminary results were not compared with other genotyping methods [166].
CRISPR-based spoligotyping, defined as a genotyping technique to identify C. diphtheriae strains at the phylogeographic level, was also described. According to the authors, this typing methodology presents a high level of discrimination and may be employed to study local epidemiology. One of the limitations is the need for expensive equipment or the use of external services, but the comparison of results between laboratories is easy, and the data generated can be compared in a database. In a study conducted with C. diphtheriae of Russia and Belarus epidemic clone, the 156 strains tested were subdivided into 45 spoligotypes. A high level of discrimination in the study of the local epidemiology of diphtheria was observed. However, the three selected CRISPR loci were not present in all C. diphtheriae tested strains, and most of them had unique spaces in the leader sequence, indicating that they evolved independently after diverging from a common ancestor [141, 167, 168].
Whole-genome sequencing (WGS) has become an essential tool for molecular epidemiology of infectious disease studies. In recent years, WGS has become the gold standard of high-resolution typing methods, allowing the understanding of the molecular epidemiology and global transmission of pathogens. Genome sequencing remains expensive to be employed in routine genotyping. Nevertheless advances in C. diphtheriae genomics concern an increasing number of complete genomes in GenBank may benefit sequence-based genotyping methods as identification of SNPS, tandem repeats (VNTRs), and CRISPR-based spoligotyping. The development of more inexpensive and discriminatory methodologies for use in epidemiological studies will be crucial in our understanding of the molecular epidemiology and carriage of C. diphtheriae and C. ulcerans [23, 87, 144].
12 Conclusions
Molecular typing methods have become essential during the analysis of studies involving epidemiology outbreaks, endemic conditions, recurrent infections, transmission, and virulence potential of C. diphtheriae and zoonotic diphtheria toxin-producing Corynebacterium species. Application of any typing method depends on the objectives of the study, the level of resolution desired (species vs. strain), and the laboratory conditions and technical expertise available. In studies of complex epidemiological situations and strain-dependent virulence mechanisms, it is recommended not to rely on a single method but to use combinations of methods for strain identification and to interpret results within the context of the epidemiological background and evolution of once acquired pathogenicity features during vaccine era [22, 23, 38, 55, 74, 97, 143, 155].
Epidemiological investigations demonstrated the prevalence of C. diphtheriae biovars gravis and mitis in Eastern Europe and most of Brazilian outbreaks, respectively. Diphtheria cases and deaths caused by C. diphtheriae biovar intermedius were also documented in previously immunized individuals in India and during the most recent diphtheria outbreak in Brazil [5, 17, 22, 169, 170]. Nowadays, molecular epidemiological investigations demonstrated the prevalence of different C. diphtheriae genotypes in specific geographic regions, including epidemic outbreak: Thailand (ST-243), South Africa (ST 379), and Malaysia (ST453) [171,172,173].
The occurrence of diphtheria among immunized persons and the increasing frequency of atypical infections caused by non-DT-producing clones indicated that other microbial factors should be used as one of the antigens in the potential vaccine development in the near future. In conclusion, molecular typing methods became a remarkable achievement in wide-ranging research to potentially DT-producing Corynebacterium species (C. diphtheriae complex), group of extremely dangerous human pathogenic species.
13 Summary
Corynebacterium diphtheriae is the leading causing agent of respiratory and cutaneous diphtheria, an acute disease with local and systemic manifestations, which remains as an important cause of morbidity and mortality in different continents. Diphtheria vaccination programs implemented in industrialized and developing countries led to an increasing number of atypical cases of diphtheria in addition to localized and systemic infections, including fully immunized adults. Changes in the clinical epidemiology and virulence features of diphtheria pathogens have been investigated. Cases of infections due to diphtheria toxin (DT)-producing and non-DT-producing C. diphtheriae and Corynebacterium ulcerans, a zoonotic pathogen, have been increasingly reported. The timely and precise diagnosis of DT-producing Corynebacterium strains is indispensable for the patient management and for establishment of surveillance and control strategy of disease. Different molecular methods, such as real-time PCR (polymerase chain reaction) and multiplex PCR, have been used for the characterization of C. diphtheriae, C. ulcerans, and C. pseudotuberculosis and the detection of the gene for DT (tox). Recent investigations by genomic sequencing and chemotaxonomic analyses reported DT-producing C. diphtheriae subsp. lausannense and Corynebacterium belfantii sp. nov. in addition to Corynebacterium rouxii sp. nov. Genotyping methods have been used as essential epidemiological tools for C. diphtheriae and C. ulcerans infection prevention and control, including pulsed-field gel electrophoresis (PFGE), random amplification of polymorphic DNA (RAPD ), and multilocus sequence typing (MLST) assays. Molecular typing methods are required in studies involving characterization, virulence potential, and susceptibility to antimicrobial agents of C. diphtheriae and C. ulcerans clinical isolates; origin, routes, and transmission of diphtheria and atypical invasive infections; and endemicity, outbreaks, recurrent infections, and trace cross-transmission caused by non-DT-producing and DT-producing Corynebacterium spp.
References
Hadfield TL, McEvoy P, Polotsky Y et al (2000) The pathology of diphtheria. J Infect Dis 181:S116–S120. https://doi.org/10.1086/315551
Holmes RK (2000) Biology and molecular epidemiology of diphtheria toxin and the tox gene. J Infect Dis 181. https://doi.org/10.1086/315554
Trost E, Blom J, de Soares SC et al (2012) Pangenomic study of Corynebacterium diphtheriae that provides insights into the genomic diversity of pathogenic isolates from cases of classical diphtheria, endocarditis, and pneumonia. J Bacteriol 194:3199–3215. https://doi.org/10.1128/JB.00183-12
Funke G, Bernard KA (2015) Coryneform Gram-positive rods. In: Manual of clinical microbiology, 11th edn. ASM Press, Washington, DC, pp 474–503
Mattos-Guaraldi AL, Moreira LO, Damasco PV, Hirata R (2003) Diphtheria remains a threat to health in the developing world – an overview. Mem Inst Oswaldo Cruz 98:987–993
De Benoist AC, White JM, Efstratiou A et al (2004) Imported cutaneous diphtheria, United Kingdom. Emerg Infect Dis 10:511–513. https://doi.org/10.3201/eid1003.030524
Pimenta FP, Hirata R, Rosa ACP et al (2008) A multiplex PCR assay for simultaneous detection of Corynebacterium diphtheriae and differentiation between non-toxigenic and toxigenic isolates. J Med Microbiol 57:1438–1439. https://doi.org/10.1099/jmm.0.2008/000414-0
Dias AA d S d O, Santos LS, Sabbadini PS et al (2011) Corynebacterium ulcerans diphtheria: an emerging zoonosis in Brazil and worldwide. Rev Saude Publica 45:1176–1191. https://doi.org/10.1590/S0034-89102011000600021
Mattos-Guaraldi AL, Duarte Formiga LC, Marques EA et al (2001) Diphtheria in a vaccinated adult in Rio de Janeiro, Brazil. Brazilian J Microbiol 32:236–239. https://doi.org/10.1590/S1517-83822001000300015
Bitragunta S, Murhekar MV, Hutin YJ et al (2008) Persistence of diphtheria, Hyderabad, India, 2003-2006. Emerg Infect Dis 14:1144–1146. https://doi.org/10.3201/eid1407.071167
Galazka A (2000) The changing epidemiology of diphtheria in the vaccine era. J Infect Dis 181:S2–S9. https://doi.org/10.1086/315533
Wagner KS, White JM, Lucenko I et al (2012) Diphtheria in the postepidemic period, Europe, 2000-2009. Emerg Infect Dis 18:217–225. https://doi.org/10.3201/eid1802.110987
WHO/UNICEF (2020) Progress Towards Global Immunization Goals - 2019 Summary presentation of key indicators.
John TJ (2008) Resurgence of diphtheria in India in the 21st century. Indian J Med Res 128:669–670
Bonmarin I, Guiso N, Le Flèche-Matéos A et al (2009) Diphtheria: a zoonotic disease in France? Vaccine 27:4196–4200. https://doi.org/10.1016/j.vaccine.2009.04.048
Perkins S, Cordery R, Nixon G et al (2010) Investigations and control measures following a non-travel-associated case of toxigenic Corynebacterium diphtheriae. London, United Kingdom, December 2009-January 2010. Eurosurveillance 15:4–6. https://doi.org/10.2807/ese.15.16.19544-en
Phalkey RK, Bhosale RV, Joshi AP et al (2013) Preventing the preventable through effective surveillance: the case of diphtheria in a rural district of Maharashtra, India. BMC Public Health 13(1):1–10
Santos AS, Ramos RT, Silva A et al (2018) Searching whole genome sequences for biochemical identification features of emerging and reemerging pathogenic Corynebacterium species. Funct Integr Genomics 18:593–610. https://doi.org/10.1007/s10142-018-0610-3
Besa NC, Coldiron ME, Bakri A et al (2014) Diphtheria outbreak with high mortality in northeastern Nigeria. Epidemiol Infect 142:797–802. https://doi.org/10.1017/S0950268813001696
World Health Organization (2018) Diphtheria vaccine-preventable diseases surveillance standards diphtheria. World Health Organization, Geneva
Organization PAHO/ WH (2018) Epidemiological update diphtheria. 1–3
Santos LS, Sant’Anna LO, Ramos JN et al (2015) Diphtheria outbreak in Maranhão, Brazil: microbiological, clinical and epidemiological aspects. Epidemiol Infect 143:791–798. https://doi.org/10.1017/S0950268814001241
Sharma NC, Efstratiou A, Mokrousov I et al (2019) Diphtheria. Nat Rev Dis Prim 5. https://doi.org/10.1038/s41572-019-0131-y
De Mattos-Guaraldi AL, Formiga LCD (1998) Bacteriological properties of a sucrose-fermenting Corynebacterium diphtheriae strain isolated from a case of endocarditis. Curr Microbiol 37:156–158. https://doi.org/10.1007/s002849900356
Bertuccini L, Baldassarri L, Von Hunolstein C (2004) Internalization of non-toxigenic Corynebacterium diphtheriae by cultured human respiratory epithelial cells. Microb Pathog 37:111–118. https://doi.org/10.1016/j.micpath.2004.06.002
Hirata R, Napoleão F, Monteiro-Leal LH et al (2002) Intracellular viability of toxigenic Corynebacterium diphtheriae strains in HEp-2 cells. FEMS Microbiol Lett 215:115–119. https://doi.org/10.1111/j.1574-6968.2002.tb11379.x
Peixoto RS, Pereira GA, dos Santos LS et al (2014) Invasion of endothelial cells and arthritogenic potential of endocarditis-associated Corynebacterium diphtheriae. Microbiol (United Kingdom) 160:537–546. https://doi.org/10.1099/mic.0.069948-0
Hirata J, Pereira GA, Filardy AA et al (2008) Potential pathogenic role of aggregative-adhering Corynebacterium diphtheriae of different clonal groups in endocarditis. Brazilian J Med Biol Res 41:986–991. https://doi.org/10.1590/s0100-879x2008001100007
Gomes DLR, Martins CAS, Faria LMD et al (2009) Corynebacterium diphtheriae as an emerging pathogen in nephrostomy catheter-related infection: evaluation of traits associated with bacterial virulence. J Med Microbiol:58. https://doi.org/10.1099/jmm.0.012161-0
Dos Santos CS, Dos Santos LS, De Souza MC et al (2010) Non-opsonic phagocytosis of homologous non-toxigenic and toxigenic Corynebacterium diphtheriae strains by human U-937 macrophages. Microbiol Immunol 54:1–10. https://doi.org/10.1111/j.1348-0421.2009.00179.x
Crosbie WE, Wright HD (1941) Diphtheria bacilli in floor dust. Lancet 237:656–659. https://doi.org/10.1016/S0140-6736(00)61019-X
Corboz L, Thoma R, Braun U, Zbinden R (1996) Isolation of Corynebacterium diphtheriae subsp. belfanti from a cow with chronic active dermatitis. Schweiz Arch Tierheilkd 138:596–599
Hall AJ, Cassiday PK, Bernard KA et al (2010) Novel Corynebacterium diphtheriae in domestic cats. Emerg Infect Dis 16:688–691. https://doi.org/10.3201/eid1604.091107
Leggett BA, De Zoysa A, Abbott YE et al (2010) Toxigenic Corynebacterium diphtheriae isolated from a wound in a horse. Vet Rec 166:656–657. https://doi.org/10.1136/vr.b4846
Detemmerman L, Rousseaux D, Efstratiou A et al (2013) Toxigenic Corynebacterium ulcerans in human and non-toxigenic Corynebacterium diphtheriae in cat. New Microbes New Infect 1:18–19. https://doi.org/10.1002/2052-2975.9
Mattos-Guaraldi AL, Sampaio JLM, Santos CS et al (2008) First detection of Corynebacterium ulcerans producing a diphtheria-like toxin in a case of human with pulmonary infection in the Rio de Janeiro metropolitan area, Brazil. Mem Inst Oswaldo Cruz 103:396–400. https://doi.org/10.1590/S0074-02762008000400014
Simpson-Louredo L, Ramos JN, Peixoto RS et al (2014) Corynebacterium ulcerans isolates from humans and dogs: fibrinogen, fibronectin and collagen-binding, antimicrobial and PFGE profiles. Antonie van Leeuwenhoek, Int J Gen Mol Microbiol 105. https://doi.org/10.1007/s10482-013-0080-5
Hacker E, Antunes CA, Mattos-Guaraldi AL, et al (2016) Corynebacterium ulcerans, an emerging human pathogen. Encycl Immunotoxicol 11:1–18. https://doi.org/10.1007/978-3-642-54596-2_200697
Bernard K, Pacheco A, Burdz T, Wiebe D (2019) Increase in detection of Corynebacterium diphtheriae in Canada: 2006–2019. Canada Commun Dis Rep 45:296–301. https://doi.org/10.14745/ccdr.v45i11a04
Yasuda I, Matsuyama H, Ishifuji T et al (2018) Severe pneumonia caused by toxigenic Corynebacterium ulcerans infection, Japan. Emerg Infect Dis 24:588–591. https://doi.org/10.3201/eid2403.171837
Dessau RB, Brandt-Christensen M, Jensen OJ, Tonnesen P (1995) Pulmonary nodules due to Corynebacterium ulcerans. Eur Respir J 8:651–653. https://doi.org/10.1183/09031936.95.08040651
Hommez J, Devriese LA, Vaneechoutte M et al (1999) Identification of nonlipophilic corynebacteria isolated from dairy cows with mastitis. J Clin Microbiol 37:954–957. https://doi.org/10.1128/jcm.37.4.954-957.1999
Hatanaka A, Tsunoda A, Okamoto M, et al (2003) Corynebacterium ulcerans diphtheria in Japan [3]. Emerg Infect Dis 9:752–753. https://doi.org/10.3201/eid0906.020645
Sing A, Hogardt M, Bierschenk S, Heesemann J (2003) Detection of differences in the nucleotide and amino acid sequences of diphtheria toxin from Corynebacterium diphtheriae and Corynebacterium ulcerans causing extrapharyngeal infections. J Clin Microbiol 41:4848–4851. https://doi.org/10.1128/JCM.41.10.4848-4851.2003
Zakikhany K, Efstratiou A (2012) Diphtheria in Europe: current problems and new challenges. Future Microbiol 7:595–607
Simpson-Lourêdo L, Silva CMF, Hacker E et al (2019) Detection and virulence potential of a phospholipase D-negative Corynebacterium ulcerans from a concurrent diphtheria and infectious mononucleosis case. Antonie van Leeuwenhoek, Int J Gen Mol Microbiol 112:1055–1065. https://doi.org/10.1007/s10482-019-01240-4
Dias AASO, Silva FC, Pereira GA, et al (2010) Corynebacterium ulcerans isolated from an asymptomatic dog kept in an animal shelter in the metropolitan area of Rio de Janeiro, Brazil. Vector-Borne Zoonotic Dis 10:743–748. https://doi.org/10.1089/vbz.2009.0132
Join-Lambert OF, Ouache M, Canioni D, et al (2006) Corynebacterium pseudotuberculosis necrotizing lymphadenitis in a twelve-year-old patient. Pediatr Infect Dis J 25:848–851. https://doi.org/10.1097/01.inf.0000234071.93044.77
Torres LFC, Ribeiro D, Hirata R et al (2013) Multiplex polymerase chain reaction to identify and determine the toxigenicity of Corynebacterium spp with zoonotic potential and an overview of human and animal infections. Mem Inst Oswaldo Cruz 108:272–279. https://doi.org/10.1590/0074-0276108032013003
Azevedo V, Lopes Bastos B, Dias Portela RW et al (2012) Corynebacterium pseudotuberculosis: immunological responses in animal models and zoonotic potential. J Clin Cell Immunol 4:5. https://doi.org/10.4172/2155-9899.S4-005
Keslin MH, McCoy EL, McCusker JJ, Lutch JS (1979) Corynebacterium pseudotuberculosis. A new cause of infectious and eosinophilic pneumonia. Am J Med 67:228–231. https://doi.org/10.1016/0002-9343(79)90395-4
Liu DTL, Chan WM, Fan DSP, Lam DSC (2005) An infected hydrogel buckle with Corynebacterium pseudotuberculosis [8]. Br J Ophthalmol 89:245–246
Wagner KS, White JM, Crowcroft NS et al (2010) Diphtheria in the United Kingdom, 1986-2008: the increasing role of Corynebacterium ulcerans. Epidemiol Infect 138:1519–1530. https://doi.org/10.1017/S0950268810001895
Pereira GA, Pimenta FP, Dos Santos FRW et al (2008) Antimicrobial resistance among Brazilian Corynebacterium diphtheriae strains. Mem Inst Oswaldo Cruz 103:507–510. https://doi.org/10.1590/S0074-02762008000500019
Fraz Z, Litt D, Duncan J et al (2019) Molecular epidemiology and antimicrobial resistance of Corynebacterium diphtheriae and Corynebacterium ulcerans strains isolated in the UK between 2004-2017. Access Microbiol 1:251. https://doi.org/10.1099/acmi.ac2019.po0110
Husada D, Soegianto SDP, Kurniawati IS et al (2019) First-line antibiotic susceptibility pattern of toxigenic Corynebacterium diphtheriae in Indonesia. BMC Infect Dis 19. https://doi.org/10.1186/s12879-019-4675-y
Otsuji K, Fukuda K, Endo T et al (2017) The first fatal case of Corynebacterium ulcerans infection in Japan. JMM Case Reports 4. https://doi.org/10.1099/jmmcr.0.005106
Galazka A, Kardymowicz B (1989) Immunity against diphtheria in adults in Poland. Epidemiol Infect 103:587–593. https://doi.org/10.1017/S0950268800030983
Walory J, Grzesiowski P, Hryniewicz W (2000) Comparison of four serological methods for the detection of diphtheria anti-toxin antibody. J Immunol Methods 245:55–65. https://doi.org/10.1016/S0022-1759(00)00273-8
Ohuabunwo C, Perevoscikovs J, Griskevica A et al (2005) Respiratory diphtheria among highly vaccinated military trainees in Latvia: improved protection from DT compared with Td booster vaccination. Scand J Infect Dis 37:813–820. https://doi.org/10.1080/00365540500262658
Pimenta FP, Vieira Damasco P, Cerbino Neto J et al (2006) Diphtheria-neutralizing antibody levels in healthy adults from Rio de Janeiro, Brazil. Mem Inst Oswaldo Cruz 101(4):459–462
Speranza FAB, Ishii SK, Hirata R et al (2010) Diphtheria toxin IgG levels in military and civilian blood donors in Rio de Janeiro, Brazil. Brazilian J Med Biol Res 43:120–123. https://doi.org/10.1590/s0100-879x2009007500032
Berger A, Huber I, Merbecks SS et al (2011) Toxigenic Corynebacterium ulcerans in woman and cat. Emerg Infect Dis 17:1767–1769
Pimenta FP, Souza MC, Pereira GA et al (2008) DNase test as a novel approach for the routine screening of Corynebacterium diphtheriae. Lett Appl Microbiol 46:307–311. https://doi.org/10.1111/j.1472-765X.2007.02310.x
Robert-Gangneux F, Belaz S, Revest M et al (2014) Diagnosis of Pneumocystis jirovecii pneumonia in immunocompromised patients by real-time PCR: a 4-year prospective study. J Clin Microbiol 52:3370–3376. https://doi.org/10.1128/JCM.01480-14
Badell E, Guillot S, Tulliez M et al (2019) Improved quadruplex real-time PCR assay for the diagnosis of diphtheria. J Med Microbiol 68:1455–1465. https://doi.org/10.1099/jmm.0.001070
Mancini F, Monaco M, Pataracchia M et al (2012) Identification and molecular discrimination of toxigenic and nontoxigenic diphtheria Corynebacterium strains by combined real-time polymerase chain reaction assays. Diagn Microbiol Infect Dis 73:111–120. https://doi.org/10.1016/j.diagmicrobio.2012.02.022
Nakao H, Popovic T (1997) Development of a direct PCR assay for detection of the diphtheria toxin gene. J Clin Microbiol 35:1651–1655
Konrad R, Berger A, Huber I et al (2010) Matrix-assisted laser desorption/ionisation time-of-flight (MALDI-TOF) mass spectrometry as a tool for rapid diagnosis of potentially toxigenic Corynebacterium species in the laboratory management of diphtheria-associated bacteria. Eurosurveillance 15:1–5. https://doi.org/10.2807/ese.15.43.19699-en
Senneby E, Nilson B, Petersson AC, Rasmussen M (2013) Matrix-assisted laser desorption ionization-time of flight mass spectrometry is a sensitive and specific method for identification of aerococci. J Clin Microbiol 51:1303–1304. https://doi.org/10.1128/JCM.02637-12
Alibi S, Ferjani A, Gaillot O et al (2015) Identification of clinically relevant Corynebacterium strains by Api Coryne, MALDI-TOF-mass spectrometry and molecular approaches. Pathol Biol 63:153–157. https://doi.org/10.1016/j.patbio.2015.07.007
Pallen MJ (1991) Rapid screening for toxigenic Corynebacterium diphtheriae by the polymerase chain reaction. J Clin Pathol 44:1025–1026. https://doi.org/10.1136/jcp.44.12.1025
Pallen MJ, Hay AJ, Puckey LH, Efstratiou A (1994) Polymerase chain reaction for screening clinical isolates of corynebacteria for the production of diphtheria toxin. J Clin Pathol 47:353–356. https://doi.org/10.1136/jcp.47.4.353
Efstratiou A, Engler KH, Dawes CS, Sesardic D (1998) Comparison of phenotypic and genotypic methods for detection of diphtheria toxin among isolates of pathogenic corynebacteria. J Clin Microbiol 36:3173–3177
Mothershed EA, Cassiday PK, Pierson K et al (2002) Development of a real-time fluorescence PCR assay for rapid detection of the diphtheria toxin gene. J Clin Microbiol 40:4713–4719. https://doi.org/10.1128/JCM.40.12.4713-4719.2002
Pacheco LGC, Pena RR, Castro TLP et al (2007) Multiplex PCR assay for identification of Corynebacterium pseudotuberculosis from pure cultures and for rapid detection of this pathogen in clinical samples. J Med Microbiol 56:480–486. https://doi.org/10.1099/jmm.0.46997-0
Pimenta FP, Matias GAM, Pereira GA et al (2008) A PCR for dtxR gene: application to diagnosis of non-toxigenic and toxigenic Corynebacterium diphtheriae. Mol Cell Probes. https://doi.org/10.1016/j.mcp.2008.01.001
Almeida S, Dorneles EMS, Diniz C et al (2017) Quadruplex PCR assay for identification of Corynebacterium pseudotuberculosis differentiating biovar Ovis and Equi. BMC Vet Res 13:290. https://doi.org/10.1186/s12917-017-1210-5
Espy MJ, Uhl JR, Sloan LM et al (2006) Real-time PCR in clinical microbiology: applications for routine laboratory testing. Clin Microbiol Rev 19:165–256
De Zoysa A, Efstratiou A, Mann G et al (2016) Development, validation and implementation of a quadruplex real-time PCR assay for identification of potentially toxigenic corynebacteria. J Med Microbiol 65:1521–1527. https://doi.org/10.1099/jmm.0.000382
Schuhegger R, Lindermayer M, Kugler R et al (2008) Detection of toxigenic Corynebacterium diphtheriae and Corynebacterium ulcerans strains by a novel real-time PCR. J Clin Microbiol 46:2822–2823. https://doi.org/10.1128/JCM.01010-08
Adékambi T, Shinnick TM, Raoult D, Drancourt M (2008) Complete rpoB gene sequencing as a suitable supplement to DNA-DNA hybridization for bacterial species and genus delineation. Int J Syst Evol Microbiol 58:1807–1814. https://doi.org/10.1099/ijs.0.65440-0
Khamis A, Raoult D, La Scola B (2004) rpoB gene sequencing for identification of Corynebacterium species. J Clin Microbiol 42:3925–3931. https://doi.org/10.1128/JCM.42.9.3925-3931.2004
Khamis A, Raoult D, La Scola B (2005) Comparison between rpoB and 16S rRNA gene sequencing for molecular identification of 168 clinical isolates of Corynebacterium. J Clin Microbiol 43:1934–1936. https://doi.org/10.1128/JCM.43.4.1934-1936.2005
Funke G, Von Graevenitz A, Clarridge JE, Bernard KA (1997) Clinical microbiology of Coryneform bacteria. Clin Microbiol Rev 10:125–159
Sangal V, Burkovski A, Hunt AC et al (2014) A lack of genetic basis for biovar differentiation in clinically important Corynebacterium diphtheriae from whole genome sequencing. Infect Genet Evol 21:54–57. https://doi.org/10.1016/j.meegid.2013.10.019
Sangal V, Hoskisson PA (2016) Evolution, epidemiology and diversity of Corynebacterium diphtheriae: new perspectives on an old foe. Infect Genet Evol 43:364–370
Hirata R, Pacheco LG, Soares SC et al (2011) Similarity of rpoB gene sequences of sucrose-fermenting and non-fermenting Corynebacterium diphtheriae strains. Antonie van Leeuwenhoek 99:733–737. https://doi.org/10.1007/s10482-010-9519-0
Mattos-Guaraldi AL, Formiga LC (1991) Relationship of biotype and source to the hemagglutination and adhesive properties of C. diphtheriae. Braz J Med Biol Res 24:399–406
Mattos-Guaraldi AL, Formiga LC, Camello TC, et al (2001) Corynebacterium diphtheriae threats in cancer patients. Rev Argent Microbiol 33:96–100
Bolt F, Cassiday P, Tondella ML, et al (2010) Multilocus sequence typing identifies evidence for recombination and two distinct lineages of Corynebacterium diphtheriae. J Clin Microbiol 48:4177–4185. https://doi.org/10.1128/JCM.00274-10
Badell E, Hennart M, Rodrigues C, et al (2020) Corynebacterium rouxii sp. nov., a novel member of the diphtheriae species complex. Res Microbiol 171:122–127. https://doi.org/10.1016/j.resmic.2020.02.003
Tagini F, Pillonel T, Croxatto A et al (2018) Distinct genomic features characterize two clades of Corynebacterium diphtheriae: proposal of Corynebacterium diphtheriae subsp. diphtheriae subsp. nov. and Corynebacterium diphtheriae subsp. lausannense subsp. nov. Front Microbiol 9. https://doi.org/10.3389/fmicb.2018.01743
Farfour E, Badell E, Dinu S et al (2013) Microbiological changes and diversity in autochthonous non-toxigenic Corynebacterium diphtheriae isolated in France. Clin Microbiol Infect 19:980–987. https://doi.org/10.1111/1469-0691.12103
Dazas M, Badell E, Carmi-Leroy A et al (2018) Taxonomic status of Corynebacterium diphtheriae biovar Belfanti and proposal of Corynebacterium belfantii sp. nov. Int J Syst Evol Microbiol 68:3826–3831. https://doi.org/10.1099/ijsem.0.003069
Windsor PA (2011) Control of caseous lymphadenitis. Vet Clin North Am – Food Anim Pract 27:193–202
Sellyei B, Bányai K, Bartha D et al (2017) Multilocus sequencing of Corynebacterium pseudotuberculosis biotype ovis strains. Biomed Res Int 2017. https://doi.org/10.1155/2017/1762162
Trost E, Al-Dilaimi A, Papavasiliou P et al (2011) Comparative analysis of two complete Corynebacterium ulcerans genomes and detection of candidate virulence factors. BMC Genomics 12. https://doi.org/10.1186/1471-2164-12-383
Otsuji K, Fukuda K, Ogawa M, Saito M (2019) Mutation and diversity of diphtheria toxin in Corynebacterium ulcerans. Emerg Infect Dis 25:2122–2123. https://doi.org/10.3201/eid2511.181455
Sunarno, Mulyastuti Y, Puspandari N, Sariadji K (2017) DNA sequence analysis of dtxR gene (partial) of Corynebacterium diphtheriae causing diphtheria in Jawa and Kalimantan Islands, Indonesia. Indones Biomed J 9:91–98. https://doi.org/10.18585/inabj.v9i2.268
Dangel A, Berger A, Konrad R, Sing A (2019) NGS-based phylogeny of diphtheria-related pathogenicity factors in different Corynebacterium spp. implies species-specific virulence transmission. BMC Microbiol 19:28. https://doi.org/10.1186/s12866-019-1402-1
Antunes CA, Dos Santos LS, Hacker E et al (2015) Characterization of DIP0733, a multi-functional virulence factor of Corynebacterium diphtheriae. Microbiol (United Kingdom) 161:639–647. https://doi.org/10.1099/mic.0.000020
Ott L, Hacker E, Kunert T et al (2017) Analysis of Corynebacterium diphtheriae macrophage interaction: dispensability of corynomycolic acids for inhibition of phagolysosome maturation and identification of a new gene involved in synthesis of the corynomycolic acid layer. PLoS One 12:1–24. https://doi.org/10.1371/journal.pone.0180105
Dias AASO, Silva FC, Santos LS et al (2011) Strain-dependent arthritogenic potential of the zoonotic pathogen Corynebacterium ulcerans. Vet Microbiol 153. https://doi.org/10.1016/j.vetmic.2011.06.007
Weerasekera D, Möller J, Kraner ME et al (2019) Beyond diphtheria toxin: cytotoxic proteins of Corynebacterium ulcerans and Corynebacterium diphtheriae. Microbiol (United Kingdom) 165:876–890. https://doi.org/10.1099/mic.0.000820
Weerasekera D, Hahn J, Herrmann M, Burkovski A (2019) Live cell imaging of macrophage/bacterium interaction demonstrates cell lysis induced by Corynebacterium diphtheriae and Corynebacterium ulcerans. BMC Res Notes 12. https://doi.org/10.1186/s13104-019-4733-y
da CA de Sá M, Gouveia GV, da Costa KC et al (2013) Distribution of PLD and FagA, B, C and D genes in Corynebacterium pseudotuberculosis isolates from sheep and goats with caseus lymphadenitis. Genet Mol Biol 36:265–268. https://doi.org/10.1590/S1415-47572013005000013
Guaraldi ALDM, Hirata R, Azevedo VADC (2013) Corynebacterium diphtheriae, Corynebacterium ulcerans and Corynebacterium pseudotuberculosis-general aspects. In: Corynebacterium diphtheriae and related toxigenic species: genomics, pathogenicity and applications. Springer Netherlands, Dordrecht, pp 15–37
Gower CM, Scobie A, Fry NK et al (2020) The changing epidemiology of diphtheria in the United Kingdom, 2009 to 2017. Eurosurveillance 25:1900462. https://doi.org/10.2807/1560-7917.ES.2020.25.11.1900462
Mattos-Guaraldi AL, Formiga LC (1986) Agglutination of sheep erythrocytes by Corynebacterium diphtheriae. Braz J Med Biol Res = Rev Bras Pesqui medicas e Biol 19:75–77
Cerdeño-Tárraga AM, Efstratiou A, Dover LG et al (2003) The complete genome sequence and analysis of Corynebacterium diphtheriae NCTC13129. Nucleic Acids Res 31:6516–6523. https://doi.org/10.1093/nar/gkg874
Silva De Souza SM, Hirata R, Moreira LO et al (2003) Influence of stannous chloride on the adhesive properties of Corynebacterium diphtheriae strains. Int J Mol Med 12:657–661. https://doi.org/10.3892/ijmm.12.4.657
Moreira LDO, Braga Andrade AF, Damasceno Vale MD et al (2003) Effects of iron limitation on adherence and cell surface carbohydrates of Corynebacterium diphtheriae strains. Appl Environ Microbiol 69:5907–5913. https://doi.org/10.1128/AEM.69.10.5907-5913.2003
dos Santos LS, Antunes CA, de Oliveira DM et al (2015) Tellurite resistance: a putative pitfall in Corynebacterium diphtheriae diagnosis? Antonie van Leeuwenhoek, Int J Gen Mol Microbiol 108:1275–1279. https://doi.org/10.1007/s10482-015-0558-4
dos Santos LS, Antunes CA, dos Santos CS, et al (2015) Corynebacterium diphtheriae putative tellurite-resistance protein (CDCE8392_0813) contributes to the intracellular survival in human epithelial cells and lethality of Caenorhabditis elegans. Mem Inst Oswaldo Cruz 110:662–668. https://doi.org/10.1590/0074-02760140479
Gomes DLR, Peixoto RS, Barbosa EAB et al (2013) SubMICs of penicillin and erythromycin enhance biofilm formation and hydrophobicity of Corynebacterium diphtheriae strains. J Med Microbiol 62:754–760. https://doi.org/10.1099/jmm.0.052373-0
Mattos-Guaraldi AL, Formiga LCD, Andrade AFB (1999) Cell surface hydrophobicity of sucrose fermenting and nonfermenting Corynebacterium diphtheriae strains evaluated by different methods. Curr Microbiol 38:37–42. https://doi.org/10.1007/PL00006769
Kaplan JB (2011) Antibiotic-induced biofilm formation. Int J Artif Organs 34:737–751
Peixoto RS, Hacker E, Antunes CA et al (2016) Pathogenic properties of a Corynebacterium diphtheriae strain isolated from a case of osteomyelitis. J Med Microbiol 65:1311–1321. https://doi.org/10.1099/jmm.0.000362
Ott L, Mckenzie A, Teresa Baltazar M et al (2012) Evaluation of invertebrate infection models for pathogenic corynebacteria. FEMS Immunol Med Microbiol 65(3):413–421. https://doi.org/10.1111/j.1574-695X.2012.00963.x
Antunes CA, Clark L, Wanuske MT, et al (2016) Caenorhabditis elegans star formation and negative chemotaxis induced by infection with corynebacteria. Microbiol (United Kingdom) 162:84–93. https://doi.org/10.1099/mic.0.000201
Broadway MM, Rogers EA, Chang C et al (2013) Pilus gene pool variation and the virulence of Corynebacterium diphtheriae clinical isolates during infection of a nematode. J Bacteriol 195:3774–3783. https://doi.org/10.1128/JB.00500-13
Ton-That H, Schneewind O (2003) Assembly of pili on the surface of Corynebacterium diphtheriae. Mol Microbiol 50:1429–1438. https://doi.org/10.1046/j.1365-2958.2003.03782.x
Mandlik A, Swierczynski A, Das A, Ton-That H (2007) Corynebacterium diphtheriae employs specific minor pilins to target human pharyngeal epithelial cells. Mol Microbiol 64:111–124. https://doi.org/10.1111/j.1365-2958.2007.05630.x
Colombo AV, Hirata R, Souza CMR et al (2001) Corynebacterium diphtheriae surface proteins as adhesins to human erythrocytes. FEMS Microbiol Lett 197:235–239. https://doi.org/10.1111/j.1574-6968.2001.tb10609.x
Ott L, Höller M, Gerlach RG, et al (2010) Corynebacterium diphtheriae invasion-associated protein (DIP1281) is involved in cell surface organization, adhesion and internalization in epithelial cells. BMC Microbiol 10:2. https://doi.org/10.1186/1471-2180-10-2
Brodzik K et al (2019) Analysis of the amino acid sequence variation of the 67–72p protein and the structural pili proteins of Corynebacterium diphtheriae for their suitability as potential vaccine antigens. Pol J Microbiol 68(2):233–246. https://doi.org/10.33073/pjm-2019-025
Sabbadini PS, Assis MC, Trost E, et al (2012) Corynebacterium diphtheriae 67-72p hemagglutinin, characterized as the protein DIP0733, contributes to invasion and induction of apoptosis in HEp-2 cells. Microb Pathog 52:165–176. https://doi.org/10.1016/j.micpath.2011.12.003
Rajamani Sekar SK, Veeraraghavan B, Anandan S et al (2017) Strengthening the laboratory diagnosis of pathogenic Corynebacterium species in the vaccine era. Lett Appl Microbiol 65:354–365
Ei PW, Aung WW, Lee JS et al (2016) Molecular strain typing of Mycobacterium tuberculosis: a review of frequently used methods. J Korean Med Sci 31:1673–1683
Kelly C, Efstratiou A (2003) Seventh International Meeting of the European Laboratory Working Group on Diphtheria - Vienna, June 2002. In: Euro surveillance: bulletin européen sur les maladies transmissibles = European communicable disease bulletin. pp 189–195
De Zoysa A, Efstratiou A (2004) Eighth international meeting of the European laboratory working group on diphtheria and the diphtheria surveillance network – June 2004: progress is needed to sustain control of diphtheria in European region. Eurosurveillance 9:13–14. https://doi.org/10.2807/esm.09.11.00489-en
Neal S, Efstratiou A, on behalf of DIPNET (2007) DIPNET – establishment of a dedicated surveillance network for diphtheria in Europe. Eurosurveillance 12:9–10. https://doi.org/10.2807/esm.12.12.00754-en
Efstratiou A, Roure C (2000) The European Laboratory Working Group on diphtheria: a global microbiologic network. J Infect Dis 181:S146–S151. https://doi.org/10.1086/315553
Dallman T, Neal S, Green J, Efstratiou A (2008) Development of an online database for diphtheria molecular epidemiology under the remit of the DIPNET project. Euro Surveill Bull Eur sur les Mal Transm = Eur Commun Dis Bull 13:
Mokrousov I (2009) Corynebacterium diphtheriae: genome diversity, population structure and genotyping perspectives. Infect Genet Evol 9:1–15. https://doi.org/10.1016/j.meegid.2008.09.011
De Zoysa A, Efstratiou A, George RC et al (1995) Molecular epidemiology of Corynebacterium diphtheriae from northwestern Russia and surrounding countries studied by using ribotyping and pulsed- field gel electrophoresis. J Clin Microbiol 33:1080–1083. https://doi.org/10.1128/jcm.33.5.1080-1083.1995
De Zoysa AS, Efstratiou A (1999) PCR typing of Corynebacterium diphtheriae by random amplification of polymorphic DNA. J Med Microbiol 48:335–340. https://doi.org/10.1099/00222615-48-4-335
Popovic T, Kombarova SY, Reeves MW et al (1996) Molecular epidemiology of diphtheria in Russia, 1985-1994. J Infect Dis 174:1064–1072. https://doi.org/10.1093/infdis/174.5.1064
Riegel P, Freitas FI, Prévost G et al (1997) Comparison of traditional and molecular methods for typing nontoxigenic strains of Corynebacterium diphtheriae. Eur J Clin Microbiol Infect Dis Off Publ Eur Soc Clin Microbiol 16:610–614. https://doi.org/10.1007/BF02447928
Mokrousov I, Narvskaya O, Limeschenko E, Vyazovaya A (2005) Efficient discrimination within a Corynebacterium diphtheriae epidemic clonal group by a novel macroarray-based method. J Clin Microbiol 43:1662–1668. https://doi.org/10.1128/JCM.43.4.1662-1668.2005
Popovic T, Mazurova IK, Efstratiou A et al (2000) Molecular epidemiology of diphtheria. J Infect Dis 181(Suppl):S168–S177. https://doi.org/10.1086/315556
De Zoysa A, Hawkey P, Charlett A, Efstratiou A (2008) Comparison of four molecular typing methods for characterization of Corynebacterium diphtheriae and determination of transcontinental spread of C. diphtheriae based on BstEII rRNA gene profiles. J Clin Microbiol 46:3626–3635. https://doi.org/10.1128/JCM.00300-08
Seth-Smith HMB, Egli A (2019) Whole genome sequencing for surveillance of diphtheria in low incidence settings. Front Public Heal 7:235. https://doi.org/10.3389/fpubh.2019.00235
Grimont PAD, Grimont F, Efstratiou A et al (2004) International nomenclature for Corynebacterium diphtheriae ribotypes. Res Microbiol 155:162–166. https://doi.org/10.1016/j.resmic.2003.12.005
Komiya T, Seto Y, De Zoysa A et al (2010) Two Japanese Corynebacterium ulcerans isolates from the same hospital: Ribotype, toxigenicity and serum antitoxin titre. J Med Microbiol 59:1497–1504. https://doi.org/10.1099/jmm.0.022491-0
De Zoysa A, Hawkey PM, Engler K et al (2005) Characterization of toxigenic Corynebacterium ulcerans strains isolated from humans and domestic cats in the United Kingdom. J Clin Microbiol 43:4377–4381. https://doi.org/10.1128/JCM.43.9.4377-4381.2005
Sekizuka T, Katsukawa C, Kuroda M et al (2020) Limitations of ribotyping as genotyping method for Corynebacterium ulcerans. Emerg Infect Dis 26:2457–2459. https://doi.org/10.3201/eid2610.200086
Kaufmann ME (1998) Pulsed-field gel electrophoresis. In: Molecular bacteriology. Humana Press, New Jersey, pp 33–50
Sulakvelidze A, Kekelidze M, Gomelauri T et al (1999) Diphtheria in the Republic of Georgia: use of molecular typing techniques for characterization of Corynebacterium diphtheriae strains. J Clin Microbiol 37:3265–3270. https://doi.org/10.1128/JCM.37.10.3265-3270.1999
Selander RK, Caugant DA, Ochman H et al (1986) Methods of multilocus enzyme electrophoresis for bacterial population genetics and systematics. Appl Environ Microbiol 51:873–884. https://doi.org/10.1128/AEM.51.5.873-884.1986
Williams JG, Kubelik AR, Livak KJ et al (1990) DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18:6531–6535. https://doi.org/10.1093/nar/18.22.6531
Nakao H, Popovic T (1998) Use of random amplified polymorphic DNA for rapid molecular subtyping of Corynebacterium diphtheriae. Diagn Microbiol Infect Dis 30:167–172. https://doi.org/10.1016/s0732-8893(97)00237-x
Vos P, Hogers R, Bleeker M et al (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414. https://doi.org/10.1093/nar/23.21.4407
Zasada AA, Formińska K, Wołkowicz T et al (2014) The utility of the PCR melting profile technique for typing Corynebacterium diphtheriae isolates. Lett Appl Microbiol 59:292–298. https://doi.org/10.1111/lam.12274
De Zoysa A, Efstratiou A (2000) Use of amplified fragment length polymorphisms for typing Corynebacterium diphtheriae. J Clin Microbiol 38:3843–3845. https://doi.org/10.1128/jcm.38.10.3843-3845.2000
Turner KME, Feil EJ (2007) The secret life of the multilocus sequence type. Int J Antimicrob Agents 29:129–135. https://doi.org/10.1016/j.ijantimicag.2006.11.002
Czajka U, Wiatrzyk A, Mosiej E et al (2018) Changes in MLST profiles and biotypes of Corynebacterium diphtheriae isolates from the diphtheria outbreak period to the period of invasive infections caused by nontoxigenic strains in Poland (1950-2016). BMC Infect Dis:18. https://doi.org/10.1186/s12879-018-3020-1
Sangal V, Blom J, Sutcliffe IC et al (2015) Adherence and invasive properties of Corynebacterium diphtheriae strains correlates with the predicted membrane-associated and secreted proteome. BMC Genomics 16:765. https://doi.org/10.1186/s12864-015-1980-8
Dangel A, Berger A, Konrad R et al (2018) Geographically diverse clusters of nontoxigenic Corynebacterium diphtheriae infection, Germany, 2016–2017. Emerg Infect Dis 24:1239–1245. https://doi.org/10.3201/eid2407.172026
Clarke KEN, MacNeil A, Hadler S et al (2019) Global epidemiology of diphtheria, 2000-2017. Emerg Infect Dis 25:1834–1842. https://doi.org/10.3201/eid2510.190271
Viguetti SZ, Pacheco LGC, Santos LS et al (2012) Multilocus sequence types of invasive Corynebacterium diphtheriae isolated in the Rio de Janeiro urban area, Brazil. Epidemiol Infect 140:617–620. https://doi.org/10.1017/S0950268811000963
König C, Meinel DM, Margos G et al (2014) Multilocus sequence typing of Corynebacterium ulcerans provides evidence for zoonotic transmission and for increased prevalence of certain sequence types among toxigenic strains. J Clin Microbiol 52:4318–4324. https://doi.org/10.1128/JCM.02291-14
Jolley KA, Bray JE, Maiden MCJJ (2018) Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res 3:124. https://doi.org/10.12688/wellcomeopenres.14826.1
Both L, Collins S, De Zoysa A et al (2015) Molecular and epidemiological review of toxigenic diphtheria infections in England between 2007 and 2013. J Clin Microbiol 53:567–572
Zasada AA, Jagielski M, Rzeczkowska M, Januszkiewicz A (2011) [The use of MLVA for Corynebacterium diphtheriae genotyping--preliminary studies]. Med Dosw Mikrobiol 63:209–218
Mokrousov I, Limeschenko E, Vyazovaya A, Narvskaya O (2007) Corynebacterium diphtheriae spoligotyping based on combined use of two CRISPR loci. Biotechnol J 2:901–906. https://doi.org/10.1002/biot.200700035
Sangal V, Fineran PC, Hoskisson PA (2013) Novel configurations of type I and II CRISPR-Cas systems in Corynebacterium diphtheriae. Microbiology 159:2118–2126. https://doi.org/10.1099/mic.0.070235-0
WHO (2017) Operational protocol for clinical management of diphtheria Bangladesh. Cox’s Bazar 25
Dravid MN, Joshi SA (2008) Resurgence of diphtheria in Malegaon & Dhule regions of north Maharashtra. Indian J Med Res 127:616–617
du Plessis M, Wolter N, Allam M et al (2017) Molecular characterization of Corynebacterium diphtheriae outbreak isolates, South Africa, March-June 2015. Emerg Infect Dis 23:1308–1315. https://doi.org/10.3201/eid2308.162039
Mohd Khalid MKN, Ahmad N, Hii SYF et al (2019) Molecular characterization of Corynebacterium diphtheriae isolates in Malaysia between 1981 and 2016. J Med Microbiol 68:105–110. https://doi.org/10.1099/jmm.0.000881
Paveenkittiporn W, Sripakdee S, Koobkratok O et al (2019) Molecular epidemiology and antimicrobial susceptibility of outbreak-associated Corynebacterium diphtheriae in Thailand, 2012. Infect Genet Evol J Mol Epidemiol Evol Genet Infect Dis 75:104007. https://doi.org/10.1016/j.meegid.2019.104007
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Vieira, V.V., Ramos, J.N., dos Santos, L.S., Mattos-Guaraldi, A.L. (2022). Corynebacterium: Molecular Typing and Pathogenesis of Corynebacterium diphtheriae and Zoonotic Diphtheria Toxin-Producing Corynebacterium Species. In: de Filippis, I. (eds) Molecular Typing in Bacterial Infections, Volume I. Springer, Cham. https://doi.org/10.1007/978-3-030-74018-4_1
Download citation
DOI: https://doi.org/10.1007/978-3-030-74018-4_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-74017-7
Online ISBN: 978-3-030-74018-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)