Abstract
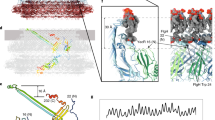
The bacterial flagellum is a large assembly of about 30 different proteins and is divided into three parts: filament, hook, and basal body. The machineries for its crucial functions, such as torque generation, rotational switch regulation, protein export, and assembly initiation, are all located around the basal body. Although high-resolution structures of the filament and hook have already been revealed, the structure of the basal body remains elusive. Recently, the purification protocol for the MS ring, which is the core ring of the basal body, has been improved for the structural study of the MS ring by electron cryomicroscopy (cryoEM) and single particle image analysis. The structure of intact basal body has also been revealed in situ at a resolution of a few nanometers by electron cryotomography (ECT) of minicells. Here, we describe the methods for the MS ring purification, Salmonella minicell culture, and cryoEM/ECT data collection and image analysis.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Berg HC (2003) The rotary motor of bacterial flagella. Annu Rev Biochem 72:19–54
Ueno T, Oosawa K, Aizawa SI (1992) M ring, S ting and proximal rod of the flagellar basal body of Salmonella typhimurium are composed of subunits of a single protein FliF. J Mol Biol 227:672–677
Berg HC (2000) Constraints on models for the flagellar rotary motor. Philos Trans R Soc London B Biol Sci 335:491–502
González-Pedrajo B, Minamino T, Kihara M et al (2006) Interactions between C ring proteins and export apparatus components: a possible mechanism for facilitating type III protein export. Mol Microbiol 60:984–998
Zhou J, Lloyd SA, Blair DF (1998) Electro-static interactions between rotor and stator in the bacterial flagellar motor. Proc Natl Acad Sci U S A 95:6436–6441
Francis NR, Irikura VM, Yamaguchi S et al (1992) Localization of the Salmonella typhimurium flagellar switch protein FliG to the cytoplasmic M-ring face of the basal body. Proc Natl Acad Sci U S A 89:6304–6308
Jones CJ, Macnab RM, Okino H et al (1990) Stoichiometric analysis of the flagellar hook-(basal-body) complex of Salmonella typhimurium. J Mol Biol 212:377–387
Suzuki H, Yonekura K, Namba K (2004) Structure of the rotor of the bacterial flagellar motor revealed by electron cryomicroscopy and single-particle image analysis. J Mol Biol 337:105–113
Thomas DR, Francis NR, Xu C et al (2006) The three-dimensional structure of the flagellar rotor from a clockwise-locked mutant of Salmonella enterica serovar typhimurium. J Bacteriol 188:7039–7048
Gao Y, Cao E, Julius D et al (2016) TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Nature 534:347–351
Taylor NM, Prokhorov NS, Guerrero-Ferreira RC et al (2016) Structure of the T4 baseplate and its function in triggering sheath contraction. Nature 533:346–352
Frauenfeld J, Löving R, Armache JP et al (2016) A saposin-lipoprotein nanoparticle system for membrane protein. Nat Methods 13:345–351
Liao M, Cao E, Julius D et al (2013) Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504:107–112
Hauer F, Gerle C, Fischer N et al (2015) GraDeR: membrane protein complex preparation for single-particle cryo-EM. Structure 23:1769–1775
Kawamoto A, Morimoto VY, Miyata T et al (2013) Common and distinct structural features of Salmonella injectisome and flagellar basal body. Sci Rep 3:3369
Yamaguchi S, Fujita H, Ishihara A et al (1986) Subdivision of flagellar gens of Salmonella typhimurium into regions responsible for assembly, rotation and switching. J Bacteriol 166:187–193
Li X, Mooney P, Zheng S et al (2013) Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat Methods 10:584–590
Mindell JA, Grigorieff N (2003) Accurate determination of local defocus and specimen tilt in electron microscopy. J Struct Biol 142:334–347. doi:10.1016/S1047-8477(03)00069-8
Scheres SH (2012) RELION: implementation of a Bayesian approach to cryo-EM structure determination. J Struct Biol 180:519–530
Kucukelbir A, Sigworth FJ, Tagare HD (2014) Quantifying the local resolution of cryo-EM density maps. Nat Methods 11:63–65
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Kremer JR, Mastronarde DN, McIntosh JR (1996) Computer visualization of three-dimensional image data using IMOD. J Struct Biol 116:71–76
Acknowledgments
The research study described in this chapter was supported by JSPS KAKENHI Grant Number JP25000013 to K.N.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media LLC
About this protocol
Cite this protocol
Kawamoto, A., Namba, K. (2017). Structural Study of the Bacterial Flagellar Basal Body by Electron Cryomicroscopy and Image Analysis. In: Minamino, T., Namba, K. (eds) The Bacterial Flagellum. Methods in Molecular Biology, vol 1593. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6927-2_9
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6927-2_9
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6926-5
Online ISBN: 978-1-4939-6927-2
eBook Packages: Springer Protocols