Abstract
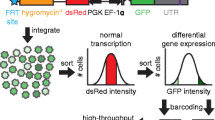
GFP reporter constructs are widely used as an expression system for studying the function of regulatory sequence motifs (cis elements) within the 3′-UTRs (3′ untranslated regions) of mRNAs. Here we provide details on the characterization of individual sequence motifs, which typically regulate mRNA decay and translation. In addition, we describe methods to identify trans factors required for the function of such elements. To facilitate efficient identification of novel functional 3′-UTR motifs, we describe a screening approach based on dual-color fluorescence reporter constructs. Such screening approaches can be used to test large collections of defined sequence or libraries of random sequences.
Similar content being viewed by others
References
Roy B, Jacobson A (2013) The intimate relationships of mRNA decay and translation. Trends Genet 29:691–699
Garneau NL, Wilusz J, Wilusz CJ (2007) The highways and byways of mRNA decay. Nat Rev Mol Cell Biol 8:113–126
Siepel A, Bejerano G, Pedersen JS, Hinrichs AS, Hou M, Rosenbloom K, Clawson H, Spieth J, Hillier LW, Richards S, Weinstock GM, Wilson RK, Gibbs RA, Kent WJ, Miller W, Haussler D (2005) Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res 15:1034–1050
Xie X, Lu J, Kulbokas EJ, Golub TR, Mootha V, Lindblad-Toh K, Lander ES, Kellis M (2005) Systematic discovery of regulatory motifs in human promoters and 3′ UTRs by comparison of several mammals. Nature 434:338–345
Andreassi C, Riccio A (2009) To localize or not to localize: mRNA fate is in 3′UTR ends. Trends Cell Biol 19:465–474
Balagopal V, Parker R (2009) Polysomes, P bodies and stress granules: states and fates of eukaryotic mRNAs. Curr Opin Cell Biol 21:403–408
Dreyfuss G, Kim VN, Kataoka N (2002) Messenger-RNA-binding proteins and the messages they carry. Nat Rev Mol Cell Biol 3:195–205
Kuersten S, Goodwin EB (2003) The power of the 3′ UTR: translational control and development. Nat Rev Genet 4:626–637
Mazumder B, Seshadri V, Fox PL (2003) Translational control by the 3′-UTR: the ends specify the means. Trends Biochem Sci 28:91–98
Mitchell P, Tollervey D (2000) mRNA stability in eukaryotes. Curr Opin Genet Dev 10:193–198
Moore MJ (2005) From birth to death: the complex lives of eukaryotic mRNAs. Science 309:1514–1518
Palacios IM, St Johnston D (2001) Getting the message across: the intracellular localization of mRNAs in higher eukaryotes. Annu Rev Cell Dev Biol 17:569–614
Bhandari D, Raisch T, Weichenrieder O, Jonas S, Izaurralde E (2014) Structural basis for the Nanos-mediated recruitment of the CCR4-NOT complex and translational repression. Genes Dev 28:888–901
Braun JE, Huntzinger E, Fauser M, Izaurralde E (2011) GW182 proteins directly recruit cytoplasmic deadenylase complexes to miRNA targets. Mol Cell 44:120–133
Chekulaeva M, Mathys H, Zipprich JT, Attig J, Colic M, Parker R, Filipowicz W (2011) miRNA repression involves GW182-mediated recruitment of CCR4-NOT through conserved W-containing motifs. Nat Struct Mol Biol 18:1218–1226
Collart MA (2016) The Ccr4-Not complex is a key regulator of eukaryotic gene expression. Wiley Interdiscip Rev RNA 7:438–454. https://doi.org/10.1002/wrna.1332
Lykke-Andersen J, Wagner E (2005) Recruitment and activation of mRNA decay enzymes by two ARE-mediated decay activation domains in the proteins TTP and BRF-1. Genes Dev 19:351–361
Leppek K, Schott J, Reitter S, Poetz F, Hammond MC, Stoecklin G (2013) Roquin promotes constitutive mRNA decay via a conserved class of stem-loop recognition motifs. Cell 153:869–881
Van Etten J, Schagat TL, Hrit J, Weidmann CA, Brumbaugh J, Coon JJ, Goldstrohm AC (2012) Human Pumilio proteins recruit multiple deadenylases to efficiently repress messenger RNAs. J Biol Chem 287:36370–36383
Geissler R, Simkin A, Floss D, Patel R, Fogarty EA, Scheller J, Grimson A (2016) A widespread sequence-specific mRNA decay pathway mediated by hnRNPs A1 and A2/B1. Genes Dev 30:1070–1085
Chen CY, Gherzi R, Ong SE, Chan EL, Raijmakers R, Pruijn GJ, Stoecklin G, Moroni C, Mann M, Karin M (2001) AU binding proteins recruit the exosome to degrade ARE-containing mRNAs. Cell 107:451–464
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Barreau C, Paillard L, Osborne HB (2005) AU-rich elements and associated factors: are there unifying principles? Nucleic Acids Res 33:7138–7150
Staals RH, Bronkhorst AW, Schilders G, Slomovic S, Schuster G, Heck AJ, Raijmakers R, Pruijn GJ (2010) Dis3-like 1: a novel exoribonuclease associated with the human exosome. EMBO J 29:2358–2367
Tomecki R, Kristiansen MS, Lykke-Andersen S, Chlebowski A, Larsen KM, Szczesny RJ, Drazkowska K, Pastula A, Andersen JS, Stepien PP, Dziembowski A, Jensen TH (2010) The human core exosome interacts with differentially localized processive RNases: hDIS3 and hDIS3L. EMBO J 29:2342–2357
Parker R, Sheth U (2007) P bodies and the control of mRNA translation and degradation. Mol Cell 25:635–646
Malecki M, Viegas SC, Carneiro T, Golik P, Dressaire C, Ferreira MG, Arraiano CM (2013) The exoribonuclease Dis3L2 defines a novel eukaryotic RNA degradation pathway. EMBO J 32:1842–1854
Lubas M, Damgaard CK, Tomecki R, Cysewski D, Jensen TH, Dziembowski A (2013) Exonuclease hDIS3L2 specifies an exosome-independent 3′-5′ degradation pathway of human cytoplasmic mRNA. EMBO J 32:1855–1868
Chaudhury A, Kongchan N, Gengler JP, Mohanty V, Christiansen AE, Fachini JM, Martin JF, Neilson JR (2014) A piggyBac-based reporter system for scalable in vitro and in vivo analysis of 3′ untranslated region-mediated gene regulation. Nucleic Acids Res 42:e86
Weingarten-Gabbay S, Elias-Kirma S, Nir R, Gritsenko A, Stern-Ginossar N, Yakhini Z, Weinberger A, Segal E (2016) Comparative genetics. Systematic discovery of cap-independent translation sequences in human and viral genomes. Science 351:aad4939
Wissink EM, Fogarty EA, Grimson A (2016) High-throughput discovery of post-transcriptional cis-regulatory elements. BMC Genomics 17:177
Chen CY, Ezzeddine N, Shyu AB (2008) Messenger RNA half-life in mammalian cells. Methods Enzymol 448:335–357
Acknowledgment
This work was supported by a Research Scholar Grant from the American Cancer Society and R01GM105668 from NIGMS (both to A.G.).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Geissler, R., Grimson, A. (2018). Characterizing mRNA Sequence Motifs in the 3′-UTR Using GFP Reporter Constructs. In: Lamandé, S. (eds) mRNA Decay. Methods in Molecular Biology, vol 1720. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7540-2_6
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7540-2_6
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7539-6
Online ISBN: 978-1-4939-7540-2
eBook Packages: Springer Protocols