Abstract
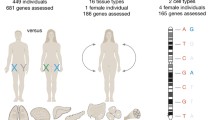
X chromosome inactivation silences one X chromosome in female mammals. However, this silencing is incomplete, and some genes escape X inactivation. We describe methods to determine the chromosome-wide X inactivation status of genes in tissues or cell lines derived from mice using a combination of skewing of X inactivation and allele-specific analyses of gene expression based on RNA-seq.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Deng X, Berletch JB, Nguyen DK et al (2014) X chromosome regulation: diverse patterns in development, tissues and disease. Nat Rev Genet 15:367–378
Carrel L, Willard HF (2005) X-inactivation profile reveals extensive variability in X-linked gene expression in females. Nature 434:400–404
Yang F, Babak T, Shendure J et al (2010) Global survey of escape from X inactivation by RNA-sequencing in mouse. Genome Res 20:614–622
Berletch JB, Yang F, Disteche CM (2010) Escape from X inactivation in mice and humans. Genome Biol 11:213
Bellott DW, Hughes JF, Skaletsky H et al (2014) Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508:494–499
Cortez D, Marin R, Toledo-Flores D et al (2014) Origins and functional evolution of Y chromosomes across mammals. Nature 508:488–493
Balaton BP, Brown CJ (2016) Escape artists of the X chromosome. Trends Genet 32:348–359
Berletch JB, Yang F, Xu J et al (2011) Genes that escape from X inactivation. Hum Genet 130:237–245
Disteche CM (2016) Dosage compensation of the sex chromosomes and autosomes. Semin Cell Dev Biol 56:9–18
Disteche CM (2012) Dosage compensation of the sex chromosomes. Annu Rev Genet 46:537–560
Lyon M (1961) Gene action in the X-chromosome of the mouse (Mus musculus L). Nature 190:372–373
Migeon BR (2014) Females are mosaic: X inactivation and sex differences in disease. Oxford University Press, Oxford
Calabrese JM, Sun W, Song L et al (2012) Site-specific silencing of regulatory elements as a mechanism of X inactivation. Cell 151:951–963
Corbel C, Diabangouaya P, Gendrel AV et al (2013) Unusual chromatin status and organization of the inactive X chromosome in murine trophoblast giant cells. Development 140:861–872
Finn EH, Smith CL, Rodriguez J et al (2014) Maternal bias and escape from X chromosome imprinting in the midgestation mouse placenta. Dev Biol 390:80–92
Berletch JB, Ma W, Yang F et al (2015) Identification of genes escaping X inactivation by allelic expression analysis in a novel hybrid mouse model. Data Brief 5:761–769
Lingenfelter PA, Adler DA, Poslinski D et al (1998) Escape from X inactivation of Smcx is preceded by silencing during mouse development. Nat Genet 18:212–213
Balaton BP, Cotton AM, Brown CJ (2015) Derivation of consensus inactivation status for X-linked genes from genome-wide studies. Biol Sex Differ 6:35
Berletch JB, Ma W, Yang F et al (2015) Escape from X inactivation varies in mouse tissues. PLoS Genet 11:e1005079
Deng Q, Ramskold D, Reinius B et al (2014) Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343:193–196
Marks H, Kerstens HH, Barakat TS et al (2015) Dynamics of gene silencing during X inactivation using allele-specific RNA-seq. Genome Biol 16:149
Benitez JA, Cheng S, Deng Q (2017) Revealing allele-specific gene expression by single-cell transcriptomics. Int J Biochem Cell Biol
Al Nadaf S, Deakin JE, Gilbert C et al (2011) A cross-species comparison of escape from X inactivation in Eutheria: implications for evolution of X chromosome inactivation. Chromosoma 121:71–78
Wu H, Luo J, Yu H et al (2014) Cellular resolution maps of X chromosome inactivation: implications for neural development, function, and disease. Neuron 81:103–119
Lee JH, Daugharthy ER, Scheiman J et al (2015) Fluorescent in situ sequencing (FISSEQ) of RNA for gene expression profiling in intact cells and tissues. Nat Protoc 10:442–458
Cotton AM, Lam L, Affleck JG et al (2011) Chromosome-wide DNA methylation analysis predicts human tissue-specific X inactivation. Hum Genet 130:187–201
Cotton AM, Price EM, Jones MJ et al (2015) Landscape of DNA methylation on the X chromosome reflects CpG density, functional chromatin state and X-chromosome inactivation. Hum Mol Genet 24:1528–1539
Filippova GN, Cheng MK, Moore JM et al (2005) Boundaries between chromosomal domains of X inactivation and escape bind CTCF and lack CpG methylation during early development. Dev Cell 8:31–42
Keown CL, Berletch JB, Castanon R et al (2017) Allele-specific non-CG DNA methylation marks domains of active chromatin in female mouse brain. Proc Natl Acad Sci U S A 114:E2882–E2890
Lister R, Mukamel EA, Nery JR et al (2013) Global epigenomic reconfiguration during mammalian brain development. Science 341:1237905
Marks H, Chow JC, Denissov S et al (2009) High-resolution analysis of epigenetic changes associated with X inactivation. Genome Res 19:1361–1373
Hoki Y, Kimura N, Kanbayashi M et al (2009) A proximal conserved repeat in the Xist gene is essential as a genomic element for X-inactivation in mouse. Development 136:139–146
Trapnell C, Williams BA, Pertea G et al (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28:511–515
Hooper M, Hardy K, Handyside A et al (1987) HPRT-deficient (Lesch-Nyhan) mouse embryos derived from germline colonization by cultured cells. Nature 326:292–295
Mouse Genome Sequencing C, Waterston RH, Lindblad-Toh K et al (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 420:520–562
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25:1105–1111
Kent WJ, Sugnet CW, Furey TS et al (2002) The human genome browser at UCSC. Genome Res 12:996–1006
Raney BJ, Dreszer TR, Barber GP et al (2014) Track data hubs enable visualization of user-defined genome-wide annotations on the UCSC Genome Browser. Bioinformatics 30:1003–1005
Kim D, Pertea G, Trapnell C et al (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14:R36
Anders S, Pyl PT, Huber W (2015) HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31:166–169
Li H, Handsaker B, Wysoker A et al (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550
Acknowledgments
This work was supported by grants GM046883 (C.M.D.), GM113943 (C.M.D., W.M.), and DK107979 (C.M.D., W.N.).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ma, W., Bonora, G., Berletch, J.B., Deng, X., Noble, W.S., Disteche, C.M. (2018). X-Chromosome Inactivation and Escape from X Inactivation in Mouse. In: Sado, T. (eds) X-Chromosome Inactivation. Methods in Molecular Biology, vol 1861. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-8766-5_15
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8766-5_15
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-8765-8
Online ISBN: 978-1-4939-8766-5
eBook Packages: Springer Protocols