Abstract
The accumulation of non-functional oxidized proteins is a hallmark of aging both in cells and in the body. This age-related build-up of proteins modified by oxidative processes results, at least in part, from an increase in reactive oxygen species and other toxic compounds from both cellular metabolism and external environmental factors. Failure of protein maintenance (i.e. oxidized protein degradation and repair) is another major contributor to the age-associated accumulation of damaged proteins. Oxidative damage to the cellular proteome, leading to the formation of carbonylated proteins derives from the direct oxidation of several amino acids side chains and also through protein adducts formation with lipid peroxidation products and dicarbonyl glycating compounds. Since the accumulation of oxidatively damaged proteins is believed to participate to the age-related decline in cellular function, their identification has been achieved in human or mammalian animal models of aging and age-related diseases as well as in human fibroblasts and myoblasts during cellular senescence and upon oxidative stress. Indeed, the identification of damaged protein targets is expected not only to define potential biomarkers of aging but also to give insight into the mechanisms by which these damaged proteins accumulate and may contribute to cellular dysfunction.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Protein oxidation
- Protein glycoxidation
- Oxidized protein degradation
- Proteasome
- Oxidized protein repair
- Methionine sulfoxide reductases
- Protein maintenance
- Cytoskeleton
- Energy metabolism
1 Introduction
A characteristic of aging, both in cells and in the body, is the accumulation of non-functional oxidized proteins. This age-related accumulation of proteins modified by oxidative processes results, at least in part, from an increase in reactive oxygen species (ROS) and other toxic compounds from both cellular metabolism and external environmental factors (Stadtman 2006; Ugarte et al. 2010). Another major contributor to the accumulation of damaged proteins with age is the decreased efficiency of the intracellular elimination (degradation and repair) of oxidized proteins (Friguet 2006; Baraibar and Friguet 2012; Chondrogianni et al. 2014).
Inside the cell ROS are formed mainly by the mitochondrial electron transport chain during aerobic respiration resulting in the production of superoxide radical (O2°−). Superoxide is then converted into hydrogen peroxide (H2O2) by superoxide dismutase, which in the presence of traces of metals (Cu+ or Fe++) is itself converted by the Fenton reaction to hydroxyl radical (OH°), the most deleterious ROS. Peroxisomes, which are organelles involved in the oxidative metabolism of organic molecules, also contribute to the endogenous formation of ROS. External stresses of a physical nature (such as ultraviolet radiation) or chemical (toxin or xenobiotic) may also participate in the intracellular production of ROS. To trap these ROS, the cell has an arsenal of antioxidant defenses, both enzymatic (superoxide dismutase, catalase, glutathione peroxidase, peroxiredoxins, etc. …) and non-enzymatic (vitamin C, vitamin E, flavonoids, carotenoids, etc. …). However, when their production becomes too large to be completely trapped by antioxidant defenses, ROS are reacting with biological macromolecules (lipids, proteins and nucleic acids), causing irreversible damage (Berlett and Stadtman 1997; Stadtman and Levine 2003).
With the notable exception of the oxidation of cysteines and methionines for which specific systems of reduction have been demonstrated, in all other situations where the amino acid modification of the polypeptide chain is considered irreversible, the problem arises as to the removal of oxidized protein by degradation (Petropoulos and Friguet, 2005; Farout and Friguet 2006; Chondrogianni et al. 2014). At the intracellular level, the degradation of oxidized proteins is mainly ensured in the nucleus and in the cytoplasm by the proteasome, a high molecular weight proteolytic multicatalytic complex. The observed decrease in proteasome activity with age plays a crucial role in the accumulation of abnormal oxidized proteins that can be toxic to the cell. Thus, depending on the cell or tissue type, the decrease in proteasome activity with age can come from the combined action of: (i) the decrease of its expression, (ii) the existence of structural modifications of the proteasome subunits, (iii) the presence in aged cells of damaged proteasome inhibitory proteins (Shringarpure and Davies 2002; Farout and Friguet 2006; Breusing and Grune 2008; Baraibar and Friguet 2012). Other protein maintenance systems are also capable of repairing oxidatively damaged proteins. Concerning the methionine sulfoxide reductases system, which has the ability to reverse the oxidation of methionine in proteins, its activity and its expression are also altered during aging and cellular senescence (Petropoulos et al. 2001; Picot et al. 2004.)
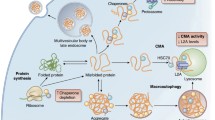
In this chapter, the different types of oxidative modifications of proteins, the incidence of which increases with age, are first described. Then, specific proteins that have been identified as increasingly carbonylated or modified by glycation/glycoxidation or conjugation with lipid peroxidation products in human or mammalian animal models of aging and age-related diseases and that may represent interesting biomarkers are presented. Intracellular degradation and repair systems for oxidized proteins, with a special attention devoted to the involvement of the proteasome and of the methionine sulfoxide reductases system (Fig. 8.1), and their impairment during the aging process are then discussed. Finally, since the accumulation of oxidatively damaged proteins is believed to participate to the age-related decline in cellular function, their previously reported identification in human fibroblasts and myoblasts during cellular senescence and upon oxidative stress is also presented. Indeed, the identification of damaged protein targets is expected not only to define potential biomarkers of aging but also to give insight into the mechanisms by which these damaged proteins accumulate and may contribute to cellular dysfunction.
Schematic representation of the alteration of the protein homeostasis network during aging and upon oxidative stress. Accumulation of proteins modified by oxidative processes with age results from an increase in reactive oxygen species and other toxic compounds from both cellular metabolism and environmental factors together with a decreased efficacy of the intracellular degradation and/or repair of oxidatively damaged proteins (Baraibar and Friguet 2012)
2 Protein Modification by Oxidation and Related Pathways
Proteins constitute the main targets for ROS mediated damage that occurs directly or indirectly through their reaction with lipids and carbohydrates and the subsequent generation of oxidized products that can react with proteins. Oxidations of proteins by ROS can be classified into those that oxidize the amino acid side chains and those that oxidize and cleave the peptide bond. Almost all amino acid side chains can react with the hydroxyl radical OH° but certain amino acids are more sensitive to oxidation with such ROS as hydrogen peroxide and superoxide. Indeed, sulfur-containing amino acids methionine and cysteine are readily oxidized by all sorts of ROS, while histidine and aromatic amino acids are also very sensitive to oxidation.
Oxidation of cysteine first results in either the formation of a disulfide bridge or a sulfenic acid. Sulfenic acid can be converted to disulfide or sulfenamide or further oxidized to sulfinic and then sulfonic acids. Both disulfide and sulfenic acid can be enzymatically reduced by different enzymatic systems such as the thioredoxin/thioredoxin reductase and the glutaredoxin/glutathione/glutathione reductase systems while sulfinic acid reduction has so far been limited to oxidized cysteines within the active site of peroxiredoxins (Biteau et al. 2003). Cysteine can also react with nitric oxide to produce S-nitrosothiol. Oxidation of methionine leads to the formation of methionine sulfoxide and further oxidation of methionine sulfoxide leads to the irreversible formation of methionine sulfone. Some oxidative modifications are rather specific in terms of oxidized residues and products generated such as the oxidation of phenylalanine to tyrosine, which can be further converted to di-tyrosine (Giulivi et al. 2003). Tyrosine residues also represent preferred targets for nitration by nitrogen dioxide and peroxynitrite and are converted to nitrotyrosine.
Oxidation of several amino acid residues such as lysine, arginine, proline and threonine results in the formation of carbonyl groups (Berlett and Stadtman 1997). Carbonyl derivatives can also originate from the fragmentation products of the peptide bond oxidative cleavage (Stadtman and Levine 2003). Aminoadipic and glutamic semi-aldehydes resulting from the oxidation of lysine and arginine, respectively, are quantitatively important products of the carbonylation reaction. Protein carbonylation has been considered as an indicator of severe oxidative damage as well as age- and disease-derived protein dysfunction since this modification often leads to a loss of protein function, as well as an increased thermosensitivity and hydrophobicity (Berlett and Stadtman 1997).
Protein carbonyls are the most commonly used marker of protein oxidation and different methods have been developed for the detection and quantification of carbonylated proteins. Most of these methods are based on immunochemical and/or spectrophotometric assays of protein carbonyls previously derivatized by 2-4-dinitrophenylhydrazine to form 2-4-dinitrophenylhydrazone protein adducts (Levine 2002). We have developed a novel application of the difference gel electrophoresis (DIGE) approach, but for the detection and quantification of carbonylated proteins, referred as to Oxi-DIGE (Baraibar et al. 2012a, 2013). In Oxi-DIGE, protein carbonyls derived from any biological sample are labeled with two spectrally resolvable fluorescent hydrazide probes that bind specifically to carbonyl groups in proteins (Fig. 8.2). The matched dyes have the same ionic and pH characteristics but absorb and/or emit light at different wavelengths, producing different color fluorescence. Recent studies by the group of J. Ros and E. Cabiscol have also demonstrated the usefulness of fluorescent hydrazides for analyzing protein carbonylation caused by oxidative stress and chronological aging in yeast (Tamarit et al. 2012). A central advantage of the use of sensitive fluorescent probes is the detection of lower abundance carbonylated proteins. For spot excision from the gel to establish its identification, either total protein stain by NHS-ester cyanines or BodipyFL-Hz would be the method of choice (Tamarit et al. 2012). In carrying out Oxi-DIGE, labeled carbonylated proteins from different groups of samples are co-resolved on a single 2D gel for direct quantification. In addition, the Oxi-DIGE method provides a significant improvement in terms of reproducibility and statistical support of the data for proteomic analysis of carbonylated proteins, which is essential for the robust identification of this modification, and can be applied to the identification of carbonylated proteins in any biological sample.
Representative Oxi-DIGE analysis for the quantitative detection of carbonylated proteins. In Oxi-DIGE, protein carbonyls are labeled with two spectrally resolvable fluorescent hydrazide probes that bind specifically to carbonyl groups in proteins. Labeled carbonylated proteins from different groups of samples are then co-resolved on a single 2D gel for direct quantification. Oxi-DIGE analysis can be performed for the quantification and identification of carbonylated proteins in any biological sample (Baraibar et al. 2013)
In addition to direct oxidation of certain aminoacid side chains, protein carbonyl derivatives can originate from the conjugation on cysteine, lysine and histidine residues of such aldehydes as malondialdehyde, acrolein and 4-hydroxy-2-nonenal (HNE). Indeed, oxygen free radicals can attack cellular membranes and induce lipid peroxidation resulting in the production of these reactive aldehydes which are precursors of advanced lipid peroxidation end products (ALE) that have been found to accumulate on proteins during aging and certain age-related diseases (Sayre et al. 1997; Szweda et al. 2003). Moreover, sugar aldehydes or ketones can also react with the amino groups of lysine and arginine through a Schiff base which is rearranged to form an Amadori product (e.g. fructosamine when the reacting sugar is glucose). These products are referred as to early stage glycation adducts that are further modified to form stable end-stage products also called advanced glycation end products (AGE) through either rearrangement, oxidation, dehydration, fragmentation and/or cyclization. Deleterious effect on protein function is observed when the modification affects critical amino acids within the protein and many proteins, including intracellular proteins, accumulate with age as AGE-modified in vivo (Horiuchi and Araki 1994).
3 Age-Associated Accumulation of Oxidatively Modified Proteins
Accumulation of damaged macromolecules , including oxidatively damaged proteins, is a hallmark of aging, both at the cellular and organismal level, which is resulting from increased oxidative stress and/or failure of protein repair and maintenance systems (Stadtman 2006; Ugarte et al. 2010). ROS are produced as byproducts of oxidative phosphorylation and aerobic metabolism. Moreover, ROS production and accumulation are usually increased during disease pathogenesis, in particular age-related diseases (Kregel and Zang 2007). Transient exposure to low concentration of ROS induces cell proliferation and regulates the activation of several signaling pathways (Apel and Hirt 2004) while excess of ROS causes oxidative damage to lipids, proteins, and nucleic acids (Mecocci et al. 1999; Avery 2011). Protein oxidation is particularly detrimental as the resulting damages to protein structures can render oxidized proteins inactive and/or prone to form protein aggregates, hence leading to cellular functional abnormalities (Picot et al. 2007; Baraibar et al. 2012b).
ROS can induce various types of protein oxidative modifications either directly or indirectly by reactions with secondary products of oxidative stress (Stadman and Levine 2003). The irreversible oxidation of residues other than cysteine and methionine most frequently leads to hydroxylated and carbonylated aminoacid side chain derivatives. The exponential accumulation of carbonylated proteins during life span both at the cellular and organismal level and their particular increase in organs affected by age-related diseases, imply that this “Oxi-proteome” (i.e. the restricted set of proteins targeted by oxidation) may be a potential molecular substratum for many of the associated cellular dysfunctions. Indeed, carbonylated proteins are generally less active, less thermostable and are exposing hydrophobic amino acids at their surface, making them prone to form protein aggregates. Since oxidative modifications that give rise to carbonyl groups generally cause loss of catalytic or structural alterations in the affected proteins, the increased level of oxidatively modified proteins observed during aging and age-related disease has been proposed to have deleterious effects on cellular and organ function.
Increased levels of protein carbonyls have been observed in age-related diseases, such as neurodegenerative diseases (amyotrophic lateral sclerosis, Alzheimer’s, Parkinson’s, and Huntington’s diseases), cataractogenesis, systemic amyloidosis, muscular dystrophy, progeria, Werner’s syndrome, rheumatoid arthritis, and respiratory distress syndrome (Berlett and Stadtman 1997; Dalle-Donne et al. 2003; Martinez et al. 2010). Elevated levels of proteins modified by lipid oxidation products reactive aldehydes are associated with neurodegenerative diseases, iron induced renal carcinogenesis, cardiovascular disease, as well as elevated levels of protein glycation/glycoxidation end products (AGE) are associated with diabetes mellitus, neurodegenerative diseases, atherosclerosis and Down’s syndrome. Significant advances in the past recent years have been made towards the identification of proteins targeted by these modifications, although their possible causative role in the pathogenesis of these diseases has yet to be elucidated.
To further address the role of modified proteins, we have performed a bibliographical search for specific proteins identified as increasingly carbonylated or modified by AGE or HNE in human or mammalian animal models of aging and age-related diseases (neurodegenerative diseases, cancer, diabetes) in articles published in peer-reviewed journals (Baraibar et al. 2012c). A total of 183 modified proteins were identified in brain, cerebellum, spinal cord, skeletal muscle, liver, eye, and cerebrospinal and bronchoalveolar fluids. Due to the high number of studies addressing the importance of protein carbonylation in the pathogenesis of neurodegenerative diseases, such as Alzheimer’s, Parkinson’s and Huntington’s diseases, most of the proteins belong from the brain (Butterfield et al. 2006). However, several proteins have been identified consistently modified in other organs such as liver and eye, indicating that the spectrum of proteins targeted by these modifications may be conserved. Among them, cytoplasmic proteins were predominant, followed by proteins from mitochondria, nucleus, endoplasmic reticulum and plasma membrane. However, since most of the studies were performed in total tissue soluble extracts, membrane and mitochondrial proteins are clearly underrepresented when compared to cytosolic proteins.
Functional grouping indicated that proteins were distributed within biological processes such as inflammatory response, cellular metabolism, free radical scavenging, protein synthesis and folding. Concerning proteins involved in the inflammatory response, inflammation is now accepted as a key factor in physiological aging, referred as to “inflamm-aging” (Franceschi et al. 2000), as well as in the development of several age-related pathologies including neurodegenerative and cardiovascular diseases. Interestingly, the inflammatory environment is highly oxidative, and increased protein oxidation has been described, generating a positive feedback process. Proteins involved in energy metabolism were also evidenced in the referenced modified proteins. The most represented canonical pathways across the entire dataset included: glycolysis/gluconeogenesis, citrate cycle, pyruvate metabolism, amino acids degradation, mitochondrial dysfunction, cell death, butanoate metabolism, nrf-2 oxidative stress response and cellular function and maintenance.
More recently, we have identified carbonylated proteins in human rectus abdominis muscle obtained from old and young healthy donors to better understand the mechanisms by which these damaged proteins build up and potentially affect muscular function (Lourenço dos Santos et al. 2015). Using a bi-dimensional gel electrophoresis-based proteomic approach coupled with the immunodetection of carbonylated proteins, 14 proteins were found to be increasingly carbonylated in biopsies from old donors compared to young counterparts. Interestingly, about half of them are already present in the above-mentioned list of 183 oxidatively modified proteins (Baraibar et al. 2012c). These proteins are involved in key cellular functional pathways such as cellular morphology and transport, muscle contraction and energy metabolism. Since, impairment of these pathways has been previously described in skeletal muscle during aging, the irreversible oxidation of these proteins, leading to their functional decline, may therefore contribute to the sarcopenic phenotype by negatively impacting on such pathways.
4 Age-Associated Impairment of Oxidized Proteins Elimination by Repair and Degradation
In contrast to DNA, for which many repair enzymes and pathways have been described for oxidative and other insults, oxidized protein repair is limited to the reduction of certain oxidation products of the sulfur-containing amino acids, cysteine and methionine. Indeed, damaged intracellular proteins are mainly eliminated be degradation by the proteasomal and the lysosomal pathways. Major systems that have been implicated in oxidized protein repair include thioredoxin/thioredoxin reductase and the glutaredoxin/glutathione/glutathione reductase systems for the reduction of sulfenic acid and disulfide bridges, the sulfiredoxin for the reduction of sulfinic acid when formed on the catalytic cysteine of peroxiredoxins, and the methionine sulfoxide reductases (Msr) for the reduction of methionine sulfoxide within proteins (Petropoulos and Friguet 2005; Lourenço dos Santos et al. 2018).
The Msr system is found in almost all organisms, from bacteria to mammals, and is composed of two enzyme families, MsrA and MsrB, that catalytically reverse the oxidation of the S-sulfoxide and R-sulfoxide diastereoisomeric forms of methionine sulfoxide, respectively (Boschi-Muller et al. 2008). Oxidized methionine sulfoxide reductases are then reduced by the thioredoxin/thioredoxin reductase system. Oxidation of methionine has been implicated in the impairment of protein structure and/or function while the reduction of methionine sulfoxide has been associated with the recovery of protein function. Hence, oxidation/reduction of methionine has been involved in redox regulation of protein-protein interactions and protein function. In combination with protein surface-exposed methionine residues, the Msr system has also been shown to be efficient as a built-in ROS scavenging system preventing further irreversible protein oxidation (Picot et al. 2005; Cabreiro et al. 2008). Since the Msr system can protect proteins from irreversible oxidation and that protein carbonyls levels are usually referred as a marker of oxidative stress in pathophysiological conditions and during aging, the implication of Msr in diseases and in aging process would be expected.
Reduced MsrA activity was found in the brains of Alzheimer’s disease patients (Gabbita et al. 1999) while it has been shown that oxidized proteins accumulate in tissues from patients exhibiting age-related diseases such as neurodegenerative diseases and cataracts (Gil-Mohapel et al. 2014; Swomley and Butterfield 2015). Amyloid ß-peptide (Aβ peptide) Met-35 oxidation is thought to be critical for its aggregation and neurotoxicity (Hou et al. 2002) and the absence of MsrA was shown to modify Aβ solubility properties and to cause mitochondrial dysfunction in a mouse model of Alzheimer’s disease (Moskovitz et al. 2016). Oxidation of α-synuclein methionine residues in Parkinson’s disease is thought to be the main reason of protein fibrillation, hence contributing to the pathology (Glaser et al. 2005). Interestingly, MsrA K.O. mice demonstrated behavioral abnormality (tip-toe walking) consistent with cerebellar dysfunction (Moskovitz et al. 2001), enhanced neurodegeneration with characteristic features of neurodegenerative diseases and increased light scattering, a common cataract symptom (Brennan et al. 2009).
Since the accumulation of oxidatively modified proteins during aging has been largely attributed to declining efficacy of the systems involved in protein homeostasis such as protein degradation and protein repair (Chondrogianni et al. 2014), impairment of the Msr system would be expected to play an important role in the aging phenotype. Indeed, our laboratory has shown that MsrA is down-regulated in aged rats (Petropoulos et al. 2001) and during replicative senescence of fibroblasts (Picot et al. 2004). Both cytosolic and mitochondrial Msr activities were found to decline upon replicative senescence (Ahmed et al. 2010) and increased MetO levels were also reported in membrane proteins of senescent erythrocytes (Brovelli et al. 1990).
The implication of the Msr system in regulating lifespan has been addressed in several studies but it remains controversial. This hypothesis has been originally tested using two different models: MsrA K.O. mice and MsrA overexpressing Drosophila (Moskovitz et al. 2001; Ruan et al. 2002). Overexpression of MsrA in Drosophila resulted in a 70% extension in their healthy lifespan (Ruan et al. 2002) while the knockout of the MsrA gene in mice reduced its lifespan by 40% (Moskovitz et al. 2001). In both studies, the MsrA-dependent lifespan modulation was related to its role in protection against oxidative stress but another study has since shown that the lack of MsrA in mice does not diminish lifespan although it does increase sensitivity to oxidative stress (Salmon et al. 2009).
Overall these studies indicate the importance of Msr system in aging and neurodegenerative diseases pointing out to its role as an antioxidant enzyme protecting cells and organisms from the deleterious effects of oxidative stress. However, the recent discovery that alternation between methionine oxidation and reduction could serve as regulator of protein function raises the interesting hypothesis that the role of the Msr system in aging and survival could also come from these intracellular signaling functions (Lourenço dos Santos et al. 2018).
Non-repairable alterations, which represent the majority of protein damage, are eliminated through protein degradation by the proteasomal or the lysosomal systems in the cytosol while oxidized proteins are degraded by the Lon protease in the mitochondrion (Ugarte et al. 2010; Hamon et al. 2015). These proteolytic systems have been documented to decline with age and during replicative senescence, hence implicating protein maintenance failure in the age-associated build-up of damaged proteins (Baraibar and Friguet 2012; Hamon et al. 2015).
The proteasomal and lysosomal pathways are the two main proteolytic cytosolic machineries by which intracellular proteins are degraded. Protein degradation by the proteasome is a key process for the maintenance of cellular protein homeostasis. In the cytosol and in the nucleus, the proteasome plays a key role in the removal of altered proteins since mildly oxidized proteins are good substrates of the proteasome in vitro and do not to require ubiquitin and ATP to be eliminated in vivo (Chondrogianni et al. 2014). However, some studies have shown that the ubiquitin-proteasome system could be implicated in the degradation of certain oxidized proteins. The increased susceptibility of oxidized proteins to degradation by the proteasome has been attributed to an increased exposure of hydrophobic amino acids at the protein surface and an increased flexibility of their C- and N-terminus extremity, making them more prone to degradation by either the 20S or 26S proteasomes (Grune et al. 2003). However, when proteins are highly oxidized or modified by glycation or conjugation by lipid peroxidation products, intra- and/or inter-molecular cross-links are formed that render these heavily modified proteins resistant to proteolysis by the proteasome (Friguet and Szweda 1997). Moreover, although proteins modified by either glycoxidation or conjugation with lipid peroxidation products, have been evidenced as ubiquitinated, suggesting that they might be substrates of the 26S proteasome, these modified proteins have also been found to be targeted to and degraded by the lysosomes (Bulteau et al. 2001; Marques et al. 2004).
Several studies have indicated that proteasome function is impaired during aging suggesting that its decreased functionality might be causally related to aging and age-associated diseases (Shringarpure and Davies 2002; Farout and Friguet 2006; Breusing and Grune 2008; Baraibar and Friguet 2012), although other studies have shown that this decline may not be universal (Cook et al. 2009; Altun et al. 2010). Pioneering studies from our group and that of Walter Ward showed that rat liver proteasome proteolytic activity is altered during aging (Conconi et al. 1996; Shibatani and Ward 1996; Shibatani et al. 1996). A decrease in proteasome peptidase activity has been since reported in aged tissues of other mammals (mouse, rat and bovine), like liver (Hayashi and Goto 1998), spinal cord (Keller et al. 2000), lens (Shang et al. 2001), heart (Bulteau et al. 2002) and retina (Louie et al. 2002). Furthermore, an age-related decline of proteasome activity has been also shown ex vivo in human lymphocytes (Ponnapan et al. 1999; Carrard et al. 2003), keratinocytes and fibroblasts (Petropoulos et al. 2000; Hwang et al. 2007) and in human primary cell cultures undergoing replicative senescence (Merker and Grune 2000; Sitte et al. 2000; Chondrogianni et al. 2003). Impairment of proteasomal activity has also been reported during aging in model organisms such as Drosophila melanogaster (Vernace et al. 2007; Tonoki et al. 2009) and Caenorhabditis elegans (Hamer et al. 2010).
Proteasome impairment has been reported at different levels, including decreased transcription of certain proteasomal subunits in mice (Huber et al. 2009), dissociation of the proteasome complex in Drosophila (Vernace et al. 2007), and reduced proteasome proteolytic capacity in different aged mammalian tissues and organs (Chondrogianni and Gonos 2005; Farout and Friguet 2006). In contrast, centenarians who represent an interesting example of successful aging, and the long-lived naked mole rats were found to exhibit elevated proteasome levels and activity (Chondrogianni et al. 2000; Perez et al. 2009). The main emphasis has long been placed on preventing protein damage (i.e. interest in antioxidants for protecting against oxidative damage) rather than potentiating the mechanisms that normally handle these damaged products. However, several lines of evidence support the idea that the main problem is not the damage per se, but rather how the cell handles this damage. These includes the better understanding of the cellular mechanisms that contribute to protein quality control, the fact that some of the genes coding for the components of these systems have been implicated in lifespan extension, and the growing evidence supporting that failure of protein homeostasis represents an early event in aging (Lopez-Otin et al. 2013). In addition, recent studies have shown that the proteasome can be activated by genetic manipulations as well as by factors that affect either its conformation and stability or the expression of its subunits and the rate of proteasome assembly. Indeed, over expression of the 20S β5 subunit extended the replicative lifespan of cultured human fibroblasts (Chondrogianni et al. 2005) and both the lifespan and healthspan of wild type Caenorhabditis elegans (Chondrogianni et al. 2015). Over expression of 19S Rpn11 prolonged Drosophila melanogaster lifespan (Tonoki et al. 2009) while overexpression of the proteasome related transcription factors Rpn4 and/or Rpn6 enhanced the replicative lifespan and resistance to proteotoxic stress of Saccharomyces cerevisiae (Kruegel et al. 2011; Yao et al. 2015) and of Caenorhabditis elegans , respectively (Vilchez et al. 2012).
5 Oxidative Protein Damage is Restricted to Specific Protein Targets upon Oxidative Stress and During Cellular Senescence
Increased protein oxidative damage during aging is well documented and is believed to play an important role in cellular aging (Berlett and Stadtman 1997). However, the identification of the damaged protein targets has been usually performed approaching only a single type of modification (e.g. carbonylation or conjugation with HNE) in different aging model systems and tissues from aged animals (Kapphahn et al. 2006; Baraibar et al. 2012c; Lourenço dos Santos et al. 2015).
Our previous studies showed an increase in proteins modified with HNE, AGE and carbonylation in senescent WI-38 human embryonic fibroblasts (Ahmed et al. 2007). The identification of proteins targeted by these modifications showed that they represent a restricted set within the total cellular proteome that fall in key functional categories, such as protein quality control, energy metabolism and cellular morphology (Ahmed et al. 2010). Since impairment of these functional pathways has been previously documented in senescent cells, the observed protein modifications may play a role in the development of the senescent phenotype.
The cytoskeletal proteins vimentin, actin and tubulin were found among the proteins identified as HNE-modified. Cytoskeletal proteins are involved in key cellular processes such as cell division, signal transduction, cell motility and protein synthesis. Follow up studies showed several structural changes of the intermediate filament protein vimentin during cellular senescence. Vimentin filaments form thick, long dense bundles in senescent cells while irregular and small fur-like networks in young or early-passage fibroblasts (Ahmed et al. 2010).
Cellular senescence is also accompanied by alterations in energy metabolism. Increased oxidative damage causes impairment of mitochondrial respiration affecting mainly the activity of complexes I, III and IV of the respiratory chain. Almost half of the modified proteins identified upon replicative senescence of human WI-38 fibroblasts were from mitochondria, which indicates a highly oxidative environment within this organelle during cellular aging (Ahmed et al. 2010). In senescent WI-38 fibroblasts, the iron-sulfur subunit of complex I and subunit α of ATP synthase, the subunit 1 of complex III, and FAD subunit of complex II have been identified as increasingly modified by HNE-, AGE-, and carbonylation, respectively. Among the modified proteins identified in senescent fibroblasts, the citric acid cycle enzymes malate dehydrogenase and 2-oxoglutarate dehydrogenase E1 component, glycerol-3-phosphate dehydrogenase, glycerol kinase and glutaminase appear to be specifically targeted by oxidation (Ahmed et al. 2010). These results suggest that modification of proteins responsible for energy metabolism may participate in the impairment of mitochondrial function observed in senescent cells.
Proteins directly linked with the regulation of protein homeostasis, such as protein folding and degradation were also identified as increasingly modified in senescent cells. Proteins with chaperone function, such as Hsc70, calreticulin, endoplasmic reticulum protein ERp29, as well as proteasome subunits linked to a decreased proteasome activity underscore this issue.
More recently, the occurrence of specific carbonylated proteins upon oxidative stress induced premature senescence of WI-38 human fibroblasts has been analyzed and their follow-up identification has been achieved (Le Boulch et al. 2018). Indeed, increased protein oxidative damage has been clearly associated to both cellular and organismal aging and accumulation of oxidatively damaged proteins has been also reported in HDFs upon both replicative and SIPS (Chondrogianni et al. 2003; Debacq-Chainiaux et al. 2005). Only carbonylated proteins were analyzed in SIPS fibroblasts and they were mainly cytosolic, either belonging to the cytoskeleton, involved in redox and energy metabolism or involved in protein maintenance. Interestingly, cytoskeleton, redox and energy metabolism as well as protein maintenance have been all reported to be impaired during cellular aging. Such carbonylated proteins accumulating in SIPS that were previously identified as increasingly modified for replicative senescence of WI-38 fibroblasts include: Actin, Vimentin, Glucose-6-phosphate dehydrogenase, Heat shock cognate 71 kDa protein, Heterogeneous nuclear ribonucleoprotein H, Tryptophanyl-tRNA synthetase and Tubulin. Other carbonylated proteins that were already found in different models of aging and age-related diseases (Baraibar et al. 2012c) include: Vimentin, Tubulin, Actin, Glyceraldehyde-3-phosphate dehydrogenase, Enolase, Pyruvate kinase, Protein disulfide-isomerase, Catalase and Heat shock cognate 71 kDa protein. From these comparisons, a similarity in the proteins targeted by oxidation emerges to a certain extent, suggesting that specific cellular functions might be affected by the build-up of this restricted set of oxidatively modified proteins due to their impaired functionality and/or altered regulation.
We have also previously characterized the proteome changes of adult human muscle stem cells (i.e. satellite cells or myoblasts) in response to oxidative stress (Baraibar et al. 2011). Using a dual proteomic approach, we intended to unravel the mechanism involved in human myoblasts dysfunction upon oxidative stress. Selective proteins either modulated at the expression level or those targeted by oxidation (carbonylated) were identified after a sub-toxic insult of hydrogen peroxide. For this purpose, a 2D gel electrophoresis-based proteomic approach coupled with immunodetection of carbonylated proteins, after their derivatization with DNPH, and identification of the spots of interest by mass spectrometry has been used. Twenty-one protein spots were evidenced, as increasingly carbonylated upon oxidative stress, indicating that only a restricted set of proteins is prone to accumulation upon oxidative stress. Most of the carbonylated proteins identified belong from the cytosol but also proteins from the nucleus, endoplasmic reticulum as well as the plasma membrane were identified. Major functional categories include energy metabolism, cellular assembly, protein synthesis, cell morphology and protein degradation. Modified proteins such as peroxiredoxins, GAPDH and alpha-enolase are involved in the antioxidant response and energy metabolism. Moreover, proteins involved either in protein degradation such as the proteasome regulatory subunit 10B, and in protein synthesis such as elongation factor 2 were found to be carbonylated. Such proteasome subunits carbonylation may explain, at least in part, the decreased proteasome activity observed, suggesting that oxidative stress do not only induce the modification of proteins but also compromise their degradation by affecting proteasome function.
Finally, we have also recently addressed the potential impact of oxidatively modified proteins on the altered metabolism of senescent human satellite cells (Baraibar et al. 2016). By using a modified 2D gels based proteomics approach, we have found that a restricted set of proteins is targeted by carbonylation and modification with advanced glycation/lipid peroxidation end products during the replicative senescence of satellite cells. 22 protein spots were shown to be increasingly carbonylated, 24 were increasingly glycated, and 8 were increasingly modified by HNE during replicative senescence (Baraibar et al. 2016). Each spot was excised from the gel and analyzed by MALDI-TOF/TOF-MS for protein identification. 28 proteins were identified as targets of one, two or the three modifications. Although similar proteomic approaches were used, this restricted sub-set of modified proteins in senescent myoblasts is different to those identified in senescent fibroblasts. Indeed, while an important number of mitochondrial proteins were found to be increasingly modified in senescent fibroblasts (Ahmed et al. 2010), this was not the case for myoblasts where most of the modified proteins are cytosolic. Nevertheless, modified proteins accumulating in senescent myoblasts that were previously identified as increasingly modified for replicative senescence of WI-38 fibroblasts include: Vimentin, Glucose-6-phosphate dehydrogenase, T-complex protein 1 subunit zeta and Heat shock cognate 71 kDa protein. The identified proteins were then analyzed and grouped by metabolic pathways and cellular functions. Major biological functions include carbohydrate metabolism, cellular morphology, migration and proliferation, as well as protein quality control, protein degradation and free radical scavenging. Interestingly, 6 enzymes involved in glycolysis were found to be increasingly modified in the senescent myoblasts: aldolase, triosephosphate isomerase, glyceraldehyde 3-phosphate dehydrogenase, phosphoglycerate mutase, enolase and pyruvate kinase (Baraibar et al. 2016). The oxidative modification of these enzymes that are involved in the glycolytic pathway is concomitant with a decreased glucose oxidation and an increased NAD+/NADH ratio, reflecting a decreased cellular reducing potential. Since the functionality of the mitochondrial respiratory chain was not affected in human myoblasts during replicative senescence, the decreased glucose oxidation we observed is most likely due to an impairment in glycolysis and/or TCA cycle (Baraibar et al. 2016). Oxidative modifications of enzymes involved in glycolysis and the TCA cycle have also been suggested to be an important pathophysiological factor in age-related diseases, such as neurodegenerative diseases (Martinez et al. 2010). Other studies have shown that the inhibition of glycolytic enzymes, such as GAPDH and PGAM by siRNA can induce premature senescence (Kondoh et al. 2005). Furthermore, metabolomic studies were indicative of a metabolic shift leading to an increased mobilization of non-carbohydrate substrates such as branched chain amino acids or long chain fatty acids. Taken together, these results are supportive of a link between oxidative protein modifications and the altered cellular metabolism associated with the senescent phenotype of human myoblasts (Fig. 8.3).
Schematic representation of the impact of the decreased quality of the cellular proteome on protein quality control and energy metabolism during replicative senescence of human satellite cells. Both proteasome and Msr activities were decreased in senescent myoblasts. Moreover, six enzymes involved in glycolysis were found to be increasingly modified in the senescent myoblasts while metabolomic studies were indicative of a metabolic shift leading to an increased mobilization of non-carbohydrate substrates such as branched chain amino acids or long chain fatty acids (Baraibar et al. 2016)
6 Summary—Conclusions
Oxidative damage to the cellular proteome, leading to the formation of carbonyl groups in proteins derives from direct oxidation of several amino acids side chains and through protein adducts formation with lipid peroxidation products and dicarbonyl glycating compounds. Failure of protein maintenance is a major contributor to the age-associated accumulation of damaged proteins that is believed to participate to the age-related decline in cellular function (Baraibar and Friguet 2012; Chondrogianni et al. 2014; Vanhooren et al. 2015). Indeed, all these damaging modifications have been implicated in cellular senescence, aging and age-related diseases (Baraibar and Friguet 2013). In most cases however, the proteins targeted by these deleterious modifications as well as their consequences have not been identified. In this context, quantitative proteomics approaches, including 2D-gel electrophoresis based methods, represent powerful tools for monitoring at the proteome level the extent of protein oxidative and related modifications and for identifying the targeted proteins. Based on 2D-Difference gel electrophoresis (2D-DIGE), we have developed a novel proteomic method called Oxi-DIGE for the detection, quantification and identification of carbonylated proteins with the potential to be used in any protein containing samples (Baraibar et al. 2013). The identification of proteins targeted by carbonylation during cellular senescence, aging and age-related diseases showed that they represent a restricted set within the total cellular proteome that fall in key functional categories, such as energy metabolism, protein quality control and cellular morphology (Ahmed et al. 2010; Baraibar et al. 2012c; Lourenço dos Santos et al. 2015; Baraibar et al. 2016; Le Boulch et al. 2018). Interestingly, cross comparison of these data sets indicates an overlap in the proteins targeted by these modifications to a certain extent. An important outcome is that several enzymes that catalyze intermediate metabolism, have been found as increasingly modified during aging and upon cellular senescence and therefore may represent functionally accurate biomarkers of aging. Finally, these studies underscore the importance of performing proteomic analyses addressing different aspects, such as expression levels and modifications by carbonylation, conjugation with lipid peroxidation products or glycoxidation, to have a broader view of the age-related changes affecting the cellular proteome.
References
Ahmed EK, Picot CR, Bulteau AL, Friguet B (2007) Protein oxidative modifications and replicative senescence of WI-38 human embryonic fibroblasts. Ann N Y Acad Sci 1119:88–96
Ahmed EK, Rogowska-Wrzesinska A, Roepstorff P, Bulteau AL, Friguet B (2010) Protein modification and replicative senescence of WI-38 human embryonic fibroblasts. Aging Cell 9:252–272
Altun M, Besche HC, Overkleeft HS, Piccirillo R, Edelmann MJ, Kessler BM, Goldberg AL, Ulfhake B (2010) Muscle wasting in aged, sarcopenic rats is associated with enhanced activity of the ubiquitin proteasome pathway. J Biol Chem 285:39597–39608
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Avery SV (2011) Molecular targets of oxidative stress. Biochem J 434:201–210
Baraibar MA, Friguet B (2012) Changes of the proteasomal system during the aging process. Prog Mol Biol Transl Sci 109:249–275
Baraibar MA, Friguet B (2013) Oxidative proteome modifications target specific cellular pathways during oxidative stress, cellular senescence and aging. Exp Gerontol 48:620–625
Baraibar MA, Hyzewicz J, Rogowska-Wrzesinska A, Ladouce R, Roepstorff P, Mouly V, Friguet B (2011) Oxidative stress-induced proteome alterations target different cellular pathways in human myoblasts. Free Radic Biol Med 51:1522–1532
Baraibar MA, Ladouce R, Friguet B (2012a) A method for detecting and/or quantifying carbonylated proteins. WO/2012/175519
Baraibar MA, Barbeito AG, Muhoberac BB, Vidal RA (2012b) A mutant light-chain ferritin that causes neurodegeneration has enhanced propensity toward oxidative damage. Free Radic Biol Med 52:1692–1697
Baraibar MA, Liu L, Ahmed EK, Friguet B (2012c) Protein oxidative damage at the crossroads of cellular senescence, ageing, and age-related diseases. Oxid Med Cell Longev. 2012:919832
Baraibar MA, Ladouce R, Friguet B (2013) Proteomic quantification and identification of carbonylated proteins upon oxidative stress and during cellular aging. J Proteomics 92:63–70
Baraibar MA, Hyzewicz J, Rogowska-Wrzesinska A, Bulteau AL, Prip-Buus C, Butler-Browne G, Friguet B (2016) Impaired energy metabolism of senescent muscle satellite cells is associated with oxidative modifications of glycolytic enzymes. Aging (Albany NY) 8:3375–3389
Berlett BS, Stadtman ER (1997) Protein oxidation in aging, disease, and oxidative stress. J Biol Chem 272:20313–20316
Biteau B, Labarre J, Toledano MB (2003) ATP-dependent reduction of cysteine-sulphinic acid by S. cerevisiae sulphiredoxin. Nature 425:980–4
Boschi-Muller S, Gand A, Branlant G (2008) The methionine sulfoxide reductases: Catalysis and substrate specificities. Arch Biochem Biophys 474:266–273
Brennan LA, Lee W, Cowell T, Giblin F, Kantorow M (2009) Deletion of mouse MsrA results in HBO-induced cataract: MsrA repairs mitochondrial cytochrome c. Mol Vis 15:985–999
Breusing N, Grune T (2008) Regulation of proteasome-mediated protein degradation during oxidative stress and aging. Biol Chem 389:203–209
Brovelli A, Seppi C, Castellana AM, De Renzis MR, Blasina A, Balduini C (1990) Oxidative lesion to membrane proteins in senescent erythrocytes. Biomed Biochim Acta 49:S218–S223
Bulteau AL, Verbeke P, Petropoulos I, Chaffotte AF, Friguet B (2001) Proteasome inhibition in glyoxal-treated fibroblasts and resistance of glycated glucose-6-phosphate dehydrogenase to 20 S proteasome degradation in vitro. J Biol Chem 276:45662–45668
Bulteau AL, Szweda LI, Friguet B (2002) Age-dependent declines in proteasome activity in the heart. Arch Biochem Biophys 397:298–304
Butterfield DA, Poon HF, St Clair D, Keller JN, Pierce WM, Klein JB, Markesbery WR (2006) Redox proteomics identification of oxidatively modified hippocampal proteins in mild cognitive impairment: insights into the development of Alzheimer’s disease. Neurobiol Dis 22:223–232
Cabreiro F, Picot CR, Perichon M, Castel J, Friguet B, Petropoulos I (2008) Overexpression of mitochondrial methionine sulfoxide reductase B2 protects leukemia cells from oxidative stress-induced cell death and protein damage. J Biol Chem 283:16673–16681
Carrard G, Dieu M, Raes M, Toussaint O, Friguet B (2003) Impact of ageing on proteasome structure and function in human lymphocytes. Int J Biochem Cell Biol 35:728–739
Chondrogianni N, Gonos ES (2005) Proteasome dysfunction in mammalian aging: steps and factors involved. Exp Gerontol 40:931–938
Chondrogianni N, Petropoulos I, Franceschi C, Friguet B, Gonos ES (2000) Fibroblast cultures from healthy centenarians have an active proteasome. Exp Gerontol 35:721–728
Chondrogianni N, Stratford FL, Trougakos IP, Friguet B, Rivett AJ, Gonos ES (2003) Central role of the proteasome in senescence and survival of human fibroblasts: induction of a senescence-like phenotype upon its inhibition and resistance to stress upon its activation. J Biol Chem 278:28026–28037
Chondrogianni N, Tzavelas C, Pemberton AJ, Nezis IP, Rivett AJ, Gonos ES (2005) Overexpression of proteasome beta5 assembled subunit increases the amount of proteasome and confers ameliorated response to oxidative stress and higher survival rates. J Biol Chem 280:11840–11850
Chondrogianni N, Petropoulos I, Grimm S, Georgila K, Catalgol B, Friguet B, Grune T, Gonos ES (2014) Protein damage, repair and proteolysis. Mol Aspects Med 35:1–71
Chondrogianni N, Georgila K, Kourtis N, Tavernarakis N, Gonos ES (2015) 20S proteasome activation promotes life span extension and resistance to proteotoxicity in Caenorhabditis elegans. FASEB J. 29:611–622
Conconi M, Szweda LI, Levine RL, Stadtman ER, Friguet B (1996) Age-related decline of rat liver multicatalytic proteinase activity and protection from oxidative inactivation by heat-shock protein 90. Arch Biochem Biophys 331:232–234
Cook C, Gass J, Dunmore J, Tong J, Taylor J, Eriksen J, McGowan E, Lewis J, Johnston J, Petrucelli L (2009) Aging is not associated with proteasome impairment in UPS reporter mice. PLoS ONE 4:e5888
Dalle-Donne I, Giustarini D, Colombo R, Rossi R, Milzani A (2003) Protein carbonylation in human diseases. Trends Mol. Med. 9:169–176
Debacq-Chainiaux F, Borlon C, Pascal T, Royer V, Eliaers F, Ninane N, Carrard G, Friguet B, de Longueville F, Boffe S, Remacle J, Toussaint O (2005) Repeated exposure of human skin fibroblasts to UVB at subcytotoxic level triggers premature senescence through the TGF-beta1 signaling pathway. J Cell Sci 118:743–758
Farout L, Friguet B (2006) Proteasome function in aging and oxidative stress: implications in protein maintenance failure. Antioxid Redox Signal 8:205–216
Franceschi C, Bonafe M, Valensin S, Olivieri F, De Luca M, Ottaviani E, De Benedictis G. Inflamm-aging (2000) An evolutionary perspective on immunosenescence. Ann N Y Acad Sci 908:244–254
Friguet B (2006) Oxidized protein degradation and repair in ageing and oxidative stress. FEBS Lett 580:2910–2916
Friguet B, Szweda LI (1997) Inhibition of the multicatalytic proteinase (proteasome) by 4-hydroxy-2-nonenal cross-linked protein. FEBS Lett 405:21–25
Gabbita SP, Aksenov MY, Lovell MA, Markesbery WR (1999) Decrease in peptide methionine sulfoxide reductase in Alzheimer’s disease brain. J Neurochem 73:1660–1666
Gil-Mohapel J, Brocardo PS, Christie BR (2014) The role of oxidative stress in Huntington’s disease: are antioxidants good therapeutic candidates? Curr Drug Targets 15:454–468
Giulivi C, Traaseth NJ, Davies KJ (2003) Tyrosine oxidation products: analysis and biological relevance. Amino Acids 25:227–232
Glaser CB, Yamin G, Uversky VN, Fink AL (2005) Methionine oxidation, alpha-synuclein and Parkinson’s disease. Biochim Biophys Acta 1703:157–169
Grune T, Merker K, Sandig G, Davies KJ (2003) Selective degradation of oxidatively modified protein substrates by the proteasome. Biochem Biophys Res Commun 305:709–718
Hamer G, Matilainen O, Holmberg CI (2010) A photoconvertible reporter of the ubiquitin-proteasome system in vivo. Nat Methods 7:473–478
Hamon MP, Bulteau AL, Friguet B (2015) Mitochondrial proteases and protein quality control in ageing and longevity. Ageing Res Rev 23:56–66
Hayashi T, Goto S (1998) Age-related changes in the 20S and 26S proteasome activities in the liver of male F344 rats. Mech Ageing Dev 102:55–66
Horiuchi S, Araki N (1994) Advanced glycation end products of the Maillard reaction and their relation to aging. Gerontology 40(Suppl 2):10–15
Hou L, Kang I, Marchant RE, Zagorski MG (2002) Methionine 35 oxidation reduces fibril assembly of the amyloid abeta-(1-42) peptide of Alzheimer’s disease. J Biol Chem 277:40173–40176
Huber N, Sakai N, Eismann T, Shin T, Kuboki S, Blanchard J, Schuster R, Edwards MJ, Wong HR, Lentsch AB (2009) Age-related decrease in proteasome expression contributes to defective nuclear factor-kappaB activation during hepatic ischemia/reperfusion. Hepatology 49:1718–1728
Hwang JS, Chang I, Kim S (2007) Age-associated decrease in proteasome content and activities in human dermal fibroblasts: restoration of normal level of proteasome subunits reduces aging markers in fibroblasts from elderly persons. J Gerontol A Biol Sci Med Sci 62:490–9
Kapphahn RJ, Giwa BM, Berg KM, Roehrich H, Feng X, Olsen TW, Ferrington DA (2006) Retinal proteins modified by 4-hydroxynonenal: identification of molecular targets. Exp Eye Res 83:165–175
Keller JN, Huang FF, Markesbery WR (2000) Decreased levels of proteasome activity and proteasome expression in aging spinal cord. Neuroscience 98:149–156
Kondoh H, Lleonart ME, Gil J, Wang J, Degan P, Peters G, Martinez D, Carnero A, Beach D (2005) Glycolytic enzymes can modulate cellular life span. Cancer Res 65:177–185
Kregel KC, Zhang HJ (2007) An integrated view of oxidative stress in aging: basic mechanisms, functional effects, and pathological considerations. Am J Physiol Regul Integr Comp Physiol 292:R18–R36
Kruegel U, Robison B, Dange T, Kahlert G, Delaney JR, Kotireddy S, Tsuchiya M, Tsuchiyama S, Murakami CJ, Schleit J, Sutphin G, Carr D, Tar K, Dittmar G, Kaeberlein M, Kennedy BK, Schmidt M (2011) Elevated proteasome capacity extends replicative lifespan in Saccharomyces cerevisiae. PLoS Genet 7:e1002253
Le Boulch M, Ahmed EK, Rogowska-Wrzesinska A, Baraibar MA, Friguet B (2018) Proteome oxidative carbonylation during oxidative stress-induced premature senescence of WI-38 human fibroblasts. Mech Ageing Dev 170:59–71
Levine RL (2002) Carbonyl modified proteins in cellular regulation, aging, and disease. Free Radic Biol Med 32:790–796
López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G (2013) The hallmarks of aging Cell 153:1194–217
Louie JL, Kapphahn RJ, Ferrington DA (2002) Proteasome function and protein oxidation in the aged retina. Exp Eye Res 75:271–284
Lourenço dos Santos S, Baraibar MA, Lundberg S, Eeg-Olofsson O, Larsson L, Friguet B (2015) Oxidative proteome alterations during skeletal muscle ageing. Redox Biol 5:267–74
Lourenço dos Santos S, Petropoulos I, Friguet B (2018) The oxidized protein repair enzymes methionine sulfoxide reductases and their roles in protecting against oxidative stress, in ageing and in regulating protein function. Antioxidants (Basel) 7(12). pii: E191
Marques C, Pereira P, Taylor A, Liang JN, Reddy VN, Szweda LI, Shang F (2004) Ubiquitin-dependent lysosomal degradation of the HNE-modified proteins in lens epithelial cells. FASEB J 18:1424–1426
Martinez A, Portero-Otin M, Pamplona R, Ferrer I (2010) Protein targets of oxidative damage in human neurodegenerative diseases with abnormal protein aggregates. Brain Pathol 20:281–297
Mecocci P, Fano G, Fulle S, MacGarvey U, Shinobu L, Polidori MC, Cherubini A, Vecchiet J, Senin U, Beal MF (1999) Age-dependent increases in oxidative damage to DNA, lipids, and proteins in human skeletal muscle. Free Radic Biol Med 26:303–308
Merker K, Grune T (2000) Proteolysis of oxidised proteins and cellular senescence. Exp Gerontol 35:779–786
Moskovitz J, Bar-Noy S, Williams WM, Requena J, Berlett BS, Stadtman ER (2001) Methionine sulfoxide reductase (MsrA) is a regulator of antioxidant defense and lifespan in mammals. Proc Natl Acad Sci U S A 98:12920–5
Moskovitz J, Du F, Bowman CF, Yan SS (2016) Methionine sulfoxide reductase a affects beta-amyloid solubility and mitochondrial function in a mouse model of Alzheimer’s disease. Am J Physiol Endocrinol Metab 310:E388–E393
Perez VI, Buffenstein R, Masamsetti V, Leonard S, Salmon AB, Mele J, Andziak B, Yang T, Edrey Y, Friguet B, Ward W, Richardson A, Chaudhuri A (2009) Protein stability and resistance to oxidative stress are determinants of longevity in the longest-living rodent, the naked mole-rat. Proc Natl Acad Sci U S A 106:3059–64
Petropoulos I, Friguet B (2005) Protein maintenance in aging and replicative senescence: a role for the peptide methionine sulfoxide reductases. Biochim Biophys Acta 1703:261–266
Petropoulos I, Conconi M, Wang X. Hoenel B, Bregegere F, Milner Y, Friguet B (2000) Increase of oxidatively modified protein is associated with a decrease of proteasome activity and content in aging epidermal cells. J Gerontol A Biol Sci Med Sci 55:B220–7
Petropoulos I, Mary J, Perichon M, Friguet B (2001) Rat peptide methionine sulphoxide reductase: cloning of the cDNA, and down-regulation of gene expression and enzyme activity during aging. Biochem J 355:819–825
Picot CR, Perichon M, Cintrat J-C, Friguet B, Petropoulos I (2004) The peptide methionine sulfoxide reductases, MsrA and MsrB (hCBS-1), are downregulated during replicative senescence of human WI-38 fibroblasts. FEBS Lett 558:74–78
Picot CR, Petropoulos I, Perichon M, Moreau M, Nizard C, Friguet B (2005) Overexpression of MsrA protects WI-38 SV40 human fibroblasts against H2O2-mediated oxidative stress. Free Radic Biol Med 39:1332–1341
Picot CR, Moreau M, Juan M, Noblesse E, Nizard C, Petropoulos I, Friguet B (2007) Impairment of methionine sulfoxide reductase during UV irradiation and photoaging. Exp Gerontol 42:859–863
Ponnappan U, Zhong M, Trebilcock GU (1999) Decreased proteasome-mediated degradation in T cells from the elderly: a role in immune senescence. Cell Immunol 192:167–174
Ruan H, Tang XD, Chen ML, Joiner ML, Sun G, Brot N, Weissbach H, Heinemann SH, Iverson L, Wu CF, Hoshi T (2002) High-quality life extension by the enzyme peptide methionine sulfoxide reductase. Proc Natl Acad Sci U S A 99:2748–2753
Salmon AB, Perez VI, Bokov A, Jernigan A, Kim G, Zhao H, Levine RL, Richardson A (2009) Lack of methionine sulfoxide reductase A in mice increases sensitivity to oxidative stress but does not diminish life span. FASEB J 23:3601–3608
Sayre LM, Zelasko DA, Harris PL, Perry G, Salomon RG, Smith MA (1997) 4-Hydroxynonenal-derived advanced lipid peroxidation end products are increased in Alzheimer’s disease. J Neurochem 68:2092–2097
Shibatani T, Ward WF (1996) Effect of age and food restriction on alkaline protease activity in rat liver. J Gerontol A Biol Sci Med Sci 51:B175–B178
Shibatani T, Nazir M, Ward WF (1996) Alteration of rat liver 20S proteasome activities by age and food restriction. J Gerontol A Biol Sci Med Sci 51:B316–22
Shringarpure R, Davies KJ (2002) Protein turnover by the proteasome in aging and disease. Free Radic Biol Med 32:1084–1089
Sitte N, Merker K, von Zglinicki T, Grune T (2000) Protein oxidation and degradation during proliferative senescence of human MRC-5 fibroblasts. Free Radic Biol Med 28:701–708
Stadtman ER (2006) Protein oxidation and aging. Free Radic Res. 40:1250–1258
Stadtman ER, Levine RL (2003) Free radical-mediated oxidation of free amino acids and amino acid residues in proteins. Amino Acids 25:207–218
Swomley AM, Butterfield DA (2015) Oxidative stress in Alzheimer disease and mild cognitive impairment: evidence from human data provided by redox proteomics. Arch Toxicol 89:1669–1680
Szweda PA, Camouse M, Lundberg KC, Oberley TD, Szweda LI (2003) Aging, lipofuscin formation, and free radical-mediated inhibition of cellular proteolytic systems. Ageing Res Rev 2:383–405
Tamarit J, de Hoogh A, Obis E, Alsina D, Cabiscol E, Ros J (2012) Analysis of oxidative stress-induced protein carbonylation using fluorescent hydrazides. J Proteomics 75:3778–3788
Tonoki A, Kuranaga E, Tomioka T, Hamazaki J, Murata S, Tanaka K, Miura M (2009) Genetic evidence linking age-dependent attenuation of the 26S proteasome with the aging process. Mol Cell Biol 29:1095–1106
Ugarte N, Petropoulos I, Friguet B (2010) Oxidized mitochondrial protein degradation and repair in aging and oxidative stress. Antioxid Redox Signal 13:539–549
Vanhooren V, Navarrete Santos A, Voutetakis K, Petropoulos I, Libert C, Simm A, Gonos ES, Friguet B (2015) Protein modification and maintenance systems as biomarkers of ageing. Mech Ageing Dev 151:71–84
Vernace VA, Arnaud L, Schmidt-Glenewinkel T, Figueiredo-Pereira ME (2007) Aging perturbs 26S proteasome assembly in Drosophila melanogaster. FASEB J. 21:2672–2682
Vilchez D, Morantte I, Liu Z, Douglas PM, Merkwirth C, Rodrigues AP, Manning G, Dillin A (2012) RPN-6 determines C. elegans longevity under proteotoxic stress conditions. Nature 489:263–268
Yao Y, Tsuchiyama S, Yang C, Bulteau AL, He C, Robison B, Tsuchiya M, Miller D, Briones V, Tar K, Potrero A, Friguet B, Kennedy BK, Schmidt M (2015) Proteasomes, Sir2, and Hxk2 form an interconnected aging network that impinges on the AMPK/Snf1-regulated transcriptional repressor. Mig1.PLoS Genet 11:e1004968
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Friguet, B., Baraibar, M.A. (2019). Oxidatively Modified Proteins and Maintenance Systems as Biomarkers of Aging. In: Moskalev, A. (eds) Biomarkers of Human Aging. Healthy Ageing and Longevity, vol 10. Springer, Cham. https://doi.org/10.1007/978-3-030-24970-0_8
Download citation
DOI: https://doi.org/10.1007/978-3-030-24970-0_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-24969-4
Online ISBN: 978-3-030-24970-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)