Abstract
Notch signaling exerts multiple important functions in various developmental processes, including cell differentiation and cell proliferation, while mis-regulation of this pathway results in a variety of complex diseases, such as cancer and developmental defects. The simplicity of the Notch pathway in Drosophila melanogaster, in combination with the availability of powerful genetics, makes this an attractive model for studying the fundamental mechanisms of how Notch signaling is regulated and how it functions in various cellular contexts. Recently, increasing evidence for epigenetic control of Notch signaling reveals the intimate link between epigenetic regulators and Notch signaling pathway. In this chapter, we summarize the research advances of Notch and CAF-1 in Drosophila development and the epigenetic regulation mechanisms of Notch signaling activity by CAF-1 as well as other epigenetic modification machineries, which enables Notch to orchestrate different biological inputs and outputs in specific cellular contexts.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Notch
- CAF-1
- Signal transduction
- Gene expression
- Epigenetic regulation
- Chromatin assembly factors
- Development
- Drosophila
Introduction
The Notch pathway is an evolutionarily conserved signaling cascade present in most of multicellular organisms and plays important roles in development and physiology . Notch signaling regulates a variety of biological processes, including cell proliferation, differentiation, and apoptosis (Radtke et al. 2005). Mis-regulation of Notch signaling activity has been associated with various complex diseases, such as cancer and neurological disorders (Salazar and Yamamoto 2018). Understanding the mechanisms of signaling regulation, as well as the outcome of signaling in various tissues, is therefore of great importance. The existence of multiple paralogues of Notch receptor (Notch 1–4) and ligands (Delta 1–4 and Jagged 1–2) in mammals and other vertebrates makes Notch-related studies more complicated in those animals. However, the situation is much simpler in Drosophila, which has only one Notch receptor and two ligands, Delta (Dl) and Serrate (Ser). All of these three proteins share highly conserved sequences with their mammalian counterparts (Muskavitch 1994). The simplicity of the Notch signaling pathway in Drosophila , in combination with the availability of well-established powerful genetic tools and materials (Zacharioudaki and Bray 2014), makes Drosophila an extremely attractive system for studying Notch pathway. Recent studies in both Drosophila and mammals provide insights into the epigenetic regulation of Notch signaling, and this chapter summarizes the current understanding of how Notch signaling is epigenetically regulated, mainly by CAF-1.
Notch Signaling
A century ago, Notch was first described as a wing margin developmental defect phenotype in Drosophila melanogaster (Bray 2016; Ntziachristos et al. 2014). Notch locus was identified as a gene that is responsible for the notched wing phenotype (Welshons 1958a, b), which gives the name to the pathway. Notch gene encodes a single-pass type I transmembrane receptor, the extracellular domain of which includes a variable number of EGF-like repeats, with the functions of ligand binding (Fig. 4.1).
Phenotype of Drosophila with Notch pathway mutations. (a) Drawing of a Notch receptor mutant fly with a notched wing tip. (b, c) Photo of a wing from a fly carrying a Notch mutation (b) and a mutation in Serrate (c). (Images are adapted from Alabi et al. (2018))
Mutants with defects in other genes that are part of the Notch pathway were later identified because they had similar phenotypes and were named Delta (Dl) and Serrate (Ser) (Siren and Portin 1989; Shepard et al. 1989; Thomas et al. 1991). Dl and Ser are also transmembrane proteins and share the similar EGF repeats with Notch. Thus, in order for Notch signaling to occur, the ligand-expressing cells (or signal sending cells) have to be in intimate contact with the receptor-expressing cells (or signal receiving cells) (Fig. 4.2).
The canonical Notch signaling pathway is rather simple. While vertebrates have several Notch receptors and ligands, the Drosophila genome only contains one Notch receptor and two ligands, Delta (Dl) and Serrate (Ser) (Klueg and Muskavitch 1999). Like many other signaling pathways, Notch signaling is initiated by receptor-ligand interaction between neighboring cells with close contact or direct contact. The Notch receptor can be activated by binding with the ligands Dl or Ser that are expressed in adjacent cells. Upon activation by this intimate binding, the Notch receptor undergoes two consecutive cleavage events, which are catalyzed sequentially by an ADAM family metalloprotease (Kuz and Tace in Drosophila) (Alabi et al. 2018) and by the γ-secretase complex (containing Presenilin/Psn, Nicastrin/Nct, PEN2, and APH1) (De Strooper et al. 1999; Yang et al. 2019), resulting in the release of the intracellular portion of the protein, called the Notch intracellular domain (NICD) , which then migrates into the nucleus and joins a protein complex directly bound to chromatin to initiate the transcription of target genes (Borggrefe and Oswald 2016). This complex includes the transcription factors Suppressor of Hairless (Su(H)), as well as other potential co-regulators, such as the transcriptional coactivator Mastermind (Mam), thereby leading to the expression of Notch-dependent target genes (Borggrefe and Liefke 2012).
As we mentioned above, Notch signaling is an evolutionally conserved pathway throughout metazoans. Over time, it became clear that it is repeatedly employed in cell fate decisions, cell differentiation, cell proliferation, and cell survival in diverse contexts and at distinct stages of development (Bray 2016). In many developmental contexts, Notch specifies cell fate decisions. In the developing vertebrate eye, for example, Notch regulates which cells develop into glial cells and which develop into optic neurons (Genethliou et al. 2009). Not surprisingly, mutations leading to dysregulated Notch signaling have also been implicated in cancer, including hematological malignancies (Bugeon et al. 2011) and solid tumors (Mutvei et al. 2015). Mis-regulation of Notch signaling in ovarian follicle cells disturbs the balance between cell proliferation and cell differentiation in Drosophila oogenesis, leading to cell death and sterility (Deng et al. 2001; Lopez-Schier and St Johnston 2001; Palmer et al. 2014). In other developmental contexts, Notch regulates the survival of cells (Giraldez and Cohen 2003). For example, loss of Notch function results in increased cell death of neuron cells in the mouse nervous system (Mason et al. 2006). Notch signaling has also been associated with cell survival in B-cell malignancies, prostate cancer cells, and myeloma cells (Zhang et al. 2018; Nefedova et al. 2008; Zweidler-McKay et al. 2005).
Notch signaling must be under extremely tight control to keep its proper activity. Emerging evidence indicates that the regulation of Notch signaling seems to be considerately complicated; multiple levels of regulation are added to the pathway via receptor-ligand internalization, posttranslational modification, protein stability, and ligand availability (Kovall et al. 2017). Productive Notch ligand-receptor binding depends on the proper posttranslational modification, such as glycosylation of the receptor (Haines and Irvine 2003). The retention time of Notch and ligands on plasm membrane is determined by the endocytosis of the receptor and ligands (Kandachar and Roegiers 2012), mediated mainly by lysosomal degradation. Mutants that stabilize NICD can cause T-cell acute lymphoblastic leukemia in humans (Grabher et al. 2006). Polarity proteins , such as Numb (Song and Lu 2012) and Crumbs (Nemetschke and Knust 2016), are also required for local distribution of Notch in the plasm membrane, which results in region-specific Notch activity. Like Notch, Dl and Ser are also subject to transmembrane domain cleavage by the γ-secretase complex, with this process called ligand processing, which may be used to downregulate the activity of Notch pathway. Alternatively, ligand processing also could generate biologically soluble ligands that may act as antagonists of Notch signaling (Masuya et al. 2002). Although the mechanisms of signal transduction from the cell surface to the nucleus are relatively simple and clear, it is not fully understood how such a straightforward pathway can result in tremendously complex outcomes in different cellular contexts. Recent studies have revealed that epigenetic modifiers play important roles in regulating Notch activity and may provide a novel angle to explain how and why the various developmental outputs occur in different contexts by a single Notch signaling pathway.
The Advantages of Drosophila as Model Organism for Notch Signaling Study
Drosophila melanogaster is an ideal model organism and has been extensively used in scientific research for over 100 years since Professor Thomas H. Morgan (1866–1945), who won the Nobel Prize in Physiology or Medicine in 1933, started to use the Drosophila for genetic studies. Owing to several practical advantages that are suitable for laboratory study, Drosophila has significantly pushed forward the development of biological research in various fields, such as developmental biology, immunobiology, and metabolism (Mirth et al. 2019). First, Drosophila has a short life cycle (about 10 days in the laboratory conditions) and has high fecundity, which allow producing large number of progenies in a short time. Besides, it is relatively easy and cost-effective to maintain the stable Drosophila stocks. Second, Drosophila has a low number of chromosomes, which make it as one of the most studied organisms in biological research, particularly in genetics and developmental biology (Perrimon 2014; Tolwinski 2017). Notably, the genome sequencing of Drosophila reveals that approximately 75% of known human disease-associated genes have counterparts in Drosophila (McGurk et al. 2015; Chen and Crowther 2012). The high similarity and conservation in genomic features between Drosophila and human enables fly to benefit the biomedical studies of human diseases. Third, a large number of genetic tools are available for Drosophila researchers mostly through stock centers, such as the Bloomington Drosophila Stock Center (BDSC) and the Vienna Drosophila Resource Center (VDRC) (Zacharioudaki and Bray 2014; Housden et al. 2014). Large-scale mutagenesis and screen projects are easy to carry out to discover the novel components or novel regulators of a classic pathway. In particular, for those genes that are humongously lethal or semilethal when mutated, somatic or germline clonal analysis based on FRT recombination system would be a good choice. Other FRT system-derived methods, such as MARCM (mosaic analysis with a repressible cell marker) (Lee and Luo 2001), are further developed to create mutant cells in a wild-type background tissue, facilitating reduction of the inter-organismal variability when analyzing mutant versus wild-type tissues.
Notch and its ligands are broadly expressed in many tissues/organs in the Drosophila. Therefore, it is of great importance to develop tools to directly examine where the pathway is activated or inhibited. The current arsenal of genetic and biological tools makes Drosophila such a valuable model to study the fundamental principles of how Notch signaling transduces the signal and how it is regulated in different cellular contexts, which can deepen the understanding of its roles in physiological and pathological conditions in humans. Table 4.1 shows the commonly used biochemical (antibodies) and genetic (transgenic flies) tools for studying Notch signaling in vitro and in vivo.
Further genetic methods include (1) conditional gene expression and silencing with the Gal4-UAS system, (2) genome-scale bioinformatics analysis, (3) genomic tagging and disruption of genes using CRISPR/Cas9 genome editing for gene activation and inactivation (Yu et al. 2013a, 2014), and (4) advanced imaging technology, such as light-sheet microscopy (Lu et al. 2019). All these methods and tools are likely to further facilitate the use of this sophisticated model to better understand the Notch signaling.
It is worth mentioning that Guangzhou Drosophila Stock Center (GDSC) , a newly established stock center for generating mutants through genome-wide gene targeting using CRISPR/Cas9 system, has generated more than 1000 mutant stocks. This resource would benefit a lot to those who use Drosophila as model for studying Notch signaling and other biological fields.
The Chromatin Assembly Factor CAF-1
Drosophila CAF-1 was first biochemically identified as a chromatin assembly factor about 30 years ago (Smith and Stillman 1989). Drosophila genome has three CAF-1-coding genes encoding three subunits, CAF-1 p180, CAF-1 p105, and CAF-1 p55, which correspond to human p150, p60, and p48, respectively (Ridgway and Almouzni 2000) (Table 4.2). Notably, there are two distinct CAF-1 complexes in Drosophila, each with three subunits of p180, p105, and p55 or p180, p75, and p55. Among them, p75 is a C-terminally truncated form of p105 in vivo. Though p105 and p75 have similar functions, p105 is predominantly expressed during embryogenesis, while p75 dominates after larval formation (Tyler et al. 2001).
CAF-1 has been biochemically well-characterized to be responsible for nucleosome assembly by guiding the histone trafficking and depositing them into chromatin by mediating H3 and H4 dimers onto newly synthesized DNA during DNA replication and DNA repair (Burgess and Zhang 2013). Reduction of CAF-1 activity in culture cells leads to reduced and delayed packaging of the DNA into chromatin, accompanied with DNA replication defects, S-phase arrest, checkpoint activation defects in cell cycle, and even cell death (Klapholz et al. 2009; Jiao et al. 2012; Krude 1995; Collins and Moon 2013). CAF-1 mutant mice exhibit developmental arrest at the embryonic stage with severe alterations in the nuclear organization of constitutive heterochromatin (Houlard et al. 2006). In Drosophila , knocking out any of the CAF-1 three subunits results in a similar lethal larval phenotype, and tissue-specific knockdown of CAF-1 p180 in the eye results in eye developmental defects, indicating the CAF-1 complex is also indispensable for the normal development in multicellular organism, including Drosophila (Jiao et al. 2012; Song et al. 2007; Huang et al. 2010; Yu et al. 2013b; Wen et al. 2012; Anderson et al. 2011).
However, genes encoding all three subunits CAF-1 are not essential in yeast (Kaufman et al. 1997; Monson et al. 1997; Enomoto and Berman 1998) and plants (Exner et al. 2006; Kirik et al. 2006; Endo et al. 2006; Schonrock et al. 2006), with the CAF-1 mutant in these species being viable, although mutants also exhibit some growth defects. The nonessential role of CAF-1 in unicellular eukaryotes (yeast) and plants appears to be inconsistent with the well-established role of CAF-1 in nucleosome assembly during DNA replication and DNA repair, an activity that might have been expected to be essential for all eukaryotic cells.
Interestingly, emerging evidence reported that CAF-1 p55, the small subunit of Drosophila CAF-1 , not only functions in the CAF-1 complex but also is a component in several chromatin-modulating complexes, such as PRC1 (Jones et al. 1998) and NuRD (Campbell et al. 2018), indicating that CAF-1 may have multiple functional roles that are not restricted to acting as a histone chaperone. Moreover, it is reasonable to propose that CAF-1 may serve as a protein platform for chromatin metabolism that integrates epigenetic regulation cues of gene transcription by interacting with chromatin modification machinery or transcriptional factors (Yu et al. 2015).
Epigenetic Regulation of Notch Signaling by CAF-1
In a search for which signaling pathway is regulated by CAF-1 in Drosophila development, Yu et al. found that tissue-specific knockdown of the Drosophila CAF-1 p105, the medium subunit of CAF-1 complex, results in a notched wing phenotype, resembling that of Notch loss-of-function mutations (Yu et al. 2013b). Moreover, the notched wing phenotype could be enhanced by combination with loss of function of Notch, revealing a synergistic genetic interaction between CAF-1 p105 Notch signaling. This study also establishes a functional connection between CAF-1 complex and Notch signaling for the first time.
Similar to wing development, eye development defects caused by eye-specific knockdown of CAF-1 p105 are also significantly enhanced in heterozygous mutant backgrounds of several Notch-positive regulatory components such as Mam (Yu et al. 2013b). Altogether, these results suggest that CAF-1 p105 synergistically interacts with the Notch signaling pathway to regulate normal tissue development.
To further confirm that CAF-1 p105 is required for the normal activity of Notch pathway, Yu et.al generated a null allele of dCAF-1 p105, namely, CAF-1 p10536, and performed clonal analyses in wing discs to investigate the effect of dCAF-1 p105 null mutation on Notch signaling, by examination of the developmental defects and the expression change of Notch target genes in the absence of CAF-1 p105. As expected, flies carrying CAF-1 p10536 clones exhibited a notched wing phenotype, and the protein expression level of cut and wg, two well-characterized Notch target genes, was also significantly decreased in CAF-1 p10536 clones, compared with control (Fig. 4.3). The transcriptional level of cut is also significantly downregulated when CAF-1 p105 is depleted. These results indicate that the activity of Notch signaling is compromised in the absence of CAF-1 p105 and thus CAF-1 p105 functions as a positive regulator of the Notch signaling pathway by promoting its target gene transcription.
CAF-1-p105 is required for the normal wing development and proper expression of Notch target genes cut and wg. (a, b) Induction of CAF-1 p10536 mutant clones leads to a notched wing (b), whereas induction of mock clones leads to wild-type wings (a). (c–f′) In CAF-1 p10536 mutant clones, the expression of Cut (e, e′, GFP-negative area, arrowheads) and Wg (f, f′, FP-negative area, arrowheads) is abolished in a cell-autonomous manner, whereas in the mock clones the expression of both Cut (c, c′) and Wg (d, d′) is unaffected. (Images are adapted from Yu et al. (2013b))
As CAF-1 functions as a histone chaperone, it is likely that its reduction may trigger dilution of newly assembled nucleosomes at key enhancer elements and loosening of chromatin structure, resulting in a more accessible chromatin structure for efficient transcription factor binding to their target loci and activation of key target genes. However, the CAF-1 p105 specifically regulates the output of Notch signaling in the wing disc, since the Hedgehog (Hh) signaling is not affected in CAF-1 p105 mutant clone.
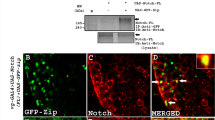
Further, it is found that CAF-1 forms a functional complex with NICD and Su(H), the core transcriptional factor for Notch target gene expression. And this complex directly binds to the enhancer region of one of the Notch target genes, E(spl)mβ. The occupancy of Su(H) at Notch target genes is highly increased to efficiently initiate gene transcription when Notch signaling is activated. In the absence of CAF-1 p105, the abundance of Su(H) at the E(spl)mβ enhancer region is dramatically decreased, proposing that CAF-1 probably regulates Notch target gene expression, at least in part, by controlling the accessibility and binding abundance of Su(H) to their enhancer regions.
It is reported that CAF-1 could act as a chromatin platform that is permissive for transcription by regulating histone modifications by forming complex with other chromatin remodeling complexes (Yu et al. 2015), and histone H4 acetylation is believed to be associated with active promoters of Notch target genes (Giaimo et al. 2018). As expected, H4ac level in the E(spl)mβ enhancer region is significantly reduced in CAF-1 p105 mutant flies. These results reveal that CAF-1 p105 promotes Notch target gene expression by maintaining a high level of histone H4 acetylation in the enhancer region of the Notch target genes to establish a local active chromatin structure. Interestingly, CAF-1 function in regulating Notch signaling is dependent on its integrity as a triple subunit complex. Knockdown of any component of CAF-1 complex causes the reduction of cut expression and the notched wing phenotype in Drosophila (Yu et al. 2013b). However, there are still open questions for how CAF-1 directs the H4 acetylation modification. One possibility is that CAF-1 recruits histone acetylation machinery, such as p300/Nejire, directly or indirectly, to change the landscape of local chromatin modification, thus enhancing target gene transcription. CAF-1 has functions beyond its classic role in histone assembly and the newly established positive role in Notch signaling in wing development and plays an essential role in proliferating cells.
However, in contrast to the positive role of CAF-1 on Notch target gene expression in wing development, a recent study reported that CAF-1 play a negative role in regulating Notch signaling in Drosophila ovarian mitotic follicle cells (Lo et al. 2019). Loss of function of either CAF-1 p105 or CAF-1 p180 caused the increased activation of Notch signaling target genes in Drosophila ovarian follicle cells. Further, Notch is functionally responsible for these phenotypes observed in both the CAF-1 p105- and CAF-1 p180-deficient follicle cells. It is still unclear how CAF-1 p180 and CAF-1 p150 suppress Notch target gene expression in mitotic follicle cells. It is likely that CAF-1 have physical interaction with Su(H), which is known to be involved in maintaining the repressive chromatin status for inhibiting Notch target gene expression when it is associated with other repressive subunits, such as Hairless, Gro, or CtBP (Yu et al. 2015; Cheloufi and Hochedlinger 2017; Yuan et al. 2016). Thus, the molecular basis for that CAF-1 play a dual role to sustain cell proliferation positively (in imaginal discs) or negatively (in ovarian follicle cells) may lie in that CAF-1 recruits different histone modification machineries in imaginal discs and follicle cells to regulate Drosophila Notch signaling in a tissue context-dependent manner.
Two recent studies in mammals confirmed the negative role of CAF-1 in retrotransposon jumping and gene expression . Hatanaka et al. reported that CAF-1 mediates repressive histone modifications to protect preimplantation mouse embryos from endogenous retrotransposons (Hatanaka et al. 2015). Multiple classes of retrotransposons are derepressed in morula embryos when CAF-1 is depleted, likely through affecting the histone methylation status, thus influencing local chromatin accessibility. The other study found that the p150 and p60, two subunits of mammalian CAF-1 complex, are the most prominent chromatin-modulating factors during transcription factor-mediated reprogramming of mouse fibroblasts to induced pluripotent stem cells (iPS cells) (Cheloufi et al. 2015). Suppression of CAF-1 leads to a more accessible chromatin structure at enhancer elements and the increased binding of Sox2 to pluripotency-specific targets and activation of associated genes during reprogramming (Cheloufi et al. 2015).
Altogether, CAF-1 functions not only as histone chaperone for nucleosome assembly but also as an epigenetic regulation switch for regulating Notch signaling target gene expression in response to integrated proliferation and differentiation signals during animal development. However, CAF-1 does not harbor the histone modification enzyme activity; thus it is likely that CAF-1 works together with the histone modification machinery (histone methylation, histone acetylation, etc.) to regulate Notch signaling activity at the chromatin level through modifying the chromatin structure.
Epigenetic Regulation of Notch Signaling by Other Epigenetic Regulators
In addition to the epigenetic regulation of Notch signaling by the CAF-1 complex, this paragraph briefly summarizes the epigenetic regulation of Notch signaling by other epigenetic regulators. Several epigenetic regulators that are involved in Notch signaling are listed in Table 4.3. Among them, histone acetylation and methylation are main executors for epigenetic regulation of gene transcription (Tchasovnikarova and Kingston 2018).
Histone deacetylases are generally associated with transcriptional repressor complexes, such as Sin3 (Barnes et al. 2018), NuRD (Feng and Zhang 2003; Ahringer 2000), and CoREST (You et al. 2001; Domanitskaya and Schupbach 2012) complexes, and have regulatory functions in various signaling pathways. It is generally accepted that HDAC1 forms a transcriptional corepressor complex to modify chromatin structure for target gene silencing. For example, HDAC1 physically interacts with CBF1 (homolog of Su(H) in Drosophila), and treatment of HDAC1 inhibitor derepresses Notch target gene ESR-1 expression in mammalian cells (Kao et al. 1998). In zebrafish, her4 and her6, two of Notch target genes, are upregulated in HDAC1 mutant fish (Cunliffe 2004; Yamaguchi et al. 2005). Furthermore, overexpression of HDAC1 represses the expression of Notch target gene Hey2 in mice (Wu et al. 2016). In these contexts, HDAC1 negatively regulates Notch signaling. Unexpectedly, opposite to the inhibitory role of HDAC1 in Notch signaling, knockdown of HDAC1 causes a notched wing phenotype and reduces Notch target gene expression in Drosophila (Wang et al. 2018), suggesting a positive role of HDAC1 in regulating the Notch pathway during Drosophila wing development, although the molecular mechanism behind this remains largely unknown. It is highly possible that HDAC1 directly regulates histone deacetylation status at the Notch target gene locus. Notably, a recent study reported that HDAC1 could activate the Notch signaling pathway to promote metastasis in a similar way (Mao et al. 2017).
The complicated regulation network by other epigenetic regulators, such as LSD1 (Mulligan et al. 2011; Lopez et al. 2016), Brms1 (Zhang et al. 2014), histone acetylase p300 (Franz Oswald et al. 2001), Brahma SWI/SNF chromatin remodeling complex (Pillidge and Bray 2019), UTX (Herz et al. 2010), H3K4 demethylase Kdm5A (Liefke et al. 2010; Dreval et al. 2019), methylcytosine dioxygenases Tet2/3 (Li et al. 2015), SAGA and Tip60 complex (Gause et al. 2006; Medgett and Langer 1984), PcG-TrxG complex (Saj et al. 2010; Tolhuis et al. 2006), Putzig-NURF complex (Kugler and Nagel 2010), and many others, may directly influence Notch-mediated gene transcription activity at the chromatin level and thus explain, at least in part, the pleiotropic effects of Notch in the complex biological processes that affect cell growth, differentiation, and cell death.
Conclusion
The Notch signaling pathway is a highly conserved molecular network that, depending on the cellular context, acts through the regulation of cell proliferation, differentiation, and apoptosis. In order to better control the expression of Notch target gene expression, the Notch signaling must be precisely regulated at different steps in a series of developmental events. Epigenetic regulation of Notch signaling by CAF-1 and other epigenetic regulators plays essential roles in fine-tuning the transcriptional output of Notch signaling to coordinate multicellular organism development. It remains an open question as to why and how different epigenetic regulators are involved in mediating different histone modifications status, leading to different transcriptional outputs of either gene repression or gene activation in one specific signal transduction pathway.
References
Ahringer J (2000) NuRD and SIN3 histone deacetylase complexes in development. Trends Genet 16(8):351–356
Alabi RO, Farber G, Blobel CP (2018) Intriguing roles for endothelial ADAM10/Notch signaling in the development of organ-specific vascular beds. Physiol Rev 98(4):2025–2061
Anderson AE et al (2011) The enhancer of trithorax and polycomb gene Caf1/p55 is essential for cell survival and patterning in Drosophila development. Development 138(10):1957–1966
Barnes VL et al (2018) Systematic analysis of SIN3 histone modifying complex components during development. Sci Rep 8(1):17048
Borggrefe T, Liefke R (2012) Fine-tuning of the intracellular canonical Notch signaling pathway. Cell Cycle 11(2):264–276
Borggrefe T, Oswald F (2016) Setting the stage for Notch: the Drosophila Su(H)-hairless repressor complex. PLoS Biol 14(7):e1002524
Bray SJ (2016) Notch signalling in context. Nat Rev Mol Cell Biol 17(11):722–735
Bugeon L et al (2011) The NOTCH pathway contributes to cell fate decision in myelopoiesis. Haematologica 96(12):1753–1760
Burgess RJ, Zhang Z (2013) Histone chaperones in nucleosome assembly and human disease. Nat Struct Mol Biol 20(1):14–22
Campbell AE et al (2018) NuRD and CAF-1-mediated silencing of the D4Z4 array is modulated by DUX4-induced MBD3L proteins. Elife 7:pii: e31023
Cheloufi S, Hochedlinger K (2017) Emerging roles of the histone chaperone CAF-1 in cellular plasticity. Curr Opin Genet Dev 46:83–94
Cheloufi S et al (2015) The histone chaperone CAF-1 safeguards somatic cell identity. Nature 528(7581):218–224
Chen KF, Crowther DC (2012) Functional genomics in Drosophila models of human disease. Brief Funct Genomics 11(5):405–415
Collins H, Moon NS (2013) The components of Drosophila histone chaperone dCAF-1 are required for the cell death phenotype associated with rbf1 mutation. G3 (Bethesda) 3(10):1639–1647
Cunliffe VT (2004) Histone deacetylase 1 is required to repress Notch target gene expression during zebrafish neurogenesis and to maintain the production of motoneurones in response to hedgehog signalling. Development 131(12):2983–2995
De Strooper B et al (1999) A presenilin-1-dependent gamma-secretase-like protease mediates release of Notch intracellular domain. Nature 398(6727):518–522
Deng WM, Althauser C, Ruohola-Baker H (2001) Notch-Delta signaling induces a transition from mitotic cell cycle to endocycle in Drosophila follicle cells. Development 128(23):4737–4746
Domanitskaya E, Schupbach T (2012) CoREST acts as a positive regulator of Notch signaling in the follicle cells of Drosophila melanogaster. J Cell Sci 125(Pt 2):399–410
Dreval K, Lake RJ, Fan HY (2019) HDAC1 negatively regulates selective mitotic chromatin binding of the Notch effector RBPJ in a KDM5A-dependent manner. Nucleic Acids Res 47(9):4521–4538
Endo M et al (2006) Increased frequency of homologous recombination and T-DNA integration in Arabidopsis CAF-1 mutants. EMBO J 25(23):5579–5590
Enomoto S, Berman J (1998) Chromatin assembly factor I contributes to the maintenance, but not the re-establishment, of silencing at the yeast silent mating loci. Genes Dev 12(2):219–232
Exner V et al (2006) Chromatin assembly factor CAF-1 is required for cellular differentiation during plant development. Development 133(21):4163–4172
Feng Q, Zhang Y (2003) The NuRD complex: linking histone modification to nucleosome remodeling. Curr Top Microbiol Immunol 274:269–290
Franz Oswald BT, Dobner T, Bourteele S, Kostezka U, Adler G, Liptay S, Schmid RM (2001) p300 acts as a transcriptional coactivator for mammalian Notch-1. Mol Cell Biol 21(22):7761–7774
Gause M et al (2006) Nipped-A, the Tra1/TRRAP subunit of the Drosophila SAGA and Tip60 complexes, has multiple roles in Notch signaling during wing development. Mol Cell Biol 26(6):2347–2359
Genethliou N et al (2009) SOX1 links the function of neural patterning and Notch signalling in the ventral spinal cord during the neuron-glial fate switch. Biochem Biophys Res Commun 390(4):1114–1120
Giaimo BD et al (2018) Histone variant H2A.Z deposition and acetylation directs the canonical Notch signaling response. Nucleic Acids Res 46(16):8197–8215
Giraldez AJ, Cohen SM (2003) Wingless and Notch signaling provide cell survival cues and control cell proliferation during wing development. Development 130(26):6533–6543
Grabher C, von Boehmer H, Look AT (2006) Notch 1 activation in the molecular pathogenesis of T-cell acute lymphoblastic leukaemia. Nat Rev Cancer 6(5):347–359
Haines N, Irvine KD (2003) Glycosylation regulates Notch signalling. Nat Rev Mol Cell Biol 4(10):786–797
Hatanaka Y et al (2015) Histone chaperone CAF-1 mediates repressive histone modifications to protect preimplantation mouse embryos from endogenous retrotransposons. Proc Natl Acad Sci U S A 112(47):14641–14646
Herz HM et al (2010) The H3K27me3 demethylase dUTX is a suppressor of Notch- and Rb-dependent tumors in Drosophila. Mol Cell Biol 30(10):2485–2497
Houlard M et al (2006) CAF-1 is essential for heterochromatin organization in pluripotent embryonic cells. PLoS Genet 2(11):e181
Housden BE, Li J, Bray SJ (2014) Visualizing Notch signaling in vivo in Drosophila tissues. Methods Mol Biol 1187:101–113
Huang H et al (2010) Drosophila CAF-1 regulates HP1-mediated epigenetic silencing and pericentric heterochromatin stability. J Cell Sci 123(Pt 16):2853–2861
Jiao R-J, Wu Q-H, Liu J-Y, Chen Y-X (2012) dCAF-1-p55 is essential for Drosophila development and involved in the maintenance of chromosomal stability. Prog Biochem Biophys 39(11):1073–1081
Jones CA et al (1998) The Drosophila esc and E(z) proteins are direct partners in polycomb group-mediated repression. Mol Cell Biol 18(5):2825–2834
Kandachar V, Roegiers F (2012) Endocytosis and control of Notch signaling. Curr Opin Cell Biol 24(4):534–540
Kao HY et al (1998) A histone deacetylase corepressor complex regulates the Notch signal transduction pathway. Genes Dev 12(15):2269–2277
Kaufman PD, Kobayashi R, Stillman B (1997) Ultraviolet radiation sensitivity and reduction of telomeric silencing in Saccharomyces cerevisiae cells lacking chromatin assembly factor-I. Genes Dev 11(3):345–357
Kirik A et al (2006) The chromatin assembly factor subunit FASCIATA1 is involved in homologous recombination in plants. Plant Cell 18(10):2431–2442
Klapholz B et al (2009) CAF-1 is required for efficient replication of euchromatic DNA in Drosophila larval endocycling cells. Chromosoma 118(2):235–248
Klueg KM, Muskavitch MA (1999) Ligand-receptor interactions and trans-endocytosis of Delta, Serrate and Notch: members of the Notch signalling pathway in Drosophila. J Cell Sci 112(Pt 19):3289–3297
Kovall RA et al (2017) The canonical Notch signaling pathway: structural and biochemical insights into shape, sugar, and force. Dev Cell 41(3):228–241
Krude T (1995) Chromatin assembly factor 1 (CAF-1) colocalizes with replication foci in HeLa cell nuclei. Exp Cell Res 220(2):304–311
Kugler SJ, Nagel AC (2010) A novel Pzg-NURF complex regulates Notch target gene activity. Mol Biol Cell 21(19):3443–3448
Lee T, Luo L (2001) Mosaic analysis with a repressible cell marker (MARCM) for Drosophila neural development. Trends Neurosci 24(5):251–254
Li C et al (2015) Overlapping requirements for Tet2 and Tet3 in Normal development and hematopoietic stem cell emergence. Cell Rep 12(7):1133–1143
Liefke R et al (2010) Histone demethylase KDM5A is an integral part of the core Notch-RBP-J repressor complex. Genes Dev 24(6):590–601
Lo PK et al (2019) Inhibition of Notch signaling by the p105 and p180 subunits of Drosophila chromatin assembly factor 1 is required for follicle cell proliferation. J Cell Sci 132(2):jcs224170
Lopez CI et al (2016) The chromatin modifying complex CoREST/LSD1 negatively regulates notch pathway during cerebral cortex development. Dev Neurobiol 76(12):1360–1373
Lopez-Schier H, St Johnston D (2001) Delta signaling from the germ line controls the proliferation and differentiation of the somatic follicle cells during Drosophila oogenesis. Genes Dev 15(11):1393–1405
Lu CH et al (2019) Lightsheet localization microscopy enables fast, large-scale, and three-dimensional super-resolution imaging. Commun Biol 2:177
Mao G, Jin H, Wu L (2017) DDX23-Linc00630-HDAC1 axis activates the Notch pathway to promote metastasis. Oncotarget 8(24):38937–38949
Mason HA et al (2006) Loss of notch activity in the developing central nervous system leads to increased cell death. Dev Neurosci 28(1–2):49–57
Masuya M et al (2002) The soluble Notch ligand, Jagged-1, inhibits proliferation of CD34+ macrophage progenitors. Int J Hematol 75(3):269–276
McGurk L, Berson A, Bonini NM (2015) Drosophila as an in vivo model for human neurodegenerative disease. Genetics 201(2):377–402
Medgett IC, Langer SZ (1984) Heterogeneity of smooth muscle alpha adrenoceptors in rat tail artery in vitro. J Pharmacol Exp Ther 229(3):823–830
Mirth CK, Nogueira Alves A, Piper MD (2019) Turning food into eggs: insights from nutritional biology and developmental physiology of Drosophila. Curr Opin Insect Sci 31:49–57
Monson EK, de Bruin D, Zakian VA (1997) The yeast Cac1 protein is required for the stable inheritance of transcriptionally repressed chromatin at telomeres. Proc Natl Acad Sci U S A 94(24):13081–13086
Mulligan P et al (2011) A SIRT1-LSD1 corepressor complex regulates Notch target gene expression and development. Mol Cell 42(5):689–699
Muskavitch MA (1994) Delta-Notch signaling and Drosophila cell fate choice. Dev Biol 166(2):415–430
Mutvei AP, Fredlund E, Lendahl U (2015) Frequency and distribution of Notch mutations in tumor cell lines. BMC Cancer 15:311
Nefedova Y et al (2008) Inhibition of Notch signaling induces apoptosis of myeloma cells and enhances sensitivity to chemotherapy. Blood 111(4):2220–2229
Nemetschke L, Knust E (2016) Drosophila crumbs prevents ectopic Notch activation in developing wings by inhibiting ligand-independent endocytosis. Development 143(23):4543–4553
Ntziachristos P et al (2014) From fly wings to targeted cancer therapies: a centennial for Notch signaling. Cancer Cell 25(3):318–334
Palmer WH, Jia D, Deng WM (2014) Cis-interactions between Notch and its ligands block ligand-independent Notch activity. Elife 3. https://doi.org/10.7554/eLife.04415
Perrimon N (2014) Drosophila developmental biology methods. Methods 68(1):1
Pillidge Z, Bray SJ (2019) SWI/SNF chromatin remodeling controls Notch-responsive enhancer accessibility. EMBO Rep 20(5):pii: e46944
Radtke F, Wilson A, MacDonald HR (2005) Notch signaling in hematopoiesis and lymphopoiesis: lessons from Drosophila. BioEssays 27(11):1117–1128
Ridgway P, Almouzni G (2000) CAF-1 and the inheritance of chromatin states: at the crossroads of DNA replication and repair. J Cell Sci 113(Pt 15):2647–2658
Saj A et al (2010) A combined ex vivo and in vivo RNAi screen for notch regulators in Drosophila reveals an extensive notch interaction network. Dev Cell 18(5):862–876
Salazar JL, Yamamoto S (2018) Integration of Drosophila and human genetics to understand notch signaling related diseases. Adv Exp Med Biol 1066:141–185
Schonrock N et al (2006) Functional genomic analysis of CAF-1 mutants in Arabidopsis thaliana. J Biol Chem 281(14):9560–9568
Shepard SB, Broverman SA, Muskavitch MA (1989) A tripartite interaction among alleles of Notch, Delta, and Enhancer of split during imaginal development of Drosophila melanogaster. Genetics 122(2):429–438
Siren M, Portin P (1989) Interaction of hairless, delta, enhancer of split and notch genes of Drosophila melanogaster as expressed in adult morphology. Genet Res 54(1):23–26
Smith S, Stillman B (1989) Purification and characterization of CAF-I, a human cell factor required for chromatin assembly during DNA replication in vitro. Cell 58(1):15–25
Song Y, Lu B (2012) Interaction of Notch signaling modulator Numb with alpha-Adaptin regulates endocytosis of Notch pathway components and cell fate determination of neural stem cells. J Biol Chem 287(21):17716–17728
Song Y et al (2007) CAF-1 is essential for Drosophila development and involved in the maintenance of epigenetic memory. Dev Biol 311(1):213–222
Tchasovnikarova IA, Kingston RE (2018) Beyond the histone code: a physical map of chromatin states. Mol Cell 69(1):5–7
Thomas U, Speicher SA, Knust E (1991) The Drosophila gene Serrate encodes an EGF-like transmembrane protein with a complex expression pattern in embryos and wing discs. Development 111(3):749–761
Tolhuis B et al (2006) Genome-wide profiling of PRC1 and PRC2 Polycomb chromatin binding in Drosophila melanogaster. Nat Genet 38(6):694–699
Tolwinski NS (2017) Introduction: Drosophila-A model system for developmental biology. J Dev Biol 5(3):9
Tyler JK et al (2001) Interaction between the Drosophila CAF-1 and ASF1 chromatin assembly factors. Mol Cell Biol 21(19):6574–6584
Wang Z et al (2018) The histone deacetylase HDAC1 positively regulates Notch signaling during Drosophila wing development. Biol Open 7(2):bio029637
Welshons WJ (1958a) A preliminary investigation of pseudoallelism at the notch locus of Drosophila melanogaster. Proc Natl Acad Sci U S A 44(3):254–258
Welshons WJ (1958b) The analysis of a pseudoallelic recessive lethal system at the notch locus of Drosophila melanogaster. Cold Spring Harb Symp Quant Biol 23:171–176
Wen P, Quan Z, Xi R (2012) The biological function of the WD40 repeat-containing protein p55/Caf1 in Drosophila. Dev Dyn 241(3):455–464
Wu LM et al (2016) Zeb2 recruits HDAC-NuRD to inhibit Notch and controls Schwann cell differentiation and remyelination. Nat Neurosci 19(8):1060–1072
Yamaguchi M et al (2005) Histone deacetylase 1 regulates retinal neurogenesis in zebrafish by suppressing Wnt and Notch signaling pathways. Development 132(13):3027–3043
Yang G et al (2019) Structural basis of Notch recognition by human gamma-secretase. Nature 565(7738):192–197
You A et al (2001) CoREST is an integral component of the CoREST- human histone deacetylase complex. Proc Natl Acad Sci U S A 98(4):1454–1458
Yu Z et al (2013a) Highly efficient genome modifications mediated by CRISPR/Cas9 in Drosophila. Genetics 195(1):289–291
Yu Z et al (2013b) CAF-1 promotes Notch signaling through epigenetic control of target gene expression during Drosophila development. Development 140(17):3635–3644
Yu Z et al (2014) Various applications of TALEN- and CRISPR/Cas9-mediated homologous recombination to modify the Drosophila genome. Biol Open 3(4):271–280
Yu Z et al (2015) Histone chaperone CAF-1: essential roles in multi-cellular organism development. Cell Mol Life Sci 72(2):327–337
Yuan Z et al (2016) Structure and function of the Su(H)-hairless repressor complex, the major antagonist of Notch signaling in Drosophila melanogaster. PLoS Biol 14(7):e1002509
Zacharioudaki E, Bray SJ (2014) Tools and methods for studying Notch signaling in Drosophila melanogaster. Methods 68(1):173–182
Zhang Q et al (2014) dBrms1 acts as a positive regulator of Notch signaling in Drosophila wing. J Genet Genomics 41(6):317–325
Zhang R, Engler A, Taylor V (2018) Notch: an interactive player in neurogenesis and disease. Cell Tissue Res 371(1):73–89
Zweidler-McKay PA et al (2005) Notch signaling is a potent inducer of growth arrest and apoptosis in a wide range of B-cell malignancies. Blood 106(12):3898–3906
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Wei, C., Phang, CW., Jiao, R. (2020). Epigenetic Regulation of Notch Signaling During Drosophila Development. In: Reichrath, J., Reichrath, S. (eds) Notch Signaling in Embryology and Cancer. Advances in Experimental Medicine and Biology, vol 1218. Springer, Cham. https://doi.org/10.1007/978-3-030-34436-8_4
Download citation
DOI: https://doi.org/10.1007/978-3-030-34436-8_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-34435-1
Online ISBN: 978-3-030-34436-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)