Abstract
Optical biosensors are powerful alternatives to the conventional analytical techniques, due to their particular high specificity, sensitivity, small size, and cost effectiveness. Although promising developments of optical biosensors are reported, new bioprobes of cheap and easy synthesis are required, for detection of eukaryotic cells or dangerous infectious agents. In this regard, silicon nanoparticles (SiNPs) can be used as nanoplatform owing to their high specific surface area, optical properties and biocompatibility. They can also be functionalized with bio-probes and used in diagnostic applications. Different methods are described to obtain a stable bond between SiNPs and probes such as nucleotides, antibodies or peptides; however, the latter show many disadvantages about folding instability and sensitivity during the functionalization. Phage Display is a technique for the screening and selection of peptide ligands, that uses an engineered filamentous bacteriophage, mostly made up of 2700 copies of a major coat protein (pVIII) displaying a foreign peptide specific for a target. The bacteriophage or its coat proteins alone can be used as probes to functionalize nanomaterials such as SiNPs. In this work, we propose a new approach to obtain fluorescent bio-probes that can be used for the realization of an optical biosensor. By pulsed laser ablation in liquid (PLAL), SiNPs are functionalized in a “one step” process with phages or isolated pVIII-engineered proteins, selective for Pseudomonas aeruginosa. This process led to complexation of SiNPs with both bioprobes proposed. The PLAL did not alter the biological function of phage probes, maintaining their binding capacity to the bacterial target.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
- Biosensor
- Phage display
- M13 pVIII engineered proteins
- Silicon nanoparticles
- Pulsed laser ablation in liquid
1 Introduction
Nowadays, nanotechnologies are applied in medical field to design new probes for obtaining innovative diagnostic and theranostic systems [1].
Furthermore, the combination of nanotechnology on microfluidic devices has allowed the execution of different analyses with time saving and cost reduction. In addition, the integration of nanoparticles arising from semiconductor materials into different microfluidic based nanosystems may offer a viable alternative to fluorescently labeled particles. In particular, silicon nanoparticles (SiNPs) can be employed in easy-to-use and cheap sensing systems, offer a lot of advantages for their physical properties, their surface state can be easily activated, have stability against photobleaching and a distinctive photoluminescence, exhibit no toxicity and present biocompatibility [2, 3].
Phage Display is a powerful tool used for the screening and selection of peptide ligands to a wide variety of targets, therefore it has been used as a valid substitute for the research of antibodies or peptides. This technique uses M13 filamentous bacteriophage (phage), which consists in a cylindrical shell, mostly made up of 2700 copies of a major coat protein (pVIII) and other four minor coat proteins (pIII; pVI; pVII; pIX), that enclose a circular single-stranded DNA molecule. The “in-frame” insertion of exogenous DNA fragments in the gene encoding the major capsid protein pVIII allows the formation of large molecular libraries, which can be used to discover new bioprobes [4,5,6,7].
Bacteriophage or its protein alone can both be used as probes to functionalize several metal (gold or silver [8]) and semiconductor nanoparticles. Furthermore, the whole phage, the isolated pVIII protein or the exogenous peptide alone can be isolated without loss of activity, maintaining their selectivity, specificity and biological activity [9] at different conditions of temperature and pH, or in the presence of acid and organic solvents.
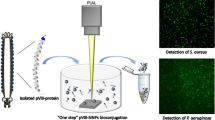
In this work, we propose a new approach for biofunctionalization of SiNPs with M13-engineered bacteriophage or isolated pVIII-engineered proteins, displaying specific peptides that selectively recognize Pseudomonas aeruginosa [10]. The “one-step” functionalization is conducted during the pulsed laser ablation in liquid (PLAL) of a silicon plate in a solution containing the bio-probes. This proposed strategy demonstrates its potential use for in vitro applications and could be exploited to realize an optical biosensor to detect a specific target.
2 Results and Discussion
The phage clone used in this work has been selected from landscape M13-pVIII -9aa peptide library. This clone (P9b) displays the foreign peptide QRKLAAKLT which recognizes and specifically binds the 42 KDa outer membrane protein (OMP) of P. aeruginosa [10], the most common agent of nosocomial infections.
The two probes (the whole bacteriophage or the isolated pVIII protein alone) have been used separately during PLAL.
The isolation and purification of the major coat protein pVIII of P9b phage clone was performed according to the protocol of Pei Liu et al. [9].
SiNPs were generated and simultaneously (one-step) functionalised by PLAL as follows. High purity (99.99%) monocrystalline silicon plate was immersed in a glass vessel filled with 2.5 mL of an aqueous solution of pVIII protein (25 μg mL−1; pVIII-SiNPs) or a phage suspension in TBS buffer (8 × 1011 PFU mL−1; phage-SiNPs). The ablation process was performed using the second harmonic (532 nm) of a neodymium-doped yttrium aluminum garnet (Nd:YAG) laser (model New Wave Mod. Tempest 300), operating at 10 Hz repetition rate with a pulse width of 5 ns [8, 11]. The silicon target was irradiated at the laser fluence of 7.5 J cm−2 and for an ablation time of 30 min.
To separate the phage–SiNPs from the unbounded phages and free SiNPs, networks were purified by centrifugation at 20,800 × g for 30 min, while to isolate the complex pVIII-SiNPs ultracentrifugation at 44.700 × g was performed, according to procedure described by Bagga et al. [12].
Preliminarily, we used the above mentioned parameters to test phage and protein stabilities. Despite the high temperatures, a sufficient amount of the phage population complexed with SiNPs and kept its structure intact, as demonstrated in our previous work [13]. In particular, it has been noticed that the phage (800 nm length and 5 nm diameter) was decorated with numerous SiNPs of different sizes as showed in SEM image (Fig. 1).
The binding occurred due to the electrostatic interaction between the charges of SiNPs surface and the phage surface [14], was mediated by ions present in buffer, forming salt bridges in phage–SiNPs network.
EDX analysis showed the presence of N (nitrogen) atomic species typical of proteins exposed on the surface of the bacteriophage; moreover, the presence of Si (silicon), O (oxygen), Na (sodium) and Cl (chloride) confirmed the functionalization of the nanoparticles with the bacteriophage.
Since the isolated pVIII protein can be easily isolated without loss of its activity, we verified whether it could be used as functionalizing agent for SiNPs during PLAL. Although parameters have still to be optimized, the STEM image (Fig. 2) results show the formation of SiNPs complexed with proteins. Furthermore, the interactions with pVIII proteins caused changes in size, shape, and aggregation state of SiNPs. The mean size of the SiNPs and complexes were estimated. The SiNPs had an average diameter of ∼17.5 nm, while the pVIII-SiNPs of ∼20 nm. Although a portion of these proteins was altered during the laser ablation, a large portion of pVIII proteins assembled with SiNPs and interacted with the target. The phage proteins are polyelectrolytes and then tend to aggregate, but during the ablation process, the thermal and electrostatic variations near the plume may initially determine the disaggregation of the proteins. As consequence, the proteins adsorbed on the SiNPs surface will be arranged as monolayer due to the negative charges on the SiNPs (Zeta potential −31 mV), creating a protein corona on every single SiNP [15].
On the other hand, STEM images showed that a fraction of pVIII-SiNPs could generate clumps due to electrostatic attractions among the exposed groups of the pVIII amino acids, leading to the formation of SiNPs-pVIII-pVIII-SiNPs complexes. Despite this, both single pVIII-SiNPs and larger aggregates are able to recognize the bacterial target. In fact, when the pVIII-SiNPs solution was tested against P. aeruginosa, STEM images (Fig. 3) showed the binding of the nanoparticles on the surface of the bacterium, whereas no binding was observed in the control (data not shown). In addition, the image highlights the lytic effect of the peptide on the membrane, confirming the maintenance of the structural and functional characteristics of the exposed peptide.
Consequently, the PLAL did not alter the structure and the properties of the functionalized bioprobes, so the specific peptide maintained its ability to recognize and interact with bacterial target.
To evaluate the possibility of using pVIII-SiNPs complexes as fluorescent probes in the identification of P. aeruginosa, a solution of pVIII-SiNPs was incubated with P. aeruginosa for 30 min at RT in rotator mixer and then washed in PBS. Finally, the samples were observed by epifluorescence microscope Leica DMRE (Excitation filter BP450-490, Suppression filter LP 515).
Figure 4a shows the presence of P. aeruginosa cells covered by the yellow-green complexes of pVIII-SiNPs, while in Fig. 4b, no fluorescence was observed when P. aeruginosa cells were treated with SiNPs alone.
These results confirm that pVIII-SiNPs were able to provide a fluorescence response through the luminescent signal to their bacterial target, then they may be used as bio-functional nanoprobes.
3 Conclusions
Fluorescent silicon-based nanoparticles can be functionalized with M13 engineered bacteriophages or their isolated pVII proteins by a “one step” process without altering their ability to bind the target.
Phages are cheap and easy to produce, and SiNPs luminescence could be a safer and valid alternative to fluorochrome labeling. Moreover, these phage-SiNPs complexes have demonstrated their potential use for in vitro applications and could be exploited as an optical biosensor to detect prokaryotic or eukaryotic targets.
The physical and biological features of these complexes offer convenient multi-functional integration within a single entity with potential for nanotechnology-based biomedical applications.
Therefore, this strategy allows to obtain low-cost and highly-specific luminescent complexes, which may be employed in LOC system for diagnostic applications.
References
Solano-Umaña, V., Vega-Baudrit, J.R., González-Paz, R.: The new field of the nanomedicine. Int. J. Appl. Sci. Technol. 5(1) (2015)
Huan, C., Shu-Qing, S.: Silicon nanoparticles: preparation, properties, and applications. In: Invited Review—International Conference on Nanoscience & Technology, China 2013, Chin. Phys. B 23(8), 088102 (2014)
Wang, G., Yau, S.-T., Mantey, K., Nayfeh, M.H.: Fluorescent Si nanoparticle-based electrode for sensing biomedical substances. Opt. Commun. 281, 1765–1770 (2008)
Petrenko, V.A., Vodyanoy, V.J.: Phage display for detection of biological threat agents. J. Microbiol. Methods 53, 253–262 (2003)
De Plano, L.M., Carnazza, S., Messina, G.M.L., Rizzo, M.G., Marletta, G., Guglielmino, S.P.P.: Specific and selective probes for Staphylococcus aureus from phage-displayed random peptide libraries. Colloids Surf. B Biointerfaces 157, 473–480 (2017)
Barbas III, C.F., Burton, D.R., Scott, J.K., Silverman, G.J.: Phage Display, A Laboratory Manual. Cold Spring Harbor Laboratory Press, Woodbury, NY (2001)
Petrenko, V.A., Smith, G.P.: Phage from landscape libraries as substitute antibodies. Protein Eng. 13, 101–104 (2000)
Scibilia, S., Lentini, G., Fazio, E., Franco, D., Neri, F., Mezzasalma, A.M., Guglielmino, S.P.P.: Self-assembly of silver nanoparticles and bacteriophage. Sens. Bio-Sens. Res. 7, 146–152 (2016)
Liu, P., Han, L., Wang, F., Petrenko, V.A., Liu, A.: Gold nanoprobe functionalized with specific fusion protein selection from phage display and its application in rapid, selective and sensitive colorimetric biosensing of Staphylococcus aureus. Biosens. Bioelectron. 82, 195–203 (2016). https://doi.org/10.1016/j.bios.2016.03.075
Carnazza, S., Foti, C., Gioffrè, G., Felici, F., Guglielmino, S.P.P.: Specific and selective probes for Pseudomonas aeruginosa from phage-displayed random peptide libraries. Biosens. Bioelectron. 23, 1137–1144 (2008)
Fazio, E., Cacciola, A., Mezzasalma, A.M., Mondio, G., Neri, F., Saija, R.: Modelling of the optical absorption spectra of PLAL prepared ZnO colloids. J. Quant. Spectrosc. Radiat. Trans. 124, 86–93 (2013)
Bagga, K., Barchanski, A., Intartaglia, R., Dante, S., Marotta, R., Diaspro, A., Sajti, C.L., Brandi, F.: Laser-assisted synthesis of Staphylococcus aureus protein-capped silicon quantum dots as bio-functional nanoprobes. Laser Phys. Lett. 10, 065603 (8 pp) (2013)
De Plano, L.M., Scibilia, S., Rizzo, M.G., Crea, S., Franco, D., Mezzasalma, A.M., Guglielmino, S.P.P.: One-step production of phage–silicon nanoparticles by PLAL as fluorescent nanoprobes for cell identification. Appl. Phys. A 124, 222 (2018)
Coen, M., Lehmann, R., Groning, P., Bielmann, M., Galli, C., Schlapbach, L.: Adsorption and bioactivity of protein A on silicon surfaces studied by AFM and XPS. J. Colloid Interface Sci. 233, 180–189 (2001)
Shemetov, A.A., Nabiev, I., Sukhanova, A.: Molecular Interaction of Proteins and Peptides with Nanoparticles, vol. 6. American Chemical Society (2012)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Rizzo, M.G. et al. (2019). One-Step Functionalization of Silicon Nanoparticles with Phage Probes to Identify Pathogenic Bacteria. In: Andò, B., et al. Sensors. CNS 2018. Lecture Notes in Electrical Engineering, vol 539. Springer, Cham. https://doi.org/10.1007/978-3-030-04324-7_21
Download citation
DOI: https://doi.org/10.1007/978-3-030-04324-7_21
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-04323-0
Online ISBN: 978-3-030-04324-7
eBook Packages: EngineeringEngineering (R0)