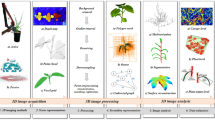
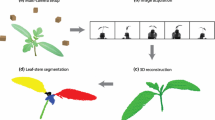
Abstract
In recent years, there has been significant progress in computer vision based plant phenotyping technologies. Due to their non-invasive and non-contact properties, imaging techniques are becoming state of the art in automated plant phenotyping analysis. There are several aspects of phenotyping, including plant growth, organ classification and tracking, disease detection, etc. This chapter presents a broad overview of computer vision based 3D plant phenotyping techniques. Some case studies of state-of-the-art techniques are described in detail. In the first case study, automated robotic systems for 3D plant phenotyping are discussed. The second study focuses on general registration techniques of point cloud and alignment of multiple view challenging plant point cloud data. Next, recently successful plant organ segmentation techniques are reviewed. Finally, some open challenges of vision-based plant phenotyping are discussed, followed by conclusion and some hands on exercises.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Introduction
Due to the increase in the world population in recent years, the demand of food/crop is increasing very fast. Automated analysis for crop production systems is of extreme demand in order to fulfil the task of mass production as well as efficient monitoring of crops. Computer vision based techniques can be very efficient because of the non-invasive and non-contact nature of image-based analysis. Also, imaging systems can be more accurate than other approaches. In recent years, there has been tremendous progress in image-based analysis of plant phenotyping (measuring biologically significant properties of plants) technologies. 3D imaging based approaches are becoming state of the art in quantifying biological properties of plants. The advantage of 3D over 2D are numerous. For example, in order to analyze the growth of a plant, representing that plant as a 3D mesh model is a very effective methodology. From the 3D mesh, it is possible to compute the 3D volume (the convex hull) and 3D surface area of the plant, in addition to analyzing other desirable properties. As we will see later in this chapter, 3D laser scanners are widely used to perform 3D plant phenotyping. Also consider the area of a leaf. If the leaf is curved, its 3D area will be significantly different from the area computed from its 2D image.
Additionally, apart from the 3D based analysis of plants, data collection is also an important aspect of an automated phenotyping system and cannot be ignored. However, building a high throughput real-time system is a challenging task. Several challenges include the following factors: communication among the hardware devices, reliable data transfer and analysis, fault tolerance, etc. Ideally, the system should be simple enough for a naive user to operate and obtain the phenotyping result as a ready-made product. An ideal system should also be able to be general enough to handle several varieties of plants and analyze their phenotypes accurately. With the advancements of recent robotic technologies and high precision mechanical devices, fully automated real-time systems are becoming possible. These systems are shown to be reliable enough to capture and analyze the data throughout the lifetime of a plant. Automation is also important to study the effect of several environmental factors (e.g. lighting, temperature, humidity, etc.) on a plant. Designing controlled chamber can fulfil the needs of restricted environmental factors. For example, if we need to study the effect of light on a specific plant under certain temperature, then we need to place the plant in a chamber where the temperature is controlled and the lighting can be turned on/off at desired time of the day. This is an example of an embedded system that can be programmatically controlled as needed. This is an integral part of a 3D plant phenotyping system.
Recently, laser scanning systems have gained popularity to capture 3D data on the surface of a plant. Laser scanning is an excellent way to perform non-invasive 3D analysis of plant phenotyping. Kinect laser scanners are now available at cheap cost and are being widely used in various fields including plant phenotyping, 3D modelling and remote sensing applications. Depending on the need of the application, the resolution of the scanner can be controlled. For example, dense spacing of points is needed in order to model a very detailed surface, whereas sparse scans might be sufficient for applications that do not need much local details to be modelled. The 3D model of the plant can be obtained by aligning multiple overlapping views of the scans taken from different directions around the plant. Then the merged point cloud data can be used to analyze different aspects of phenotyping.
Phenotyping refers to measuring (or quantifying) various biologically interesting properties of plants. There are several aspects of phenotyping. State-of-the-art techniques in computer vision are adopted for different application areas. For example, consider tracking a particular organ over time. To make the system fully automated, first we need to set up a robotic system that will be able to place the laser scanner at a suitable position, scan the plant and extract the point cloud data. Having the raw point cloud data, computer vision algorithms need to be designed to perform shape matching of point clouds in order to identify the organ of interest. As the plant grows, the robot should dynamically change its position to perform the scanning. There are many challenges involved in this type of phenotyping. As the plant grows bigger, the leaves occlude each other and the organ of interest might become partially or fully occluded by some other organs. In these cases, a single scan might not be sufficient to recognize the organ and the robot needs to be moved at suitable positions around the occlusion and take multiple scans of the occluded object. This is just an example of a challenging 3D phenotyping problem. Other types of phenotyping include the growth analysis of living plants. In that case, multi-view points cloud data needs to be aligned into a single point cloud, which can be triangulated to obtain a 3D mesh model of the plant. This is a challenging task since aligning multiple views of a complex plant structure is a difficult problem. Also, efficient triangulation is very important to retain the detailed structure of the plant. From the 3D mesh model, several properties like volume and surface area can be computed. Segmentation of plant organs has also gained attention for phenotyping tasks. Estimation of leaf area, stem diameter, etc. requires proper segmentation of the organs. Different types of shape primitives are often used to extract the required structure. For example, tubular shape fitting is a common technique to extract the round-shaped stem of a plant.
3D plant phenotyping is highly inspired from computer vision algorithms. Throughout this chapter, we will see some applications of 3D plant phenotyping. Although there are several components of phenotyping, we will discuss certain key areas, which are the most popular, as well as challenging. Initially we will discuss about automated systems for 3D plant phenotyping. Then registration of multi-view point cloud data are described. Next, some popular techniques of plant organ segmentation are studied. We also discuss in brief about related phenotyping problems to give an overview of the recent focus of the phenotyping research community.
The organization of the chapter is as follows. In the next section, related literature is summarized. Then we discuss the key techniques in 3D plant phenotyping, followed by the current challenges and concluding remarks. We end the chapter with some hands-on exercises related to plant phenotyping techniques.
2 Related Work
A large body of work has been reported on computer vision based plant phenotyping in the last decade. In recent literature, several aspects of plant phenotyping are discussed. Tremendous progress in automated plant phenotyping and imaging technologies have created a mini-renaissance [1]. Software products are becoming available for building high throughout phenotyping systems [2, 3]. Automated data collection systems [4, 5] are becoming prevalent over tedious manual techniques [6]. However, most of the automated systems have some limitations on the type, size or geometrical properties of the plants that can be processed. The ultimate goal of computer vision-based automation techniques is to generalize these type of systems [7,8,9]. Recently, a fully automated robotic system has been proposed [10]. The system works in a fully automated way throughout the lifetime of a plant and analyzes the growth pattern from the reconstructed 3D point cloud data. The system is general and can be customized to perform different types of automated tasks related to 3D plant phenotyping. We discuss the related literature on different aspects of plant phenotyping in the next subsections.
2.1 Organ Tracking
Detection and tracking of plant organ is a well-studied problem. Jimenez et al. [11] proposed a fruit harvesting system that can detect fruits of a plant from their colour and morphological properties. It was one of the first methods where laser scanners were used in plant phenotyping analysis. These types of systems are of extreme demand in agricultural applications. Chattopadhyay et al. [12] presented 3D reconstruction of apple trees for dormant pruning (cutting off certain primary branches to improve the yield and crop quality of the plant) applications in an automated manner. These techniques can be very helpful in the pruning process as a part of intelligent agricultural robotic application. Paulus et al. [13, 14] performed organ segmentation of wheat, grapevine and barley plants using a surface feature-based histogram analysis of 3D point cloud data. Klodt et al. [15] performed segmentation to monitor the growth of grapevines. Similar type of work on plant organ segmentation was proposed in [16] via unsupervised clustering technique. Paproki et al. [17] measured plant growth in the vegetative stage. They generated 3D point cloud from 2D images. Golbach et al. [18] setup a multiple camera system and the 3D model of the plant was reconstructed using a shape-from-silhouette method. Then the geometric properties (e.g. area, length) of the leaves and stems are computed by segmenting these organs. The final results are validated by comparing with the ground truth data obtained by destructing the plant by hand. Dellen et al. [19] built up a system to analyze leaf growth of tobacco plants by tracking the leaves over time from time-lapsed video. In each video frame, leaves are detected by assuming a circular leaf shape model. A graph-based method is employed to track leaves through the temporal sequence of video frames.
2.2 Plant Health Monitoring
Another type of plant phenotyping that has gained attention is determining the condition of a plant from specific patterns of its leaves. Usually, the texture properties of the leaves are exploited to perform the analyses, and then the leaves are tracked over time. A challenging task to perform this type of analysis is to segment the leaves in different imaging conditions [20]. Active contour model was used in [21] to detect lesions in Zea Mayes. This crop is widely used and lesion detection can be very helpful in disease detection at the early stages. Xu et al. [22] detected nitrogen-deficient tomatoes from the texture of the leaves. Tracking the leaves of rosette plants can be very useful for growth rate measurement. Similar type of work was presented in [23].
2.3 3D Reconstruction
3D reconstruction from multiple views is a quintessential part of many 3D phenotyping applications. This is a very challenging problem. The complex geometrical structure of the plants makes the problem extremely difficult to handle. Pound et al. [24, 25] reconstructed the 3D model of a plant using level set based technique. Santos et al. [26] performed a structure from motion technique to reconstruct the 3D point cloud model of the plant surface, and then a spectral clustering technique is used to segment the leaves. Similar type of approach was used for visual odometry applications in [27]. Kumar et al. [28] used a mirror-based system to obtain multiple views of the plant. A visual hull algorithm is used to perform the reconstruction. The setup alleviates the need for camera calibration due to the use of the mirrors. Recently, Gibbs et al. [29] proposed to improve the image acquisition that results in improving the overall 3D reconstruction. Instead of using a fixed camera position for all types of plants, they proposed to change the camera position dynamically, depending on the geometry of the plant. This type of approach can be embedded in the intelligent robotic systems. Simek et al. [30] modelled spatial smoothness of the branches of plant by Gaussian Process. Their method is designed to estimate the thin structures of a plant from monocular images. Brophy et al. [31] presented an approach to align multiple views of a plant into a single point cloud. The approach exploits recently successful Gaussian Mixture Model registration and mutual nearest neighbour techniques. However, the method needs a good initial guess, which can be rectified by automatic feature matching of junction points [32].
2.4 Rhythmic Pattern Detection
Rhythmic pattern of plant growth is a well-known phenomenon [33]. There have been attempts to capture the circadian rhythm of plant movements using imaging techniques [34]. Plant leaves are known to be affected by various lighting conditions. Dornbusch et al. [35] captured the effect of rhythmic leaf movements by lighting via laser scanning system. Tracking and growth analysis of seedling was studied in [36]. Corn seedling growth was studied by Barron and Liptay [37,38,39]. They demonstrated that the growth is well correlated with room temperature.
2.5 Structural Analysis
Structural analysis of plants is also studied in the literature [40]. Augustin et al. [41] extracted geometric features of Arabidopsis plant for phenotypic analysis. Li et al. [42] performed a 4D analysis to track budding and bifurcation events of plants from point cloud data.
3 Key Techniques
In this section, we will focus on some specific aspects of 3D plant phenotyping and explain the state-of-the-art techniques. Before we explain the key techniques, we briefly discuss some terminologies that will be used throughout this section. The flow of the section is as follows. First, we give an overview of some key terms. Then we discuss different types of automated systems for 3D plant phenotyping. These systems aim at collecting data without (or minimal) manual intervention. We focus on 3D model building of plants, and that is why we discuss about aligning multiple datasets to obtain a 3D point cloud model of a plant. Although the problem is basically the general point cloud registration and alignment problem, we will discuss certain variations of the standard algorithms related to plant structures. Finally, some segmentation algorithms are discussed.
3.1 Terminologies
3.1.1 Point Cloud
A point cloud is simply a set of data points. Point clouds are typically generated by range scanners, which record the point coordinates at the surface of an object. The density of points depends on the scanner settings. In the general case, high density of points encodes fine geometry of the object, and requires high computation time to process the data. On the contrary, low density of points encodes less local geometry and mostly keep the global shape of the object, and usually requires less computational time to process. 3D point clouds are usually stored as raw coordinate values (x, y, z). However, the fourth attribute can be the colour or intensity information, depending on the type of scanner used. Among different file formats for storing the point cloud, the most commonly used extensions are: .xyz, .pcd, .asc, .pts, and .csv formats.
3.1.2 3D Mesh
A 3D mesh or a polygonal mesh is a data structure that connects the points in the cloud by means of a set of vertices (which are the points themselves), a set of edges, and polygonal elements (e.g. triangles for triangular mesh). Polygon meshes are also referred as surface meshes which represent both the surface and the volumetric structure of the object. The process of making a triangular mesh is also called the triangulation. The efficient rendering of the triangles can produce a realistic representation of a synthetic object, which is a center of attention in the computer graphics research community. Among different types of triangulation technique, most commonly used are the Delaunay and Alpha shape triangulation algorithms. Let us consider a set of points \(P = \{p_1,\ldots ,p_n\} \subset \mathbb {R}^d\). Let’s call these as sites. A Voronoi diagram is a decomposition of \(\mathbb {R}^d\) into convex polyhedra. Each region or Voronoi cell \(\mathcal {V}(p_i)\) for \(p_i\) is defined to be the set of points x that are closer to \(p_i\) than to any other site. Mathematically,
where || . || denotes the Euclidean distance. The Delaunay triangulation of P is defined as the dual of the Voronoi diagram.
The \(\alpha \)-complex of P is defined as the Delaunay triangulation of P having an empty circumscribing sphere with a squared radius equal to or smaller than \(\alpha \). The Alpha shape is the domain covered by alpha complex. If \(\alpha = 0\), the \(\alpha \)-shape is the point set P, and for \(0 \le \alpha \le \infty \), the boundary \(\partial \mathcal {P}_\alpha \) of the \(\alpha \)-shape is a subset of the Delaunay triangulation of P.
3.1.3 Registration of Point Cloud
In the general sense, registration of two point clouds refers to aligning one point cloud to the other. One of the point cloud is called the model point set, which remains “fixed” in space. The other point cloud, referred as the data, is the “moving” point set. We seek to find the transformation parameters (typically the rotation, translation and scaling) of the data point cloud, that best aligns it to the model point cloud. By best, we mean the alignment that has the minimal error with respect to the ground truth. There are usually two cases in this regard: rigid and non-rigid. Rigid point cloud registration problems are usually easier to handle, since estimation of the transformation parameters is relatively less complicated. On the other hand, non-rigid point cloud registration problems are hard in nature, and typically a single set of transformation parameters are not sufficient to align the data to the model point cloud. Among various types of challenges associated with non-rigid point cloud registration, the following are the most prevalent ones: occlusion, deformation and minimal overlap between the two point clouds.
3.1.4 Viewing Software
There are a variety of software available to visualize the point clouds and meshes. The following software are widely used in the computer vision and graphics community: Meshlab,Footnote 1 CloudCompare,Footnote 2 Point Cloud LibraryFootnote 3 (also offers lots of functionalities for point cloud processing), etc.
3.2 Automated Systems for 3D Phenotyping
The goal of a high throughput plant phenotyping system is to monitor a mass crop production system and analyze several phenotypic parameters related to growth, yield and stress tolerance in different environmental conditions. An automated green house system looks like the one in Fig. 14.1. The plants are placed on conveyor belts and the image acquisition devices capture images in different time frames.
A typical green house system [2]. Plants are placed on conveyor belts and images are taken automatically as the belt moves around (licensed under the Creative Commons Attribution 4.0 International License)
In many applications, the phenotyping demands high precision results. In order to obtain high precision quality, robots can be used to perform the task in more efficient manner. Subramanian et al. [5] developed a high throughput robotic system in order to quantify seedling development. A 3-axis gantry robot system is used to move the robot in the vertical X-Z plane. The movement of each axis is controlled by linear servo motors. Two cameras are attached to the robot, one of them is of high resolution and the other one is of low resolution. Perpendicular to the optical axis of the cameras, a series of petri dish containing plant seedlings are attached to a sample fixture.
The robot periodically moves along the gantries and captures images of each petri dish. A probabilistic localization is performed to locate the seedling. Focusing is also performed automatically. As the seedlings grow over time, the system dynamically analyzes the images with high accuracy. This type of automated system is very useful in studying growth of mini-seedlings (a young plant grown from seed).
However, the system described above is not designed to monitor a whole plant throughout its lifetime. Also, the robot system does not have enough degrees of freedom (DOF) to move anywhere around a plant to perform real-time 3D data capture. Recently, a machine vision system has been proposed in order to perform real-time 3D system [10]. The system is fully automated, including the growth chamber, robot operation, data collection and analysis. A naive user can obtain phenotyping results with a few mouse clicks.
The system comprises of an adjustable pedestal and a 2-axis overhead gantry which carries a 7-DOF robotic arm. A near-infrared laser scanner is attached at the end of the arm, which can measure dense depth map of the surface of an object. The arm provides high level of flexibility for controlling the position and orientation of the scanner attached at the end. The plant is placed on a pedestal, which can be moved vertically to adjust the room for different plant sizes. The whole setup is housed in a programmable chamber, which is fully controllable in terms of lighting, temperature, humidity, etc. Different components of the system are integrated together as a single system. The setup is shown in Fig. 14.2.
During an experiment, the chamber is programmed according to the requirements. The plant is placed on the pedestal, and other parameters such as number of scans, resolution, timings, etc., are provided by the user. Initially the robot remains at the home position. When the scan starts, the robot moves to the scan position and takes an initial scan. This initial scan is performed to compute the bounding box comprising the whole plant. The bounding box calculation is needed to dynamically change the robot position as the plant grows over time. When the bounding box is determined, actual scanning is performed. The laser scanner records point cloud data in xyz format on the surface of the plant. If the scanner field of view is not able to enclose the whole plant due to size constraint, multiple overlapping partial scans are taken. After the first scan, the robot moves to the next scanning position and performs a similar scanning routine. When the scan sets are complete, the robot goes back to the home position until the next set of scan is scheduled. Captured data are transferred to the server automatically and processing gets started immediately (aligning multiple views are discussed in the next subsection).
The system is designed for general use of phenotyping applications, and can be customized according to the need. The robot arm can be exploited for tracking specific organs by exploiting the high degree of flexibility of the arm. To handle the case of occlusion of the organs, the robot arm can be programmed to move to specific coordinates in order to obtain full view of the occluded organ. In recent years, automated data collection through robotic system is performed in outdoor environment for agricultural applications [43, 44].
3.3 Multiple-View Alignment
Although registration of 3D point cloud data has been studied extensively in the literature, registration of plant structures is a challenging task. The self recursive and thin structure makes the problem of pairwise registration extremely complicated and non-rigid. Although different types of approaches exist for solving the pairwise registration and multiple view alignment problem, recently probabilistic methods have been successful in many applications. We first discuss the background of general registration problem and then end by discussing the adaptation of the technique for registration of plant structures.
Recently, Gaussian Mixture Models (GMM) have been very successful in the registration of non-rigid point sets. Let us consider two overlapping views (point clouds) of a plant. One point cloud is called the model point set, and the other is called the data point set. The target is to transform the data point set to the model point set in order to obtain the merged point cloud. Mathematically, let’s say the model point set is denoted as \(\mathcal {M}=(x_1,x_2,...,x_M)^T\), and the observed data point set is denoted as \(\mathcal {S}=(y_1,y_2,...,y_N)^T\). The model point set undergoes a non-rigid transformation \(\mathcal {T}\), and our goal is to estimate \(\mathcal {T}\) so that the two point sets become aligned. Then the GMM probability density function can be written as,
where \(z_n\) are latent variables that assign an observed data point \(y_n\) to a GMM centroid. Usually all the GMM components are modelled as having equal covariances \(\sigma ^2\), and the outlier distribution is considered as uniform, i.e. 1/a, where a is usually set as the number of points in the model point set. The unknown parameter \(\omega \in [0,1]\) is the percentage of the outliers. The membership probabilities \(\pi _{mn}\) are assumed to be equal for all GMM components. Denoting the set of unknown parameters \(\varvec{\theta } = \{\mathcal {T}, \sigma ^2, \omega \}\), the mixture model can be written as
The goal is to find the transformation \(\mathcal {T}\). Sometimes a prior is used to estimate the transformation parameters. A common form of prior [45] is
where \(\phi (\mathcal {T})\) is the smoothness factor, and \(\lambda \) is a positive real number. The parameters \(\varvec{\theta }\) are estimated using the Bayes’ rule. The optimal parameters can be obtained as
which is equivalent to minimizing the negative log-likelihood:
Jian and Vemuri [46] followed this approach and represented the point set by Gaussian mixtures. They proposed an approach to minimize the discrepancy between two Gaussian mixtures by minimizing the \(L_2\) distance between two mixtures.
The Coherent Point Drift (CPD) registration method was proposed by Myronenko et al. [47, 48]. Their method is based on GMM, where the centroids are moved together. Given two point clouds, \(\mathcal {M}=(x_1,x_2,...,x_M)^T\) and \(\mathcal {S}=(y_1,y_2,...,y_N)^T\), in general for a point x, the GMM probability density function will be \(p(x) = \sum _{i=1}^{M+1} P(i)p(x|i)\), where:
They minimize the following negative log-likelihood function to obtain the optimal alignment:
There are many ways to estimate the parameters, such as gradient descent, Expectation Maximization (EM) algorithm and variational inference. EM is a standard and widely used technique to optimize the cost function. Basically the E-step (or the Expectation) computes the posterior probability, and the M-step (or the Maximization) computes the new parameter values from the likelihood function. The aim is to find the parameters \(\varvec{\theta }\) and \(\sigma ^2\).
Let us denote the initial and updated probability distributions as \(P^{old}\) and \(P^{new}\), respectively. The E-step basically computes the “old” parameter values, and then computes the posterior probability distributions \(P^{old}(i|x_j)\). In the M-step, the new parameter values are computed by minimizing the log-likelihood function:
which can be rewritten as
where
Now the current parameter values \(\varvec{\theta }^{old}\) is used to find the posterior probabilities:
Although pairwise registration works reasonably well, aligning multiple views is problematic. The reason is that, the errors from pairwise registrations accumulate during multiple alignment and as a result, the merging does not yield good results. To handle this problem, Brophy et al. [31] proposed a solution based on the Mutual Nearest Neighbour (MNN) [49] algorithm. The algorithm is based on CPD which can align many views with minimal error. More specifically, it is a drift-free algorithm for merging non-rigid scans, where drift is the build-up of alignment error caused by sequential pairwise registration. Although CPD alone is effective in registering pairs with a fair amount of overlap, when registering multiple scans, especially scans that have not been pre-aligned; this method achieves a much better fit both visually and quantitatively than CPD by itself, utilizing sequential pairwise registration.
First, the scans are aligned sequentially, and then a global method is used to refine the result. The global method involves registering each scan \(X_i\) to an “average” shape, which we construct using the centroids of the mutual nearest neighbors (MNN) [49] of each point. For \(X_i\), we use scans \(X_j\) where \(j \ne i\) to obtain the average shape \(Y_{cent}\) from the centroids, and \(X_i\) is then registered to this average shape. This is repeated for every scan until the result converges.
For a point x, the density function is written as
where \(\hat{y}_i \in Y_{ cent }\) are the points in the target scan \(Y_ cent \), which is constructed from all scans other than itself.
For a pair of scans X and Y, a point \(x_i \in X\) and \(y_j \in Y\) is called MNN if \(x_i = x_{i_n}\) and \(y_{j_n} = y_j\), where
and
For each point \(x_j\) in scan \(X_i\), the set of points \(\{ x_k | x_k \in X_l \wedge MNN ( x_k,x_j ) \}\) are found, where \(l \ne i\), i.e. all scans other than \(X_i\). For each of these sets of points \(x_k\), the centroid is computed as
\(X_{cent}\), the set of centroids calculated for each \(x_j\), is registered to scan \(X_i\).
3.3.1 Approximate Alignment
In general, to reconstruct a 3D model of a plant, a set of scans are captured around the plant at specific increments. After acquiring them, the idea is to solve for the rigid transformation \(T_0 = (R_0, \mathbf {t}_0)\) (where R is a rotation matrix and \(\mathbf {t}\) is a translation vector) between the first scan (\(X_0\)) and the second scan (\(X_1\)) using the rigid version of CPD. After we solve for \(T_0\) only once, for each scan \(X_i\), the transformation is applied i times
The new set of transformed scans \(\hat{X}\) should now be roughly aligned in the coordinate system of the last scan. This method is used to obtain a rigid registration. The initial registration is important when the pair of scans to be registered has minimal overlap. The result of approximately aligned scans on some real plant data is shown in Fig. 14.3.
12 scans of the Arabidopsis plant, prior to registration, but with rotation and translation pre-applied. Different colours indicate different scans [31]
3.3.2 Global Non-rigid Registration via MNN
Once the initial registration is complete, CPD is used in conjunction with MNN to recover the non-rigid deformation field that the plant undergoes between the capture of each scan. At this point, the scans should be approximately aligned to one another. The centroid/average scan is constructed and the scan is registered to it.
3.3.3 Global Registration
Algorithm 14.1 is used to merge all scans, where MNN(\(\cdot \)) computes the mutual nearest neighbour for each point in scans \(X_i\) and \(X_j\) and the centroids function likewise takes the centroids computed for each point in each scan and combines them into one average scan using Eq. 14.13. For each point in scan \(X_i\), the single nearest neighbour from all other scans is found and we use this set of distances to compute the \(L^2\)-norm.

Figure 14.4 shows all 12 scans, merged into a single point cloud after subsampling each scan. Each colour in the point cloud represents a different scan.
12 scans captured in \(30^\circ \) increments about the plant and then merged into a single point cloud using MNN. Shown from two viewpoints, are the front facing scans on the left and the above facing scans on the right. Different colours indicate different scans [31]
A problem with GMM based registration is that the views need to have been approximately aligned before the registration. For large rotation angle differences, the algorithm fails drastically. In the literature, there has been significant work on feature matching of two point cloud datasets. For example, Fast Feature Point Histogram (FPFH) [50] is a popular technique for feature matching in point cloud. However, these type of descriptors exploit surface normal information to uniquely characterize an interest point. For thin structured plant data, accurately computing surface normal is an extremely difficult and error-prone task. Traditional descriptors fail to produce reasonable results for plant feature correspondence.
Bucksch et al. [51] presented a method to register two plant point clouds. Their method performs skeletonization of the input point cloud and then estimates the transformation parameters by minimizing point-to-line distances. The idea is to map a point \(p_0\) from one point cloud to the line joining two nearest neighbour points \(p_1\) and \(p_2\) in the skeletonized second point cloud. That is, the mapping condition is the following:
where \(p'_0 = R p_0 + t\) is the transformed point, R is the rotation matrix and t is the translation vector. However, the algorithm needs the point clouds to be roughly aligned in order to obtain good registration results.
A remedy to the above problem can be obtained by exploiting the junction points as features, as proposed by Chaudhury et al. [32]. The advantage of using junctions as feature points is that, even if there is deformation and non-rigidity in the point cloud data, a junction point will not be affected by these factors. Initially, the neighbourhood of each 3D point is transformed into 2D by performing the appropriate 3D coordinate transformations. The method is two step. First, a statistical dip test of multi-modality is performed to detect non-linearity of the local structure. Then each branch is approximated by sequential RANSAC line fitting and an Euclidean clustering technique. The straight line parameters of each branch are extracted using Total Least Squares (TLS) estimation. Finally, the straight line equations are solved to determine if they intersect in the local neighbourhood. Such junction points are good candidates for subsequent correspondence algorithms. Using these detected junction points, the correspondence algorithm is formulated as an optimized sub-graph matching problem.
3.3.4 Coordinate Transformation
Using a kd-tree algorithm, the nearest neighbour points of a point within a certain radius can be obtained. Given such points in a local neighbourhood about some 3D point, the data is transformed so that the surface normal of the plane fitting the data is a line-of-sight vector (0, 0, 1). More specifically, the center of mass \((x_{cm},y_{cm},z_{cm})\) of the neighbourhood 3D points is computed first. To reformulate as a 2D problem, the following steps are performed: translate the origin to the center of mass by \(-(x_{cm},y_{cm},z_{cm})\), rotate about the x-axis onto the \(x-z\) plane by some Euler angle \(\alpha \), rotate about the y-axis onto the longitudinal axis (0, 0, 1) by some Euler angle \(\beta \) and finally transform the origin back to the previous location by \((x_{cm},y_{cm},z_{cm})\). The detailed calculations are shown below.
A plane of the form \(ax+by+cz+d=0\) is fitted to the neighbourhood data and the parameters are obtained. Consider 3 points on a planar surface: \(P_1(x_1,y_1,z_1)\), \(P_2(x_2,y_2,z_2)\) and \(P_3(x_3,y_3,z_3)\). Compute the vectors \(\mathbf {V_1}\) and \(\mathbf {V_2}\) (see Fig. 14.5) as
Then \(\mathbf {V_1} \times \mathbf {V_2}\) is the normal to the surface \(ax+by+cz+1=0\). That is, \(\mathbf {V_1} \times \mathbf {V_2}\) and \((a, b, c)^T\) are in the same direction.
We aim to nullify the effect of z-coordinates, which require the following steps. First we translate the origin to the center of mass (CM) \((-x_m, -y_m, -z_m)\) so that the origin coincides with the CM. We use 4D homogeneous coordinates to perform all the matrix multiplications. In 3D heterogeneous coordinates, translation is specified as vector addition but in the equivalent 4D homogeneous coordinates it is now specified by matrix multiplication, as are all the other operations, allowing matrix concatenation of all matrices to be performed by one matrix. The 4D homogeneous transformation matrix has the following form:
Thus, \(T(-x_m, -y_m, -z_m)\) does the translation to the center of mass (the new origin). Next we project the rotation axis onto the z-axis. This requires two steps: rotate by some unknown \(\alpha \) angle about x-axis so that the vector \(\hat{u}\) is in the xz-plane, and then rotate by some unknown \(\beta \) angle about the y-axis to bring vector \(\hat{u}\) onto the z-axis. We show how to calculate \(\alpha \) and \(\beta \) in the next 2 subsections. Finally we re-translate back the origin to the previous location by the inverse translation \(T(x_m,y_m,z_m)\).
Let us consider rotation about the z-axis. In that case, \(\mathbf {V}\) is the rotation axis with endpoints \((x_1,y_1,z_1)\) and \((x_2,y_2,z_2)\). We rotate about \(\mathbf {V}\) (see Fig. 14.6), given by
In this case, \(\hat{u} = \frac{\mathbf {V}}{||\mathbf {V}||_2} = (a,b,c)\) is the unit vector in \(\mathbf {V}\)’s direction. The direction cosines of \(\mathbf {V}\) (the Euler angles) are given by
We use the following convention: \(\hat{u}\) is the normal vector, and \(\mathbf {u}\) is the unnormalized vector (of the projection of \(\hat{u}\) onto \(y-z\) plane).
3.3.5 Rotate \(\hat{u}\) into the XZ-Plane
Let \(\alpha \) be the rotation angle between the projection of \(\mathbf {u}\) in the yz-plane and the positive z-axis and \(\mathbf {u}^{\prime }\) be the projection of \(\hat{u}\) in the yz-plane (Fig. 14.7). That is, \(\hat{u}=(a,b,c)^T \implies \mathbf {u}^{\prime }=(0,b,c)^T\).
Then the angle \(\alpha \) can be obtained simply from the equation,
Let \(\hat{k}=(0,0,1)\) is the unit vector in the z-direction, i.e. \(||\hat{k}||_2=1\). Then
Thus,
The vector product can also be used to compute \(\sin \alpha \). Note that \(\mathbf {u}^{\prime } \times \hat{k}\) is a vector in x’s direction, i.e., \(\hat{i}\). Then
and
Then \(b\hat{i} = \hat{i} \sqrt{b^2+c^2}\sin \alpha \), or \(\sin \alpha = \frac{b}{\sqrt{b^2+c^2}}\).
Given \(\sin \alpha \) and \(\cos \alpha \), we can specify the 4D homogeneous rotation matrix for rotation about the x-axis as
This matrix rotates \(\hat{u}\) onto the xz-plane.
3.3.6 Align \(\hat{u}_{xz}\) Along Z-Axis
As shown in Fig. 14.8, we need to compute \(\sin \beta \) and \(\cos \beta \) in this case.
Using the dot product we can write
Also, using the vector product, \(\hat{k} \times \hat{u}_{xz}\) is a vector in the direction of the y-axis, thus resulting
and
Thus, \(-a\hat{j} = \hat{j}\sin \beta \), or \(\sin \beta = -a\). Then the 4D homogeneous rotation matrix about the y-axis can be specified as
which aligns \(\hat{u}_{xz}\) with the z-axis. Thus we apply the transformations:
to all 3D points. If we wish to undo this transformation we could use
where \(R^T_x(\alpha ) \equiv R^{-1}_y(\alpha )\) and \(R^T_y(\beta ) \equiv R^{-1}_y(\beta )\) because rotation matrices are unitary and orthogonal.
Next, a plane of the form \(ax+by+cz+d=0\) is fit to the neighbourhood data using Cramer’s rule. The parameters \(\mathbf {n}=(a,b,c)\) are the plane’s surface normal. Since these transformations result in vertical surface normals we need only be concerned with the structure in the \(x-y\) plane, i.e. the problem is now 2D.
3.3.7 Dip Test for Multi-modality
The detection of multi-modality in numeric data is a well-known problem in statistics. A probability density function having more than one mode is denoted as a multi-modal distribution. Hartigan et al. [52] proposed a dip test for unimodality by maximizing the difference between the empirical distribution function and the unimodal distribution function. In the case of a unimodal distribution, the value for the dip should asymptotically approach 0, while for the multi-modal case it should yield a positive floating point number. Zhao et al. [53] exploited this idea to detect bifurcations in the coronary artery. A similar idea is applicable in this case too. Points having non-linear local neighbourhood are potential candidates for a junction point. The idea is to perform the dip test for a local neighbourhood of a point. If it is a stem or a leaf, the data should be uniform and the distribution should only be unimodal. For a junction point likely due to a bifurcation, it should exhibit multi-modality. The dip value can thus be used as a measure of multi-modality (Fig. 14.9).
Distribution of data: column 1 are the neighbourhood point clouds under consideration, column 2 are the histograms of the x coordinate distribution and column 3 are the histograms of the y coordinate distribution for a single stem (first row), a leaf (second row) and a stem with 2 branches (last row). The later is potentially a junction point
The dip test is performed along the x and y directions (note that as we have reduced the dimensionality from 3D to 2D the z-coordinates can be ignored) and obtain the maximum dip value. The neighbourhood is determined to be multi-modal if the dip value is over some threshold. However, the threshold value of the dip value is highly dependent on the data and should be tuned carefully (done visually for now).
The dip measurement is used for initial filtering of non-junction neighbourhood data. Note that non-linearity and high dip values in local neighbourhood do not guarantee that those points are junction points. For some leaf and stem data, sometimes the data shows high dip values. Instead of relying blindly on dip test results, further processing is needed in order to confirm the presence of junction in the neighbourhood.
3.3.8 RANSAC Fitting and TLS Approximation
Consider the case of a maximum three branches at an intersection point (which is typically the case in real life): the branches may intersect at a single point (the red dot in Fig. 14.10c) or at two different points (the red dots in Fig. 14.10d).
Examples of detected junction points (red dots) on the real Arabidopsis plant data [32]
Assuming the fact that the main stem will be thicker than the branches, the thick stem can be extracted simply by using RANSAC straight line fitting using a high distance threshold for inliers. Other branches can be estimated by sequential RANSAC fitting. However, there may be other points due to additional branches, a leaf or a noise event (or some combination of the three). After removing the RANSAC fitted main stem, Euclidean clustering is performed on the rest of the data to choose the biggest connected component(s) to extract the sub-branches. Two sets of points, \(\mathcal {X}_i = \big \{ p_i \in \mathcal {P}\big \}\) and \(\mathcal {X}_j = \big \{ p_j \in \mathcal {P}\big \}\) form two different clusters, if the following condition holds
where \(\tau \) is the distance threshold. The branches may be straight or curved, but by using RANSAC we can estimate the principal direction of the branch [54]. A criterion is imposed to estimate a broken branch shape (due to occlusion): two branches are merged if they are spatially close to each other and have the roughly same direction.
After estimating the points for each branch, we need to know the straight line parameters in order to estimate their intersection (if any). We use TLS to approximate the straight line represented by a set of points in a branch and extract the parameters. Consider a set of points \((x_1,y_1),\ldots ,(x_n,y_n)\) and the normal line equation \(ax+by+c=0\). [Note that a is \(\cos (\theta )\) and b is \(\sin (\theta )\) where \(\theta \) is the angle of the normal line with respect to the positive x axis and c is minus the magnitude of the line from (x, y) to (0,0).] To fit all the points to the line, we have to find parameters a, b and c to minimize the sum of perpendicular distances, i.e. we minimize
(as \(\cos ^2\theta +\sin ^2\theta = 1\)). Equating the first order derivative to zero we get
Replacing c in Eq. (14.19) with its value in Eq. (14.20) we obtain
To minimize the above equation we rewrite it in the following form:
The expression in the right-hand side of the above equation can be written as \((UN)^T(UN)\), where U is an \(n\times 2\) matrix having rows \((x_i - \bar{x}, y_i - \bar{y})\) and N is \((a,b)^T\). Setting \(\frac{dE}{dN} = 0\), we obtain \(2(U^TU)N = 0\), the solution of which (subject to \(||N||^2 = 1\)), is the eigenvector of \(U^TU\) associated with the smallest eigenvalue. We extract the parameters a, b, \(\bar{x}\) and \(\bar{y}\) from the equation \(a(x_i-\bar{x}) + b(y_i-\bar{y}) = 0\).
After approximating the straight lines, we solve the equations given below to determine if these lines intersect or not. Recall that two branches are approximated by two straight lines, and the presence of junction is confirmed if the lines intersect. For two straight line equations of the form \(ax+by+c=0\) and \(dx+ey+f=0\), the intersection point can be obtained as
If the straight lines are parallel, the discriminant \((ae-db)\) will be equal to zero. If the lines are non-parallel, we check if the intersection point is contained in the local neighbourhood or not. Note that the obtained intersection point is 2D, so we apply the reverse transformation to find the actual 3D point. Finally non-maximal suppression is performed based on the highest dip value to reduce the number of points.
3.3.9 Correspondence Matching
The detected junction points from the last phase are potential candidates for correspondences and can be used as features points for matching. For raw 3D point cloud data, local surface normals, neighbourhood information, etc., are typically used for encoding the local structure and points are matched based on the descriptor similarities. This idea typically fails for plant data because the thin structures do not allow for good local surface normal calculations and because of deformations, the local structure can change abruptly in adjacent images. An approach to solve the problem is to exploit sub-graph matching theory as discussed below.
First, the data is triangulated using Delaunay triangulation in 3D (note that we converted the problem temporarily to 2D just for detection of junction points). Using the vertex information from triangulation, we can construct a graph connecting all the points. To handle the cases of missing or occluded data, the points to the nearest triangle vertex can be connected so that all the points are included in a single graph. Then, for each junction point, Dijkstra’s shortest path algorithm can compute geodesic distance to all other junction points. The same procedure is followed for the second point cloud as well. Then the pairwise distances will be used to be the criteria for graph matching.
Consider two graphs \(G_1=(V_1,E_1)\) and \(G_2=(V_2,E_2)\). Each junction point is considered to be a node of the graph. Each node stores the geodesic distances to all other nodes. In the end, this yields a set of edges. Compatibility of two nodes in \(G_1\) and \(G_2\) are defined as a closest distance match. For example, let us suppose two graphs \(G_1\) and \(G_2\) have \(n_1\) and \(n_2\) nodes. Each node \(V_{1_i}\) in \(G_1\) stores all distances to all other nodes. We denote this is as the set of attributes of node \(V_{1_i}\): \(\mathcal {D}_{v_{1_i}} =\{d_{v_{1_i}v_{1_j}}\}, \forall j \in n_1\). Similarly in \(G_2\), the set of attributes of node \(V_{2_i}\) is defined as \(\mathcal {D}_{v_{2_i}} =\{d_{v_{2_i}v_{2_k}}\}, \forall k \in n_2\). The compatibility of two nodes, \(V_{1_i}\) and \(V_{2_i}\), are formulated as the sum of the squares of the difference of nearest distances, multiplied by the number of matches. Suppose \(G_1\) and \(G_2\) contain 5 and 7 nodes, respectively. Let the attributes of a node \(V_{1_i}\) contain the following distances: \(\{d_1, d_2, d_3, d_4\}\) (ignoring self distance). Similarly, \(V_{2_i}\) contains the distances \(\{d'_1, d'_2, d'_3, d'_4, d'_5, d'_6\}\). We use a threshold \(\epsilon \) (\(=0.2\)) for the match of two distances. Suppose there are 3 distance matches given by \(d_1 \sim d'_4\), \(d_3 \sim d'_2\), \(d_4 \sim d'_1\). Then the affinity of the two vertices is computed as
The logic for using this kind of distance matching is that any outlier is likely to be eliminated by a lower number of matches. On the other hand, compatible points will only have the maximum number of distance matches.
Using the compatibility of two vertices, we can obtain the initial node correspondence by using the Hungarian algorithm [55]. The outliers are likely to get rejected by unmatched distance attributes. However, there still may be non-optimal matches of the vertices. Cour et al. [56] proposed a graph matching technique, which is shown to be robust and unambiguous. Given two graphs, \(G_1=(V_1,E_1,A_1)\) and \(G_2=(V_2,E_2,A_2)\), where each edge \(e=V_iV_j \in E\) has an attribute \(A_{ij}\). The objective is to find \(\mathcal {N}\) pairs of correspondences \((V_i,V_j)\) where \(V_i \in V_1\) and \(V_j \in V_2\). The affinity \(A_{ij}\) (Eq. 14.24) defines the quality of the match between nodes \(V_i\) and \(V_i'\). Denoting the similarity function of pairwise affinity as \(f(\cdot ,\cdot )\), the matching score can be computed as:
Representing \(\mathcal {N}\) as a binary vector x so that \(x(ii^\prime )=1\) if \(ii^\prime \in \mathcal {N}\), the above equation can be written as
where \(W_{ii',jj'}=f(\mathcal {A}_{ij}, \mathcal {A'}_{i'j'})\). The optimal solution of the above equation is given by
The permutation matrix provides the correspondence among the vertices (or the junction points in this case). Finally the outliers (or wrong matches) can be pruned out using RANSAC.
3.4 Organ Segmentation
Classification of different plant organs from point cloud data is an important plant phenotyping. Segmenting leaves, stems, fruits and other plant parts can help in tracking specific organ over time. There are different approaches of organ segmentation in the literature. Paulus et al. [13] presented a surface feature-based histogram technique to segment stems and leaves of grapevines, and wheat ears. The method is based on Fast Point Feature Histogram (FPFH) [50]. The idea is to build histogram for each point in the cloud and classify the histograms using Support Vector Machine (SVM). For every point p in the cloud, the surface normal is computed by considering a local neighbourhood within radius \(r_H\) around the point. For each point \(p_n\) in the neighbourhood, three types of features are computed. Let us say \(n_p\) and \(n_{pn}\) are the estimated normals at p and \(p_n\), respectively. A coordinate frame uvw is defined as follows:
Then for each p and \(p_n\), the following features are computed,
Then for every point, a histogram is built, where the index of the histogram bin is calculated by using the following formula: \(\sum _{i=0}^2 (\frac{f_i \cdot b}{f_{i(max)}-f_{i(min)}}) \cdot b^i\), where b is the division factor for the histogram size.
Next, the histogram for each point is represented as the normalized weighted sum of the histograms of the neighbouring points. For k neighbours around a point p having their histograms as \(h_n(p_k)\), the weighted histogram of a point is expressed as
\(wb = 1 - (0.5 + \frac{d}{r_H} \cdot 0.5)\), d is the distance from the source to the target point. These histograms encode primitive shapes like plane, sphere, cone and cylinder, which will be able to classify plant organs like flat leaf surface, cylinder-shaped stems, etc. The histograms are classified using SVM.
Wahabzada et al. [16] developed an unsupervised clustering method as an extension of the histogram-based method discussed above. The idea is to compare the histograms by some efficient metric, and then perform clustering like k-means using Euclidean distance measure. However, Euclidean distance metric performs poorly in presence of noise. As other alternatives, two different types of distance measures are used for histogram comparison. The first one is the standard Kullback–Leibler (KL) divergence, which uses Hellinger distance for computing the distance between two histograms. This is basically a probabilistic analog of the Euclidean distance. For two histograms \(\mathbf {x}\) and \(\mathbf {y}\), the Hellinger distance is given by
The other metric is to use the Aitchison distance given by
where \(g(\cdot )\) is the geometric mean. The k-means objective function is iteratively optimized by the standard EM algorithm.
In a different approach, Li et al. [42] formulated the organ segmentation task as an energy minimization problem. The goal of their work was to detect events (such as budding and bifurcation) from time lapse range scans. A key stage to detect these events is to segment the plant point cloud into leaves and stems. The problem is formulated as a two stage binary labelling problem. In the first stage of labelling, leaves and stems are classified, and in the second stage, individual leaves are classified separately. An organ hypothesis \(\mathcal {H}^t\) is formulated as, \(\mathcal {H}^t := L_l^t \cup S_s^t\) for frame \(F^t\), where L and S are leaf and stem categories, \(L_l^t\) is the l-th leaf, \(S_s^t\) is the s-th stem. For any point, \(p^t\) in the point cloud \(\mathcal {P}^t\) in the current frame \(F^t\), the aim is to find a labelling that maps \(p^t\) into \(\mathcal {H}^t\). The first stage finds a binary labeling \(f_B\) that maps \(\mathcal {P}^t\) to \(\{L^t,S^t\}\), and the second stage consists of two labellings \(f_L\) and \(f_S\) that decompose \(L^t\) and \(S^t\) into individual leaves \(L_l^t\) and \(S_s^t\).
The energy function to find the labelling \(f_B\) is formulated as
where \(\mathcal {N}_{\mathcal {P}^t}\) is the neighbourhood around a point. The data term \(D_{p^t}\) penalizes the cost of classifying \(p^t\) as leaf or stem, and the smoothness term \(V (f_B(p^t),f_B(q^t))\) ensures spatial coherence.
The data term is formulated based on the curvature values, considering the fact that leaves are generally flatter than stems. For the smoothness term, penalty of labelling is designed as high for neighbouring points to different organs, but less near organ borders. The term is defined as
where \(C(p^t)\) is the curvature of \(p^t\), obtained from the eigenvalues from principal component analysis of neighbourhood points.
Similar approach is followed for labelling in the second stage, with some modifications. To segment the individual leaves (which might be touching each other, thus forming a single connected component), adjacent frames are looked at simultaneously to confirm the hypothesis. Similar hypothesis is built for the data term, and short stems are trimmed out based on a threshold. The energy is minimized by the well-known \(\alpha \)-expansion algorithm [57, 58].
A more recent work on segmentation can be found in the work of Jin et al. [59]. They performed segmentation of stem and leaf of Maize plants on 3D Light detection and ranging (LiDAR) data.
4 Main Challenges
There are several challenging problems associated with vision-based plant phenotyping. First of all, no efficient registration algorithm exists that can handle every dataset. In the presence of occlusion and non-rigidity, most of the existing algorithms fail to generate good results. Incorporation of prior knowledge about the plant structure in the registration process might be worth studying. Environmental factors such as wind make a plant to jitter constantly. This can make the pairwise registration problem extremely challenging. Occlusion is still an unsolved problem. Handling these cases are open research problems. Also, the optimal number of scans needed to capture the geometric details of a plant has not been studied.
In general, Delaunay or alpha shape triangulations are widely used to polygonize 3D point cloud data. However, in order to retain the thin structure, perfect tuning of the parameters is very crucial in these cases. However, if the application demands very tiny details to be visible in the polygonized mesh, more efficient triangulation algorithms will be more demanding.
Regarding 3D point cloud segmentation methods, although it has been studied widely in the literature, the problem is still challenging for different scenarios with complex background. Also, segmentation in the case of highly occluded point cloud structure is a challenging problem. In fact, the problem is more complicated in terms of generalizing the algorithm for the sheer variety of phenotypes presented by plants.
5 Conclusion
This chapter has summarized the basic concepts of some recently successful 3D plant phenotyping techniques. Emphasis has been put on an automated system for 3D phenotyping, pairwise registration and alignment of plant point cloud data and organ segmentation techniques. Vision-based plant phenotyping is becoming more demanding these days, dedicated conferences and workshops are getting organized frequently.Footnote 4 \(^,\)Footnote 5 Challenging datasets are getting released also. Although we have not covered the recent deep learning techniques in plant phenotyping, interested readers are encouraged to read some recent work like [60] for 3D segmentation.
6 Further Reading
An interesting mathematical aspect of plant structures can be found in the book by Prusinkiewicz and Lindenmayer [61]. For an overview of recent plant phenotyping technologies, the readers are invited to read [1] and a more recent review [62]. Other reviews of imaging techniques can be found in [9, 63]. A detailed description of the setup, procedure and experiments of annotated datasets is available in [64]. The details of plant organ segmentation in energy minimization framework can be found in [42].
7 Exercises
-
1.
VascusynthFootnote 6 is a software for generating synthetic vascular (tree-like) structures. Generate some custom data using the software. Then perform CPD registration and report the average Root Mean Square (RMS) error from ground truth data. Source code of CPD is available both in MatlabFootnote 7 and Python.Footnote 8
-
2.
Add some Gaussian noise in the data above and perform CPD registration again. Increase the level of noise and report the threshold beyond which the registration algorithm fails (you can consider error up to say \(5\%\) as acceptable).
-
3.
Add some deformation (e.g. applying random rotation to some random parts) to the data and repeat the registration task.
-
4.
Extend the pairwise registration into a multi-view alignment problem. Align multiple views to obtain a single point cloud by exploiting the idea of pairwise registration in a sequential manner.
-
5.
Obtain the ground truth junction points in the point cloud data in Vascusynth. Assume that the matching of junction points are available. Now apply large amount of rotation to one of the views and perform pairwise registration. The results might not be good at this stage. In order to improve the registration result, we will test if initial rough alignment of the junction points help or not. Using the ground truth matching of the junction points, retrieve the transformation parameters (rotation, translation and scaling), and apply reverse transformation to the data. Now perform the registration. Report the effect of pre-alignment on the registration error.
-
6.
Select some random points in the 3D point cloud above. Extract small neighbourhoods (say \(50 \times 50\)) around these points. These neighbourhoods will also be 3D. Then apply the coordinate transformation as described in Sect. 14.3.3.4 to convert the data into 2D. Now compare the original 3D data of these neighbourhood structures and the transformed point cloud. If you are getting all the z-coordinate values of the transformed point cloud as almost the same, then the result is correct. Also, plot both 3D and 2D data and see if the transformation has preserved the original structure or not.
-
7.
Perform Principal Component Analysis (PCA) of the above 3D neighbourhood structures. Look at the eigenvalues and eigenvectors. Can you tell anything about the local structure from these quantities?
-
8.
Obtain the challenging vegetation dataset from ASL database.Footnote 9 Apply state-of-the-art point cloud feature matching algorithms and report their limitations on this type of data.
-
9.
Perform CPD registration of the same dataset as above.
-
10.
Triangulate the point cloud data using some standard algorithms like Delaunay triangulationFootnote 10 or alpha shape algorithm.Footnote 11 Adjust the parameters to get the best result. Do you think that these algorithms are efficient enough to triangulate the dataset?
-
11.
Obtain the dataset of Li et al. [42].Footnote 12 Implement the segmentation method as described in detail in the paper.
Notes
- 1.
- 2.
- 3.
- 4.
- 5.
- 6.
- 7.
- 8.
- 9.
- 10.
- 11.
- 12.
References
Spalding, E.P., Miller, N.D.: Image analysis is driving a renaissance in growth measurement. Current Opinion Plant Biol. 16(1), 100–104 (2013)
Hartmann, A., Czauderna, T., Hoffmann, R., Stein, N., Schreiber, F.: HTpheno: An image analysis pipeline for high-throughput plant phenotyping. BMC Bioinform. 12, Article number: 148 (2011)
Klukas, C., Chen, D., Pape, J.M.: Integrated analysis platform: an open-source information system for high-throughput plant phenotyping. Plant Physiol. 165(2), 506–518 (2014)
Scanalyzer-HTS, http://www.lemnatec.com/products/hardware-solutions/scanalyzer-hts/
Subramanian, R., Spalding, E., Ferrier, N.: A high throughput robot system for machine vision based plant phenotype studies. Mach. Vis. Appl. 24(3), 619–636 (2013)
Paulus, S., Schumann, H., Kuhlmann, H., Leon, J.: High-precision laser scanning system for capturing 3D plant architecture and analysing growth of cereal plants. Biosyst. Eng. 121 (2014)
Fiorani, F., Schurr, U.: Future scenarios for plant phenotyping. Ann. Rev. Plant Biol. 64 (2013)
Furbank, R.T., Tester, M.: Phenomics - technologies to relieve the phenotyping bottleneck. Trends Plant Sci. 16(12), 635–644 (2011)
Li, L., Zhang, Q., Huang, D.: A review of imaging techniques for plant phenotyping. Sensors 14(11) (2014)
Chaudhury, A., Ward, C., Talasaz, A., Ivanov, A.G., Brophy, M., Grodzinski, B., Hüner, N.P.A., Patel, R.V., Barron, J.L.: Machine vision system for 3D plant phenotyping. In: IEEE/ACM Transactions on Computational Biology and Bioinformatics (2018)
Jiménez, A.R., Ruíz, R.C., Rovira, J.L.P.: A vision system based on a laser rangefinder applied to robotic fruit harvesting. Mach. Vis. Appl. 11(6), 321–329 (2000)
Chattopadhyay, S., Akbar, S.A., Elfiky, N.M., Medeiros, H., Kak, A.C.: Measuring and modeling apple trees using time-of-flight data for automation of dormant pruning applications. In: Proceedings of IEEE Winter Conference on Applications of Computer Vision (WACV) (2016)
Paulus, S., Dupuis, J., Mahlein, A.K., Kuhlmann, H.: Surface feature based classification of plant organs from 3D laser scanned point clouds for plant phenotyping. BMC Bioinform. 14(238) (2013)
Paulus, S., Dupuisemail, J., Riedelemail, S., Kuhlmann, H.: Automated analysis of barley organs using 3D laser scanning: an approach for high throughput phenotyping. Sensors 14(7), 12670–12686 (2014)
Klodt, M., Herzog, K., Tïpfer, R., Cremers, D.: Field phenotyping of grapevine growth using dense stereo reconstruction. BMC Bioinform. 16, 143 (2015)
Wahabzada, M., Paulus, S., Kersting, K., Mahlein, A.-K.: Automated interpretation of 3d laserscanned point clouds for plant organ segmentation. BMC Bioinform. 16, 248 (2015)
Paproki, A., Sirault, X., Berry, S., Furbank, R., Fripp, J.: A novel mesh processing based technique for 3d plant analysis. BMC Plant Biol. 12(1) (2012)
Golbach, F., Kootstra, G., Damjanovic, S., Otten, G., Zedde, R.: Validation of plant part measurements using a 3D reconstruction method suitable for high-throughput seedling phenotyping. Mach. Vis. Appl. (2015)
Dellen, B., Scharr, H., Torras, C.: Growth signatures of rosette plants from timelapse video. IEEE/ACM Trans. Comput. Biol. Bioinform. 12(6), 1470–1478 (2015)
Scharr, H., Minervini, M., French, A.P., Klukas, C., Kramer, D.M., Liu, X., Luengo, I., Pape, J., Polder, G., Vukadinovic, D., Yin, X., Tsaftaris, S.A.: Leaf segmentation in plant phenotyping: a collation study. Mach. Vis. Appl. 27(4), 585–606 (2016)
Kelly, D., Vatsa, A., Mayham, W., Kazic, T.: Extracting complex lesion phenotypes in zea mays. Mach. Vis. Appl. 27(1), 145–156 (2016)
Xu, G., Zhang, F., Shah, S.G., Ye, Y., Mao, H.: Use of leaf color images to identify nitrogen and potassium deficient tomatoes. Pattern Recognit. Lett. 32(11), 1584–1590 (2011)
Aksoy, E.E., Abramov, A., Wörgötter, F., Scharr, H., Fischbach, A., Dellen, B.: Modeling leaf growth of rosette plants using infrared stereo image sequences. Comput. Electron. Agricult. 110, 78–90 (2015)
Pound, M.P., French, A.P., Fozard, J.A., Murchie, E.H., Pridmore, T.P.: A patch based approach to 3D plant shoot phenotyping. Mach. Vis. Appl. (2016)
Pound, M.P., French, A.P., Murchie, E.H., and Pridmore, T.P.: Surface reconstruction of plant shoots from multiple views. In: Proceedings of ECCV Workshops (2014)
Santos, T.T., Koenigkan, L.V., Barbedo, J.G.A., Rodrigues, G.C.: 3D plant modeling: localization, mapping and segmentation for plant phenotyping using a single hand-held camera. In: Proceedings of ECCV 2014 Workshops, Lecture Notes in Computer Science, vol. 8928
Santos, T.T., Rodrigues, G.C.: Flexible three-dimensional modeling of plants using low-resolution cameras and visual odometry. Mach. Vis. Appl. 27(5), 695–707 (2016)
Kumar, P., Cai, J., Miklavcic, S.: High-throughput 3D modelling of plants for phenotypic analysis. In: Proceedings of of 27th Conference on Image and Vision Computing New Zealand (IVCNZ) (2012)
Gibbs, J.A., Pound, M., French, A.P., Wells, D.M., Murchie, E., Pridmore, T.: Plant phenotyping: an active vision cell for three-dimensional plant shoot reconstruction. Plant Physiol. 178, 524–534 (2018)
Simek, K., Palanivelu, R., Barnard, K.: Branching gaussian processes with applications to spatiotemporal reconstruction of 3d trees. In: Proceedings of European Conference on Computer Vision (ECCV) (2016)
M. Brophy, A. Chaudhury, S. S. Beauchemin, and J. L. Barron, “A method for global non-rigid registration of multiple thin structures”, Proceedings of Conference on Computer and Robot Vision (CRV), 2015
Chaudhury, A., Brophy, M., Barron, J.L.: Junction-based correspondence estimation of plant point cloud data using subgraph matching. IEEE Geosci. Remote Sens. Lett. 13(8), 1119–1123 (2016)
Cox, M.C., Millenaar, F.F., van Berkel, Y.E.D.J., Peeters, A.J., Voesenek, L.A.: Plant movement. submergence-induced petiole elongation in Rumex palustris depends on hyponastic growth. Plant Physiol. 132, 282–291 (2003)
Navarro, P.J., Ferníndez, C., Weiss, J., Egea-Cortines, M.: Development of a configurable growth chamber with a computer vision system to study circadian rhythm in plants. Sensors 12(11), 15356 (2012)
Dornbusch, T., Lorrain, S., Kuznetsov, D., Fortier, A., Liechti, R., Xenarios, I., Fankhauser, C.: Measuring the diurnal pattern of leaf hyponasty and growth in Arabidopsis a novel phenotyping approach using laser scanning. Funct. Plant Biol. 39(11), 860–869 (2012)
Benoit, L., Rousseau, D., Belin, E., Demilly, D., Chapeau-Blondeau, F.: Simulation of image acquisition in machine vision dedicated to seedling elongation to validate image processing root segmentation algorithms. Comput. Electron. Agricult. 104 (2014)
Barron, J.L., Liptay, A.: Optic flow to measure minute increments in plant growth. BioImaging (BI1994) 2(1), 57–61 (1994)
Barron, J.L., Liptay, A.: Measuring 3d plant growth using optical flow. BioImaging (BI1997) 5(2), 82–86 (1997)
Liptay, A., Barron, J.L., Jewett, T., Wesenbeeck, I.V.: Oscillations in corn seedling growth as measured by optical flow. J. Am. Soc. Hort. Sci. 120(3) (1995)
Godin, C., Ferraro, P.: Quantifying the degree of self-nestedness of trees: application to the structural analysis of plants. IEEE/ACM Trans. Comput. Biol. Bioinform. 7(4), 688–703 (2010)
Augustin, M., Haxhimusa, Y., Busch, W., Kropatsch, W.G.: A framework for the extraction of quantitative traits from 2D images of mature Arabidopsis thaliana. Mach. Vis. Appl. (2015)
Li, Y., Fan, X., Mitra, N.J., Chamovitz, D., Cohen-Or, D., Chen, B.: Analyzing growing plants from 4d point cloud data. In: ACM Transactions on Graphics (Proceedings of SIGGRAPH Asia 2013), vol. 32 (2013)
Medeiros, F., Kim, D., Sun, J., Seshadri, H., Akbar, S.A., Elfiky, N.M., Park, J.: Modeling dormant fruit trees for agricultural automation. J. Field Robot. 34(7), 1203–1224 (2016)
Sa, I., Lehnert, C., English, A., McCool, C., Dayoub, F., Upcroft, B., Perez, T.: Peduncle detection of sweet pepper for autonomous crop harvesting-combined color and 3D information. IEEE Robot. Autom. Lett. 2(2) (2017)
Yuille, A.L., Grzywacz, N.M.: A computational theory for the perception of coherent visual motion. Nature 333(6168) (1988)
Jian, B., Vemuri, B.C.: Robust point set registration using gaussian mixture models. IEEE Trans. Pattern Anal. Mach. Intell. 33(8) (2011)
Myronenko, A., Song, X., Carreira-Perpinan, M.A.: Non-rigid point set registration: coherent point drift. In: Advances in Neural Information Processing Systems (NIPS) (2006)
Myronenko, A., Song, X.: Point set registration: coherent point drift. IEEE Trans. Pattern Anal. Mach. Intell. 32(12) (2010)
Toldo, R., Beinat, A., Crosilla, F.: Global registration of multiple point clouds embedding the generalized procrustes analysis into an ICP framework. In: Proceedings of 3DPVT 2010 (2010)
Rusu, R.B., Blodow, N., Beetz, M.: Fast point feature histograms (FPFH) for 3D registration (2009)
Bucksch, A., Khoshelham, K.: Localized registration of point clouds of botanic trees. IEEE Geosci. Remote Sens. Lett. 10(3) (2013)
Hartigan, J.A., Hartigan, P.M.: The dip test of unimodality. Ann. Stat. 13(1), 70–84 (1985)
Zhao, F., Bhotika, R.: Coronary artery tree tracking with robust junction detection in 3D CT angiography. In: Proceedings of the 8th IEEE International Symposium on Biomedical Imaging (ISBI), pp. 2066–2071 (2011)
Uhercik, M., Kybic, J., Liebgott, H., Cachard, C.: Model fitting using RANSAC for surgical tool localization in 3D ultrasound images. IEEE Trans. Biomed. Eng. 57(8), 1907–1916 (2010)
Kuhn, H.: The Hungarian method for the assignment problem. Naval Res. Logist. Quart. 2(1–2), 83–97 (1955)
Cour, T., Srinivasan, P., Shi, J.: Balanced graph matching. In: Proceedings of Advances in Neural Information Processing Systems (NIPS), pp. 313–320 (2007)
Boykov, Y., Kolmogorov, V.: An experimental comparison of min-cut/max-flow algorithms for energy minimization in vision. IEEE Trans. Pattern Anal. Mach. Intell. 26(9), 1124–1137 (2004)
Boykov, Y., Veksler, O., Zabih, R.: Fast approximate energy minimization via graph cuts. IEEE Trans. Pattern Anal. Mach. Intell. 23(11), 1222–1239 (2001)
Jin, Y., Su, Y., Wu, F., Pang, S., Gao, S., Hu, T., Liu, J., Guo, Q.: Stem-leaf segmentation and phenotypic trait extraction of individual maize using terrestrial LiDAR data. IEEE Trans. Geosci. Remote Sens. 57(3), 1336–1346 (2018)
Jin, S., Su, Y., Gao, S., Wu, F., Hu, T., Liu, J., Li, W., Wang, D., Chen, S., Jiang, Y., Pang, S., Guo, Q.: Deep learning: individual maize segmentation from terrestrial lidar data using faster R-CNN and regional growth algorithms. Frontiers Plant Sci. (2018)
Prusinkiewicz, P., Lindenmayer, A.: The Algorithmic Beauty of Plants. Springer, Berlin (1990)
Gibbs, J.A., Pound, M., French, A.P., Wells, D.M., Murchie, E., Pridmore, T.: Approaches to three-dimensional reconstruction of plant shoot topology and geometry. Funct. Plant Biol. 44(1), 62–75 (2017)
Dhondt, S., Wuyts, N., Inzé, D.: Cell to whole-plant phenotyping: the best is yet to come. Trends Plant Sci. 18(8), 428–439 (2013)
Minervini, M., Fischbach, A., Scharr, H., Tsaftaris, S.A.: Finely-grained annotated datasets for image-based plant phenotyping. Pattern Recognit. Lett. 81, 80–89 (2016)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Chaudhury, A., Barron, J.L. (2020). 3D Phenotyping of Plants. In: Liu, Y., Pears, N., Rosin, P.L., Huber, P. (eds) 3D Imaging, Analysis and Applications. Springer, Cham. https://doi.org/10.1007/978-3-030-44070-1_14
Download citation
DOI: https://doi.org/10.1007/978-3-030-44070-1_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-44069-5
Online ISBN: 978-3-030-44070-1
eBook Packages: Computer ScienceComputer Science (R0)