Abstract
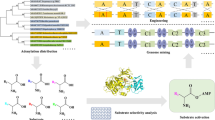
Many amino acid-containing natural products are biosynthesized by large, multifunctional enzymes known as non-ribosomal peptide synthetases (NRPSs). Adenylation (A) domains in NRPSs are responsible for the incorporation of amino acid building blocks and can be considered as engineering domains; therefore, advanced techniques are required to not only rapidly verify expression and folding, but also accelerate the functional prediction of the A-domains in lysates from native and heterologous systems. We recently developed activity-based protein profiling (ABPP) of NRPSs that offers a simple and robust analytical platform for A-domains and provides insights into their enzyme–substrate specificity. In this chapter, we describe the design and synthesis of these ABPP probes and provide a summary of our work on the development of a series of protocols for labeling, visualizing, and analyzing endogenous NRPSs in complex biological systems.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Süssmuth RD, Mainz A (2017) Nonribosomal peptide synthesis-principles and prospects. Angew Chem Int Ed 56:3770–3821
Hur GH, Vickery CR, Burkart MD (2012) Explorations of catalytic domains in non- ribosomal peptide synthetase enzymology. Nat Prod Rep 10:1074–1098
Zhang B, Tian W, Wang S, Yan X, Jia X, Pierens GK, Chen W, Ma H, Deng Z, Qu X (2017) Activation of natural products biosynthetic pathways via a protein modification level regulation. ACS Chem Biol 12:1732–1736
Kwok R (2010) Five hard truths for synthetic biology. Nature 463:286–290
Minandri F, Imperi F, Frangipani E, Bonchi C, Visaggio D, Facchini M, Pasquali P, Bragonzi A, Visca P (2016) Role of iron systems in Pseudomonas aeruginosa virulenceand airway infection. Infect Immun 84:2324–2335
Imperi F, Massai F, Facchini M, Frangipani E, Visaggio D, Leoni L, Bragonzi A, Visca P (2013) Repurposing the antimycotic drug flucytosine for suppression of Pseudomonas aeruginosa pathogenicity. Proc Natl Acad Sci U S A 110:7458–7463
Kirienko DR, Revtovich AV, Kirienko NV (2016) A high-content, phenotypic screen identifies fluorouridine as an inhibitor of pyoverdine biosynthesis and Pseudomonas aeruginosa virulence. mSphere 1:e00217–e00216
Kaneko Y, Thoendel M, Olakanmi O, Britigan BE, Singh PK (2007) The transition metal gallium disrupts Pseudomonas aeruginosa iron metabolism and has antimicrobial and antibiofilm activity. J Clin Invest 117:877888–877888
Kirienko DR, Kang D, Kirienko NV (2019) Novel pyoverdine inhibitors mitigate Pseudomonas aeruginosa pathogenesis. Front Microbiol 9:3317
Konno S, Ishikawa F, Suzuki T, Dohmae N, Burkart MD, Kakeya H (2015) Active site-directed proteomic probes for adenylation domains in nonribosomal peptide synthetases. Chem Commun 51:2262–2265
Ishikawa F, Konno S, Suzuki T, Dohmae N, Kakeya H (2015) Profiling nonribosomal peptide synthetase activities using chemical proteomic probes for adenylation domains. ACS Chem Biol 10:1989–1997
Finking R, Neumüller A, Solsbacher J, Konz D, Kretzschmar G, Schweitzer M, Krumm T, Marahiel MA (2003) Aminoacyl adenylate substrate analogues for the inhibition of adenylation domains of nonribosomal peptide synthetases. Chembiochem 4:903–906
Kasai S, Konno S, Ishikawa F, Kakeya H (2015) Functional profiling of adenylation domains in nonribosomal peptide synthetases by competitive activity-based protein profiling. Chem Commun 51:15764–15767
Ishikawa F, Konno S, Uchida C, Suzuki T, Takashima K, Dohmae N, Kakeya H, Tanabe G (2022) Chemoproteomics profiling of surfactin-producing nonribosomal peptide synthetases in living bacterial cells. Cell Chem Biol 29:145–156.e8
Cravatt BF, Wright AT, Kozarich JW (2008) Activity-based protein profiling: from enzyme chemistry to proteomic chemistry. Annu Rev Biochem 77:383–414
Sanman LE, Bogyo M (2014) Activity-based profiling of proteases. Annu Rev Biochem 83:249–273
Ishikawa F, Kakeya H (2014) Specific enrichment of nonribosomal peptide synthetase module by an affinity probe for adenylation domains. Bioorg Med Chem Lett 24:865–869
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Augenstein DC, Thrasher KD, Sinskey AJ, Wang DI (1974) Optimization in the recovery of a labile intracellular enzyme. Biotechnol Bioeng 16:1433–1447
Acknowledgments
This work was partly supported by Grants-in-Aid for Research (C) (19 K05722 to F.I.) from JSPS and by grants from the Noda Institute for Scientific Research (F.I.), Research Foundation for Pharmaceutical Sciences (F.I.), Japan Foundation for Applied Enzymology (F.I.), and Institute Foundation for Science, Osaka (F.I.). We are also thankful for the financial support provided by the Antiaging Project for Private Universities.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ishikawa, F., Tanabe, G. (2023). Chemoproteomic Profiling of Adenylation Domain Functions in Gramicidin S-Producing Non-ribosomal Peptide Synthetases. In: Burkart, M., Ishikawa, F. (eds) Non-Ribosomal Peptide Biosynthesis and Engineering. Methods in Molecular Biology, vol 2670. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3214-7_4
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3214-7_4
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3213-0
Online ISBN: 978-1-0716-3214-7
eBook Packages: Springer Protocols