Abstract
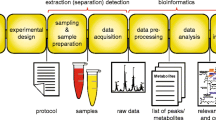
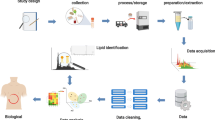
Lipids have been analyzed in applications including drug discovery, disease etiology elucidation, and natural products. The chemical and structural diversity of lipids requires a tailored lipidomics workflow for each sample type. Therefore, every protocol in the lipidomics workflow, especially those involving sample preparation, should be optimized to avoid the introduction of bias. The coupling of ultra-high-performance liquid chromatography (UHPLC) with high-resolution mass spectrometry (HRMS) allows for the separation and identification of lipids based on class and fatty acid acyl chain. This work provides a comprehensive untargeted lipidomics workflow that was optimized for various sample types (mammalian cells, plasma, and tissue) to balance extensive lipid coverage and specificity with high sample throughput. For identification purposes, both data-dependent and data-independent tandem mass spectrometric approaches were incorporated, providing more extensive lipid coverage. Popular open-source feature detection, data processing, and identification software are also outlined.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Anonymous (2015) Major fats and oils industry overview. In: Chemical economics handbook. IHS Markit, London
Batenburg JJ (1992) Surfactant phospholipids: synthesis and storage. Am J Phys 262(4 Pt 1):L367–L385
Bhuyan S, Sundararajan S, Yao L, Hammond EG, Wang T (2006) Boundary lubrication properties of lipid-based compounds evaluated using microtribological methods. Tribol Lett 22(2):167–172. doi:10.1007/s11249-006-9076-x
Brennan L, Owende P (2010) Biofuels from microalgae—a review of technologies for production, processing, and extractions of biofuels and co-products. Renew Sust Energ Rev 14(2):557–577. doi:10.1016/j.rser.2009.10.009
Piazza GJ, Foglia TA (2001) Rapeseed oil for oleochemical usage. Eur J Lipid Sci Technol 103(7):450–454. doi:10.1002/1438-9312(200107)103:7<450::AID-EJLT450>3.0.CO;2-D
Uner M, Wissing SA, Yener G, Muller RH (2005) Skin moisturizing effect and skin penetration of ascorbyl palmitate entrapped in solid lipid nanoparticles (SLN) and nanostructured lipid carriers (NLC) incorporated into hydrogel. Pharmazie 60(10):751–755
Fernandis AZ, Wenk MR (2009) Lipid-based biomarkers for cancer. J Chromatogr B Anal Technol Biomed Life Sci 877(26):2830–2835. doi:10.1016/j.jchromb.2009.06.015
Kwan BCH, Kronenberg F, Beddhu S, Cheung AK (2007) Lipoprotein metabolism and lipid management in chronic kidney disease. J Am Soc Nephrol 18(4):1246–1261. doi:10.1681/asn.2006091006
Lukyanov AN, Torchilin VP (2004) Micelles from lipid derivatives of water-soluble polymers as delivery systems for poorly soluble drugs. Adv Drug Deliv Rev 56(9):1273–1289. doi:10.1016/j.addr.2003.12.004
Sutphen R, Xu Y, Wilbanks GD, Fiorica J, Grendys EC Jr, LaPolla JP, Arango H, Hoffman MS, Martino M, Wakeley K, Griffin D, Blanco RW, Cantor AB, Xiao YJ, Krischer JP (2004) Lysophospholipids are potential biomarkers of ovarian cancer. Cancer Epidemiol Biomarkers Prev 13(7):1185–1191
Yetukuri L, Ekroos K, Vidal-Puig A, Oresic M (2008) Informatics and computational strategies for the study of lipids. Mol BioSyst 4(2):121–127. doi:10.1039/b715468b
Lintonen TP, Baker PR, Suoniemi M, Ubhi BK, Koistinen KM, Duchoslav E, Campbell JL, Ekroos K (2014) Differential mobility spectrometry-driven shotgun lipidomics. Anal Chem 86(19):9662–9669. doi:10.1021/ac5021744
Lin J, Snyder L, McKeon TA (1998) Prediction of relative retention times of triacylglycerols in non-aqueous reversed-phase high-performance liquid chromatography. J Chromatogr 808(1):43–49. doi:10.1016/S0021-9673(98)00134-4
Ulmer CZ, Yost RA, Chen J, Mathews CE, Garrett TJ (2015) Liquid chromatography-mass spectrometry metabolic and lipidomic sample preparation workflow for suspension-cultured mammalian cells using Jurkat t lymphocyte cells. J Proteomics Bioinform 8(6):126–132. doi:10.4172/jpb.1000360
Reis A, Rudnitskaya A, Blackburn GJ, Mohd Fauzi N, Pitt AR, Spickett CM (2013) A comparison of five lipid extraction solvent systems for lipidomic studies of human LDL. J Lipid Res 54(7):1812–1824. doi:10.1194/jlr.M034330
Silva LP, Lorenzi PL, Purwaha P, Yong V, Hawke DH, Weinstein JN (2013) Measurement of DNA concentration as a normalization strategy for metabolomic data from adherent cell lines. Anal Chem 85(20):9536–9542. doi:10.1021/ac401559v
Matyash V, Liebisch G, Kurzchalia TV, Shevchenko A, Schwudke D (2008) Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. J Lipid Res 49(5):1137–1146. doi:10.1194/jlr.D700041-JLR200
Patterson RE, Ducrocq AJ, McDougall DJ, Garrett TJ, Yost RA (2015) Comparison of blood plasma sample preparation methods for combined LC-MS lipidomics and metabolomics. J Chromatogr B Anal Technol Biomed Life Sci 1002:260–266. doi:10.1016/j.jchromb.2015.08.018
Vorkas PA, Isaac G, Anwar MA, Davies AH, Want EJ, Nicholson JK, Holmes E (2015) Untargeted UPLC-MS profiling pipeline to expand tissue metabolome coverage: application to cardiovascular disease. Anal Chem 87(8):4184–4193. doi:10.1021/ac503775m
Smith CA, Want EJ, O’Maille G, Abagyan R, Siuzdak G (2006) XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal Chem 78(3):779–787. doi:10.1021/ac051437y
Pluskal T, Castillo S, Villar-Briones A, Orešič M (2010) MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics 11(1):1–11. doi:10.1186/1471-2105-11-395
Tsugawa H, Cajka T, Kind T, Ma Y, Higgins B, Ikeda K, Kanazawa M, VanderGheynst J, Fiehn O, Arita M (2015) MS-DIAL: data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat Methods 12(6):523–526. doi:10.1038/nmeth.3393
Kind T, Liu KH, Lee DY, DeFelice B, Meissen JK, Fiehn O (2013) LipidBlast in silico tandem mass spectrometry database for lipid identification. Nat Methods 10(8):755–758. doi:10.1038/nmeth.2551
Xia J, Wishart DS (2011) Web-based inference of biological patterns, functions and pathways from metabolomic data using MetaboAnalyst. Nat Protoc 6(6):743–760. doi:10.1038/nprot.2011.319
Xia J, Mandal R, Sinelnikov IV, Broadhurst D, Wishart DS (2012) MetaboAnalyst 2.0--a comprehensive server for metabolomic data analysis. Nucleic Acids Res 40(Web Server issue):W127–W133. doi:10.1093/nar/gks374
Kessler N, Neuweger H, Bonte A, Langenkamper G, Niehaus K, Nattkemper TW, Goesmann A (2013) MeltDB 2.0-advances of the metabolomics software system. Bioinformatics 29(19):2452–2459. doi:10.1093/bioinformatics/btt414
Wu Z, Li D, Meng J, Wang H (2010) Introduction to SIMCA-P and its application. In: Esposito V, Chin W, Henseler J, Wang H (eds) Handbook of partial least squares: concepts methods and applications. Springer, Berlin, Heidelberg, pp 757–774
Fahy E, Subramaniam S, Murphy RC, Nishijima M, Raetz CR, Shimizu T, Spener F, van Meer G, Wakelam MJ, Dennis EA (2009) Update of the LIPID MAPS comprehensive classification system for lipids. J Lipid Res 50(Suppl):S9–14. doi:10.1194/jlr.R800095-JLR200
Wishart DS, Tzur D, Knox C, Eisner R, Guo AC, Young N, Cheng D, Jewell K, Arndt D, Sawhney S, Fung C, Nikolai L, Lewis M, Coutouly MA, Forsythe I, Tang P, Shrivastava S, Jeroncic K, Stothard P, Amegbey G, Block D, Hau DD, Wagner J, Miniaci J, Clements M, Gebremedhin M, Guo N, Zhang Y, Duggan GE, Macinnis GD, Weljie AM, Dowlatabadi R, Bamforth F, Clive D, Greiner R, Li L, Marrie T, Sykes BD, Vogel HJ, Querengesser L (2007) HMDB: the human metabolome database. Nucleic Acids Res 35(Database issue):D521–D526. doi:10.1093/nar/gkl923
Smith CA, O’Maille G, Want EJ, Qin C, Trauger SA, Brandon TR, Custodio DE, Abagyan R, Siuzdak G (2005) METLIN: a metabolite mass spectral database. Ther Drug Monit 27(6):747–751
Taguchi R, Nishijima M, Shimizu T (2007) Basic analytical systems for lipidomics by mass spectrometry in Japan. Methods Enzymol 432:185–211. doi:10.1016/s0076-6879(07)32008-9
Taguchi R, Ishikawa M (2010) Precise and global identification of phospholipid molecular species by an Orbitrap mass spectrometer and automated search engine lipid search. J Chromatogr A 1217(25):4229–4239. doi:10.1016/j.chroma.2010.04.034
Koelmel JP, Kroeger NM, Gill EL et al (2017) Expanding lipidome coverage using LC-MS/MS data-dependent acquisition with automated exclusion list generation. J Am Soc Mass Spectrom 1–10. doi:10.1007/s13361-017-1608-0
Acknowledgments
This work was supported by the Southeast Center for Integrated Metabolomics (SECIM)—NIH Grant #U24 DK097209.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Ulmer, C.Z., Patterson, R.E., Koelmel, J.P., Garrett, T.J., Yost, R.A. (2017). A Robust Lipidomics Workflow for Mammalian Cells, Plasma, and Tissue Using Liquid-Chromatography High-Resolution Tandem Mass Spectrometry. In: Bhattacharya, S. (eds) Lipidomics. Methods in Molecular Biology, vol 1609. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6996-8_10
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6996-8_10
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6995-1
Online ISBN: 978-1-4939-6996-8
eBook Packages: Springer Protocols