Abstract
Nonribosomal peptide synthetases (NRPSs) are multifunctional enzymes consisting of catalytic domains. The substrate specificities of adenylation (A) domains determine the amino-acid building blocks to be incorporated during nonribosomal peptide biosynthesis. The A-domains mediate ATP-dependent activation of amino-acid substrates as aminoacyl-O-AMP with pyrophosphate (PPi) release. Traditionally, the enzymatic activity of the A-domains has been measured by radioactive ATP–[32P]-PPi exchange assays with the detection of 32P-labeled ATP. Recently, we developed a colorimetric assay for the direct detection of PPi as a yellow 18-molybdopyrophosphate anion ([(P2O7)Mo18O54]4−). [(P2O7)Mo18O54]4− was further reduced by ascorbic acid to give a more readily distinguishable blue coloration. Here we demonstrate the lab protocols for the colorimetric assay of PPi released in A-domain reactions.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Key words
- Nonribosomal peptide synthetase
- Adenylation domain
- Colorimetric assay
- Poly anion
- ATP–[32P]-PPi exchange assay
1 Introduction
Nonribosomal peptide s constitute a major class of secondary metabolites produced in microorganisms and are synthesized by nonribosomal peptide synthetases (NRPSs). Unlike post-ribosomal peptide synthesis, NRPSs can accept nonproteinogenic amino-acid building blocks as substrates, thereby offering greater structural diversity. NRPSs are multifunctional enzymes consisting of catalytic domains [1–3]. The amino-acid substrate is activated as an aminoacyl-O-AMP by an adenylation (A) domain and subsequently loaded onto the 4′-phosphopantetheine (4′-PP) arm of the adjacent thiolation (T) domain with AMP and pyrophosphate (PPi) releases, resulting in the formation of an aminoacyl-S-enzyme (Fig. 1a). A condensation (C) domain catalyzes a peptide-bond formation between two amino-acid substrates activated as the aminoacyl-S-enzyme. The substrate specificities of A-domains determine the amino-acid building blocks to be incorporated during nonribosomal peptide biosynthesis. Traditionally, the enzymatic activity of A-domains has been measured by radioactive ATP–[32P]-PPi exchange assays through the detection of 32P-labeled ATP produced by a reversible reaction of the A-domain [4, 5]. In 2009, McQuade et al. reported a nonradioactive high-throughput assay for the screening and characterization of A-domains [6]. Their assay uses malachite green to measure orthophosphate (Pi) concentrations after degradation by inorganic pyrophosphatase of the PPi released during aminoacyl-O-AMP formation. However, this method seems to be inadequate for A-domains that have high catalytic rates of the reverse reaction, because the released PPi should be immediately converted to ATP, particularly in the reaction mixture without a T-domain (Fig. 1a).
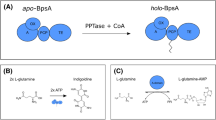
Detection of A-domain adenylation activity. (a) Traditionally, the enzymatic activity of A-domains has been measured by radioactive ATP–[32P]-PPi exchange assays with the detection of 32P-labeled ATP. (b) Addition of hydroxylamine into the A-domain reaction mixture enhances PPi release. PPi is directly detected as a yellow 18-molybdopyrophosphate anion ([(P2O7)Mo18O54]4−). [(P2O7)Mo18O54]4− is further reduced by ascorbic acid to give a more readily distinguishable blue coloration
Recently, we developed a colorimetric assay for the direct detection of PPi as a yellow 18-molybdopyrophosphate anion ([(P2O7)Mo18O54]4−) [7, 8]. [(P2O7)Mo18O54]4− was further reduced by ascorbic acid to give an eight-electron reduced species,which shows a more readily distinguishable blue coloration. Using this assay, the enzymatic activity was successfully measured in acetyl-CoA synthetase that forms AMP + PPi. However, we were unable to detect the enzymatic activities in A-domains, probably due to these enzymes’ PPi-consuming reverse reaction. Although the addition of a T-domain to the reaction mixture should facilitate PPi release, a large amount of T-domain is needed to achieve this. To address this problem, we explored the use of nucleophilic reagents instead of T-domains. Our recent study demonstrated that the aminoacyl-O-AMPs produced by A-domains are converted to hydroxamate derivatives in an enzyme reaction containing hydroxylamine [8]. In addition, the resulting PPi was detected by our colorimetric assay (Fig. 1b). Here we demonstrate the lab protocols for the colorimetric assay of A-domains.
2 Materials
Prepare all solutions using analytical-grade reagents. Prepare and store all reagents at room temperature (unless otherwise described).
2.1 A Domain Reaction Mixture
-
1.
Tris buffer: 1 M Tris (pH 9.0) (see Note 1 ) in water.
-
2.
Magnesium solution: 100 mM MgCl2 in water.
-
3.
ATP (pH 7.0): Weigh 551 mg adenosine 5′-triphosphate (ATP) disodium salt anhydrate and transfer it to a test tube. Add water to a volume of 7 mL and mix. Adjust pH with NaOH. Make up to 10 mL with water. Store in suitable aliquots at −70 °C. Final concentration 100 mM.
-
4.
Hydroxylamine (pH 7.2): Weigh 1.4 g hydroxylamine hydrochloride and transfer it to a glass beaker. Add water to a volume of 80 mL and mix. Adjust pH with KOH. Make up to 100 mL with water. Store at 4 °C (see Note 2 ). Final concentration 200 mM.
-
5.
Amino-acid substrates: 20–100 mM amino-acid solutions are prepared and used for A-domain reaction mixtures (see Note 3 ).
-
6.
A-domains (enzymes): The recombinant enzyme of an A-domain, which is purified to homogeneity by affinity chromatography, is required (see Note 4 ).
2.2 Colorimetric Assay of PPi
-
1.
Concentrated hydrochloric acid: 5 M HCl in water.
-
2.
Acetonitrile (anhydrous, 99.8 %).
-
3.
1 M Na2MoO4: Weigh 24.2 g disodium molybdate(VI) dihydrate and transfer it to a glass beaker. Make up to 100 mL with water.
-
4.
Mo(VI) solution: Mix 6 mL of concentrated hydrochloric acid and 30 mL of acetonitrile. Add water to a volume of 45 mL. Add 1 mL of 1 M Na2MoO4 slowly to the solution while stirring. Make up to 50 mL with water to give a working solution containing 20 mM Na2MoO4, 0.6 M HCl, and 60 % acetonitrile. This solution should be freshly prepared for use.
-
5.
50 mM bis(triphenylphosphoranylidene)ammonium chloride (BTPPACl): Weigh 1.44 g BTPPACl and transfer it to a glass beaker. Add acetonitrile (not water) to a volume of 50 mL.
-
6.
Ascorbic acid solution: Mix 2 mL of 5 M HCl and 3 mL of acetonitrile. Add 0.44 g l-ascorbic acid to the solution. This solution should be freshly prepared for use.
3 Methods
3.1 A Domain Reaction
-
1.
In a 1.5-mL microfuge tube, mix the solution components using the values given in Table 1 (see Note 5 ). Mix in the order shown.
Table 1 Solutions for preparing A domain assay mixture -
2.
Incubate the reaction mixture for 10–60 min at 30 °C.
3.2 PPi Detection by Colorimetric Assay
-
1.
Transfer 50 μL of the reaction mixture to a fresh 1.5-mL microfuge tube containing 500 μL of Mo(VI) solution. Mix thoroughly and incubate at room temperature for 2–5 min (see Note 6 ).
-
2.
Add 10 μL of 50 mM BTPPACl, mix thoroughly, and incubate at room temperature for 5 min (see Note 7 ).
-
3.
Centrifuge the resulting cloudy solution at 20,000 × g for 15 min and remove all liquid.
-
4.
Dissolve the precipitant in 100 μL of acetonitrile by mixing vigorously.
-
5.
Spin briefly to collect the contents at the bottom of the tube.
-
6.
Transfer 100 μL of the solution to a 96-well plate and add 10 μL of ascorbic acid solution.
-
7.
After mixing the solution by pipetting, incubate at room temperature for 10 min.
-
8.
Measure the absorbance of the well at 620 nm on a plate reader (Fig. 2).
Fig. 2 Colorimetric assay of PPi released in an A-domain reaction. The purified rORF 19 (100 μg/mL) is incubated with (a) and without (b) 2 mM β-lysine in a reaction mixture containing 32 mM hydroxylamine for 30 min at 30 °C (see Notes 5 and 8 ). The PPi released in the reaction is detected by the colorimetric assay
3.3 Standard Curves of PPi Concentration
-
1.
Instead of the enzyme reaction mixture, 50 μL of 0–1000 μM Na4P2O7 solution is used for the PPi colorimetric assay.
-
2.
The PPi colorimetric assay is carried out by the same method described above (steps 1–8 in Subheading 3.2).
-
3.
Obtain a standard curve from the results (Fig. 3).
3.4 Determination of A-Domain–Specific Activity
-
1.
In a 1.5-mL microfuge tube, mix the solution components (except for the amino-acid substrate) using the values given in Table 1. Mix in the order shown (see Note 8 ).
-
2.
Incubate the mixture for 3 min at 30 °C (preincubation).
-
3.
Add 2 μL of the amino-acid solution or water (control) to start the enzyme reaction (see Note 9 ).
-
4.
Incubate the mixture for 5–30 min at 30 °C. Terminate the enzyme reaction by adding 500 μL of Mo(VI) solution.
-
5.
Carry out the PPi colorimetric assay using the method described above (steps 1–8 in Subheading 3.2). Determine the concentration of PPi using the PPi standard curve (Subheading 3.3).
-
6.
Determine the specific activity based on the PPi production (Fig. 4).
4 Notes
-
1.
Buffers and their pH should be optimized for A-domains. Tris(hydroxymethyl)aminomethane (Tris), 3-morpholinopro-panesulfonic acid (MOPS), and N-tris(hydroxymethyl)methyl-3-aminopropanesulfonic acid (TAPS) seem to give good results at pH 7–9. Phosphate buffer is not recommended, because it inhibits the activity of the A domains and also gives a high background in the following PPi assay.
-
2.
Hydroxylamine solution can be stored for up to 3 days at 4 °C. However, the pH should be checked before use.
-
3.
When an amino-acid substrate dissolved in a diluted HCl or NaOH is used for the enzyme reaction, the pH of the reaction mixture should be checked. Water-insoluble amino acids can be dissolved in dimethyl sulfoxide (DMSO).
-
4.
Cell-free extract will give a high background in the colorimetric assay of PPi, probably due to the hydrolysis of the ATP by phosphatases. Therefore, a highly purified A-domain should be used for the enzyme reaction. His-tagged recombinant enzymes give good results in our laboratory. NaCl and imidazole, which are used for the purification steps in Ni-affinity chromatography, do not interfere with the colorimetric assay of PPi.
-
5.
In the enzyme reaction, the concentration of hydroxylamine should be optimized. 10–60 mM hydroxylamine gives good results in a large number of A domains. For example, 32 mM is the optimum concentration for a recombinant enzyme of ORF 19 (rORF 19), which is a stand-alone A-domain involved in the biosynthesis of streptothricin antibiotics [9]. In addition, 50 mM Tris–HCl (pH 9.0) and 2 mM β-lysine are used for the rORF 19 reaction as the buffer and substrate, respectively. As a control reaction, the enzyme reaction should be performed without an amino-acid substrate (Fig. 2).
-
6.
The addition of Mo(VI) solution terminates the enzyme reaction and forms a yellow 18-molybdopyrophosphate anion ([(P2O7)Mo18O54]4−). Prolonged incubation (more than 5 min) increases the background in the PPi colorimetric assay.
-
7.
The [(P2O7)Mo18O54]4− anion is precipitated with the BTPPA+ cation.
-
8.
The enzyme concentration should be optimized. For example, in rORF 19 (see Note 5 ), 100 μg/mL enzyme is good for determining the specific activity (Fig. 4).
-
9.
The substrate concentration should be optimized.
References
Marahiel MA, Stachelhaus T, Mootz HD (1997) Modular peptide synthetases involved in nonribosomal peptide synthesis. Chem Rev 97:2651–2674
Mootz HD, Schwarzer D, Marahiel MA (2002) Ways of assembling complex natural products on modular nonribosomal peptide synthetases. ChemBioChem 3:490–504
Schwarzer D, Finking R, Marahiel MA (2003) Nonribosomal peptides: from genes to products. Nat Prod Rep 20:275–287
Bryce GF, Brot N (1972) Studies on the enzymatic synthesis of the cyclic trimer of 2,3-dihydroxy-N-benzoyl-L-serine in Escherichia coli. Biochemistry 11:1708–1715
Rusnak F, Faraci WS, Walsh CT (1989) Subcloning, expression, and purification of the enterobactin biosynthetic enzyme 2,3-dihydroxybenzoate-AMP ligase: demonstration of enzyme-bound (2,3-dihydroxy-benzoyl)adenylate product. Biochemistry 28:6827–6835
McQuade TJ, Shallop AD, Sheoran A et al (2009) A nonradioactive high-throughput assay for screening and characterization of adenylation domains for nonribosomal peptide combinatorial biosynthesis. Anal Biochem 386:244–250
Katano H, Tanaka R, Maruyama C et al (2012) Assay of enzymes forming AMP + PPi by the pyrophosphate determination based on the formation of 18-molybdopyrophosphate. Anal Biochem 421:308–312
Katano H, Watanabe H, Takakuwa M et al (2013) Colorimetric determination of pyrophosphate anion and its application to adenylation enzyme assay. Anal Sci 29:1095–1098
Maruyama C, Toyoda J, Kato Y et al (2012) A stand-alone adenylation domain forms amide bonds in streptothricin biosynthesis. Nat Chem Biol 8:791–797
Acknowledgments
This work was supported in part by KAKENHI (25108720), the Asahi Glass Foundation, and the Japan Foundation for Applied Enzymology.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Maruyama, C., Niikura, H., Takakuwa, M., Katano, H., Hamano, Y. (2016). Colorimetric Detection of the Adenylation Activity in Nonribosomal Peptide Synthetases. In: Evans, B. (eds) Nonribosomal Peptide and Polyketide Biosynthesis. Methods in Molecular Biology, vol 1401. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-3375-4_5
Download citation
DOI: https://doi.org/10.1007/978-1-4939-3375-4_5
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3373-0
Online ISBN: 978-1-4939-3375-4
eBook Packages: Springer Protocols