Abstract
Poly(ADP-ribose) polymerases (PARP) participate in diverse biological processes contributing to cellular homeostasis or exacerbating injury. PARP catalyzes the addition of ADP-ribose molecules (pADPr) to the target proteins, a process termed poly-ADP-ribosylation. Overactivation of PARP, as reflected by increased poly-ADP-ribosylation, accumulation of pADPr-modified proteins or free pADPr, contributes to depletion of NAD+ and mitochondrial dysfunction, potentially leading to cell death. Since PARP overactivation and increases in free pADPr have been identified as key contributors to the pathobiology of many diseases, monitoring PARP-1 activation by detecting and quantifying pADPr may provide valuable mechanistic insights as well as facilitating therapeutic drug monitoring for PARP inhibitors.
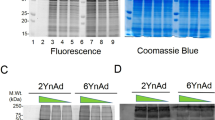
Several non-isotopic immunodetection methods for quantifying pADPr are discussed: western blotting of poly-ADP-ribosylated proteins, cellular localization of pADPr by immunohistochemistry, quantification of pADPr by enzyme-linked immunoassay and small scale two-dimensional gel electrophoresis.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Chiarugi A, Moskowitz MA (2003) Poly(ADP-ribose) polymerase-1 activity promotes NF-kappa B-driven transcription and microglial activation: implication for neurodegenerative disorders. J Neurochem 85:306–317
De Murcia JM, Niedergang C, Trucco C et al (1997) Requirement of poly(ADP-ribose) polymerase in recovery from DNA damage in mice and in cells. Proc Natl Acad Sci U S A 94:7303–7307
Ferro AM, Olivera BM (1984) Poly(ADP-ribosylation) of DNA topoisomerase I from calf thymus. J Biol Chem 259:547–554
Ha HC, Hester LD, Snyder SH (2002) Poly(ADP-ribose) polymerase-1 dependence of stress-induced transcription factors and associated gene expression in glia. Proc Natl Acad Sci U S A 99:3270–3275
Oei SL, Shi Y (2001) Poly(ADP-ribosyl)ation of transcription factor Yin Yang 1 under conditions of DNA damage. Biochem Biophys Res Commun 285:27–31
Satoh MS, Lindahl T (1992) Role of poly(ADP-ribose) formation in DNA repair. Nature 356:356–358
Scovassi AI, Mariani C, Negroni M et al (1993) ADP-ribosylation of nonhistone proteins in HeLa cells: modification of DNA topoisomerase II. Exp Cell Res 206:177–181
Tulin A, Spradling A (2003) Chromatin loosening by poly(ADP)-ribose polymerase (PARP) at drosophila puff loci. Science 299:560–562
Valenzuela MT, Guerrero R, Nunez MI et al (2002) PARP-1 modifies the effectiveness of p53-mediated DNA damage response. Oncogene 21:1108–1116
Alano CC, Ying W, Swanson RA (2004) Poly(ADP-ribose) polymerase-1-mediated cell death in astrocytes requires NAD+ depletion and mitochondrial permeability transition. J Biol Chem 279:18895–18902
Cozzi A, Cipriani G, Fossati S et al (2006) Poly(ADP-ribose) accumulation and enhancement of postischemic brain damage in 110-kDa poly(ADP-ribose) glycohydrolase null mice. J Cereb Blood Flow Metab 26:684–695
Endres M, Wang ZQ, Namura S et al (1997) Ischemic brain injury is mediated by the activation of poly(ADP-ribose)polymerase. J Cereb Blood Flow Metab 17:1143–1151
Lai Y, Chen Y, Watkins SC et al (2008) Identification of poly-ADP-ribosylated mitochondrial proteins after traumatic brain injury. J Neurochem 104:1700–1711
Oliver FJ, Menissier-De Murcia J, Nacci C et al (1999) Resistance to endotoxic shock as a consequence of defective NF-kappa B activation in poly (ADP-ribose) polymerase-1 deficient mice. EMBO J 18:4446–4454
Satchell MA, Zhang X, Kochanek PM et al (2003) A dual role for poly-ADP-ribosylation in spatial memory acquisition after traumatic brain injury in mice involving NAD+ depletion and ribosylation of 14-3-3gamma. J Neurochem 85:697–708
Zingarelli B, Hake PW, O'connor M et al (2004) Differential regulation of activator protein-1 and heat shock factor-1 in myocardial ischemia and reperfusion injury: role of poly(ADP-ribose) polymerase-1. Am J Physiol Heart Circ Physiol 286:H1408–H1415
Szabo C, Dawson VL (1998) Role of poly(ADP-ribose) synthetase in inflammation and ischaemia-reperfusion. Trends Pharmacol Sci 19:287–298
Zhang J, Dawson VL, Dawson TM et al (1994) Nitric oxide activation of poly(ADP-ribose) synthetase in neurotoxicity. Science 263:687–689
Nishizuka Y, Ueda K, Honjo T et al (1968) Enzymic adenosine diphosphate ribosylation of histone and poly adenosine diphosphate ribose synthesis in rat liver nuclei. J Biol Chem 243:3765–3767
Du X, Matsumura T, Edelstein D et al (2003) Inhibition of GAPDH activity by poly(ADP-ribose) polymerase activates three major pathways of hyperglycemic damage in endothelial cells. J Clin Invest 112:1049–1057
Krietsch J, Rouleau M, Pic E et al (2013) Reprogramming cellular events by poly(ADP-ribose)-binding proteins. Mol Asp Med 34:1066–1087
Andrabi SA, Umanah GK, Chang C et al (2014) Poly(ADP-ribose) polymerase-dependent energy depletion occurs through inhibition of glycolysis. Proc Natl Acad Sci U S A 111:10209–10214
Wang Y, Kim NS, Haince JF et al (2011) Poly(ADP-ribose) (PAR) binding to apoptosis-inducing factor is critical for PAR polymerase-1-dependent cell death (parthanatos). Sci Signal 4:ra20
Andrabi SA, Kim NS, Yu SW et al (2006) Poly(ADP-ribose) (PAR) polymer is a death signal. Proc Natl Acad Sci U S A 103:18308–18313
Yu SW, Andrabi SA, Wang H et al (2006) Apoptosis-inducing factor mediates poly(ADP-ribose) (PAR) polymer-induced cell death. Proc Natl Acad Sci U S A 103:18314–18319
Besson VC, Margaill I, Plotkine M et al (2003) Deleterious activation of poly(ADP-ribose)polymerase-1 in brain after in vivo oxidative stress. Free Radic Res 37:1201–1208
Kanai Y, Miwa M, Matsushima T et al (1974) Studies on anti-poly(adenosine diphosphate ribose) antibody. Biochem Biophys Res Commun 59:300–306
Sairanen T, Szepesi R, Karjalainen-Lindsberg ML et al (2009) Neuronal caspase-3 and PARP-1 correlate differentially with apoptosis and necrosis in ischemic human stroke. Acta Neuropathol 118:541–552
Fink EL, Lai Y, Zhang X et al (2008) Quantification of poly(ADP-ribose)-modified proteins in cerebrospinal fluid from infants and children after traumatic brain injury. J Cereb Blood Flow Metab 28:1523–1529
Sarnaik AA, Conley YP, Okonkwo DO et al (2010) Influence of PARP-1 polymorphisms in patients after traumatic brain injury. J Neurotrauma 27:465–471
Acknowledgements
This work was supported by P50 NS30318 (RSC) and T32 HD40686 (YL, MAS).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Lai, YC., Aneja, R.K., Satchell, M.A., Clark, R.S.B. (2017). Detecting and Quantifying pADPr In Vivo. In: Tulin, A. (eds) Poly(ADP-Ribose) Polymerase. Methods in Molecular Biology, vol 1608. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6993-7_3
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6993-7_3
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6992-0
Online ISBN: 978-1-4939-6993-7
eBook Packages: Springer Protocols