Abstract
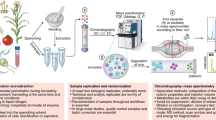
Metabolomics has emerged in the past decade as a highly attractive and impactful technique for phenotype-level profiling in diverse biological applications. Most recently, the dual developments of high-throughput analytical techniques along with dramatically increased sensitivity of high-resolution mass spectrometers have enabled the routine analysis of hundreds of unique samples per day. We have previously reported a robust 3 min isocratic metabolomics platform for the quantification of amino acids and the key pathways of central carbon and nitrogen metabolism. Building on this work, we describe here a 5 min reverse phase gradient followed by global, untargeted profiling of the hydrophilic metabolome. In addition to observing those metabolites measured in the 3 min run, the use of the longer gradient run here also allows for coverage of less polar compounds such as fatty acids and acylcarnitines, both key players in mitochondrial and lipid metabolism, without a significant sacrifice in throughput.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, Smith HO, Yandell M, Evans CA, Holt RA, Gocayne JD, Amanatides P, Ballew RM, Huson DH, Wortman JR, Zhang Q, Kodira CD, Zheng XH, Chen L, Skupski M, Subramanian G, Thomas PD, Zhang J, Gabor Miklos GL, Nelson C, Broder S, Clark AG, Nadeau J, McKusick VA, Zinder N, Levine AJ, Roberts RJ, Simon M, Slayman C, Hunkapiller M, Bolanos R, Delcher A, Dew I, Fasulo D, Flanigan M, Florea L, Halpern A, Hannenhalli S, Kravitz S, Levy S, Mobarry C, Reinert K, Remington K, Abu-Threideh J, Beasley E, Biddick K, Bonazzi V, Brandon R, Cargill M, Chandramouliswaran I, Charlab R, Chaturvedi K, Deng Z, Di Francesco V, Dunn P, Eilbeck K, Evangelista C, Gabrielian AE, Gan W, Ge W, Gong F, Gu Z, Guan P, Heiman TJ, Higgins ME, Ji RR, Ke Z, Ketchum KA, Lai Z, Lei Y, Li Z, Li J, Liang Y, Lin X, Lu F, Merkulov GV, Milshina N, Moore HM, Naik AK, Narayan VA, Neelam B, Nusskern D, Rusch DB, Salzberg S, Shao W, Shue B, Sun J, Wang Z, Wang A, Wang X, Wang J, Wei M, Wides R, Xiao C, Yan C, Yao A, Ye J, Zhan M, Zhang W, Zhang H, Zhao Q, Zheng L, Zhong F, Zhong W, Zhu S, Zhao S, Gilbert D, Baumhueter S, Spier G, Carter C, Cravchik A, Woodage T, Ali F, An H, Awe A, Baldwin D, Baden H, Barnstead M, Barrow I, Beeson K, Busam D, Carver A, Center A, Cheng ML, Curry L, Danaher S, Davenport L, Desilets R, Dietz S, Dodson K, Doup L, Ferriera S, Garg N, Gluecksmann A, Hart B, Haynes J, Haynes C, Heiner C, Hladun S, Hostin D, Houck J, Howland T, Ibegwam C, Johnson J, Kalush F, Kline L, Koduru S, Love A, Mann F, May D, McCawley S, McIntosh T, McMullen I, Moy M, Moy L, Murphy B, Nelson K, Pfannkoch C, Pratts E, Puri V, Qureshi H, Reardon M, Rodriguez R, Rogers YH, Romblad D, Ruhfel B, Scott R, Sitter C, Smallwood M, Stewart E, Strong R, Suh E, Thomas R, Tint NN, Tse S, Vech C, Wang G, Wetter J, Williams S, Williams M, Windsor S, Winn-Deen E, Wolfe K, Zaveri J, Zaveri K, Abril JF, Guigó R, Campbell MJ, Sjolander KV, Karlak B, Kejariwal A, Mi H, Lazareva B, Hatton T, Narechania A, Diemer K, Muruganujan A, Guo N, Sato S, Bafna V, Istrail S, Lippert R, Schwartz R, Walenz B, Yooseph S, Allen D, Basu A, Baxendale J, Blick L, Caminha M, Carnes-Stine J, Caulk P, Chiang YH, Coyne M, Dahlke C, Mays A, Dombroski M, Donnelly M, Ely D, Esparham S, Fosler C, Gire H, Glanowski S, Glasser K, Glodek A, Gorokhov M, Graham K, Gropman B, Harris M, Heil J, Henderson S, Hoover J, Jennings D, Jordan C, Jordan J, Kasha J, Kagan L, Kraft C, Levitsky A, Lewis M, Liu X, Lopez J, Ma D, Majoros W, McDaniel J, Murphy S, Newman M, Nguyen T, Nguyen N, Nodell M, Pan S, Peck J, Peterson M, Rowe W, Sanders R, Scott J, Simpson M, Smith T, Sprague A, Stockwell T, Turner R, Venter E, Wang M, Wen M, Wu D, Wu M, Xia A, Zandieh A, Zhu X (2001) The sequence of the human genome. Science 291(5507):1304–1351
Scolamiero E, Cozzolino C, Albano L, Ansalone A, Caterino M, Corbo G, di Girolamo MG, Di Stefano C, Durante A, Franzese G, Franzese I, Gallo G, Giliberti P, Ingenito L, Ippolito G, Malamisura B, Mazzeo P, Norma A, Ombrone D, Parenti G, Pellecchia S, Pecce R, Pierucci I, Romanelli R, Rossi A, Siano M, Stoduto T, Villani GRD, Andria G, Salvatore F, Frisso G, Ruoppolo M (2015) Targeted metabolomics in the expanded newborn screening for inborn errors of metabolism. Mol BioSyst 11(6):1525–1535
Danaei G, Finucane MM, Lu Y, Singh GM, Cowan MJ, Paciorek CJ, Lin JK, Farzadfar F, Khang Y-H, Stevens GA, Rao M, Ali MK, Riley LM, Robinson CA, Ezzati M (2011) National, regional, and global trends in fasting plasma glucose and diabetes prevalence since 1980: systematic analysis of health examination surveys and epidemiological studies with 370 country-years and 2·7 million participants. Lancet 378(9785):31–40
Odom SR, Howell MD, Silva GS, Nielsen VM, Gupta A, Shapiro NI, Talmor D (2013) Lactate clearance as a predictor of mortality in trauma patients. J Trauma Acute Care Surg 74(4):999–1004
Lusczek ER, Muratore SL, Dubick MA, Beilman GJ (2017) Assessment of key plasma metabolites in combat casualties. J Trauma Acute Care Surg 82(2):309–316
D’Alessandro A, Gevi F, Zolla L (2011) A robust high resolution reversed-phase HPLC strategy to investigate various metabolic species in different biological models. Mol BioSyst 7(4):1024–1032
Zampieri M, Sekar K, Zamboni N, Sauer U (2017) Frontiers of high-throughput metabolomics. Curr Opin Chem Biol 36:15–23
Nemkov T, Hansen KC, D’Alessandro A (2017) A three-minute method for high-throughput quantitative metabolomics and quantitative tracing experiments of central carbon and nitrogen pathways. Rapid Commun. Mass Spectrom 31(8):663–673
Lewis MR, Pearce JTM, Spagou K, Green M, Dona AC, Yuen AHY, David M, Berry DJ, Chappell K, Horneffer-van der Sluis V, Shaw R, Lovestone S, Elliott P, Shockcor J, Lindon JC, Cloarec O, Takats Z, Holmes E, Development NJK (2016) Application of ultra-performance liquid chromatography-TOF MS for precision large scale urinary metabolic phenotyping. Anal Chem 88(18):9004–9013
Miura D, Fujimura Y, Yamato M, Hyodo F, Utsumi H, Tachibana H, Wariishi H (2010) Ultrahighly sensitive in situ metabolomic imaging for visualizing spatiotemporal metabolic behaviors. Anal Chem 82(23):9789–9796
Gasilova N, Yu Q, Qiao L, Girault HH (2014) On-Chip spyhole mass spectrometry for droplet-based microfluidics. Angew Chem Int Ed 53(17):4408–4412
Wouters S, De Vos J, Dores-Sousa JL, Wouters B, Desmet G, Eeltink S (2017) Prototyping of thermoplastic microfluidic chips and their application in high-performance liquid chromatography separations of small molecules. J Chromatogr A 1523:224–233
Gehrke S, Reisz JA, Nemkov T, Hansen KC, D’Alessandro A (2017) Characterization of rapid extraction protocols for high-throughput metabolomics. Rapid Commun Mass Spectrom 31(17):1445–1452
Dias DA, Koal T (2016) Progress in metabolomics standardisation and its significance in future clinical laboratory medicine. EJIFCC 27(4):331–343
Liquid-handling Lego robots and experiments for STEM education and research. [date unknown]. http://journals.plos.org/plosbiology/article?id=10.1371/journal.pbio.2001413. Accessed 19 July 2017
Nemkov T, D’Alessandro A, Hansen KC (2015) Three-minute method for amino acid analysis by UHPLC and high-resolution quadrupole orbitrap mass spectrometry. Amino Acids 47(11):2345–2357
Reisz JA, Wither MJ, Dzieciatkowska M, Nemkov T, Issaian A, Yoshida T, Dunham AJ, Hill RC, Hansen KC, D’Alessandro A (2016) Oxidative modifications of glyceraldehyde 3-phosphate dehydrogenase regulate metabolic reprogramming of stored red blood cells. Blood 128(12):e32–e42
D’alessandro A, Dzieciatkowska M, Nemkov T, Hansen KC (2017) Red blood cell proteomics update: is there more to discover? Blood Transfus 15(2):182–187
Conflicts of Interest
A.D. and T.N. are part of Omix Technologies, Inc., and A.D. is a consultant for Hemanext, Inc.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Nemkov, T., Reisz, J.A., Gehrke, S., Hansen, K.C., D’Alessandro, A. (2019). High-Throughput Metabolomics: Isocratic and Gradient Mass Spectrometry-Based Methods. In: D'Alessandro, A. (eds) High-Throughput Metabolomics. Methods in Molecular Biology, vol 1978. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9236-2_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9236-2_2
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9235-5
Online ISBN: 978-1-4939-9236-2
eBook Packages: Springer Protocols