Abstract
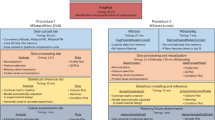
Data-independent acquisition (DIA) mode of mass spectrometry, such as the SWATH-MS technology, enables accurate and consistent measurement of proteins, which is crucial for comparative proteomics studies. However, there is lack of free and easy to implement data analysis protocols that can handle the different data processing steps from raw spectrum files to peptide intensity matrix and its downstream analysis. Here, we provide a data analysis protocol, named diatools, covering all these steps from spectral library building to differential expression analysis of DIA proteomics data. The data analysis tools used in this protocol are open source and the protocol is distributed at Docker Hub as a complete software environment that supports Linux, Windows, and macOS operating systems.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Aebersold R, Mann M (2003) Mass spectrometry-based proteomics. Nature 422:198–207
Gillet LC, Navarro P, Tate S et al (2012) Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteomics 11:O111.016717
Huang Q, Yang L, Luo J et al (2015) SWATH enables precise label-free quantification on proteome scale. Proteomics 15:1215–1223
Schubert OT, Gillet LC, Collins BC et al (2015) Building high-quality assay libraries for targeted analysis of SWATH MS data. Nat Protoc 10:426–441
Röst HL, Rosenberger G, Navarro P et al (2014) OpenSWATH enables automated, targeted analysis of data-independent acquisition MS data. Nat Biotechnol 32:219–223
Röst HL, Liu Y, D’Agostino G et al (2016) TRIC: an automated alignment strategy for reproducible protein quantification in targeted proteomics. Nat Methods 13:777–783
Merkel D (2014) Docker: lightweight Linux containers for consistent development and deployment. Linux J
Chambers MC, Maclean B, Burke R et al (2012) A cross-platform toolkit for mass spectrometry and proteomics. Nat Biotechnol 30:918–920
Sturm M, Bertsch A, Gröpl C et al (2008) OpenMS—an open-source software framework for mass spectrometry. BMC Bioinformatics 9:163
Deutsch EW, Mendoza L, Shteynberg D et al (2010) A guided tour of the trans-proteomic pipeline. Proteomics 10:1150–1159
Suomi T, Corthals GL, Nevalainen OS et al (2015) Using peptide-level proteomics data for detecting differentially expressed proteins. J Proteome Res 14:4564–4570
Huber W, Carey VJ, Gentleman R et al (2015) Orchestrating high-throughput genomic analysis with bioconductor. Nat Methods 12:115–121
Elo LL, Filén S, Lahesmaa R et al (2008) Reproducibility-optimized test statistic for ranking genes in microarray studies. IEEE/ACM Trans Comput Biol Bioinform 5:423–431
Suomi T, Elo LL (2017) Enhanced differential expression statistics for data-independent acquisition proteomics. Sci Rep 7:5869
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Pietilä, S., Suomi, T., Aakko, J., Elo, L.L. (2019). A Data Analysis Protocol for Quantitative Data-Independent Acquisition Proteomics. In: Wang, X., Kuruc, M. (eds) Functional Proteomics. Methods in Molecular Biology, vol 1871. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8814-3_27
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8814-3_27
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8813-6
Online ISBN: 978-1-4939-8814-3
eBook Packages: Springer Protocols