Abstract
Machine learning is as growing as fast as concepts such as Big data and the field of data science in general. The purpose of the systematic review was to analyze scholarly articles that were published between 2015 and 2018 addressing or implementing supervised and unsupervised machine learning techniques in different problem-solving paradigms. Using the elements of PRISMA, the review process identified 84 scholarly articles that had been published in different journals. Of the 84 articles, 6 were published before 2015 despite their metadata indicating that they were published in 2015. The existence of the six articles in the final papers was attributed to errors in indexing. Nonetheless, from the reviewed papers, decision tree, support vector machine, and Naïve Bayes algorithms appeared to be the most cited, discussed, and implemented supervised learners. Conversely, k-means, hierarchical clustering, and principal component analysis also emerged as the commonly used unsupervised learners. The review also revealed other commonly used algorithms that include ensembles and reinforce learners, and future systematic reviews can focus on them because of the developments that machine learning and data science is undergoing at the moment.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Supervised machine learning
- Unsupervised machine learning
- Machine learning
- Systematic review
- Data science
- Supervised and unsupervised learners
1 Introduction
The demand for advanced data analytics leading to the use of machine learning and other emerging techniques can be attributed to the advent and subsequent development of technologies such as Big Data, Business Intelligence, and the applications that require automation. As Sandhu [1] explains, machine learning is a subset of artificial intelligence, which uses computerized techniques to solve problems based on historical data and information without unnecessarily requiring modification in the core process. Essentially, artificial intelligence involves creation of algorithms and other computation techniques that promote smartness of machines. It encompasses algorithms that think, act, and implement tasks using protocols that are otherwise beyond human’s reach.
Machine learning is a component of artificial intelligence although it endeavors to solve problems based on historical or previous examples [2]. Unlike artificial intelligence applications, machine learning involves learning of hidden patterns within the data (data mining) and subsequently using the patterns to classify or predict an event related to the problem [3]. Simply, intelligent machines depend on knowledge to sustain their functionalities and machine learning offers such a knowledge. In essence, machine learning algorithms are embedded into machines and data streams provided so that knowledge and information are extracted and fed into the system for faster and efficient management of processes. It suffices to mention that all machine learning algorithms are also artificial intelligence techniques although not all artificial intelligence methods qualify as machine learning algorithms.
Machine learning algorithms can either be supervised or unsupervised although some authors also classify other algorithms as reinforcement, because such techniques learn data and identify pattern for the purposes of reacting to an environment. However, most articles recognize supervised and unsupervised machine learning algorithms. The difference between these two main classes is the existence of labels in the training data subset. According to Kotsiantis [4], supervised machine learning involves predetermined output attribute besides the use of input attributes. The algorithms attempt to predict and classify the predetermined attribute, and their accuracies and misclassification alongside other performance measures is dependent on the counts of the predetermined attribute correctly predicted or classified or otherwise. It is also important to note the learning process stops when the algorithm achieves an acceptable level of performance [5]. According to Libbrecht and Noble [2], technically, supervised algorithms perform analytical tasks first using the training data and subsequently construct contingent functions for mapping new instance of the attribute. As stated previously, the algorithms require prespecifications of maximum settings for the desired outcome and performance levels [2, 5]. Given the approach used in machine learning, it has been observed that training subset of about 66% is rationale and helps in achieving the desired result without demanding for more computational time [6]. The supervised learning algorithms are further classified into classification and regression algorithms [3, 4].
Conversely, unsupervised data learning involves pattern recognition without the involvement of a target attribute. That is, all the variables used in the analysis are used as inputs and because of the approach, the techniques are suitable for clustering and association mining techniques. According to Hofmann [7], unsupervised learning algorithms are suitable for creating the labels in the data that are subsequently used to implement supervised learning tasks. That is, unsupervised clustering algorithms identify inherent groupings within the unlabeled data and subsequently assign label to each data value [8, 9]. On the other hand, unsupervised association mining algorithms tend to identify rules that accurately represent relationships between attributes.
1.1 Motivation and Scope
Even though both supervised and unsupervised algorithms are widely used to accomplish different data mining tasks, the discussion of the algorithms has been mostly done singly or grouped depending on the need of learning tasks. More importantly, literature reviews that have been conducted to account for supervised and unsupervised algorithms either handle supervised techniques or unsupervised ones with limited focus on both approaches in the same. For instance, Sandhu [1] wrote a review article on machine learning and natural language processing but focused on supervised machine learning. The author did not conduct a systematic review and, as such, the article does not focus on any specific period or target any given database. Baharudin et al. [10] also conducted a literature review on machine learning techniques though in the context of text data mining and did not implement any known systematic review methodology. Praveena [11] also conducted a review of papers that had implemented supervised learning algorithms and, as such, did implement any of the known systematic review approaches. However, Qazi et al. [12] conducted a systematic review although with a focus on the challenges that different authors encountered while implementing different classification techniques in sentimental analysis. The authors reviewed 24 papers that were published between 2002 and 2014 and concluded that most review articles published during the period focused on eight standard machine learning classification techniques for sentimental analysis along with other concept learning algorithms. Unlike these reviews, the systematic review here conducted focused on all major stand-alone machine learning algorithms, both supervised and unsupervised published during the 2015–2018 period.
1.2 Novelty and Review Approach
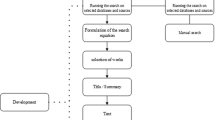
The systematic review relied on Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) tool to review studies that have used different supervised and unsupervised learning algorithms to address different issues [13]. The approach used in the search was such that different papers published between 2013 and 2018 dealing with the use of machine learning algorithms as methods of data analysis were included. The identification and subsequent inclusion and exclusion of the articles reviewed was based on whether the paper is peer-reviewed, scholarly, full-text, and year of publication that ranges between 2015 and 2018 [13,14,15]. The search was conducted on EBSCO and ProQuest Central Databases. The search queries used are as follows, and they were implemented in the two databases. In conventional PRISMA review, it is a requirement to check and identify the search criteria in the title and the structure of the abstract alongside introduction (rationale and objectives) and methods including information sources, data items, summary measures, and synthesis results [16]. However, such an approach was adopted, and applied to published articles instead of being implemented on review articles. Table 1.1 summarizes the search queries that were run in the two databases.
The inclusion criteria deferred for both databases with EBSCO relying on date of publication and full-text to narrow the search, while ProQuest Central search filters included Abstract (AB), Document Text (FT), Document Title (TI), and Publication Title (PUB). An instance of search implemented in ProQuest Central with some of the above criteria is as shown below.
ft(Supervised machine learning) AND ft(Unsupervised machine learning) OR ti(Supervised machine learning) AND ti(Unsupervised machine learning) OR pub(Supervised machine learning) AND pub(Unsupervised machine learning)
2 Search Results
The search and screening results based on PRISMA and elements of meta-analysis are presented in the following section. The major steps used to arrive at the final articles and subsequent analysis included screening (rapid title screening), full test screening, data extraction including extraction of the characteristics of the study, and meta-analysis based on specific check lists and aspects of the machine learning algorithm used.
2.1 EBSCO and ProQuest Central Database Results
The search results obtained from the two databases before the commencement of the review process were as follows. The EBSCO search identified 144 articles that were published between 2015 and 2018. Of the 144 documents, 74 had complete information including name of authors, date of publication, name of journal, and structured abstracts. However, only 9 of the 74 articles had full-text and, as such, selected for inclusion in the review process. As for the search results from ProQuest Central, the initial search yielded over 19,898 results, but application of the filters reduced 3301 articles, of which 42 were reviews and 682 covered classification techniques, while 643 covered or had information related to algorithms in general. However, the subject alignment of the research papers was not considered because of the wide spectrum of application of the algorithms such that both supervised and unsupervised methods were also applied in other subjects. The distribution the search result based on top ten journals is as shown in Fig. 1.1.
Figure 1.1 shows that PloS One had the highest number of articles published on supervised and unsupervised machine learning. Sensors and Scientific Reports (Nature Publisher Group) had 213 and 210 articles. Multimedia Tools and Applications (172), Remote Sensing (150), and International Journal of Computer Vision (124) had over 100 articles. Even though Mathematics Problems in Engineering and Internal Computer Vision had 61 and 58 articles, the two publications were better placed at exploring the mathematical and algorithmic aspects of supervised and unsupervised machine learning algorithms. The inclusion and exclusion criteria focused on the algorithms as well as their mathematical discourse and application in different fields.
Based on the PRISMA checklist, a total of 84 articles were included in the study and their content analyzed for the implementation of supervised and unsupervised machine learning techniques.
The final number of articles used in the review is 84, although 20 of them underwent meta-analysis when each study was vetted for clarity of the objectives and study questions. Regarding study questions and the effectiveness of the approached used to implement the chosen machine learning algorithms resulted in exclusion of 1290 articles (Fig. 1.2). The rest (1985) met the required study question criteria but also screened for the comprehensiveness of the literature search, data abstraction, evaluation of the results, and the applicability of results [17,18,19]. It is imperative to note that publication bias and disclosure of funding sources were not considered as part of the screen process. The 84 articles met these meta-analysis requirements and were subsequently included in the analysis (Fig. 1.2).
It is crucial to note that of the 84 articles that were included in the study, 3 were published in 2013 and 3 were published in 2014 but were not filtered out by the data of publication restriction.
2.2 Distribution of Included Articles
The articles used in the study consisted of Feature, Journal Articles, General Information, Periodical, and Review types with a distribution represented in the following chart.
From Fig. 1.3, 78 articles were published between 2015 and 2018, while the missing articles were published in 2013 [20,21,22] and 2014 [23,24,25] and their inclusion can be associated to publication biasness, which is also observed in the type of documents or study. According to the search, inclusion, and inclusion criteria, the final results ought to have only journal articles, but others were features, general information, periodicals, and reviews. The six papers that were published between 2013 and 2014 were included, because they met all the criteria required for meta-analysis and the indexed meta-data showed that the papers were published in 2015. Regarding the misinformation, we can deduce that the publications had an inaccuracy of about 7.2%.
3 Discussion
The 84 articles discussed different supervised and unsupervised machine learning techniques without necessarily making the distinction. According to Praveena [11], supervised learning requires an assistance born out of experience or acquired patterns within the data and, in most cases, involves a defined output variable [26,27,28,29,30]. The input dataset is segregated into train and test subsets, and several papers address the concept of training datasets based on the desired outcome [31,32,33,34]. All the algorithms that use supervised learning approach acquire patterns within the training dataset and subsequently apply them to the test subset with the object of either predicting or classifying an attribute [35,36,37]. Most of the authors described the workflow of a supervised machine learning and, as it also emerged from the review, decision tree, Naïve Bayes, and Support Vector Machines are the most commonly used algorithms [8, 38,39,40,41,42].
3.1 Decision Tree
It is important to recall that supervised learning can either be based on a classification or regression algorithm, and decision tree algorithm can be used as both although it is mainly used for classification as noted in these articles [20, 43,44,45]. The algorithm emulates a tree, and it sorts attributes through groupings based on data values [46]. Just like a conventional tree, the algorithm has branches and nodes with nodes representing variable group for classification and branches, assuming the values that the attribute can take as part of the class [47, 48]. The pseudocode illustrating the decision tree algorithm is as shown below. In the algorithm, D is the dataset, while x and y are the input and target variables, respectively [49, 50].
Algorithm 1.1: Decision Tree
Protocol DT Inducer (D, x, y)
-
1.
T = Tree Growing (D, x, y)
-
2.
Return Tree Pruning (D, T)
Method Tree Growing (D, x, y)
-
1.
Create a tree T
-
2.
if at least one of the Stopping Criteria is satisfied then;
-
3.
label the root node as a leaf with the most frequent value of y in D as the correct class.
-
4.
else;
-
5.
Establish a discrete function f(x) of the input variable so that splitting D according to the functions outcomes produces the best splitting metric
-
6.
if the best metric is greater or equal to the threshold then;
-
7.
Mark the root node in T as f(x)
-
8.
for each outcome of f(x) at the node do;
-
9.
\( \boldsymbol{Subtree}=\boldsymbol{Tree}\ \boldsymbol{Growing}\ \left({\boldsymbol{\delta}}_{\boldsymbol{f}\left(\boldsymbol{x}\right)={\boldsymbol{t}}_{\mathbf{1}}},\boldsymbol{D},\boldsymbol{x},\boldsymbol{y}\right) \)
-
10.
Connect the root of T to Subtree and label the edge t 1
-
11.
end for
-
12.
else
-
13.
Label the root node T for a leaf with the frequent value of y in D as the assigned class
-
14.
end if
-
15.
end if
-
16.
Return T
Protocol Tree Pruning (D, T, y)
-
1.
repeat
-
2.
Select a node t in T to maximally improve pruning evaluation procedure
-
3.
if t ≠ 0 then;
-
4.
T = pruned (T, t)
-
5.
end if
-
6.
until t = 0
-
7.
Return T
As illustrated in the pseudocode, Decision Tree achieves classification in three distinct steps. Firstly, the algorithm induces both tree growing and tree pruning functionalities [51]. Secondly, it grows the tree by assigning each data value to a class based on the value of the target variable that is the most common one at the instance of iteration [52, 53]. The final step deals with pruning the grown tree to optimize the performance of the resultant model [19, 53, 54]. Most of the reviewed studies involved application of decision trees for different applications, although most involved classification cancer and lung cancer studies, clinical medicine especially diagnosis of conditions based on historical data as well as some rare forms of artificial intelligence applications [40, 52, 55,56,57]. Most of the studies have also recognized decision tree algorithms to be more accurate when dealing with data generated using the same collection procedures [43, 44, 52].
3.2 Naïve Bayes
The Naïve Bayes algorithm has gained its fame because of its background on Bayesian probability theorem. In most texts, it is considered a semisupervised method, because it can be used either in clustering or classification tasks [58, 59]. When implemented as a technique for creating clusters, Naïve Bayes does not require specification of an outcome and it uses conditional probability to assign data values to classes and, as such, is a form of unsupervised learning [47, 60,61,62]. However, when used to classify data, Naïve Bayes requires both input and target variables and, as such, is a supervised learning technique [55, 63, 64]. As a classifier, the algorithm creates Bayesian networks, which are tree generated based on the condition probability of an occurrence of an outcome based on probabilities imposed on it by the input variables [65, 66]. The pseudocode for the Naïve Bayes algorithm is presented below [49, 67, 68].
Algorithm 1.2: Naïve Bayes Learner
Input: training set T s, Hold-out set H s, initial components, I c, and convergence thresholds ρ EM and ρ add
Initial M using one component
I ← I c.
repeat
Add I components to M thereby initializing M using random components drawn from the training set T s
Remove the I initialization instances from T s
repeat
E-step: Proportionally assign examples in T s to resultant mixture component using M
M-Step: Calculate maximum likelihood parameters using the input data.
if log P (H s/M) is the best maximum probability, then save M in M best
every 5 cycles of the two steps, prune low-weight components of M
until P (H s/M) fails to increase by the ratio ρ EM
M←M best
Prune low weight components of M
I ← 2 I.
until P (H s/M) fails to increase by the ratio ρ add
Execute both E: step and M: step twice on M best using examples from H s and T s
Return M←M best
As the pseudocode illustrates, Naïve Bayes algorithm relies on Bayes’ theorem represented mathematical below to assign independent variables to classes based on probability [31, 58].
In Eq. (1.1), the probability of H when the probability of D is known is defined in terms of the product probability of H, probability of D given the probability of H divided by the probability of D. The H and D are events with defined outcome and they can represent Heads and Tails in coil tossing experiments [12, 45, 69, 70]. The extension of the theorem in supervised learning is of the form represented in Eq. (1.2).
In the above equation, x i, … , x n represents the input attribute, for which conditional probabilities are computed based on the known probabilities of the target variables in the training dataset [71,72,73]. The algorithm has been discussed in different contexts and its application is mainly attributed to the creation of data labels for subsequent unsupervised learning verifications [16, 74, 75].
3.3 Support Vector Machine
The support vector machines (SVMs) algorithm was also common among the search results articles. The articles that explored the applications of SVM did so with the objective of evaluating its performance in different scenarios [30, 58, 73, 76]. All the applications of SVM are included toward classification and the tenet of the algorithm is computation of margins [53, 77, 78]. Simply, SVM draws margins as boundary between the classes in the provided dataset. Its principle is to create the margins such that the distance between each class and the nearest margin is maximized and in effect leading to the minimum possible classification error [68, 78,79,80]. The margins are defined as the distance between two supporting vectors separated by a hyperplane. The pseudocode for the SVM algorithm is as demonstrated below. The algorithm assumes that the data are linearly separable so that the weight associated with support vectors can be drawn easily and the margin computed [62, 70]. The assumption makes regularization possible [49, 81].
Algorithm 1.3: Support Vector Machine
Input: S, λ, T, k
Initialize: Choose w 1 such that \( \left\Vert {\boldsymbol{w}}_{\mathbf{1}}\right\Vert \boldsymbol{\le}\sqrt{\boldsymbol{\lambda}} \)
FOR t = 1 , 2 … ,T
Select A t ⊆ S, in which |A t| = k
Set \( {\boldsymbol{A}}_{\boldsymbol{t}}^{+}=\left\{\left(\boldsymbol{x},\boldsymbol{y}\right)\boldsymbol{\in}{\boldsymbol{A}}_{\boldsymbol{t}}:\boldsymbol{y}\left({\boldsymbol{w}}_{\boldsymbol{t}},\boldsymbol{x}\right)<\mathbf{1}\right\} \)
Set \( {\boldsymbol{\delta}}_{\boldsymbol{t}}=\frac{\mathbf{1}}{\boldsymbol{\lambda} \boldsymbol{t}} \)
Set \( {\boldsymbol{w}}_{\boldsymbol{t}+\mathbf{0.5}}=\left(\mathbf{1}-{\boldsymbol{\delta}}_{\boldsymbol{t}}\boldsymbol{\lambda} \right){\boldsymbol{w}}_{\boldsymbol{t}}+\frac{{\boldsymbol{\delta}}_{\boldsymbol{t}}}{\boldsymbol{k}}\sum_{\left(\boldsymbol{x},\boldsymbol{y}\right)\boldsymbol{\in}{\boldsymbol{A}}_{\boldsymbol{t}}^{+}}\boldsymbol{yx} \)
Set \( {\boldsymbol{w}}_{\boldsymbol{t}+\mathbf{1}}=\left\{\mathbf{1},\frac{\mathbf{1}/\sqrt{\boldsymbol{\lambda}}}{\left\Vert {\boldsymbol{w}}_{\boldsymbol{t}+\mathbf{0.5}}\right\Vert}\right\}{\boldsymbol{w}}_{\boldsymbol{t}+\mathbf{0.5}} \)
Output: w T + 1
The implementation of the algorithm and its accuracy is dependent on its ability to margin violations and subsequent misclassification of classes on either side of the vectors. The margin is based on the following set of equations:
In Eq. (1.3), the three sets of equation describe the hyperplane separating two linear support vectors W T x + b = 1 and W T x + b = − 1, and all the classes within the two support vectors are classified accurately, while those outside the support vectors violate the margin [25, 81, 82]. Consequently, the larger the distance between the support vectors, the higher the chances that points are correctly classified.
As for unsupervised learning algorithms, most of the studies either discussed, cited, or implemented k-means, hierarchical clustering, and principal component analysis, among others [20, 55, 73, 83, 84]. Unlike supervised learning, unsupervised learning extract limited features from the data, and it relies on previously learned patterns to recognize likely classes within the dataset [85, 86]. As a result, unsupervised learning is suitable for feature reduction in case of large dataset and clustering tasks that lead to the creation of new classes in unlabeled data [80, 87, 88]. It entails selection and importation of data into appropriate framework followed by selection of an appropriate algorithm, specification of thresholds, review of the model, and subsequent optimization to produce desired outcome [89, 90]. Of the many unsupervised learners, k-means was widely discussed among the authors and as such was also previewed in the review.
3.4 k-Means Algorithms
The algorithm has been used in different studies to create groups or classes in unlabeled datasets based on the mean distance between classes [91, 92]. The technique initiates and originates the classes or labels that are subsequently used in other prospective analysis [69]. A pseudocode for the k-means algorithm is as shown in the illustration below [15, 61].
Algorithm 1.4: k-Means Learner
Function k-means ()
Initialize k prototypes (w 1…, w k) so that the weighted distance between the clusters becomes w j = i l ,j ∈ {1, … , k},l ∈ {1, … , n}
Associate each cluster C j with the prototype weight w j
Repeat
for each input vector i l;,l ∈ {1, … , n}
do
Assign i l to cluster C j∗ with the nearest w j∗
for each cluster C j∗ : j ∈ {1, … , k}, do;
Update the prototype w j to be centroid of the sample
observations in the current C j∗; \( {\boldsymbol{w}}_{\boldsymbol{j}}=\sum_{{\boldsymbol{i}}_{\boldsymbol{l}\boldsymbol{\in }{\boldsymbol{c}}_{\boldsymbol{j}}}}{\boldsymbol{i}}_{\boldsymbol{l}}/\left|{\boldsymbol{C}}_{\boldsymbol{j}}\right| \)
Calculate the error function
until E becomes constant or does not change significantly.
The pseudocode demonstrates the process of assigning data values to classes based on their proximity to the nearest mean with the least error function [93,94,95,96]. The error function is computed as the difference between the mean and the assigned cluster mean [97, 98].
3.5 Semisupervised and Other Learners
Even though the search was focused and narrowed down to supervised and unsupervised learning techniques, it emerged that research preferred using different methods for the purposes of comparing the results and verification of the classification and prediction accuracy of the machine learning models [75, 99, 100]. Some of the studies used supervised and unsupervised machine learning approaches alongside reinforcement learning techniques such as generative models, self-training algorithms, and transudative SVM [101,102,103]. Other studies focused on ensemble learning algorithms such as boosting and bagging, while other studies defined different perceptions related to neural networks. [59, 66, 104,105,106,107]. Finally, some of the studies addressed algorithms such as k-Nearest Neighbor as an instance-based learning but could not categorize it as either supervised or unsupervised machine learning algorithm because of the limitations of the applications [41, 108,109,110].
4 Conclusion and Future Work
Even though the search results yielded over 3300 qualified papers, the filtering processes based on title screening, abstract screening, full text screening, and data extraction coupled with meta-analysis reduced the number of articles to 84. Despite the narrowing the search results to supervised and unsupervised machine learning as key search words, the results contained articles that addressed reinforced learners and ensembled learners among other techniques that review did not focus. The trend is understandable, because machine learning and data science is evolving and most of the algorithms are undergoing improvements, hence the emergence of categories such as reinforced and ensembled learner. Hence, future systematic review prospect should focus on these emerging aggregations of learners and assess through research progress based on authorship, regions, and applications to identify the major driving forces behind the growth.
References
Sandhu, T. H. (2018). Machine learning and natural language processing—A review. International Journal of Advanced Research in Computer Science, 9(2), 582–584.
Libbrecht, M. W., & Noble, W. S. (2015). Machine learning applications in genetics and genomics. Nature Reviews Genetics, 16(6), 321–332.
Alpaydın, E. (2014). Introduction to machine learning. Cambridge, MA: MIT Press.
Kotsiantis, S. B. (2007). Supervised machine learning: A review of classification techniques. Informatica, 31, 249–268.
MathWorks. (2016). Applying supervised learning. Machine Learning with MATLAB.
Ng, A. (2012). 1. Supervised learning. Machine Learning, 1–30.
Hofmann, T. (2001). Unsupervised learning by probabilistic latent semantic analysis. Machine Learning, 42, 177–196.
Dougherty, J., Kohavi, R., & Sahami, M. (1995). Supervised and unsupervised discretization of continuous features. In Machine Learning Proceedings.
Marshland, S. (2015). Machine learning: An algorithm perspective. Boca Raton, FL: CRC Press.
Baharudin, B., Lee, L. H., & Khan, K. (2010). A review of machine learning algorithms for text-documents classification. Journal on Advance in Information Technology, 1(1), 4–20.
Praveena, M. (2017). A literature review on supervised machine learning algorithms and boosting process. International Journal of Computer Applications, 169(8), 975–8887.
Qazi, A., Raj, R. G., Hardaker, G., & Standing, C. (2017). A systematic literature review on opinion types and sentiment analysis techniques: Tasks and challenges. Internet Research, 27(3), 608–630.
Hutton, B., et al. (2015). The PRISMA extension statement for reporting of systematic reviews incorporating network meta-analyses of health care interventions: Checklist and explanations. Annals of Internal Medicine, 163(7), 566–567.
Zorzela, L., Loke, Y. K., Ioannidis, J. P., Golder, S., Santaguida, P., Altman, D. G., et al. (2016). PRISMA harms checklist: Improving harms reporting in systematic reviews. BMJ (Online), 352, i157.
Shamseer, L., et al. (2015). Preferred reporting items for systematic review and meta-analysis protocols (prisma-p) 2015: Elaboration and explanation. BMJ (Online), 349, g7647.
Moher, D., et al. (2015). Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015 statement. Systematic Reviews, 4, 1.
Stroup, D. F., et al. (2000). Meta-analysis of observational studies in epidemiology: A proposal for reporting. Meta-analysis Of Observational Studies in Epidemiology (MOOSE) group. JAMA, 283(15), 2008–2012.
Bloch, M. H., Landeros-Weisenberger, A., Rosario, M. C., Pittenger, C., & Leckman, J. F. (2008). Meta-analysis of the symptom structure of obsessive-compulsive disorder. The American Journal of Psychiatry, 165(12), 1532–1542.
Fujimoto, M. S., Suvorov, A., Jensen, N. O., Clement, M. J., & Bybee, S. M. (2016). Detecting false positive sequence homology: A machine learning approach. BMC Bioinformatics, 17, 101.
Mani, S., et al. (2013). Machine learning for predicting the response of breast cancer to neoadjuvant chemotherapy. Journal of the American Medical Informatics Association, 20(4), 688–695.
Kovačević, A., Dehghan, A., Filannino, M., Keane, J. A., & Nenadic, G. (2013). Combining rules and machine learning for extraction of temporal expressions and events from clinical narratives. Journal of the American Medical Informatics Association, 20(5), 859–866.
Klann, J. G., Anand, V., & Downs, S. M. (2013). Patient-tailored prioritization for a pediatric care decision support system through machine learning. Journal of the American Medical Informatics Association, 20(e2), e267–e274.
Gultepe, E., Green, J. P., Nguyen, H., Adams, J., Albertson, T., & Tagkopoulos, I. (2014). From vital signs to clinical outcomes for patients with sepsis: A machine learning basis for a clinical decision support system. Journal of the American Medical Informatics Association, 21(2), 315–325.
Mani, S., et al. (2014). Medical decision support using machine learning for early detection of late-onset neonatal sepsis. Journal of the American Medical Informatics Association, 21(2), 326–336.
Nguyen, D. H. M., & Patrick, J. D. (2014). Supervised machine learning and active learning in classification of radiology reports. Journal of the American Medical Informatics Association, 21(5), 893–901.
Deo, R. C. (2015). Machine learning in medicine HHS public access. Circulation, 132(20), 1920–1930.
Mullainathan, S., & Spiess, J. (2017). Machine learning: An applied econometric approach. The Journal of Economic Perspectives, 31(2), 87–106.
Wu, M.-J., et al. (2017). Identification and individualized prediction of clinical phenotypes in bipolar disorders using neurocognitive data, neuroimaging scans and machine learning. NeuroImage, 145, 254–264.
Oudah, M., & Henschel, A. (2018). Taxonomy-aware feature engineering for microbiome classification. BMC Bioinformatics, 19, 227.
Palma, S. I. C. J., Traguedo, A. P., Porteira, A. R., Frias, M. J., Gamboa, H., & Roque, A. C. A. (2018). Machine learning for the meta-analyses of microbial pathogens’ volatile signatures. Scientific Reports, 8, 1–15.
Jaspers, S., De Troyer, E., & Aerts, M. (2018). Machine learning techniques for the automation of literature reviews and systematic reviews in EFSA. EFSA Supporting Publications, 15(6), 1427E.
Crawford, M., Khoshgoftaar, T. M., Prusa, J. D., Richter, A. N., & Al Najada, H. (2015). Survey of review spam detection using machine learning techniques. Journal of Big Data, 2(1), 1–24.
Dinov, I. D. (2016). Methodological challenges and analytic opportunities for modeling and interpreting Big Healthcare Data. Gigascience, 5, 12.
Dimou, A., Vahdati, S., Di Iorio, A., Lange, C., Verborgh, R., & Mannens, E. (2017). Challenges as enablers for high quality Linked Data: Insights from the Semantic Publishing Challenge. PeerJ Computer Science, 3, e105.
Trilling, D., & Boumans, J. (2018). Automatische inhoudsanalyse van Nederlandstalige data. Tijdschrift voor Communicatiewetenschap, 46(1), 5–24.
Van Nieuwenburg, E. P. L., Liu, Y., & Huber, S. D. (2017). Learning phase transitions by confusion. Nature Physics, 13(5), 435–439.
Hoyt, R., Linnville, S., Thaler, S., & Moore, J. (2016). Digital family history data mining with neural networks: A pilot study. Perspectives in Health Information Management, 13, 1c.
Dobson, J. E. (2015). Can an algorithm be disturbed? Machine learning, intrinsic criticism, and the digital humanities. College Literature, 42(4), 543–564.
Downing, N. S., et al. (2017). Describing the performance of U.S. hospitals by applying big data analytics. PLoS One, 12(6), e0179603.
Hoang, X. D., & Nguyen, Q. C. (2018). Botnet detection based on machine learning techniques using DNS query data. Future Internet, 10(5), 43.
Kothari, U. C., & Momayez, M. (2018). Machine learning: A novel approach to predicting slope instabilities. International Journal of Geophysics, 2018, 9.
Thompson, J. A., Tan, J., & Greene, C. S. (2016). Cross-platform normalization of microarray and RNA-seq data for machine learning applications. PeerJ, 4, e1621.
Ahmed, M. U., & Mahmood, A. (2018). An empirical study of machine learning algorithms to predict students’ grades. Pakistan Journal of Science, 70(1), 91–96.
Carifio, J., Halverson, J., Krioukov, D., & Nelson, B. D. (2017). Machine learning in the string landscape. Journal of High Energy Physics, 2017(9), 1–36.
Choudhari, P., & Dhari, S. V. (2017). Sentiment analysis and machine learning based sentiment classification: A review. International Journal of Advanced Research in Computer Science, 8(3).
Lloyd, S., Garnerone, S., & Zanardi, P. (2016). Quantum algorithms for topological and geometric analysis of data. Nature Communications, 7, 10138.
Pavithra, D., & Jayanthi, A. N. (2018). A study on machine learning algorithm in medical diagnosis. International Journal of Advanced Research in Computer Science, 9(4), 42–46.
Krittanawong, C., Zhang, H., Wang, Z., Aydar, M., & Kitai, T. (2017). Artificial intelligence in precision cardiovascular medicine. Journal of the American College of Cardiology, 69(21), 2657–2664.
Kaytan, M., & Aydilek, I. B. (2017). A review on machine learning tools. 2017 International Artificial Intelligence and Data Processing Symposium, 8(3), 1–4.
Lynch, C. M., van Berkel, V. H., & Frieboes, H. B. (2017). Application of unsupervised analysis techniques to lung cancer patient data. PLoS One, 12(9), e0184370.
Beck, D., Pfaendtner, J., Carothers, J., & Subramanian, V. (2017). Data science for chemical engineers. Chemical Engineering Progress, 113(2), 21–26.
Heylman, C., Datta, R., Sobrino, A., George, S., & Gratton, E. (2015). Supervised machine learning for classification of the electrophysiological effects of chronotropic drugs on human induced pluripotent stem cell-derived cardiomyocytes. PLoS One, 10(12), e0144572.
Torkzaban, B., et al. (2015). Machine learning based classification of microsatellite variation: An effective approach for Phylogeographic characterization of olive populations. PLoS One, 10(11), e0143465.
Guo, Z., Shao, X., Xu, Y., Miyazaki, H., Ohira, W., & Shibasaki, R. (2016). Identification of village building via Google earth images and supervised machine learning methods. Remote Sensing, 8(4), 271.
Xia, C., Fu, L., Liu, Z., Liu, H., Chen, L., & Liu, Y. (2018). Aquatic toxic analysis by monitoring fish behavior using computer vision: A recent progress. Journal of Toxicology, 2018, 11.
Fuller, D., Buote, R., & Stanley, K. (2017). A glossary for big data in population and public health: Discussion and commentary on terminology and research methods. Journal of Epidemiology and Community Health, 71(11), 1113.
Gibson, D., & de Freitas, S. (2016). Exploratory analysis in learning analytics. Technology, Knowledge and Learning, 21(1), 5–19.
Cuperlovic-Culf, M. (2018). Machine learning methods for analysis of metabolic data and metabolic pathway modeling. Metabolites, 8(1), 4.
Tan, M. S., Chang, S.-W., Cheah, P. L., & Yap, H. J. (2018). Integrative machine learning analysis of multiple gene expression profiles in cervical cancer. PeerJ, 6, e5285.
Meenakshi, K., Safa, M., Karthick, T., & Sivaranjani, N. (2017). A novel study of machine learning algorithms for classifying health care data. Research Journal of Pharmacy and Technology, 10(5), 1429–1432.
Dey, A. (2016). Machine learning algorithms: A review. International Journal of Computer Science and Information Technology, 7(3), 1174–1179.
Zhao, C., Wang, S., & Li, D. (2016). Determining fuzzy membership for sentiment classification: A three-layer sentiment propagation model. PLoS One, 11(11), e0165560.
Mossotto, E., Ashton, J. J., Coelho, T., Beattie, R. M., MacArthur, B. D., & Ennis, S. (2017). Classification of paediatric inflammatory bowel disease using machine learning. Scientific Reports, 7, 1–10.
Lau, O., & Yohai, I. (2016). Using quantitative methods in industry. Political Science and Politics, 49(3), 524–526.
Qiu, J., Wu, Q., Ding, G., Xu, Y., & Feng, S. (2016). A survey of machine learning for big data processing. EURASIP Journal on Advances in Signal Processing, 2016, 1–16.
Parreco, J. P., Hidalgo, A. E., Badilla, A. D., Ilyas, O., & Rattan, R. (2018). Predicting central line-associated bloodstream infections and mortality using supervised machine learning. Journal of Critical Care, 45, 156–162.
Wuest, T., Irgens, C., & Thoben, K.-D. (2016). Changing states of multistage process chains. Journal of Engineering, 2016, 1.
Tarwani, N. (2017). Survey of cyberbulling detection on social media big-data. International Journal of Advanced Research in Computer Science, 8(5).
Martinelli, E., Mencattini, A., Daprati, E., & Di Natale, C. (2016). Strength is in numbers: Can concordant artificial listeners improve prediction of emotion from speech? PLoS One, 11(8), e0161752.
Liu, N., & Zhao, J. (2016). Semi-supervised online multiple kernel learning algorithm for big data. TELKOMNIKA, 14(2), 638–646.
Goh, K. L., & Singh, A. K. (2015). Comprehensive literature review on machine learning structures for Web spam classification. Procedia Computer Science, 70, 434–441.
Mishra, C., & Gupta, D. L. (2017). Deep machine learning and neural networks: An overview. IAES International Journal of Artificial Intelligence, 6(2), 66–73.
Yan, X., Bai, Y., Fang, S., & Luo, J. (2016). A kernel-free quadratic surface support vector machine for semi-supervised learning. The Journal of the Operational Research Society, 67(7), 1001–1011.
Yared, R., & Abdulrazak, B. (2016). Ambient technology to assist elderly people in indoor risks. Computers, 5(4), 22.
Osborne, J. D., et al. (2016). Efficient identification of nationally mandated reportable cancer cases using natural language processing and machine learning. Journal of the American Medical Informatics Association, 83(5), 605–623.
Kolog, E. A., Montero, C. S., & Tukiainen, M. (2018). Development and evaluation of an automated e-counselling system for emotion and sentiment analysis. Electronic Journal of Information Systems Evaluation, 21(1), 1–19.
Rafiei, M. H., Khushefati, W. H., Demirboga, R., & Adeli, H. (2017). Supervised deep restricted Boltzmann machine for estimation of concrete. ACI Materials Journal, 114(2), 237–244.
Almasre, M. A., & Al-Nuaim, H. (2017). Comparison of four SVM classifiers used with depth sensors to recognize Arabic sign language words. Computers, 6(2), 20.
Hashem, K. (2018). The rise and fall of machine learning methods in biomedical research. F1000Research, 6, 2012.
Torshin, I. Y., & Rudakov, K. V. (2015). On the theoretical basis of metric analysis of poorly formalized problems of recognition and classification. Pattern Recognition and Image Analysis, 25(4), 577–587.
Petrelli, M., & Perugini, D. (2016). Solving petrological problems through machine learning: The study case of tectonic discrimination using geochemical and isotopic data. Contributions to Mineralogy and Petrology, 171(10), 1–15.
Min-Joo, K., & Kang, J.-W. (2016). Intrusion detection system using deep neural network for in-vehicle network security. PLoS One, 11(6). https://doi.org/10.1371/journal.pone.0155781
Alicante, A., Corazza, A., Isgrò, F., & Silvestri, S. (2016). Unsupervised entity and relation extraction from clinical records in Italian. Computers in Biology and Medicine, 72, 263–275.
Shanmugasundaram, G., & Sankarikaarguzhali, G. (2017). An investigation on IoT healthcare analytics. International Journal of Information Engineering and Electronic Business, 9(2), 11.
Huang, G., Song, S., Gupta, J. N. D., & Wu, C. (2014). Semi-supervised and unsupervised extreme learning machines. IEEE Transactions on Cybernetics, 44(12), 2405–2417.
Rastogi, R., & Saigal, P. (2017). Tree-based localized fuzzy twin support vector clustering with square loss function. Applied Intelligence, 47(1), 96–113.
Muscoloni, A., Thomas, J. M., Ciucci, S., Bianconi, G., & Cannistraci, C. V. (2017). Machine learning meets complex networks via coalescent embedding in the hyperbolic space. Nature Communications, 8, 1–19.
Saeys, Y., Van Gassen, S., & Lambrecht, B. N. (2016). Computational flow cytometry: Helping to make sense of high-dimensional immunology data. Nature Reviews. Immunology, 16(7), 449–462.
Gonzalez, A., Pierre, & Forsberg, F. (2017). Unsupervised machine learning: An investigation of clustering algorithms on a small dataset (pp. 1–39).
Necula, S.-C. (2017). Deep learning for distribution channels’ management. Informatica Economică, 21(4), 73–85.
Munther, A., Razif, R., AbuAlhaj, M., Anbar, M., & Nizam, S. (2016). A preliminary performance evaluation of K-means, KNN and em unsupervised machine learning methods for network flow classification. International Journal of Electrical and Computer Engineering, 6(2), 778–784.
Alalousi, A., Razif, R., Abualhaj, M., Anbar, M., & Nizam, S. (2016). A preliminary performance evaluation of K-means, KNN and EM unsupervised machine learning methods for network flow classification. International Journal of Electrical and Computer Engineering, 6(2), 778–784.
Alanazi, H. O., Abdullah, A. H., & Qureshi, K. N. (2017). A critical review for developing accurate and dynamic predictive models using machine learning methods in medicine and health care. Journal of Medical Systems, 41(4), 1–10.
Almatarneh, S., & Gamallo, P. (2018). A lexicon based method to search for extreme opinions. PLoS One, 13(5), e0197816.
Assem, H., Xu, L., Buda, T. S., & O’sullivan, D. (2016). Machine learning as a service for enabling Internet of things and people. Personal and Ubiquitous Computing, 20(6), 899–914.
Azim, M. A., & Bhuiyan, M. H. (2018). Text to emotion extraction using supervised machine learning techniques. TELKOMNIKA, 16(3), 1394–1401.
Sirbu, A. (2016). Dynamic machine learning for supervised and unsupervised classification ES. Machine Learning.
Wahyudin, I., Djatna, T., & Kusuma, W. A. (2016). Cluster analysis for SME risk analysis documents based on pillar K-means. TELKOMNIKA, 14(2), 674.
Davis, S. E., Lasko, T. A., Chen, G., Siew, E. D., & Matheny, M. E. (2018). Calibration drift in regression and machine learning models for acute kidney injury. Journal of the American Medical Informatics Association, 24, 1052–1061.
Wallace, B. C., Noel-Storr, A., Marshall, I. J., Cohen, A. M., Smalheiser, N. R., & Thomas, J. (2017). Identifying reports of randomized controlled trials (RCTs) via a hybrid machine learning and crowdsourcing approach. Journal of the American Medical Informatics Association, 24(6), 1165–1168.
Biamonte, J., Wittek, P., Pancotti, N., Rebentrost, P., Wiebe, N., & Lloyd, S. (2017). Quantum machine learning. Nature, 549(7671), 195–202.
Bisaso, K. R., Anguzu, G. T., Karungi, S. A., Kiragga, A., & Castelnuovo, B. (2017). A survey of machine learning applications in HIV clinical research and care. Computers in Biology and Medicine, 91, 366–371.
Bauder, R., Khoshgoftaar, T. M., & Seliya, N. (2017). A survey on the state of healthcare upcoding fraud analysis and detection. Health Services and Outcomes Research Methodology, 17(1), 31–55.
Bashiri, A., Ghazisaeedi, M., Safdari, R., Shahmoradi, L., & Ehtesham, H. (2017). Improving the prediction of survival in cancer patients by using machine learning techniques: Experience of gene expression data: A narrative review. Iranian Journal of Public Health, 46(2), 165–172.
Breckels, L. M., Mulvey, C. M., Lilley, K. S., & Gatto, L. (2018). A bioconductor workflow for processing and analysing spatial proteomics data. F1000Research, 5, 2926.
Saad, S. M., et al. (2017). Pollutant recognition based on supervised machine learning for indoor air quality monitoring systems. Applied Sciences, 7(8), 823.
Fiorini, L., Cavallo, F., Dario, P., Eavis, A., & Caleb-Solly, P. (2017). Unsupervised machine learning for developing personalised behaviour models using activity data. Sensors, 17(5), 1034.
Bunn, J. K., Hu, J., & Hattrick-Simpers, J. R. (2016). Semi-supervised approach to phase identification from combinatorial sample diffraction patterns. JOM, 68(8), 2116–2125.
Cárdenas-López, F. A., Lamata, L., Retamal, J. C., & Solano, E. (2018). Multiqubit and multilevel quantum reinforcement learning with quantum technologies. PLoS One, 13(7), e0200455.
Chen, R., Niu, W., Zhang, X., Zhuo, Z., & Lv, F. (2017). An effective conversation-based botnet detection method. Mathematical Problems in Engineering, 2017, 4934082.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Alloghani, M., Al-Jumeily, D., Mustafina, J., Hussain, A., Aljaaf, A.J. (2020). A Systematic Review on Supervised and Unsupervised Machine Learning Algorithms for Data Science. In: Berry, M., Mohamed, A., Yap, B. (eds) Supervised and Unsupervised Learning for Data Science . Unsupervised and Semi-Supervised Learning. Springer, Cham. https://doi.org/10.1007/978-3-030-22475-2_1
Download citation
DOI: https://doi.org/10.1007/978-3-030-22475-2_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-22474-5
Online ISBN: 978-3-030-22475-2
eBook Packages: EngineeringEngineering (R0)