Abstract
Helicobacter pylori exhibit remarkable survival even in the vulnerable environments such as acidic, peristalsis, phagocytosis and oxidative stress. These stresses on the pathogen in the host induce damage of DNA in the pathogen. H. pylori acquired the ability to survive DNA damage by transformation-mediated recombination DNA repair. This repair mechanism helps the pathogen in successfully infecting the host. While many pathogens are competent for transformation only in certain environmental conditions such as starvation, H. pylori is competent throughout the growth. H. pylori may acquire the genetic material from the surrounding environment and contribute to evolution and genetic diversity. The mechanism in acquiring genetic material is ‘horizontal gene transfer’, the major contributing factor in the development of bacterial diversity. Horizontal gene transfer may help the pathogen H. pylori in acquiring antigenic determinants, genes of antibiotic resistance and virulence factors from other organisms to alter and influence pathogenicity. In this chapter, we review and discuss the association between horizontal gene transfer and adaptation of gastric human pathogen H. pylori to the host.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Antibiotics resistance
- Evolution
- Horizontal gene transfer
- H. pylori
- Macro-diversity
- Multidrug resistance
- Nickel-binding proteins
- Nickel transporter genes
1 Introduction
Helicobacter pylori was discovered in human stomach, dental plaque, oral lesions, saliva, tonsil and adenoid tissue. H. pylori was known for causing gastrointestinal disorders like gastritis, ulcers and gastric cancer (Neelapu et al. 2014; Neelapu 2018). Sometimes H. pylori may trigger some other diseases like otitis, sinusitis, phyrangitis, laryngitis and glossitis (Kurtaran et al. 2008). Microorganisms survive in nature either as individuals or in a community known as biofilm (Challa et al. 2018). H. pylori uses biofilm lifestyle to survive in unfavourable environmental conditions such as pH, antibiotics, immune defences, disinfectants, nutritional changes and high temperatures (Challa and Neelapu 2018). Biofilm provides a strong platform for interaction and communication among the individuals present in the colony (Mohana Sheela et al. 2018; Neelapu et al. 2018). Till date research to prevent bacterial infections involved identification of drug targets, drugs (Neelapu et al. 2013, 2015, 2016; Neelapu and Pavani 2013; Nammi et al. 2016, 2017), vaccines (Pasupuleti et al. 2017) and antibiofilm agents (Challa and Neelapu 2018). This review discusses how bacterium H. pylori acquire traits via horizontal gene transfer (HGT) and adapt to the particular niche.
2 Role of HGT and Mechanisms of H. pylori Adaptation to the Host
The “selective pressures on the invading H. pylori bacteria would expose it to environment (e.g., exposure to antibiotics and changes in gastric pH or mucosal defences) and host factors (e.g., specific and nonspecific defence mechanisms) (Ferrero and Jenks 2001). These pressures are harmful damaging DNA of H. pylori and sometimes may also prevent the colonization of H. pylori strain (O’Rourke et al. 2003). H. pylori are competent enough to pick DNA from the surroundings either from other H. pylori strains, or from other bacteria in the gut of the host (via HGT) or sometimes even in from the host (Fig. 1). Then H. pylori use acquired transformation-mediated recombination DNA repair for successful infection of the pathogen” (Dorer et al. 2010). This transformation is helping the bacteria to adapt itself in the gastric niche of the host (Schuster et al. 2008). Literature also reports changes in the genomic material of H. pylori when transmitted between individuals of the host. Burst of mutations will be induced when exposed to selective pressures mentioned above (O’Rourke et al. 2003). These bacteria (H. pylori) harbour genes which are affected and/or not mutated changing the surface components of bacteria (Linz et al. 2013). This becomes disadvantage to the pathogen, where it is indirectly recognized by the host. During evolution some of the genes will be deleted and some genes will be imported via HGT from the already adapted bacteria which are coexisting in the gut of the host altering the surface components (Linz et al. 2013). This importation helps the bacteria to shape its genome and adapt to the host of the genome (Schuster et al. 2008; Eppinger et al. 2006). This demonstrates the role of HGT in shaping the genome of bacteria to adapt it to the new host.
H. pylori are competent enough to pick DNA from the surroundings either from other H. pylori strains or from other bacteria (Campylobacter jejuni) in the gut of the host (via HGT) or sometimes even from the host (Source: Fernandez-Gonzalez and Backert 2014)
HGT, the “key evolutionary force”, transferred genetic material between genomes and thereby shape the genome of bacteria. This helped many bacteria to gain genes and selectively provided advantages to the bacteria (Fernandez-Gonzalez and Backert 2014; Garcia-Aljaro et al. 2017). Literature reports that adaptation of H. pylori to the gastric niche (Vinella et al. 2015; Fischer et al. 2016), micro- and macrodiversity in H. pylori (Alm et al. 1999; Hofreuter and Haas 2002) and antibiotic resistance in H. pylori (Von Wintersdorff et al. 2016; Lood et al. 2017) are due to HGT. This section discusses in detail (a) HGT of nickel-binding proteins, nickel transporter genes and their role; (b) macrodiversity in H. pylori and HGT; and (c) antibiotic resistance in H. pylori and HGT. This section further discusses how HGT has shaped the genome of H. pylori in due course of evolution.
2.1 HGT of Nickel-Binding Proteins, Nickel Transporters Genes and Their Role
H. pylori utilizes specific enzymes or Ni proteins like urease and [NiFe] hydrogenase for colonization of gastric tract in humans (Fischer et al. 2016). The pH in the stomach is acidic and urease (Ni protein) of H. pylori helps in changing/converting the acidic pH in the stomach to neutral pH. Urease needs a cofactor nickel to convert urea into CO2 and NH3 (Neelapu et al. 2014; Fischer et al. 2016). These compounds are used by the bacteria to maintain the pH in the bacterium cytoplasm near to neutral. [NiFe] hydrogenase (Ni protein) is another enzyme where a bacterium utilizes molecular hydrogen as a source of energy (Fischer et al. 2016). Nickel is scarcely or meagrely available in the human body. So, H. pylori requires nickel transporter genes for acquisition of nickel and colonization of H. pylori. Thus, “…acquisition of nickel transporters and Ni-binding proteins by gastric Helicobacter species was a key event for the emergence of one of the most successful bacterial pathogens, H. pylori…” (Vinella et al. 2015; Fischer et al. 2016). Transporters (NixA, NiuBDE, NikABCDE and NikZOppBCDE), Ni-dependent enzymes (urease, hydrogenase) or Ni-binding proteins (Hpn and Hpn-2) were reported in all Helicobacter species (Vinella et al. 2015; Fischer et al. 2016).
2.1.1 HGT of Nickel-Binding Proteins Histidine-Rich Proteins
Histidine-rich proteins, Hpn and Hpn-2, are known to protect gastric Helicobacter species against nickel overload. They also accumulate intracellular nickel and store this nickel indirectly helping them to colonize the stomach of the host. Vinella et al. (2015) revealed that histidine-rich proteins (Hpn) emerged in the last common ancestor (LCA) of gastric Helicobacter species. Hpn and hpn-2 genes are specific to the gastric Helicobacter species and are not in enterohepatic species (Vinella et al. 2015). Hpn plays a major role in the protection of H. pylori against nickel overload and participates in the accumulation of intracellular nickel storage, while Hpn-2 is not required for both these functions (Fig. 2) (Vinella et al. 2015). Hpn interacts with the UreA urease subunit, while Hpn and Hpn-2 interact with the HypAB hydrogenase maturation proteins (Fig. 2) (Vinella et al. 2015). Hpn and Hpn-2 are essential for colonization of gastric Helicobacter species in the host stomach (Vinella et al. 2015). Vinella et al. (2015) proved that hpn and hpn mutants of H. pylori were not able to colonize the stomach in the mouse model, whereas hpn and hpn mutants of H. pylori when complemented with wild genes were able to establish and colonize in the mouse model (Fig. 2). This allowed the Helicobacter gastric species to thrive in the stomach by protecting them against nickel overload, participating in the accumulation of intracellular nickel storage and colonization of the host stomach. Thus acquisition of Ni-binding proteins (Hpn and Hpn-2) via HGT followed by a “…decisive evolutionary event allowed the bacteria to adapt the human stomach a niche that no other bacterium colonized and helped in the emergence of Helicobacter species … .”
Role of Hpn and Hpn-2 in nickel trafficking, protection against nickel overload, urease activity and colonization of the host stomach (Source: Vinella et al. 2015)
2.1.2 HGT of Nickel Transporters Genes
Emergence of Ni-binding proteins (Hpn and Hpn-2) in gastric Helicobacter species was further supported by HGT of nickel transporter genes NixA and NiuBDE. Gastric Helicobacter species were able to pick up nickel-binding proteins and nickel transporter genes via HGT and adapted itself to the gastric niche. Fischer et al. (2016) revealed that LCA of gastric Helicobacter species and H. pylori acquired Ni-binding proteins and nickel transporter genes via HGT to survive in the stomach (Fig. 3). The successful acquisition of nickel transporters genes NixA and NiuBDE via HGT allowed the bacteria to utilize nickel transporter genes for urease activity (nickel-dependent urease activity) by a decisive evolutionary event. This evolutionary event can be considered as a significant change in the genome of gastric Helicobacter species allowing the bacteria to adapt the human stomach a niche that no other bacterium colonized and helped in the emergence of Helicobacter species.
Distribution, phylogeny and evolutionary history on acquisition of nickel transporter genes by gastric Helicobacter species (Source: Fischer et al. 2016)
The key role of nickel transporter genes and Ni-binding proteins shows that nickel plays a very important role in the colonization of H. pylori. Campanale et al. (2014) carried out a pilot study by supplementing H. pylori-infected patients with the nickel-free diet for 1 month and found that the nickel-free diet was able to enhance the efficiency of eradication therapy. This study recommends nickel-free diets for those patients who are infected with H. pylori, and further clinical trial studies are also required to prove the safety of the diet.
2.2 Macrodiversity in H. pylori and HGT
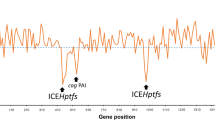
Macrodiversity between H. pylori strains is due to intragenomic rearrangements like deletion, inversion, or translocation (Alm et al. 1999). H. pylori possess insertion sequences like IS605 and IS606 and several plasticity zones in strains like Hp26695 and HpJ99. Plasticity zones are not limited to these H. pylori strains, but were also present and reported in other strains of H. pylori. These plasticity zones differ in GC content when compared to chromosomal GC content. For example, Hp26695 contains five plasticity zones with GC contents of 33% (zone 1), 35% (zone 2), 33% (zone 3), 43% (zone 4) and 33% (zone 5), which differ from the chromosomal GC content of 39% (Tomb et al. 1997). These plasticity zones in H. pylori might have been received via horizontal gene transfer. Conjugation and natural transformation are the mechanisms of HGT identified in H. pylori.
Nedenskov-Sorensen et al. (1990) first described the natural transformation in H. pylori, and several genes were identified in H. pylori which are acquired via transformation process (Schmitt et al. 1995; Hofreuter et al. 1998; Ando et al. 1999; Smeets et al. 2000). Natural transformation in H. pylori is mediated by type IV secretion system (Hofreuter et al. 2001). H. pylori encodes four T4SSs including cagPAI (mediates injection of CagA protein and induces proinflammatory signalling), comB (system involved in the uptake of DNA from the environment) and tfs3 and tfs4 genes (role not yet known). H. pylori also contain diverse genetic modules “…due to the modular structure, plasmids might either pick up chromosomal genes of H. pylori or integrate sequence modules from foreign plasmids, which are taken up by the bacteria during its natural transformation competence (gene shuffling) leading to macrodiversity among H. pylori strains and rapid generation of substrains (Hofreuter and Haas 2002)…”.
2.3 Antibiotic Resistance in H. pylori and HGT
H. pylori has developed antibiotic resistance to proton-pump inhibitors, clarithromycin, metronidazole, macrolide, amoxicillin, levofloxacin, etc., or combinations of them (Savarino et al. 1997; Bardhan et al. 2001; Torres et al. 2001; Osaki et al. 2006; Zullo et al. 2007; Ndip et al. 2008; Boyanova et al. 2009; Gao et al. 2010; Sun et al. 2010; Wüppenhorst et al. 2011; Bolor-Erdene et al. 2017; Lee et al. 2018). Multidrug resistance (MDR) or antibiotic resistance in H. pylori can be eradicated by identifying new or alternative drug targets, developing new drug combinations (Neelapu et al. 2013, 2015, 2016; Neelapu and Pavani 2013; Nammi et al. 2016, 2017; Pasupuleti et al. 2017) and using Chinese herbs (Huang et al. 2015). The new drug combinations developed for H. pylori in view of MDR are levofloxacin or moxifloxacin (novel class of fluoroquinolones) with amoxicillin, rifabutin and furazolidone. The Chinese herbs, namely, emodin, baicalin, schizandrin and berberine, can also be used to treat MDR in H. pylori (Huang et al. 2015).
Literature reports interspecies and intraspecies gene transfer of metronidazole and clarithromycin resistance between Helicobacter species (Table 1). Pot et al. (2001) proved interspecies transfer of antibiotic resistance genes between H. pylori and Helicobacter acinonychis. To prove these Kusters and group demonstrated that “…H. acinonychis is competent for natural transformation and H. pylori can acquire antibiotic resistance by uptake of DNA (HGT) from other Helicobacter species and vice versa… .” (Pot et al. 2001). Pot et al. (2001) isolated DNA from H. acinonychis isolate NCTC12686 (NCTC12686 MtzR) and H. acinonychis isolate Sheeba (Sheeba MtzR) metronidazole-resistant strains. This isolated DNA was used for natural transformation of two metronidazole-sensitive H. pylori as per the standard protocol of Wang et al. (1993). Upon transformation metronidazole-resistant transformants were obtained for both H. pylori strains. Similarly, H. acinonychis strains were readily transformed to clarithromycin resistance strains by uptake of PCR product via natural transformation. The above two experiments demonstrate that bacterium like H. pylori can acquire antibiotic resistance genes like metronidazole and clarithromycin via HGT contributing to the antibiotic resistance of the pathogen H. pylori. This also shows that H. pylori naturally has a way to successfully infect the host even in the presence of harmful antibiotics.
3 Conclusion
Helicobacter pylori survives even in the vulnerable environments such as acidic, peristalsis, phagocytosis and oxidative stress. These stresses induce damage in pathogen DNA and H. pylori had acquired the ability to survive DNA damage by transformation-mediated recombination DNA repair. H. pylori is competent throughout the growth which may help in acquiring the genetic material via HGT from the surrounding environment and contribute to evolution and genetic diversity especially macro-diversity. H. pylori has acquired nickel-binding proteins (Hpn and Hpn-2) and nickel transporter genes (NixA and NiuBDE) via HGT which helped the pathogen to establish itself as gastric species during the course of evolution. This further helped the pathogen H. pylori to adapt itself and survive in the gastric niche. H. pylori also has the capability to acquire genes of antibiotic resistance (metronidazole and clarithromycin) in addition to antigenic determinants and virulence factors via HGT from other organisms to alter and influence pathogenicity. This review clearly reveals the role of horizontal gene transfer in gastric human pathogen H. pylori to adapt itself to the host.
References
Alm RA, Ling LS, Moir DT, King BL, Brown ED, Doig PC, Smith DR, Noonan B, Guild BC, deJonge BL, Carmel G, Tummino PJ, Caruso A, Uria-Nickelsen M, Mills DM, Ives C, Gibson R, Merberg D, Mills SD, Jiang Q, Taylor DE, Vovis GF, Trust TJ (1999) Genomic-sequence comparison of two unrelated isolates of the human gastric pathogen Helicobacter pylori. Nature 397:176–180
Ando TD, Israel A, Kusugami K, Blaser MJ (1999) HP0333, a member of the dprA family, is involved in natural transformation in Helicobacter pylori. J Bacteriol 181:5572–5580
Bardhan KD, Morton D, Perry MJ, Sanders DS, Morris P, Rowland A, Thompson M, Mitchell TR, Roberts PM (2001) Ranitidine bismuth citrate with clarithromycin alone or with metronidazole for the eradication of Helicobacter pylori. Aliment Pharmacol Ther 15(8):1199–1204
Bolor-Erdene M, Namdag B, Yamaoka Y, Jav S (2017) Antibiotic resistance of Helicobacter pylori in Mongolia. J Infect Dev Ctries 11:887–894
Boyanova L, Ilieva J, Gergova G, Spassova Z, Nikolov R, Davidkov L, Evstatiev I, Kamburov V, Katsarov N, Mitov I (2009) Evaluation of clinical and socio-demographic risk factors for antibacterial resistance of Helicobacter pylori in Bulgaria. J Med Microbiol 58:94–100
Campanale M, Nucera E, Ojetti V, Cesario V, Di Rienzo TA, D’Angelo G, Pecere S, Barbaro F, Gigante G, De Pasquale T, Rizzi A, Cammarota G, Schiavino D, Franceschi F, Gasbarrini A (2014) Nickel free-diet enhances the Helicobacter pylori eradication rate: a pilot study. Dig Dis Sci 59:1851–1855. 24595654. https://doi.org/10.1007/s10620-014-3060-3
Challa C, Neelapu NRR (2018) Quorum sensing in Helicobacter pylori: role of biofilm and its implications for antibiotic resistance and immune evasion. In: Veera Bramha Chari P (ed) Implication of quorum sensing system in biofilm formation and virulence. Springer Nature, Switzerland, pp 361–381
Challa S, Mohana Sheela G, Neelapu NRR (2018) Understanding the bacterial biofilm resistance to antibiotics and immune evasion. In: Veera Bramha Chari P (ed) Implication of quorum sensing system in biofilm formation and virulence. Springer Nature, Switzerland, pp 369–381
Dorer MS, Fero J, Salama NR (2010) DNA damage triggers genetic exchange in Helicobacter pylori. PLoS Pathog 6:e1001026
Eppinger M, Baar C, Linz B, Raddatz G, Lanz C, Keller H, Morelli G, Gressmann H, Achtman M, Schuster SC (2006) Who ate whom? Adaptive Helicobacter genomic changes that accompanied a host jump from early humans to large felines. PLoS Genet 2:e120. https://doi.org/10.1371/journal.pgen.0020120
Fernandez-Gonzalez E, Backert S (2014) DNA transfer in the gastric pathogen Helicobacter pylori. J Gastroenterol 49:594–604
Ferrero RL, Jenks PJ (2001) Invivo adaptation to the host. In: HLT M, Mendz GL, Hazell SL (eds) Helicobacter pylori: physiology and genetics, Chap. 46. ASM Press, Washington, DC. https://www.ncbi.nlm.nih.gov/books/NBK2450/
Fischer F, Robbe-Saule M, Turlin E, Mancuso F, Michel V, Richaud P, Veyrier FJ, De Reuse H, Vinella D (2016) Characterization in Helicobacter pylori of a nickel transporter essential for colonization that was acquired during evolution by gastric Helicobacter species. PLoS Pathog 12(12):e1006018. https://doi.org/10.1371/journal.ppat.1006018
Gao W, Cheng H, Hu F, Li J, Wang L, Yang G, Xu L, Zheng X (2010) The evolution of Helicobacter pylori antibiotics resistance over 10 years in Beijing, China. Helicobacter 15:460–466
Garcia-Aljaro C, Balleste E, Muniesa M (2017) Beyond the canonical strategies of horizontal gene transfer in prokaryotes. Curr Opin Microbiol 38:95–105
Hofreuter D, Haas R (2002) Characterization of two cryptic Helicobacter pylori plasmids: a putative source for horizontal gene transfer and gene shuffling. J Bacteriol 184(10):2755–2766
Hofreuter D, Odenbreit S, Henke G, Haas R (1998) Natural competence for DNA transformation in Helicobacter pylori: identification and genetic characterization of the comB locus. Mol Microbiol 28:1027–1038
Hofreuter D, Odenbreit S, Haas R (2001) Natural transformation competence in Helicobacter pylori is mediated by the basic components of a type IV secretion system. Mol Microbiol 41:379–391
Huang YQ, Huang GR, Wu MH, Tang HY, Huang ZS, Zhou XH, Yu WQ, Su JW, Mo XQ, Chen BP, Zhao LJ (2015) Inhibitory effects of emodin, baicalin, schizandrin and berberine on hefA gene: treatment of Helicobacter pylori-induced multidrug resistance. World J Gastroenterol 21:4225
Kurtaran H, Uyar ME, Kasapoglu B, Turkay C, Yilmaz T, Akcay A, Kanbay M (2008) Role of Helicobacter pylori in pathogenesis of upper respiratory system diseases. J Natl Med Assoc 100:1224
Lee SM, Kim N, Kwon YH, Nam RH, Kim JM, Park JY, Lee YS, Lee DH (2018) Rdxa, frxa, and efflux pump in metronidazole-resistant Helicobacter pylori: their relation to clinical outcomes. J Gastroenterol Hepatol 33:681–688
Linz B, Windsor HM, Gajewski JP, Hake CM, Drautz DI, Schuster SC, Marshall BJ (2013) Helicobacter pylori genomic microevolution during naturally occurring transmission between adults. PLoS One 8(12):e82187. https://doi.org/10.1371/journal.pone.0082187
Lood R, Erturk G, Mattiasson B (2017) Revisiting antibiotic resistance spreading in wastewater treatment plants—bacteriophages as a much neglected potential transmission vehicle. Front Microbiol 8:2298
Mohana Sheela G, Prathyusha AMVN, Neelapu NRR, Bramhachari PV (2018) Intra and inter-species communication in microbes: living with complex and sociable neighbors. In: Veera Bramha Chari P (ed) Implication of quorum sensing system in biofilm formation and virulence. Springer Nature, Switzerland, pp 7–16
Nammi D, Srimath-Tirumala-Peddinti RCPK, Neelapu NRR (2016) Identification of drug targets in Helicobacter pylori by in silico analysis: possible therapeutic implications for gastric cancer. Curr Cancer Drug Targets 16:79–98
Nammi D, Yarla NS, Chubarev VN, Tarasov VV, Barreto GE, Pasupulati CAM, Aliev G, Neelapu NRR (2017) A systematic in-silico analysis of Helicobacter pylori pathogenic islands for identification of novel drug target candidates. Curr Genomics 18:450–465
Ndip RN, Malange Takang AE, Ojongokpoko JE, Luma HN, Malongue A, Akoachere JF, Ndip LM, MacMillan M, Weaver LT (2008) Helicobacter pylori isolates recovered from gastric biopsies of patients with gastroduodenal pathologies in Cameroon: current status of antibiogram. Tropical Med Int Health 13:848–854
Nedenskov-Sorensen P, Bukholm G, Bovre K (1990) Natural competence for genetic transformation in Campylobacter pylori. J Infect Dis 161:365–366
Neelapu RR (2018) Role and regulation of transcriptional factors in gastric cancer. In: Nagaraju GP, Bramhachari PV (eds) Role of transcription factors in gastrointestinal malignancies. Springer, Heidelberg, pp 107–130
Neelapu NRR, Pavani T (2013) Identification of novel drug targets in HpB38, HpP12, HpG27, Hpshi470, HpSJM180 strains of Helicobacter pylori: an insilico approach for therapeutic intervention. Curr Drug Targets 14:601–611
Neelapu NRR, Srimath-Tirumala-Peddinti RCPK, Nammi D, Pasupuleti ACM (2013) New strategies and paradigm for drug target discovery: a special focus on infectious diseases tuberculosis, malaria, leishmaniasis, trypanosomiasis and gastritis. Infect Disord Drug Targets 13(5):352–364
Neelapu NRR, Nammi D, ACM P, Surekha C (2014) Helicobacter pylori induced gastric inflammation, ulcer, and cancer: a pathogenesis perspective. Interdiscip J Microinflammation 1:113
Neelapu NRR, Mutha NVR, Akula S (2015) Identification of potential drug targets in Helicobacter pylori strain HPAG1 by in silico genome analysis. Infect Disord Drug Targets 15:106–117
Neelapu NRR, Nammi D, Pasupuleti AMC, Challa S (2016) Targets against Helicobacter pylori and other tumor-producing bacteria. In: Villa TG, Vinas M (eds) New weapons to control bacterial growth. Springer, Heidelberg, pp 239–279
Neelapu NRR, Titash D, Surekha C (2018) Quorum sensing and its role in agrobacterium mediated gene transfer. In: Chari PVB (ed) Implication of quorum sensing system in biofilm formation and virulence. Springer Nature, Switzerland, pp 259–275
O’Rourke EJ, Chevalier C, Pinto AV, Thiberge JM, Ielpi L, Labigne A, Radicella JP (2003) Pathogen DNA as target for host-generated oxidative stress: role for repair of bacterial DNA damage in Helicobacter pylori colonization. Proc Natl Acad Sci U S A 100:2789–2794
Osaki T, Hanawa T, Manzoku T, Fukuda M, Kawakami H, Suzuki H, Yamaguchi H, Yan X, Taguchi H, Kurata S, Kamiya S (2006) Mutation of luxS affects motility and infectivity of Helicobacter pylori in gastric mucosa of a mongolian gerbil model. J Med Microbiol 55:1477–1485
Pasupuleti AMP, Nammi D, Neelapu NRR (2017) Screening and identification of drug targets and vaccine candidates for Helicobacter pylori strain Hp26695. Int J Recent Sci Res 8(4):16384–16395
Pot RG, Kusters JG, Smeets LC, Van Tongeren W, Vandenbroucke-Grauls CM, Bart A (2001) Interspecies transfer of antibiotic resistance between Helicobacter pylori and Helicobacter acinonychis. Antimicrob Agents Chemother 45(10):2975–2976
Savarino V, Mansi C, Mele MR, Bisso G, Mela GS, Saggioro A, Caroli M, Vigneri S, Termini R, Olivieri A, Tosatto R, Celle G (1997) A new 1-week therapy for Helicobacter pylori eradication: ranitidine bismuth citrate plus two antibiotics. Aliment Pharmacol Ther 11(4):699–703
Schmitt W, Odenbreit S, Heuermann D, Haas R (1995) Cloning of the Helicobacter pylori recA gene and functional characterization of its product. Mol Gen Genet 248:563–572
Schuster SC, Wittekindt NE, Linz B (2008) Molecular mechanisms of host-adaptation in Helicobacter. In: Yamaoka Y (ed) Helicobacter pylori: molecular genetics and cellular biology. Horizon Scientific Press, Wymondham, pp 193–204
Smeets LC, Bijlsma JJ, Boomkens SY, Vandenbroucke-Grauls CM, Kusters JG (2000) comH, a novel gene essential for natural transformation of Helicobacter pylori. J Bacteriol 182:3948–3954
Sun QJ, Liang X, Zheng Q, Gu WQ, Liu WZ, Xiao SD, Lu H (2010) Resistance of Helicobacter pylori to antibiotics from 2000 to 2009 in Shanghai. World J Gastroenterol 16:5118
Tomb JF, White O, Kerlavage AR, Clayton RA, Sutton GG, Fleischmann RD, Ketchum KA, Klenk HP, Gill S, Dougherty BA, Nelson K, Quackenbush J, Zhou L, Kirkness EF, Peterson S, Loftus B, Richardson D, Dodson R, Khalak HG, Glodek A, McKenney K, Fitzegerald LM, Lee N, Adams MD, Hickey EK, Berg DE, Gocayne JD, Utterback TR, Peterson JD, Kelley JM, Cotton MD, Weidman JM, Fujii C, Bowman C, Watthey L, Wallin E, Hayes WS, Borodovsky M, Karp PD, Smith HO, Fraser CM, Venter JC (1997) The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature 388:539–547
Torres J, Camorlinga-Ponce M, Pérez-Pérez G, Madrazo-De la Garza A, Dehesa M, González-Valencia G, Muñoz O (2001) Increasing multidrug resistance in Helicobacter pylori strains isolated from children and adults in Mexico. J Clin Microbiol 39:2677–2680
Vinella D, Fischer F, Vorontsov E, Gallaud J, Malosse C, Michel V, Cavazza C, Robbe-Saule M, Richaud P, Chamot-Rooke J, Brochier-Armanet C, De Reuse H (2015) Evolution of Helicobacter: acquisition by gastric species of two histidine-rich proteins essential for colonization. PLoS Pathog 11(12):e1005312. https://doi.org/10.1371/journal.ppat.1005312
Von Wintersdorff CJ, Penders J, van Niekerk JM, Mills ND, Majumder S, van Alphen LB, Savelkoul PHM, Wolffs PFG (2016) Dissemination of antimicrobial resistance in microbial ecosystems through horizontal gene transfer. Front Microbiol 7:173
Wang Y, Roos KP, Taylor DE (1993) Transformation of Helicobacter pylori by chromosomal metronidazole resistance and by a plasmid with a selectable chloramphenicol resistance marker. J Gen Microbiol 139:2485–2493
Wüppenhorst N, Lenze F, Ross M, Kist M (2011) Isolation and eradication of a clinical isolate of Helicobacter pylori resistant to five antimicrobials in Germany. J Antimicrob Chemother 66:222–223
Zullo A, De Francesco V, Hassan C, Morini S, Vaira D (2007) The sequential therapy regimen for Helicobacter pylori eradication: a pooled-data analysis. Gut 56(10):1353–1357
Acknowledgements
CS and NNR are grateful to GITAM (Deemed to be University) for providing necessary facilities to carry out the research work and for extending constant support.
Authors Contribution
CS and NNR initiated the review, participated in writing and revised the manuscript.
Conflict of Interest
The authors declare that there is no potential conflict of interest.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Challa, S., Neelapu, N.R.R. (2019). Association Between Horizontal Gene Transfer and Adaptation of Gastric Human Pathogen Helicobacter pylori to the Host. In: Villa, T., Viñas, M. (eds) Horizontal Gene Transfer. Springer, Cham. https://doi.org/10.1007/978-3-030-21862-1_10
Download citation
DOI: https://doi.org/10.1007/978-3-030-21862-1_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-21861-4
Online ISBN: 978-3-030-21862-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)