Abstract
Extreme acidophilic bacteria are a phylogenetically diverse group of microorganisms that grow optimally at pH values below 3. They thrive in natural or man-made environments where life is challenged by extreme acidity, low availability of organic matter, and high concentrations of heavy metals. Most acidophilic bacteria are chemolitho(auto)trophs, obtaining energy from the oxidation of metal sulfides, one of the most abundant mineral classes on earth. Bacterial attachment on mineral surface and the subsequent biofilm development plays critical role in mineral dissolution, which is directly related with ecologic phenomena and biotechnological applications, such as acid mine drainage, biogeochemical cycles, and bioleaching processes. In contrast to well-studied neutrophilic bacterial strains, the understanding of cyclic di-GMP signaling in extreme acidophilic bacteria is still incipient. However, significant progress has been made in the last several years through global genomic analysis on acidophilic communities, and genetic work on species belonging to the most iconic acidophilic genus, Acidithiobacillus. This chapter presents an overview of molecular insights into cyclic di-GMP signaling obtained from At. ferrooxidans, At. caldus, and At. thioooxidans. In addition, it describes the cyclic di-GMP signaling network as a widespread but highly diverse mechanism used by acidophilic bacteria to transduce environmental signals into biofilm-related responses mainly driven by cyclic di-GMP effector proteins involved in swarming motility and the production of exopolymeric substances.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Acidophilic Microorganisms
Acidophilic microorganisms are defined as those that have pH optima below 7. They have been classified as acid-tolerant (7 ≥ pH ≥ 5), moderate acidophiles (5 > pH > 3), and extreme acidophiles (pH ≤ 3) [1]. Acidophilic microorganisms exist in natural acidic environments such as acid sulfate soils, volcanic and geothermal areas where sulfur gases occur in association with water vapor (solfataras), hydrogen sulfide caves, as well as man-made environments such as biomining operations and bioreactors for wastewaters treatment. These environments are characterized by low pH, high metal, and low nutritional conditions, which result in a relatively low phylogenetic diversity of microorganisms [2], dominated by chemolithoautotrophic bacteria and archaea capable of oxidizing inorganic electron donors, generally ferrous iron and reduced inorganic sulfur compounds (RISCs), coupled to the reduction of oxygen or ferric iron. Chemolithoautotrophic microorganisms provide organic materials to some acidophilic mixotrophs and heterotrophs, such as bacteria belonging to Alicyclobacillus and Acidiphilium genus, respectively, that in turn, consume these compounds potentially toxic for chemolithoautotrophs maintaining the community structure [3]. Along with pH, a key physical–chemical parameter for the occurrence of particular microorganism is temperature, ranging from near to zero in acid mine drainages to near to 90 °C at solfataric springs. While most of the extreme thermophilic acidophiles correspond to archaea, mesophilic and moderate thermophilic bacteria dominate acidic environments below 50 °C.
Molecular mechanisms that allow the recognition of environmental cues and the coordination of suitable responses are important to survive in these challenging environments. Recently, through bioinformatics analysis, we have described components of different nucleotide second messenger-based signaling in extreme acidophilic bacteria of the Acidithiobacillus genus [4]. An extended analysis, comprising 201 prokaryotic genomes [106 Bacteria (38 completes, 36 drafts >91% completeness and 16S sequence complete predicted), 95 Archaea (44 completes, 35 draft)] from extreme, moderate and acid-tolerant acidophiles, gave us a general scenery of nucleotide transduction pathways in acidophilic microorganism (Fig. 21.1).
Distribution of proteins related with nucleotide second messenger systems in acidophilic bacteria. (a) Phylogenetic tree of acidophilic bacterial genus based on 16S rRNA. The number of sequenced genomes from different strains belonging to the same species is indicated in brackets. (b) Number of second messenger-related proteins in each acidophilic species. Proteins or domains involves in synthesis, degradation, and transduction of signaling nucleotides are written in green, red, and purple letters, respectively. SYNTH, RelA_SpoT (synthesis domain); HYD, HD (Hydrolase Domain); NC, Nucleotidyl cyclase. (c) Proportion of partner domains associated with the identified cyclic di-GMP metabolizing domains (synthesis: GGDEF; degradation: EAL and HD-GYP)
2 General Overview of Nucleotide Second Messenger Metabolism in Acidophilic Microorganisms
Among the most important signaling nucleotides are guanosine 3′,5′-bispyrophosphate (ppGpp) and guanosine 3′-diphosphate, 5′-triphosphate (pppGpp), known as (p)ppGpp [5], cyclic adenosine 3′,5′-monophosphate (cAMP) [6], cyclic dimeric guanosine 3′,5′-monophosphate (cyclic di-GMP) [7], and cyclic dimeric adenosine 3′,5′-monophosphate (cyclic di-AMP) [8]. (p)ppGpp, cAMP, cyclic di-GMP, and cAMP are synthetized, respectively, by (p)ppGpp synthetases, adenylyl cyclases (ACs), diguanylate cyclases (DGCs), and diadenylate cyclases (DACs). Based on amino acid sequences, nucleotidyl cyclases (NCs) have been classified into six classes (I–VI) [9]. Class III is the most diverse group of cyclases, including ACs and DGCs, while NCs belonging to classes I, II, IV, V, and VI are all AC enzymes.
The repertoire of enzymes related with the metabolism of nucleotide second messengers is very different between acidophilic Bacteria and Archaea. Just a few enzymes for the synthesis and degradation of signaling nucleotides are encoded in acidophilic archaeal genomes. Genes coding for (p)ppGpp metabolizing enzymes are mainly absent in the Archaea domain, excepting a few genes likely acquired by horizontal gene transfer from Bacteria [10]. Indeed, acidophilic Archaea have no genes for (p)ppGpp metabolism (personal communication), and therefore, they probably do not use this signaling molecule. A public database of cyclic di-GMP related proteins (http://www.ncbi.nlm.nih.gov/Complete_Genomes/c-di-GMP.html), shows that three out of 105 archaeal genomes only encode one protein containing the GGDEF domain related to cyclic di-GMP synthesis. On the other hand, the acidophilic archaeal genome of Candidatus Micrarchaeum acidiphilum ARMAN-2 encodes a predicted inactive GGDEF domain containing protein because it does not possess the key amino acids for catalytic activity neither for cyclic di-GMP binding at the inhibition sites location. Then, altogether this data indicate that acidophilic archaea do not use cyclic di-GMP either. Moreover, besides moderate acidophilic marine archaea belonging to the Aciduliprofundum genus, no other acidophilic archaea possess homologues of DAC for synthesis of cyclic di-AMP. The most common component found in 80% of the acidophilic archaeal genomes is a single putative AC belonging to class IV NCs. These proteins are formed by a single CYTH domain whose name was coined from the first two identified members, the CyaB adenylyl cyclase from Aeromonas hydrophila and the human thiamine triphosphatase (ThTPase).
Only few bacterial members of class IV ACs have been biochemically characterized so far. CyaB (A. hydrophila) and YpAC-IV (Yersinia pestis), two bacterial homologues of archaeal CYTH-containing ACs, show optimum activity at 65 °C and 50 °C, respectively [11, 12].
In acidophilic bacteria, cAMP synthesis could be achieved by CyaB orthologs (class IV NC), as well as through class III NCs. In Betaproteobacteria and Acidithiobacillia classes, the CYTH domain of ACs is fused with a “conserved histidine alpha-helical domain” (CHAD), and therefore, probably are able to interact with polyphosphate (polyP) [13], a key player in metal resistance/tolerance of acidophilic bacteria [14]. Notably, Candidatus Koribacter, Candidatus Soilbacter, and Leptospirillum class III NCs possess extracellular or membrane sensory domains like CHASE, dCache, or HAMP, suggesting that cAMP synthesis may be regulated by direct sensing of extracellular signals such as cytokinin-like adenine derivatives or peptides [15], polyamines [16], and also by signals not determined yet [17] that may be inherent to its acidophile habitat. To date, just one acidophilic bacterial genome code for one protein belonging to class I NC (Anthrax_toxA), Acidiphilium angustum ATCC 35903. The presence of a canonical secretion signal indicates that this protein may be secreted, suggesting an extracellular role, as it happens in class I NCs from other bacteria. The outcome of cAMP signaling may be the regulation of gene expression through direct binding to transcription regulator proteins such as well-known CRP (cAMP receptor protein) [18]. Class II, V, and VI NCs were not found in acidophilic bacterial genomes.
Thirteen out of twenty acidophilic bacterial genera encode DACs for cyclic di-AMP synthesis. Putative DACs are present in every Gram-positive acidophilic bacterium but are absent in 7 out of 13 Gram-negative genera, including Acidithiobacillus. Acidophilic bacteria possess one or two copies of DAC genes in their chromosomes. The DAC encoded as a single copy corresponds to a DacZ ortholog, a cytoplasmic protein with a stand-alone DisA_N domain generally found in Archaea [8]. Instead, when two DAC encoding genes are present in a single genome, one of them code for an orthologue of the membrane protein DacA, while the second one is a Thermotoga maritima DisA ortholog (DisA-DisA_linker-HhH) which couple chromosome integrity state with cyclic di-AMP synthesis [19]. Except those involved in cyclic di-AMP synthesis, none homologs genes coding for enzymes that catalyze the synthesis of other signaling nucleotides have been identified in the extremely acidophilic methanotroph Methylacidiphilum infernorum V4, suggesting that cyclic di-AMP pathway could be the exclusive nucleotide signaling system in this microorganism.
Excepting M. infernorum V4, every acidophilic bacterial genus encodes proteins for (p)ppGpp and cyclic di-GMP metabolism. All acidophilic bacteria encode one long (p)ppGpp synthetase with a classical domain configuration: synthesis (RelA_SpoT, pfam04607), hydrolysis (HD, pfam13328), TGS (ThrRS, GTPase, and SpoT), ZFD (zinc-finger domain), and RRM (RNA recognition motif) domains [10, 20]. The conservation of these accessory domains suggests that (p)ppGpp synthesis could be activated by amino acid and/or fatty acid starvation as it occurs in well-studied model microorganisms [20, 21]. As it occurs in non-acidophilic bacteria (e.g., E. coli), (p)ppGpp signaling may work through global control of gene expression by direct binding to the RNA polymerase omega subunit (RpoZ), whose gene product is encoded together with (p)ppGpp synthase gene by the same operon in many acidophilic bacteria. On the other hand, the multiplicity of DGCs and phosphodiesterases (PDEs) enzymes involved in cyclic di-GMP synthesis and degradation, respectively, present in acidophilic bacteria suggests that these microorganisms probably integrate a wide range of signals into cyclic di-GMP pathway for targeting different cellular functions.
3 Cyclic di-GMP in Acidophilic Bacteria
Globally, acidophilic bacteria encode near to the same amount of putative DGC proteins (481 GGDEF domains) and PDEs (446) including EAL (304) and HD_GYP (142) domains. However, the distribution of cyclic di-GMP metabolism related genes is not balanced in individual chromosomes, some of them having more DGCs than PDEs, while in others the opposite distribution occurs. GGDEF and HDGYP containing proteins possess sensor domains in almost the half of cases, 47% and 42%, respectively, meanwhile only one fifth (21%) of EAL PDEs does.
As shown in Fig. 21.1c, most common partners domains in GGDEF-containing proteins are PAS (Period circadian protein, Aryl hydrocarbon receptor nuclear translocator protein and Single-minded protein) and GAF (cGMP-specific phosphodiesterases, Adenylyl cyclases, and FhlA), followed from afar by other domains such as REC (response regulator receiver domain) and Protoglobin [or globin-coupled sensor (GCS)]. On the other hand, most partner domains present in EAL and HD-GYP putative PDEs belong to metal-dependent phosphohydrolase [HD (histidine and aspartate)] family. Recently, the regulation of DGC/PDE activities through the binding of small nucleotides (such as cAMP, GDP) to GAF domain have been characterized [22, 23], suggesting that the GAF domain may establish a functional link between signaling pathways based in different nucleotides. PAS and GCS domains are able to sense two key environmental factors for energetic requirements of chemolithoautotrophic acidophilic bacteria: redox potential [24] and O2 [25, 26]. Energetically, molecular oxygen is the most favorable electron acceptor during RISCs and iron oxidation, while redox potential, which is conditioned by ferrous/ferric iron ratios, drives the composition of the community [27]. In some acidophilic bacteria, such as Acidithiobacillus and Leptospirillum, REC-GGDEF-EAL proteins are encoded together with a histidine kinase in a characteristic configuration of a two-component system [28]. In these cases, phosphorylated and unphosphorylated state of REC domain may modulate the rates of cyclic di-GMP synthesis or degradation by these enzymes. Since acidophilic bacteria thrive in metal-rich environments, it is interesting to note that signaling transduction through chemoreception of metals like zinc may perform through CZB (Chemoreceptor Zn-Binding) domain coupled to GGDEF-EAL (7%), GGDEF-only (2%), or EAL-only proteins (3%).
The output of cyclic di-GMP signaling in acidophilic bacteria appears to mainly occur by PilZ receptor proteins. Excepting for M. infernorum V4, every acidophilic bacterial genera [19] encode at least one PilZ domain protein (Fig. 21.1b). However, there is no evident correlation between the quantity of PilZ proteins and cyclic di-GMP turnover proteins. This could be explained by the presence of other cyclic di-GMP receptor proteins in acidophilic bacteria such as FleQ, PelD and catalytically degenerate and inactive GGDEF and EAL domains also able to bind cyclic di-GMP allosterically (personal communication). PilZ domain occurs alone or in conjunction with other domains in the same polypeptide chain. The most frequent configuration is PilZ domain alone, but it includes proteins long enough to contain non-characterized domains. These proteins are generally related with the formation of type 4 pili (T4P) and twitching motility [29], which has been implicated in irreversible attachment to surfaces, microcolony grouping, and structural development of biofilm [30, 31]. Common partner domains of PilZ proteins of acidophiles are YcgR (pfam07317) and Glycosyl_transferase_2 (pfam00535), which are related with flagellar-based motility [32] and synthesis of different types of exopolysaccharide (EPS) [33, 34], respectively. Among the later, the most recognizable PilZ proteins correspond to A subunit of bacterial cellulose synthase (BcsA), which in some acidophiles is fused with BcsB, forming a fused membrane protein predicted to produce a cellulose-like EPS. An interesting subgroup of enhancer-binding proteins (EBPs) containing PilZ domains are present in the “professional” iron oxidizer genus, Leptospirillum, which have been related with lipopolysaccharide and flagellar biosynthesis [35]. Besides PilZ, a few acidophilic bacteria encode PelD-like effector proteins (see below).
4 Cyclic di-GMP Pathway in Acidithiobacillus
Acidithiobacillus genus members are the most studied acidophilic microorganism. They are chemolithoautotrophic Gram-negative bacteria involved in bio-oxidation of metal sulfides in natural and mining environments. Acidithiobacillus obtain energy from the oxidation of RISCs, producing sulfuric acid as a byproduct. Therefore, all of them are also extreme acidophiles, with optimal growing at pH values below 3. However, there are some important differences among Acidithiobacillus species. Besides RISCs, At. ferrooxidans, At. ferrivorans, At. ferridurans, and At. ferriphilus catalyze the oxidation of ferrous to ferric iron (Fe3+). While most of acidithiobacilli are mesophilic, showing an optimum growth temperature near to 30 °C, At. ferrivorans has psychrotolerant characteristics, being able to grow at 4 °C [36]. On the other hand, At. caldus is the only moderate thermophilic species of the genus, growing between 30 and 50 °C [37]. Several acidithiobacilli are able to express a single polar flagellum for swimming and/or swarming motility. However, strains of At. albertensis express a tuft of polar flagella [38], meanwhile, some At. ferrooxidans strains have no genes required for production and functioning of the flagellum machinery [39]. These variations may be very important in the colonization and dominance at different micro-niches in natural environments, as well as in bioleaching operations, since bio-oxidation activity depends largely on the physiological state of the cell, which in turn is intimately associated with the different bacterial lifestyles such as the single-cell planktonic state and multicellular biofilms. The attachment of acidithiobacilli to the substrate surface and the subsequent biofilm development (Fig. 21.2) plays essential role in bio-oxidation processes for the acidithiobacilli. This is due to the creation of a reaction space that concentrates leaching chemicals at the cell/mineral interface, accelerating the leaching activity [40, 41].
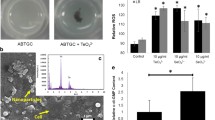
Acidithiobacillus caldus biofilms. (a) Macrobiofilm on tetrathionate plate obtained from a 3 μl-culture inoculum. Red bar, 1 mm. (b) SEM imaging of a 48 h old biofilm developed on elemental sulfur surface. Zoom (right panel) is pointing out the network of filamentous projection connecting biofilm cells. Red bar, 5 μm (left) and 1 μm (right). (c) Fluorescence microscopy (Concanavalin A-FitC) of biofilm dynamics. Sulfur surface is quickly colonized by cells that develop a flat biofilm during 72 h. Then, dispersion of cells conforming the biofilm structure begins (96 h). White bar, 10 μm
Recently, some DGCs and cyclic di-GMP receptor proteins of Acidithiobacillus type strains of At. ferrooxidans (ATCC 23270) [42], At. thiooxidans (ATCC 19377) [43, 44], and At. caldus (ATCC 51756) [45, 46] have been characterized experimentally, validating the functionality of a cyclic di-GMP based signaling in these microorganisms and its relationship with the classical motility and biofilm phenotypes.
The number and the nature of cyclic di-GMP related proteins among Acidithiobacillus species differ notably, ranging from near to 40 metabolism proteins in At. thiooxidans to just 5 (4 GGDEF-EAL, 1 EAL) proteins in both collection strains of At. ferrooxidans (ATCC 23270 and ATCC 53993) (Fig. 21.1). This variability is widespread in acidophile bacteria and it is not related with genome size (Fig. 21.3). In part, this asymmetry could be explained by the presence of transferable genetic elements that contain cyclic di-GMP related genes. Different plasmids and mobile genetic elements containing GGDEF/EAL and PilZ genes have been identified in complete genomes of Acidithiobacillus species, suggesting that they constitute a coherent cyclic di-GMP control module [4]. For instance, the integrative and conjugative element ICEAcaTY2, widely present in At. caldus genomes, carries genes predicted to encode cyclic di-GMP metabolism enzymes (dgc1879, pde1853), and cyclic di-GMP effector proteins (pilz1908, ycgR and fleQ). Besides the inherently biofilm-related functions encode in ICEs, such as assembly factors of T4P and the conjugation machinery, ICEAcaTY2 encodes accessory genes for assembly and functioning of flagellar apparatus. Such elements are potential direct or indirect targets of cyclic di-GMP signaling as it has been observed in other bacteria [47].
Compared with other Acidithiobacillus species, the cyclic di-GMP signaling pathway in At. ferrooxidans ATCC 23270 has the lowest complexity and comprise four GGDEF-EAL, one single EAL, and only two PilZ proteins. Moreover, the single EAL protein and one of the two PilZ proteins do not possess key amino acid motifs for cyclic di-GMP interaction. The use of different electron donors, such as sulfur and iron, supports the expression of all four GGDEF-EAL domain proteins. Furthermore, induction of the cellulose-production phenotype in heterologous complementation assays suggests a net DGC activity of these enzymes [42]. Notably, three GGDEF-EAL proteins, as well as the second PilZ protein are encoded on the ICEAfeTY2 (also present in At. ferrooxidans ATCC 53993). These four genes are clustered in a cyclic di-GMP genetic hot spot together with Tra conjugation and transposase genes, while the fourth GGDEF-EAL protein is encoded next to von Willebrand factor type A domain-containing protein. The genetic contexts strongly suggest the association of the cyclic di-GMP pathway with cellular attachment and clustering. Consequently, cyclic di-GMP levels in biofilm cells are near to 10 times higher than planktonic cells harvested from the same culture [42].
The bioinformatic analysis of At. caldus ATCC 51756 genome allowed the identification of 18 genes related to cyclic di-GMP turnover (9 single GGDEF, 6 GGDEF-EAL, and 3 single EAL) and 10 putative cyclic di-GMP effector proteins (9 PilZ and 1 PelD). Based on key amino acid conservation and cellulose-production phenotype assays in E. coli and Salmonella Typhimurium, the single GGDEF protein ACAty1319 was identified as a functional DGC in this bacterium and selected for mutagenesis experiment [46].
A null-mutant strain ΔACAty1319 was developed by using a suicide vector harboring a kanamycin cassette and both 5′ and 3′ ends of ACAty1319 encoding gene [46]. Then, by comparing mutant and wild-type strains it was reported that this DGC is mainly responsible for 85–93% of the global cyclic di-GMP intracellular levels and plays significant roles on (1) early stages of biofilm development (Fig. 21.4a) and (2) swarming motility (Fig. 21.4b). The immediate gene context of ACAty1319 that contains fliL, motA, and motB orthologous suggests that it may affect flagellar motor performance [46].
Attachment (a) and motility (b) phenotypes in At. caldus ATCC 51756 wild type (WT) and DGC null-mutant (ΔACAty1319) strains. Attached cells on elemental sulfur were visualized by fluorescence microscopy (upper panel a) and SEM (bottom panel a). Swarming motility patterns were observed on semi-solid media [phytagel 0.2%, pH 4.7 (upper panel b), and phytagel 0.1% pH 2.5 (bottom panel b)] with tetrathionate as an energy source
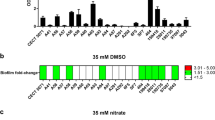
The At. thiooxidans ATCC 19377 strain encodes an extended cyclic di-GMP signaling network with 25 cyclic di-GMP metabolism genes (9 GGDEF, 12 GGDEF-EAL, 3 EAL, 1 HD-GYP) plus 9 PilZ and 1 PelD. Most of GGDEF-containing proteins were characterized as functional DGCs through induction of the rdar (rough, dry and red) biofilm morphotype in S. Typhimurium and high intracellular levels of cyclic di-GMP were reported in attached cells [44]. Like At. caldus, At. thiooxidans encode a PelD cyclic di-GMP effector in a complete pel operon (Fig. 21.5a) [48, 49] that includes an additional gene, wcaG [uridine diphosphate (UDP)-glucose-4-epimerase], downstream pelG gene and it is probably involved in the synthesis of PEL exopolysaccharide precursor. Transcription levels of pelA, pelD, and wcaG genes increase in At. thiooxidans biofilm cells. In cells attached to sulfur surface, the deletion of pelD gene induces a decrease of EPS production (sixfold less of total carbohydrates fraction), and an increase of the total protein fraction. SEM imaging reveals that ΔpelD null-mutant cells attached to sulfur overexpress a filamentous structure (Fig. 21.5b) that could have a proteinaceous nature [44]. Intriguingly, Pel biosynthesis operon does not fallow the phylogenetic distribution, and are not carried on an apparent mobile genetic element. The Pel exopolysaccharide cluster is present in Sulfurihydrogenibium [moderate acidophile (pH 6) thermophilic (68 °C)], Acidihalobacter prosperus [extreme acidophilic (pH 2) mesophile (37 °C)], Ferrovum myxofaciens [moderate acidophile (pH 2.5–4.8) mesophile (30 °C)], At. caldus [extreme acidophilic (pH 2.5), moderate thermophilic (45 °C)] and At. thiooxidans [extreme acidophilic (pH 2.5), mesophile (30 °C)]. Interestingly, PelB, a component that spans the periplasm and the outer membrane, seems to be the most variable element of the Pel biosynthesis gene products, probably due to adaptation to different extracellular conditions (Fig. 21.5a).
(a) Organization and conservation of the pel gene clusters in acidophilic bacteria. Optima growth pH and temperature for pel operon containing acidophilic bacteria are noted. Pel operon structure from P. aeruginosa PA14 is presented as a reference. The percentage of identity between adjacent gene clusters is represented as gray bars. PelD gene is depicted in purple. Note that wcaG is part of pel gene cluster in several acidophiles. (b) SEM imaging of biofilm structure produced by the At. thiooxidans ΔpelD null-mutant strain overexpressing a filamentous structure
5 Concluding Remarks
Signal transduction mechanisms are key molecular processes that allow diverse microorganisms to adapt and respond to their surroundings. This is particularly important in extremophilic microorganisms which thrive in very harsh environments. As pointed out in this chapter, acidophilic microorganisms have the capability to create signaling circuits based on diverse nucleotide second messengers. Most acidophilic Archaea probably use cAMP-based signaling, meanwhile several acidophilic bacteria use cAMP, cyclic di-AMP, (p)ppGpp, and cyclic di-GMP, with the last two being the most prevalent second messenger signaling molecules. In acidophilic bacteria, (p)ppGpp and cAMP seem to regulate gene expression in response to starvation and polyP-related signal, respectively. On the other hand, cyclic di-GMP signaling integrates several input signals through different metabolism components. DGCs from acidophilic bacteria detect environmental signals through protein domains related with critical cues for chemolithoautotrophic bacteria which dedicate large quantity of energy to carbon fixation. Oxidation of RISCs and iron II offers little amount of energy, leaving oxygen as the most favorable electron acceptor [27]. Besides, the redox potential and consequently the energy output has a tremendous impact on the growth rate of acidophilic bacteria such as Acidithiobacillus and Leptospirillum due their different iron oxidizing machinery [27]. Then, DGCs containing domains able to sense redox potential and/or oxygen, such as PAS and protoglobin may be vital to transduce these cues to a particular response, such as biofilm formation on an oxidizable substrate. On the other hand, metal sensing through CZB domain appears as an important cue for regulation of acidophiles DGCs, probably impeding biofilm formation through DGC activity inhibition, as has been shown for other bacteria [50]. Besides, acidophiles DGCs couple GGDEF domain with poorly characterized N-terminal signaling domains involved in the perception of signals at extracellular (7TMR-DISMED2 and dCache_1), membrane (CHASE, MASE1 and MHYT), and periplasmic (Reg_prop and Y_Y_Y) level. They may help to transduce specific signals from acidic ecological niche and then have to be targeted for further studies.
Key behaviors for energy acquisition, such as attachment and subsequent biofilm formation, may be regulated by cyclic di-GMP through flagellar motility inhibition and EPS synthesis, specially, cellulose-like and PEL expolysaccharides, which in turn is achieved through effector proteins such as PelD and PilZ domains containing proteins.
The physiological characterization of cyclic di-GMP pathway in acidophilic bacteria is still incipient but the current knowledge obtained from bioinformatics analysis and biological experiments in Acidithiobacillus species introduces some clues about the regulation of biofilm formation in this kind of microorganisms. Thus, it opens new ways to regulate this key physiological behavior for industrial or environmental applications, either to increase precious metals release from leaching ores or to control unwanted acid production in natural habitats.
References
Johnson DB, Quatrini R (2016) Acidophile microbiology in space and time. In: Quatrini R, Johnson DB (eds) Acidophiles: life in extremely acidic environments. Caister Academic Press, London, pp 1–16. https://doi.org/10.21775/9781910190333.16
Garcia-Moyano A, González-Toril E, Aguilera A, Amils R (2007) Prokaryotic community composition and ecology of floating macroscopic filaments from an extreme acidic environment, Río Tinto (SW, Spain). Syst Appl Microbiol 30:601–614
Nancucheo I, Johnson DB (2010) Production of glycolic acid by chemolithotrophic iron- and sulfur-oxidizing bacteria and its role in delineating and sustaining acidophilic sulfide mineral-oxidizing consortia. Appl Environ Microbiol 76:461–467
Moya-Beltrán A, Rojas C, Díaz M, Guiliani N, Quatrini R, Castro M (2019) Nucleotide second messenger-based signaling in extreme acidophiles of the Acidithiobacillus species complex: partition between the core and variable gene complements. Front Microbiol 10:381. https://doi.org/10.3389/fmicb.2019.00381
Hauryliuk V, Atkinson GC, Murakami KS, Tenson T, Gerdes K (2015) Recent functional insights into the role of (p)ppGpp in bacterial physiology. Nat Rev Microbiol 13(5):298–309. https://doi.org/10.1038/nrmicro3448
McDonough KA, Rodriguez A (2011) The myriad roles of cyclic AMP in microbial pathogens: from signal to sword. Nat Rev Microbiol 10(1):27–38. https://doi.org/10.1038/nrmicro2688
Römling U, Galperin MY, Gomelsky M (2013) Cyclic di-GMP: the first 25 years of a universal bacterial second messenger. Microbiol Mol Biol Rev 77(1):1–52. https://doi.org/10.1128/MMBR.00043-12
Corrigan RM, Gründling A (2013) Cyclic di-AMP: another second messenger enters the fray. Nat Rev Microbiol 11(8):513–524. https://doi.org/10.1038/nrmicro3069
Sinha SC, Sprang SR (2006) Structures, mechanism, regulation and evolution of class III nucleotidyl cyclases. Rev Physiol Biochem Pharmacol 157:105–140
Atkinson GC, Tenson T, Hauryliuk V (2011) The RelA/SpoT homolog (RSH) superfamily: distribution and functional evolution of ppGpp synthetases and hydrolases across the tree of life. PLoS One 6(8):e23479. https://doi.org/10.1371/journal.pone.0023479
Sismeiro O, Trotot P, Biville F, Vivares C, Danchin A (1998) Aeromonas hydrophila adenylyl cyclase 2: a new class of adenylyl cyclases with thermophilic properties and sequence similarities to proteins from hyperthermophilic archaebacteria. J Bacteriol 180(13):3339–3344
Smith N, Kim SK, Reddy PT, Gallagher DT (2006) Crystallization of the class IV adenylyl cyclase from Yersinia pestis. Acta Crystallogr Sect F Struct Biol Cryst Commun 62(Pt 3):200–204
Tumlirsch T, Jendrossek D (2017) Proteins with CHADs (conserved Histidine α-helical domains) are attached to polyphosphate granules in vivo and constitute a novel family of polyphosphate-associated proteins (Phosins). Appl Environ Microbiol 83(7):e03399–e03316. https://doi.org/10.1128/AEM.03399-16
Navarro C, von Bernath D, Jerez CA (2013) Heavy metal resistance strategies of acidophilic bacteria and their acquisition: importance for biomining and bioremediation. Biol Res 46:363–371
Anantharaman V, Aravind L (2001) The CHASE domain: a predicted ligand-binding module in plant cytokinin receptors and other eukaryotic and bacterial receptors. Trends Biochem Sci 26(10):579–582
Gavira JA, Ortega Á, Martín-Mora D, Conejero-Muriel MT, Corral-Lugo A, Morel B, Matilla MA, Krell T (2018) Structural basis for polyamine binding at the dCACHE domain of the McpU chemoreceptor from pseudomonas putida. J Mol Biol 430(13):1950–1963. https://doi.org/10.1016/j.jmb.2018.05.008
Finkbeiner M, Grischin J, Seth A, Schultz JE (2019) In search of a function for the membrane anchors of class IIIa adenylate cyclases. Int J Med Microbiol S1438-4221(19):30021–30029. https://doi.org/10.1016/j.ijmm.2019.03.006
Green J, Stapleton MR, Smith LJ, Artymiuk PJ, Kahramanoglou C, Hunt DM, Buxton RS (2014) Cyclic-AMP and bacterial cyclic-AMP receptor proteins revisited: adaptation for different ecological niches. Curr Opin Microbiol 18:1–7. https://doi.org/10.1016/j.mib.2014.01.003
Witte G, Hartung S, Büttner K, Hopfner KP (2008) Structural biochemistry of a bacterial checkpoint protein reveals diadenylate cyclase activity regulated by DNA recombination intermediates. Mol Cell 30(2):167–178. https://doi.org/10.1016/j.molcel.2008.02.020
Winther KS, Roghanian M, Gerdes K (2018) Activation of the stringent response by loading of RelA-tRNA complexes at the ribosomal A-site. Mol Cell 70(1):95–105.e4. https://doi.org/10.1016/j.molcel.2018.02.033
Battesti A, Bouveret E (2009) Bacteria possessing two RelA/SpoT-like proteins have evolved a specific stringent response involving the acyl carrier protein-SpoT interaction. J Bacteriol 191(2):616–624. https://doi.org/10.1128/JB.01195-08
da Costa Vasconcelos FN, Maciel NK, Favaro DC, de Oliveira LC, Barbosa AS, Salinas RK et al (2017) Structural and enzymatic characterization of a cAMP-dependent diguanylate cyclase from pathogenic Leptospira species. J Mol Biol 429(15):2337–2352. https://doi.org/10.1016/j.jmb.2017.06.002
Chen HJ, Li N, Luo Y, Jiang YL, Zhou CZ, Chen Y et al (2018) The GDP-switched GAF domain of DcpA modulates the concerted synthesis/hydrolysis of c-di-GMP in Mycobacterium smegmatis. Biochem J 475(7):1295–1308. https://doi.org/10.1042/BCJ20180079
Qi Y, Rao F, Luo Z, Liang ZX (2009) A flavin cofactor-binding PAS domain regulates c-di-GMP synthesis in AxDGC2 from Acetobacter xylinum. Biochemistry 48(43):10275–10285. https://doi.org/10.1021/bi901121w
Chang AL, Tuckerman JR, Gonzalez G, Mayer R, Weinhouse H, Volman G, Amikam D, Benziman M, Gilles-Gonzalez MA (2001) Phosphodiesterase A1, a regulator of cellulose synthesis in Acetobacter xylinum, is a heme-based sensor. Biochemistry 40(12):3420–3426
Tuckerman JR, Gonzalez G, Sousa EH, Wan X, Saito JA, Alam M, Gilles-Gonzalez MA (2009) An oxygen-sensing diguanylate cyclase and phosphodiesterase couple for c-di-GMP control. Biochemistry 48(41):9764–9774. https://doi.org/10.1021/bi901409g
Rawlings DE (2005) Characteristics and adaptability of iron- and sulfur-oxidizing microorganisms used for the recovery of metals from minerals and their concentrates. Microb Cell Factor 4:13. https://doi.org/10.1186/1475-2859-4-13
Zschiedrich CP, Keidel V, Szurmant H (2016) Molecular mechanisms of two- component signal transduction. J Mol Biol 428:3752–3775
Guzzo CR, Salinas RK, Andrade MO, Farah CS (2009) PILZ protein structure and interactions with PILB and the FIMX EAL domain: implications for control of type IV pilus biogenesis. J Mol Biol 393(4):848–866. https://doi.org/10.1016/j.jmb.2009.07.065
O’Toole GA, Kolter R (1998) Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol Microbiol 30(2):295–304
Semmler AB, Whitchurch CB, Mattick JS (1999) A re-examination of twitching motility in Pseudomonas aeruginosa. Microbiology 145(10):2863–2873
Ryjenkov DA, Simm R, Römling U, Gomelsky M (2006) The PilZ domain is a receptor for the second messenger c-di-GMP: the PilZ domain protein YcgR controls motility in enterobacteria. J Biol Chem 281(41):30310–30314
Merighi M, Lee VT, Hyodo M, Hayakawa Y, Lory S (2007) The second messenger bis-(3′-5′)-cyclic-GMP and its PilZ domain-containing receptor Alg44 are required for alginate biosynthesis in Pseudomonas aeruginosa. Mol Microbiol 65(4):876–895
Morgan JL, McNamara JT, Zimmer J (2014 May) Mechanism of activation of bacterial cellulose synthase by cyclic di-GMP. Nat Struct Mol Biol 21(5):489–496. https://doi.org/10.1038/nsmb.2803
Francke C, Groot Kormelink T, Hagemeijer Y, Overmars L, Sluijter V, Moezelaar R, Siezen RJ (2011) Comparative analyses imply that the enigmatic sigma factor 54 is a central controller of the bacterial exterior. BMC Genomics 12(1):385. https://doi.org/10.1186/1471-2164-12-385
Hallberg K, González-Toril E, Johnson D (2010) Acidithiobacillus ferrivorans, sp. nov.; facultatively anaerobic, psychrotolerant iron-, and sulfur-oxidizing acidophiles isolated from metal mine-impacted environments. Extremophiles 14:9–19. https://doi.org/10.1007/s00792-009-0282-y
Hallberg KB, Lindström EB (1994) Characterization of Thiobacillus caldus sp. nov., a moderately thermophilic acidophile. Microbiology 140(12):3451–3456
Castro M, Moya-Beltrán A, Covarrubias PC, Gonzalez M, Cardenas JP, Issotta F, Nuñez H, Acuña LG, Encina G, Holmes DS, Johnson DB, Quatrini R (2017) Draft genome sequence of the type strain of the sulfur-oxidizing acidophile, Acidithiobacillus albertensis (DSM 14366). Stand Genomic Sci 12:77. https://doi.org/10.1186/s40793-017-0282-y
Valdés J, Pedroso I, Quatrini R, Holmes DS (2008) Comparative genome analysis of Acidithiobacillus ferrooxidans, A. thiooxidans and A. caldus: insights into their metabolism and ecophysiology. Hydrometallurgy 94:180–184
Schippers A, Sand W (1999) Bacterial leaching of metal sulfides proceeds by two indirect mechanisms via thiosulfate or via polysulfides and súlfur. Appl Environ Microbiol 65(1):319–321
Vera M, Schippers A, Sand W (2013) Progress in bioleaching: fundamentals and mechanisms of bacterial metal sulfide oxidation--part A. Appl Microbiol Biotechnol 97(17):7529–7541. https://doi.org/10.1007/s00253-013-4954-2
Ruiz LM, Castro M, Barriga A, Jerez CA, Guiliani N (2012) The extremophile Acidithiobacillus ferrooxidans possesses a c-di-GMP signalling pathway that could play a significant role during bioleaching of minerals. Lett Appl Microbiol 54:133–139
Diaz M, Copaja S, Guiliani N (2013) Functional analysis of c-di-GMP pathway in biomining bacteria Acidithiobacillus thiooxidans. Adv Mater Res 825:133–136
Díaz M, Castro M, Copaja S, Guiliani N (2018) Biofilm formation by the Acidophile bacterium Acidithiobacillus thiooxidans involves c-di-GMP pathway and Pel exopolysaccharide. Genes (Basel) 9(2):E113. https://doi.org/10.3390/genes9020113
Castro M, Ruíz LM, Barriga A, Jerez CA, Holmes DS, Guiliani N (2009) C-di-GMP pathway in biomining bacteria. Adv Mater Res 71–73:223–226. https://doi.org/10.4028/www.scientific.net/AMR.71-73.223
Castro M, Deane S, Ruiz L, Rawlings DE, Guiliani N (2015) Diguanylate Cyclase null mutant reveals that C-Di-GMP pathway regulates the motility and adherence of the extremophile bacterium Acidithiobacillus caldus. PLoS One 10(2):e0116399. https://doi.org/10.1371/journal.pone.0116399
Ryan RP, Tolker-Nielsen T, Dow JM (2012) When the PilZ don’t work: effectors for cyclic di-GMP action in bacteria. Trends Microbiol 20(5):235–242
Friedman F, Kolter R (2004) Genes involved in matrix formation in Pseudomonas aeruginosa PA14 biofilms. Mol Microbiol 51(3):675–690
Lee VT, Matewish JM, Kessler JL, Hyodo M, Hayakawa Y, Lory S (2007) A cyclic-di-GMP receptor required for bacterial exopolysaccharide production. Mol Microbiol 65(6):1474–1484
Zähringer F, Lacanna E, Jenal U, Schirmer T, Boehm A (2013) Structure and signaling mechanism of a zinc-sensory diguanylate cyclase. Structure 21(7):1149–1157. https://doi.org/10.1016/j.str.2013.04.026
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Castro, M., Díaz, M., Moya Beltrán, A., Guiliani, N. (2020). Cyclic di-GMP Signaling in Extreme Acidophilic Bacteria. In: Chou, SH., Guiliani, N., Lee, V., Römling, U. (eds) Microbial Cyclic Di-Nucleotide Signaling. Springer, Cham. https://doi.org/10.1007/978-3-030-33308-9_21
Download citation
DOI: https://doi.org/10.1007/978-3-030-33308-9_21
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-33307-2
Online ISBN: 978-3-030-33308-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)