Abstract
The present study was taken up to test the soil sample isolated from an automobile workshop located at Coimbatore for the presence of biosurfactant producing bacteria. Ten isolates were initially screened from the soil collected from the petrochemical exposed site. Isolation was done in the minimal salt medium, and this was followed by screening for the biosurfactant production through three different methods, namely emulsification index, oil displacement activity and hemolytic assay. In comparison with all the three screening methods, two best isolates were chosen and was then subjected to mass production in the mineral salt medium for biosurfactants generation. The isolate identified as Serratia marcescens through 16S rRNA sequencing. Pinkish red-coloured occurrence was observed during the mass production, which may be possibly due to pigmentation. The extracted biosurfactants were analyzed with Fourier transform infrared spectroscopy spectra. The potent nature of the soil collected from the contaminated petrochemical site for the production of biosurfactants has a remarkable growth prospective, sourcing as a better replacement to the increasing environmental concern associated with the chemical surfactants.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
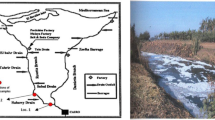
Biosurfactants refer to any biomolecules which exhibit characteristics like surface-active and are found to be amphipathic in nature, which can be produced extracellularly by various microbes including bacteria, fungi and yeasts. They form aggregates at the interfaces amid different fluid polarities, wherein decreasing the surface (Saravanan and Vijayakumar 2012). The unique features of microbial surfactants include biodegradability, emulsifying and demulsifying ability, wetting and penetrating agents, tolerance to temperature, pH, ionic strength, low toxicity and viscosity reducing agents make them most suitable for remediation of environmental contamination, environmental safety (Akbari et al. 2018) and finds application in food, pharmaceutical and allied industries (Shekhar et al. 2015; Sharma et al. 2018). Oil spills and various other hydrocarbon contaminants in the soil and aquatic environment is becoming a significant focus of attention in the today world. Absorbents and synthetic organic products like lime, plastic polymers and cellulose-based materials, polyurethane foams, polypropylene and elastomers were preferred as the commercial sorbents in the oil spill cleanup considering the properties of good hydrophobicity, uptake capacity and oil recovery (Adebajo et al. 2003; Seo et al. 2018), but then it faced the problems of low degradability comparing to the naturally occurring mineral and vegetable products (Teas et al. 2001). Such disadvantages has to be mitigated via microbialy generated surfactants. Microorganisms exposed to oil-contaminated sites, develop the potential to harness hydrocarbons as carbon and energy for its metabolic activities. Biosurfactants increase the substrate bioavailability for microorganisms and interacts with the cell surface, thereby increasing the surface hydrophobicity, thus allowing substrates to accumulate faster within the bacterial cells (Liu et al. 2017). In comparison with its chemical counterparts, biosurfactants are very particular, potent and competent under an extensive range of oil and receptacle environment and may only require meagre quantities. Because of these attributes, they are preferred in different industrial processes and physicochemical phenomena, wherein they attribute enhanced solubility, increased mobility (Pacwa-Płociniczak et al. 2011; Rufino et al. 2014). Microbial biosurfactant with all these unique properties is considered to be a best green alternative potent source when comparing to other methods for clearing oil spills and plays a promising role in bioremediation of hydrocarbons contaminated sites, proving to be better than the chemical surfactants in many aspects including degradability (Sammarco et al. 2013; Matvyeyeva et al. 2014). Serratia marcescens, which is a gram-negative bacillus, belonging to Enterobacteriaceae, is better known for the production of red pigment prodigiosin and biosurfactant serrawettin (Muthukumar et al. 2016; Sunaga et al. 2016). At the time when the cell growth rate is sluggish under unfavourable conditions, the pigment biosynthesis has a role of being the protective mechanism (Li et al. 2005). Serratia produces chitinolytic enzymes that have a practical part in the biological degradation of chitin (Someya et al. 2000) and its related environmental applications along with environmental protection (Brzezinska et al. 2014). Biosurfactants with the current light of focus are the potent tool for developing a sustainable environment. This study accounts for the isolation and screening of biosurfactants producing indigenous bacteria, that has been identified as Serratia marcescens, isolated from a soil sample collected from contaminated site identified in Coimbatore district, Tamilnadu, India (Fig. 1).
2 Materials and Method
2.1 Sample Collection
The soil sample was collected from an automobile workshop located at Saravanampatti, Coimbatore district and used for further studies.
2.2 Isolation of Bacterial Colonies
Soil sample of 5 g was inoculated in 100 ml of Minimal Salt Medium (MSM) with the composition containing 1.5 gL−1 of NaNO3, 1 gL−1 of KH2PO4, 1.5 gL−1 of (NH4)2SO4, 0.5 gL−1 of MgSO4, 0.01 gL−1 of FeSO4 and 0.002 gL−1 of CaCl2 was added along with 3 ml kerosene oil as the carbon source in a 250 ml conical flask. Incubation carried out for 72 h at a temperature of 30 ℃. These samples serially diluted up to 10ˉ 6 dilution. From this, 1 ml of each dilution transferred to nutrient agar for spread culture, and these plates were at incubated at 37 ℃ for about 72 h (Dewaliya and Jasodani 2013). Incubation is followed by single colony isolation and based on the morphology; ten distinct isolates selected for further studies. The selected isolates were screened for biosurfactant production using a modified Mineral salt medium with trace elements such as FeSO4 0.05 gL−1, H3BO3 0.5 gL−1, MnSO4.4H2O 0.008 gL−1, C2H7NO2 0.05 gL−1 were dissolved in 250 ml of distilled water to which 1 M phosphate buffer of pH 6.8 of 20 mlL−1 with major salts such as KNO3 1 gL−1, H14MgO11S 1 gL−1, CaCl2H12O6 0.1 gL−1, kerosene 20 mlL−1 were added and finally made up to 1000 ml. Best screened isolates tested for biosurfactant production used for further considered for mass production and subjected to characterization studies. All the experiments were adequately replicated, and the result values expressed in average or mean values.
2.3 Screening of Biosurfactant Producing Bacteria
2.3.1 Emulsification Index
Emulsification activity of the biosurfactant was checked, wherein 2 ml of kerosene oil was added to 1 ml of cellfree extract that had been obtained by centrifugation (Abd-alridha 2014). This mixture was then vortexed well for about 2–3 min at high speed. The emulsion activity was observed after 24 h and calculated as Eq. 1.
Oil displacement activity is a measure of the ability of the biosurfactants to alter the contact angle of the oil, which is studied on oil-water interphase. Also, it is a measure of the reduction of surface tension by the biosurfactant (Freitas et al. 2016). The oil displacement activity was checked by adding 20 µl of kerosene oil to the surface of the Petri plate containing 50 ml of distilled water. Upon this 20 µl of culture supernatant was added to check for the formation of clear zones formed in the presence of the biosurfactants.
2.4 Blood Hemolysis Activity
The single isolated colonies were streaked on the blood agar plates and incubated for about 48–72 h under the temperature of 37 ℃ (Shanks et al. 2012), and then observed for the type of clear zone based on which the presence of biosurfactant producing bacteria was determined (Pacwa-Płociniczak et al. 2014).
2.5 Extraction of Biosurfactant
The extraction of the biosurfactant was done with the initial step of bacterial cell removal by centrifugation at 12,000 rpm for 20 min, the cultured supernatant adjusted to pH 2.0 with the aid of 0.1 M HCl. The biosurfactants extracted with a solvent mixture containing chloroform and methanol in the ratio 2:1v/v in a separation funnel. After shaken vigorously, and allowed for phase separation, the lower organic phase concentrated, and anhydrous sodium sulfate added to remove water, and the residue was dried to obtain biosurfactants (Pacwa-Płociniczak et al. 2011).
2.6 Molecular Characterization of Biosurfactant Producing Bacteria–16S RRNA Gene Sequencing
The DNA was isolated from the bacterial culture and was used in PCR to amplify the bacterial 16 s rRNA using 16 s rRNA PCR kit (800). Using the primers from the kit, amplification of 800 bp amplicon done, and no amplicon was visible in the negative control. The expected sized amplicon of 800 bp observed in the positive control. The test amplicon of 800 bp was purified using magnetic beads, and the product sequenced by Sanger’s method of DNA sequencing. The sequencing results were assembled and compared with the NCBI database.
2.7 Characterization of Biosurfactants from Serratia Marcescens Using Thin Layer Chromatography
Silica gel plate prepared on the ratio 1:2, with 1 mm thickness, i.e., 10 g in 20 ml distilled water which was then allowed it to stay for 30 min. The silica gel plate was kept in the hot-air oven for 1 h at 100 ℃ to activate the absorption. Chloroform-methanol of ratio 85:15 used as the developing solvent. 20 µl of biosurfactant spotted on the silica gel plate (Sivagamasundari and Jeyakumar 2016), and Ninhydrin reagent sprayed. The retention factor Rf (eq. 2), which defined as the distance travelled by the compound to the distance travelled by the solvent calculated.
2.8 Fourier Transform Infrared Spectroscopy
For identifying types of chemical bonds (functional groups), Fourier transform infrared spectroscopy (FT-IR) analysis used to elucidate some components of an unknown mixture (Suryawanshi et al. 2014). One milligram of partially purified biosurfactant was wholly dried and then analyzed in an FTIR (Shimadzu), device that obtains the spectrum with high S/N ratio 30,000:1, 1-minute accumulation and a maximum resolution of 0.5 cm−1. All data corrected for the background spectrum.
3 Results and Discussion
Soil samples inoculated in minimal salt medium and serial diluted to obtain bacterial isolates and subcultured for the isolation of the single colonies, as shown in the Fig. 2.
3.1 Screening of Biosurfactant Producing Bacteria
All the ten isolated strains screened for biosurfactant production. The bacterial isolates named as Petrochemical Contaminated Bacteria (PCB); PCB1, PCB2, PCB3, PCB4, PCB5, PCB6, PCB7, PCB8, PCB9 and PCB10. These were subjected to the following screening assays for the selecting the biosurfactant producing bacterial isolates.
3.2 Emulsification Index
When kerosene used as the oil source for testing the emulsification activity of isolated bacteria, positive results were noted and tabulated in Table 1. The emulsification activity was noted after 24 h and emulsification index was calculated to screen for the biosurfactant producing bacteria among the ten bacterial isolates. Among the studied strains, the highest emulsification index of 25 obtained in the two isolates, namely PCB5 and PCB8. It was also noted that the bacterial isolates PCB2, PCB3, PCB7 and PCB10 had emulsification index greater than 20. Similar range of emulsification index reported in the literature, when tested for seven different isolates of Bacillus sp., and Actinomyces sp., according to Bento et al. (2005) and Wang et al. (2014).
3.3 Oil Displacement Activity
Oil displacement activity helps in testing the ability of the biosurfactant to alter the contact angle at the oil-water interface when the supernatant added to the kerosene oil. Among the obtained results, the isolates namely PCB3, PCB7 and PCB10 showed good oil displacement activity indicated by ++ within the ten isolated tested for oil displacement activity. Rest of the strains only showed partial displacement activity reported by + as noted in Table 1.
3.4 Blood Hemolysis Activity
Blood hemolysis activity is tested for checking the ability of bacterial colonies to induce hemolysis when it was grown on blood agar. When the agar under the colony turns darkish green, α hemolysis is said to occur, and when it turns light yellow with transparent nature it is noted to be β hemolysis, and γ hemolysis observed with no change in the agar. All the ten isolates showed positive hemolytic activity in reference to the results obtained in Table 1. The presence of hemolytic activity in Serratia marcescens demonstrated via Fig. 3.
Based on the overall screening results of biosurfactant producing bacterial isolates, Serratia marcescens was selected among the ten isolates to narrow down the research for the mass production of biosurfactants with reference to emulsification activity, oil displacement and hemolytic activity. The screened isolates of Serratia marcescens and PCB10 were chosen for the mass production of biosurfactants in the mineral salt medium where the biosurfactants were formed as a separate layer with pigmentation. Confirmation test for emulsification activity produced good results (Fig. 4).
3.5 Extraction of Biosurfactant
The biosurfactants extracted with a solvent mixture containing chloroform and methanol in the ratio 2:1v/v in a separation funnel (Sukirtha and Usharani 2013; Yap et al. 2016). The lower organic phase concentrated, and anhydrous sodium sulphate added to remove water and dried to obtain biosurfactants as shown in the Fig. 5.
3.6 16S RRNA Gene Sequencing
The DNA isolated from the bacterial culture was used in PCR to amplify the bacterial 16 s rRNA. The molecular analysis for the isolated bacterial strain showed 99% identity to the sequence of Serratia marcescens. The phylogenetic tree generated for the isolated bacteria shown in Fig. 5. and 99% identity to the sequence of Serratia marcescens. Based upon the sequence analyses, the isolate is likely to be Serratia marcescens. The phylogenetic tree generated for the isolated bacteria provided in Fig. 6.
3.6.1 Thin Layer Chromatography
The retention factor for the compounds separated, calculated using the Formula (3),
The results of TLC shown in Fig. 7.
3.7 Thin Layer Chromatography
The Rf value of biosurfactant obtained from Serratia marcescens as shown in Fig. 6 established to be 0.76 and 0.96 which was similar to the results reported with a red pigment isolated from Serratia marcescens (Vora et al. 2014). The quantitative analysis of the biosurfactant from Serratia marcescens is yet to be done.
3.8 Fourier Transform Infrared Spectroscopy
The FTIR studies for the extracted biosurfactant obtained from the bacterial isolate Serratia marcescens showed C–H stretching bands of –CH2 and –CH3 groups observed in the region 3000–2700 cm−1 in Fig. 8. The deformation vibrations at 1467 and 1379 cm−1 also confirmed the presence of alkyl groups. The wavenumber 1066.64 cm−1 indicated the presence of C-O bonds. The peaks observed at 1066.64 cm−1, 1226.73 cm−1, 1539.2 cm−1, 1645.28 cm−1, 2096.62 cm−1, 2956.87 cm−1, 3286.7 cm−1, 3379.29 cm−1 indicated the presence lipid moieties. In this, the absorbance of N–H stretching bond at 3132–3379 cm−1 was also observed (Vigneshwaran et al. 2018).
4 Conclusion
Among the ten bacterial isolates, Serratia marcescens, which was identified through 16 s rRNA sequencing, isolated from the soil sample that was collected from an automobile workshop located in Coimbatore district showed good activity in oil displacement, emulsification index proving to be a potent source for biosurfactant production. This could be further analyzed with the pigmentation studies and could be used for the potential application of environmental remediation and protection.
References
Abd-alridha FT (2014) Isolation and screening of biosurfactant producing bacteria from oil contaminated soils in Iraq. IOSR J Pharm Biol Sci 9(6):10–15
Adebajo MO, Frost RL, Kloprogge JT, Carmody O, Kokot S (2003) Porous materials for oil spill cleanup: a review of synthesis. J Porous Mater 10(10):159–170
Akbari S, Abdurahman NH, Yunus RM, Fayaz F, Alara OR (2018) Biosurfactants-a new frontier for social and environmental safety: a mini review. Biotechnol Res Innov 2:81–90
Bento FM, Camargo FAO, Okeke BC, Frankenberger WT (2005) Diversity of biosurfactant prodcuing microorganism isolated from soils contaminated with diesel oil. Microbiol Res 160(3):249–255
Brzezinska MS, Jankiewicz U, Burkowska A, Walczak M (2014) Chitinolytic microorganisms and their possible application in environmental protection. Current Microbiol 68:71–81
Dewaliya V, Jasodani R (2013) Isolation and identification of Bacillus licheniformis for biosurfactant production. CIBTech J Microbiol 2(4):14–19
Freitas BG, Brito JGM, Brasileiro PPF, Rufino RD, Luna JM, Santos VA, Sarubbo LA (2016) Formulation of a commercial biosurfactant for application as a dispersant of petroleum and by-products spilled in Oceans. Front Microbiol 7:1–5
Li H, Tanikawa T, Sato Y, Nakagawa Y, Matsuyama T (2005) Serratia marcescens gene required for surfactant serrawettin W1 production encodes putative aminolipid synthetase belonging to nonribosomal peptide synthetase family. Microbiol Immunol 49(4):303–310
Liu Z, Li Z, Zhong H, Zeng G, Liang Y, Chen M, Wu Z, Zhou Y, Yu M, Shao B (2017) Recent advances in the environmental applications of biosurfactant saponins: a review. J Environ Chem Eng 5:6030–6038
Matvyeyeva OL, Vasylchenko OA, Aliieva OR (2014) Microbial biosurfactants role in oil products biodegradation. Int J Environ Bioremed Biodegrad 2(2):69–74
Muthukumar A, Pradeep P, Thigale I, Mohanasrinivasan V, Jemimah NS, Subathra Devi C (2016) Exploring the bioactive potential of Serriatia marcescens VITAPI isolated from soil. Front Biol (Beijing) 11(6):476–480
Pacwa-Płociniczak M, Płaza GA, Piotrowska-seget Z (2011) Environmental applications of biosurfactants: recent advances. Int J Mol Sci 12(1):633–654
Pacwa-Płociniczak M, Płaza GA, Poliwoda A, Piotrowska-Seget Z (2014) Characterization of hydrocarbon degrading and biosurfactant-producing Pseudomonas sp. P-1 strain as a potential tool for bioremediation of petroleum-contaminated soil. Environ Sci Pollut Res 21(15):9385–9395
Rufino RD, de Luna JM, Takaki GMC, Sarubbo LA (2014) Characterization and properties of the biosurfactant produced by Candida lipolytica UCP 0988. Electron J Biotechnol 17:34–38
Sammarco PW, Kolian SR, Warby RAF, Bouldin JL, Subra WA, Porter SA (2013) Distribution and concentrations of petroleum hydrocarbons associated with the BP/Deepwater Horizon Oil Spill. Gulf of Mexico Mar Pollut Bull 73(1):129–143
Saravanan V, Vijayakumar S (2012) Isolation and screening of biosurfactant producing microorganisms. J Acad Ind Res 1(5):1–5
Seo S, Mastiani M, Mosavati B, Peters DM, Mandin P, Kim M (2018) Performance evaluation of environmentally benign nonionic biosurfactant for enhanced oil recovery. Fuel 234:48–55
Shanks RMQ, Stella NA, Lahr RM (2012) Serratamolide is a hemolytic factor produced by Serratia marcescens. PLoS ONE 7(5):1–8
Sharma R, Singh J, Verma N (2018) Production, Characterization and environmental applications of biosurfactants from Bacillus amyloliquefaciens and Bacillus subtilis. Biocatal Agric Biotechnol 16:132–139
Shekhar S, Sundaramanickam A, Balasubramanian T (2015) Biosurfactant producing microbes and their potential applications: a review. Crit Rev Environ Sci Technol 45(14):1522–1554
Sivagamasundari T, Jeyakumar N (2016) Biosurfactant production using diesel oil degrading bacteria. Int J Adv Res Sci Eng Technol 3(12):3039–3046
Someya N, Kataoka N, Komagata T, Hirayae K, Hibi T, Akutsu K (2000) Biological control of cyclamen soilborne diseases by Serratia marcescens strain B2. Plant Dis 84(3):334–340
Sukirtha T, Usharani MV (2013) Production and qualitative analysis of biosurfactant and biodegradation of the organophosphate by Nocardia mediterranie. J Bioreme Biodegrad 4(6):4–11
Sunaga S, Li H, Sato Y, Nakagawa Y, Matsuyama T (2016) Identification and characterization of the pswP gene required for the parallel production of prodigiosin and serrawettin W1 in Serratia marcescens. Microbiol Immunol 48(10):723–728
Suryawanshi RK, Patil CD, Borase HP, Salunke BK, Patil SV (2014) Studies on production and biological potential of prodigiosin by Serratia marcescens. Appl Biochem Biotechnol 173(5):1209–1221
Teas Ch, Kalligeros S, Zanikos F, Stournas S, Lois E, Anastopoulos G (2001) Investigation of the effectiveness of absorbent materials in oil spills clean up. Desalination 140(3):259–264
Vigneshwaran C, Sivasubramanian V, Vasantharaj K, Krishnanand N, Jerold M (2018) Potential of Brevibacillus sp. AVN 13 isolated from crude oil contaminated soil for biosurfactant production and its optimization studies. J Environ Chem Eng 6:4347–4356
Vora JU, Jain NK, Modi HA (2014) Extraction, Characterization and Application studies of red pigment of halophile Serratia marcescens KH1R KM035849 isolated from Kharaghoda soil. Int J Pure Appl Biosci 2(6):160–168
Wang W, Cai B, Shao, Z (2014) Oil degradation and biosurfactant production by the deep sea bacterium Dietzia maris As-13–3. Front Microbial 1-11
Yap AC, Chan KG, Choo YM (2016) Isolation and identification of metabolites from the gram-negative proteobacteria of Burkholderia cenocepacia and Serratia marcescens. Sains Malaysiana 45(7):1073–1077
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Vijayalekshmi, V., Muthukumaran, P., Jeyaseelan, A. (2021). Screening, Isolation and Molecular Characterization of Biosurfactants Producing Serratia Marcescens from Petrochemical Exposed Site. In: Marimuthu, P.D., Sundaram, R., Jeyaseelan, A., Kaliannan, T. (eds) Bioremediation and Green Technologies. Environmental Science and Engineering(). Springer, Cham. https://doi.org/10.1007/978-3-030-64122-1_17
Download citation
DOI: https://doi.org/10.1007/978-3-030-64122-1_17
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-64121-4
Online ISBN: 978-3-030-64122-1
eBook Packages: Earth and Environmental ScienceEarth and Environmental Science (R0)