Abstract
In this chapter the state of the art of live cell microarrays for high-throughput biological assays are reviewed. The fabrication of novel microarrays with respect to material science and cell patterning methods is included. A main focus of the chapter is on various aspects of the application of cell microarrays by providing selected examples in research fields such as biomaterials, stem cell biology and neuroscience. Additionally, the importance of microfluidic technologies for high-throughput on-chip live-cell microarrays is highlighted for single-cell and multi-cell assays as well as for 3D tissue constructs.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Key words
1 Introduction
The implementation of micromachining and micro-scale technologies for biomedical applications enables the advanced in vitro cell analysis using cellular microarrays, microfluidic systems, and micro-scale diagnostics. The greatest benefit of miniaturized cell analysis systems is the ability to provide quantitative data in real time with high reliability and sensitivity, which are key parameters for any cell-based assay. An additional advantage of cell-based microarrays is their inherent high-throughput capability, which allows for large-scale screening of single cells, multi-cell populations, and spheroids. The interest in cell microarrays is also reflected in a rapid increase in the number of publications over a period of 10 years. Figure 1 provides an overview of number of manuscripts published between 2000 and 2013 based on article search in the ScienceDirect database using keywords such as microarray, cell microarrays, and 3D and single-cell microarrays.
Miniaturization of cell assays using cell microarrays increases not only throughput but also significantly reduces the consumption of reagents and the requirement of cellular material, which are key criteria when using clinical grade reagents and primary cell cultures. The proven cost reduction of cell-based microarrays makes them a highly attractive tool for a wide range of applications in pharmacology, toxicology and stem cell research [1]. Consequently, cell microarrays have been explored for pharmacological applications to determine gene expression, cell-to-surface interaction, extracellular matrix (ECM) production, cell migration and proliferation [2]. Additionally a variety of cell microarrays have been used to study alterations of intracellular/extracellular biochemistry, cell morphology, motility and adhesion, survival/apoptosis, and proliferative properties.
Generally, there are two strategies for the fabrication of microarrays for cell analysis and they involve either direct or indirect cell patterning approaches. The indirect method involves the placing of cells on top of pre-modified surfaces that allow for cell attachment. The most important application of indirect patterning is the so-called “reverse transfection,” developed by the Sabbatini group in 2001, where small spots of vector constructs are printed on a slide, and a cell layer is then cultivated on the slide [3]. Functional assays are subsequently performed to identify effects of gene and protein overexpression or knockdown. Indirect cell patterning requires proper surface chemistry and applied functionalization procedures become a determining factor in the successful fabrication of cell microarrays. On the other hand, the direct cell pattering approach is mainly based on integrated geometric features within the microdevice including channels, grooves, wells, and pillars that allow for efficient cell capture. A recent technological advancement of cell-based microarrays involves their combination with microfluidics to enable nutrient supply and waste removal for optimum cell culture conditions. Microfluidics is considered to be a technology that allows for the precise manipulation and control of very small fluid volumes down to pL scale. The main advantages of integrating microfluidic channels to cell-based microarrays is the ability to regulate and transport fluids, soluble factors, drug candidates, and bioactive substances at specific solution concentrations and gradients. The application of microfluidics has already shown to create new opportunities for the spatial and temporal control of cell proliferation and stimuli [4]. For instance, microfluidic cell assays have been used for conducting fast screening experiments, evaluating drug-related toxicity, and elucidating optimal cell culture conditions. The current trend towards the integration of sensory systems into cell-based microsystems leads to the creation of fully automated, highly integrated multifunctional Lab-on-a-Chip (LOC ) systems for biomedical, biomaterial, and pharmaceutical research [5–9].
2 Live-Cell Microarrays
Understanding the impact of bioactive substances on cell cultures is a fundamental aspect of many biomedical research fields ranging from cell biology studies to drug testing and development of optimized cell cultivation strategies. Originating from the DNA microarray technology, patterns of small molecules and peptides were initially used to assess their suitability as cell adhesion promoters in large-scale screening efforts. For instance, screening of cell adhesion-promoting proteins was investigated by Ito et al. in 2005 using photochemistry for microarray patterning [10].
In another early study, different cell types were seeded on a peptide array to analyze cellular activities including cell adhesion and functional phosphorylation in response to different stimuli [11]. In a similar study, a large range of peptides was screened for their potential of binding Jurkat cells on top of predefined patterns [12]. There were other functional screening studies which investigated immune responses of antigen-specific T-cells mediated by major histocompatibility complex (MHC) using the peptide-MHC arrays [13, 14]. In addition to peptides, another substance of great interest for cell interaction studies is the glycans, which are known to play important cell biological roles including cell–cell signaling and immune responses. As an example, selective binding of CD4+ T cells to carbohydrates was demonstrated by microarray screening [15]. Additionally, cell–membrane interactions have been studied in the microarray format where a large number of cell adhesion molecules were characterized in the presence of different membrane compositions [16, 17]. Furthermore, cell microarrays have also been used for functional analysis of polymers for application in cell culture handling. In 2004, Anderson et al. screened a polymer library for transfection efficiency in cancer cells with the goal to find a nonviral DNA vector for gene therapy in cancer treatment [18]. Another interesting microarray screening for polymers was aimed at identifying relationships between surface chemical structure and related protein adsorption), and the effect of polymer on cell adhesion [19].
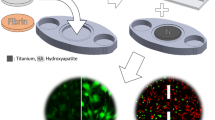
In this respect, cell microarrays demonstrated their usefulness to screen for cellular response to synthetic and natural biomaterials such as synthetic polymers and ECM-derived adhesion promoters, respectively [20–22]. The focus of these studies is based on surface chemistry and surface topography as well as biological interactions with surface-patterned biomolecules. For instance, heterogenic polymer-based microarrays can be used to identify new biomaterials that support adsorption of ECM-derived adhesion promoters, thus promoting cell adhesion [23]. Another approach implemented switchable thermo-responsive surface microarrays to support cell adhesion with consecutive nonenzymatic sample release upon temperature decrease [24, 25]. An alternative application of cell microarrays is the screening of antifouling surfaces that are able to resist bacterial adhesion, which is important to prevent biofilm formation at the surfaces of biomaterials [26–28]. Furthermore, microarrays containing different topographic patterns have been used to elucidate the interplay between surface topography and cellular behavior, and to find improved biomaterials for cell culture applications [29, 30].
The use of antibody microarrays as a platform for high-throughput screening of immune cells in blood to detect specific surface markers on immune cells constitutes a promising approach. Here, single cells and whole cell populations can readily be captured on the patterned antibody regions and functionalized microstructures because of specific cell-to-surface interactions [31, 32]. One prominent example involves the use of antibody arrays for phenotyping of blood cells to identify subpopulations of CD19+ B lymphocytes, CD16+ neutrophils, CD36+ monocytes as well as CD4+/ CD8+ T cells [33–35]. In a similar manner, antibody arrays have been applied for serotyping prokaryotic cells [36, 37]. Overall speaking, the antibody-based microarray technology can advance state-of-the-art monitoring of disease and pathological conditions (HIV, leukemia, circulating tumor cells, etc.) by increasing the sample throughput as well as the throughput of antigens to be tested.
In recent years, live-cell microarrays have mainly been employed for parallelization and high-throughput analyses in the field of cell biology, tissue engineering and tissue regeneration to gain a deeper understanding of dynamic cell response. For instance, cell-based microarrays have been applied for investigation of cell proliferation and morphology changes [38], protein expression [39], transfection of cell cultures; imaging of (single) cells and tissue constructs [40], as well as for cell–cell, cell–surface, and cell–substance interaction studies. We especially want to emphasize the importance of cell microarray technology in the field of neuroscience, stem cell research as well as cell-to-surface interaction studies. Figure 2 shows a schematic overview of life-cell microarray technology based on planar spots, cavities, and microwells as well as 3D microstructures for cell analysis applications.
With the advent of cell-based personalized therapies, cell microarrays have been increasingly applied in stem cell research for the analysis of the fate of cellular processes and of biomaterial interaction, and for the screening of cell-specific stimuli [41]. As for the fate of cellular processes, where the spatiotemporal control of the cellular microenvironment plays a major role, microarrays have been applied to control cell morphology as well as migration, differentiation, proliferation and the health status of stem cells [42–44]. As an example, the responses of human embryonic stem cells (hES) to biomaterials were investigated using microarrays revealing the impact of individual compositions of biomaterials on stem cell [20, 21]. Furthermore, ECM microarrays have been employed for high-throughput analysis of environmental factors that guide stem cell function and fate. To account for the natural 3D microenvironment of cocultures that is present in native tissues, spheroid microarrays [45] have been recently developed to improve the differentiation efficiency of multipotent mesenchymal stem cells (MSCs). In this study, the osteogenic and adipogenic differentiation efficiency as well as phenotype maintenance (see Fig. 3) was significantly increased by introduction of 100 μm-sized cell spheroid microarrays.
Comparison between monolayer culture and spheroid in the live-cell microarray technology on the capacity of mesenchymal stem cells (MSCs) to differentiate in osteogenic and adipogenic lineage (adapted from ref. 45, with permission from Elsevier)
Another prominent application of live-cell microarrays is in the field of neuroscience, where the analysis of single neurons as well as large neuronal populations is of highest interest for understanding brain development including the onset and progression of degenerative diseases. Here, live-cell microarrays are able to overcome limitations of conventional analysis technologies that mainly record neuronal data based on the activity of cellular clusters, thus only providing information on subpopulations of neurons [46]. It is important to note that spatially resolved analysis of neuronal populations have highlighted that local as well as global cell densities play a crucial role in neural network activity [47, 48]. Consequently, cell patterning approaches combined with multi-electrode arrays (MEA) containing 4,096 single electrodes have shown to retain the key properties of random neuronal networks such as transmission, short-term plasticity as well as bulk network activity [49]. In a similar manner, MEAs with an even higher spot density of 11,011 microelectrodes has been used to visualize network topography and action potential propagation of neurons upon stimulation and record dynamic responses with single-cell resolution [50]. Figure 4 shows examples of microarray layouts for the investigation of neural networks.
Live-cell microarray for applications in neuroscience. (a) Microdevice with 60 electrodes for the analysis of multiple cellular populations (A–E) (adapted from ref. 46 with permission from Elsevier). (b) Multi-electrode array MEA biosensor featuring 4096 electrodes for high-resolution measurements of neuronal networks. (c) Micro-contact printed PLL/agarose surface patterns for establishment of a symmetric network of primary neurons. Scale bar, 200 μm
3 Microfluidic Live-Cell Microarrays
As outlined in the previous section, cell-based microarrays have enabled deeper insights into important cell biological aspects such as cell-to-cell and cell-to-surface interactions, and have mainly been applied for toxicological screenings of pharmaceutical compounds and nanomaterials. Recent advancements of live-cell microarrays included the integration of microfluidic channels that allow for further miniaturization and automation of cell-based assays. Developed in the early 1980s, microfabrication and MEMS technology were revolutionized by Whitesides and colleagues in the late 1990s with the introduction of soft lithography, which made microfluidic technology available to a broad scientific community [51]. The main advantage of microfluidics for cell analysis is the incorporation of automated fluid handling routines, which allows for reproducible cell culture conditions. Some of the many benefits of microfluidics for cell culture handling include control over surface chemistry, topography, and geometry as well as the precise transport of fluids and soluble factors (growth factors etc.). Therefore, the increasing effort on combining microfluidics with various cell patterning methods has led to the development of next-generation live-cell microarrays over the last decade [52–55]. For instance, one of the major issues addressed by microfluidic live-cell microarrays is the inherent heterogeneity of cell populations by developing high-throughput single-cell microarrays that recapitulate the biological situation, thus providing in vitro data that are more relevant to in vivo situations [56].
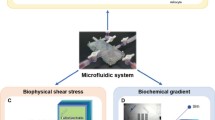
Based on the relevance of microfluidic live-cell microarrays for industrial and clinical applications, the various advancements of these microarrays for single-cell and three-dimensional assays (see Fig. 5) are described in more detail in the following two sections.
3.1 Microfluidic Single-Cell Microarrays
Practical applications of microfluidic single-cell arrays include tumor biology, stem cell biology, antibiotic resistance screening, as well as single-cell immune-typing. As indicated above, microfluidic single-cell microarrays are ideally suited to assess the heterogeneity within a cell population by analyzing the responses of a large number of individual cells to provide information on subpopulation distribution, cellular activities, and the ratio of responding and non-responding cells. It is important to highlight that the intrinsic cellular heterogeneity of single circulating tumor cells (CTCs) determines the metastatic potential, thus further highlighting the importance of single-cell analysis approaches. The main emphasis in tumor biology, therefore, is concerned with understanding the formation and growth of primary tumors, local tumor cell invasion, migration and extravasation, and finally, tumor cell metastasis. However, one of the major limitations of assessing for example the metastatic potential of CTCs is the inherent difficulty of isolating 10–100 rare cells in blood within high background of normal blood cells (109 to 1010 cells per mL). Microfluidic single-cell arrays can facilitate the study of CTCs by providing the diagnostic tools capable of isolating and analyzing CTCs using surface marker-based and marker-free methods. While surface marker-based methods predominantly employ magnetic beads for cell capture [57, 58], marker-free microfluidic isolation methods use pillars and flow focusing approaches [59–61]. A prominent example of single-cell analysis using microfluidic single-cell microarrays is the application of genotyping and mechano-typing, also called “deformability cytometry,” which has been established for identification of malignant and benign cells [62–64]. Another example of a microfluidic single-cell microarray integrates an array of PDMS-based cell-capture pockets (see Fig. 6a) that can be used to detect tumor proliferation and apoptosis following the administration of anticancer agents [65]. More recently, single-cell gene profiling has been used to identify different populations of CTCs thus highlighting that cellular heterogeneity is a major factor in cell-based assays [66]. Furthermore, it could be shown that expression profiles of CTCs diverged distinctly from those of well-established cancer cell lines, thus questioning the suitability of conventional in vitro models for drug discovery and cancer therapy research.
Similarly microfluidic single-cell microfluidic microarrays with integrated cell-capture pockets have been applied for analysis of signaling dynamics of hematopoietic stem cells as well as their cell division, time-resolved viability, and cell migration and motility analysis [67]. Another approach uses micro-arrayed 4.1 nL nano-pockets (see Fig. 6b) for investigation of rare hematopoietic stem cells, where proliferation studies of single cells have been conducted [68]. Based on these technological advances, single-cell microfluidic approaches can potentially be applied for bacterial pathogenesis research. For instance, to investigate how antibiotic resistance can arise in bacteria, the microfluidic technology has been employed that provided evidence on the establishment of resistant populations from one single bacterium [69]. In another microfluidic approach, the impact of paracrine signaling on clotting capabilities of blood was investigated using single perfused pockets inside microchannels [70].
Alternative approaches for single-cell immuno-typing are based on single-cell nanowell arrays. In one application, T cells were captured by gravity sedimentation within the nanowells and subsequently stimulated. Cell analysis was accomplished using ELISA and immunofluorescence staining to provide biological information with single-cell resolution [71, 72]. Similarly, microarrayed nanowells were applied for on-chip analysis of the secretome of CD4+ T cells [73]. In another approach implemented by electrophysiological sensor recordings on single cells with high spatiotemporal resolution, the neuronal network activity was investigated [74]. Moreover, single live-cell microarrays in combination with an array of cantilevers have been established to measure the mass of cancer cells, thus revealing information on individual cells under different physiological conditions in a noninvasive manner [75].
3.2 Microfluidic 3D Cell Microarrays
While microfluidic single-cell assays provide information on heterogeneity within a cell population, microfluidic 3D cell microarrays are used to further investigate cell-to-cell interaction and communication of heterotypic cultures including the impact of cellular secretome of individual cells on bulk cell cultures. Additional advantages of employing microfluidic 3D cell microarrays for cell culture applications include the improved in vivo-like microenvironment where cells are surrounded by ECM, are in direct contact with each other (either in a homotypic or heterotypic way) and are in combination with controlled nutrient supply and waste removal. Various studies have shown that 3D culture techniques based on aggregates, spheroids and tissue scaffolds or hydrogels help cells to retain their natural functionality. For instance, it has been shown that hepatocytes can exclusively maintain their physiologically relevant phenotype within a 3D context [76]. Additionally, mesenchymal stem cells (MSC)-derived hepatocytes exhibited key functions including urea synthesis and metabolite clearance only when cultured within a three-dimensional cellular microenvironment [77]. Moreover, the 3D spheroid systems have been applied extensively for immune-activation, as well as for engineering of native tissues including cartilage, lung, liver, kidney, gut, bone, brain, pancreatic, and cardiac organoids in vitro [78–86]. Furthermore, various spheroid cultures have been presented as viable and novel tools to establish vascular structures and to investigate network formation in vascularization research [87, 88]. More recently, there are developments that incorporate microstructures in microfabricated systems to increase the variety of functional spheroid geometries (see Fig. 7), and these microstructures include stripes, triangles, and star-shaped objects [89, 90]. Based on these advances, the microfluidic 3D cell microarrays represent a valuable tool for high-throughput screening applications such as improved drug screening and tissue engineering.
Similar to the microfluidic systems containing integrated pockets, on-chip U-shaped microstructure arrays (see Fig. 8a) have been employed as an effective method for the generation of multicellular spheroids (MCS) [91]. It is demonstrated that in situ fabrication can replace an expensive cleanroom setup for creating the PEG-based microstructures (pockets) within microchannels. The epithelial HepG2 tumor cell spheroids do respond to doxorubicin treatment, and this response could effectively demonstrate a 3D liver tissue construct is superior to the conventional 2D cultures. In a similar study, an increased chemotherapeutic resistance has been reported for spheroids generated from cells obtained from patients with terminal epithelial ovarian carcinoma, which was related to an enhanced expression of kallikrein-related peptidases in the spheroid cell culture [92]. Approaches based on the well-known hanging drop technique have also been developed for microfluidic live-cell microarray (see Fig. 8b). For instance, parallel formation of spheroids of different cell types was achieved on the hanging drop for consecutive in-line bioactivation and pharmaceutical compound evaluation assays [93]. PDMS-silicon hybrid devices containing integrated pyramid-like micro-cavity arrays were used for short-term MCF-7 breast cancer and long-term HepG2 liver spheroid culture analysis [94]. Using this microfluidic 3D cell microarray, cell viability, albumin secretion, and respiratory activity were recorded in a high-throughput manner. Another study reported a three-layer PDMS/PC membrane microfluidic system featuring integrated cell capture chambers (see Fig. 8c) capable of forming prostate cancer co-culture spheroids to recapitulate the growth behavior of PC-3 cancer cells within a bone metastatic prostate cancer microenvironment [95]. Results of study showed that spheroid culture of CD133+ PC-3 cells remained in the quiescent state and as an undifferentiated phenotype, thus preserving the relevant surface markers of cancer stem cells (CSCs). These cells are believed to play a major role in metastasis and may become a promising avenue for anticancer therapy. Other microfluidic devices as shown in Fig. 8d have utilized 3D cocultures that were embedded within a basement membrane hydrogel, rather than organoid structures [96]. Such systems have been used for chemotherapeutic drug testing using a three-dimensional hydrogel-based MCF-7 breast cancer spheroid model [97].
Microfluidic approaches to establish 3D cell cultures. (a) Microchannel with integrated U-shaped pockets for capture of cell populations and consecutive organoid formation. (b) Microfluidic devices based on formation of organoids using hanging drop techniques. (c) Microfluidic chamber arrays with cell capture pockets for organoid formation within microchannels. (d) Microfluidic devices based on cell-laden hydrogels for 3D organoid construct formation
4 Conclusion and Future Prospects
This chapter reviews live-cell microarrays as a versatile platform for high-throughput cell analysis, where cells are exposed to a range of stimuli in a highly parallelized manner. The ability to obtain a large amount of data from a single experiment using live-cell microarray technology represents an ideal approach to gain deeper insights into cellular phenotypes, which is of special relevance in the context of system biology, disease modeling and personalized medicine. In general, two major fields of applications can be defined, which can be associated with (a) screening of small substance (chemical) and genomic libraries and (b) evaluation of cell–microenvironment interactions. Consequently, live-cell microarrays have proven to be useful for a wide range of biomedical applications including investigation of cell signaling in healthy and diseased systems, cell–cell and cell–matrix interaction studies, drug screening, cell sorting, and cell phenotype characterization.
In the light of an increasing demand of robust, reliable, and reproducible cytotoxicity screening assays for disease modeling, pharmaceutical compound testing and cell-based therapies, cell microarrays are expected to play a major role in future biomedical science. The necessity for advanced in vitro cell analysis systems has therefore provided the opportunity to develop automated cell culture systems capable of monitoring single cells, multi cell populations and spheroids. Here, the combination of microfluidics with microarray technology allows for further miniaturization, automation and large volume testing even using complex biological systems. For instance, microfluidic 3D cell microarrays containing live tissue analogues can be envisioned for high-throughput drug screening with in vivo relevance. This latest trend of combining microarrays, microfluidics and 3D cell culture technology includes the reliable establishment of multi-organs-on-a-chip and human-on-a-chip systems that mimics the complex interplay of multiple organs in one single device. One prominent multi-layer multi-organ-chip system for long-term cultivation of liver and skin organoids has been applied for long-term monitoring of cellular metabolic activity including glucose consumption, lactate dehydrogenase (LDH) and lactate production in the presence of a direct long-term exposure to fluid flow [98]. In addition, the impact of troglitazole (an antidiabetic drug) on the metabolic activity was investigated over a 6-day exposure period. However, practical application of organ-on-a-chip technology for clinical testing requires to address the limitations of various components associated with systems integration such as micropumps, microheaters, microdegassers, and microsensor arrays, automation, and miniaturization. Figure 9 highlights key aspects of the need to further develop microfluidic cell microarray technology for high-throughput and high-content testing of living cell cultures.
References
Fernandes TG, Diogo MM, Clark DS, Dordick JS, Cabral JMS (2009) High-throughput cellular microarray platforms: applications in drug discovery, toxicology and stem cell research. Trends Biotechnol 27:342–349
Angres B (2005) Cell microarrays. Expert Rev Mol Diagn 5:769–779
Ziauddin J, Sabatini DM (2001) Microarrays of cells expressing defined cDNAs. Nature 411:107–110
West J, Becker M, Tombrink S, Manz A (2008) Micro total analysis systems: latest achievements. Anal Chem 80:4403–4419
Dittrich PS, Manz A (2006) Lab-on-a-chip: microfluidics in drug discovery. Nat Rev Drug Discov 5:210–218
Richter L, Charwat V, Jungreuthmayer C, Bellutti F, Brueckl H, Ertl P (2011) Monitoring cellular stress responses to nanoparticles using a lab-on-a-chip. Lab Chip 11:2551–2560
Charwat V, Rothbauer M, Tedde SF, Hayden O, Bosch JJ, Muellner P, Hainberger R, Ertl P (2013) Monitoring dynamic interactions of tumor cells with tissue and immune cells in a lab-on-a-chip. Anal Chem 85:11471–11478
Novak R, Wartmann D, Mathies RA, Dostálek J, Ertl P (2015) Microfluidic platform for multiplexed cell sampling and time-resolved spr-based cytokine sensing. In: Lacković I, Vasic D (eds.) 6th European conference of the international federation for medical and biological engineering—MBEC 2014, 7–11 September 2014, Dubrovnik, Croatia, Springer. pp. 785–788
Ertl P, Sticker D, Charwat V, Kasper C, Lepperdinger G (2014) Lab-on-a-chip technologies for stem cell analysis. Trends Biotechnol 32:245–253
Ito Y, Nogawa M, Takeda M, Shibuya T (2005) Photo-reactive polyvinylalcohol for photo-immobilized microarray. Biomaterials 26:211–216
Falsey JR, Renil M, Park S, Li S, Lam KS (2001) Peptide and small molecule microarray for high throughput cell adhesion and functional assays. Bioconjug Chem 12:346–353
Xu QC, Miyamoto S, Lam KS (2004) A novel approach to chemical microarray using ketone-modified macromolecular scaffolds: application in micro cell-adhesion assay. Mol Divers 8:301–310
Soen Y, Chen DS, Kraft DL, Davis MM, Brown PO (2003) Detection and characterization of cellular immune responses using peptide-MHC microarrays. PLoS Biol 1:E65
Stone JD, Demkowicz WE Jr, Stern LJ (2005) HLA-restricted epitope identification and detection of functional T cell responses by using MHC-peptide and costimulatory microarrays. Proc Natl Acad Sci U S A 102:3744–3749
Nimrichter L, Gargir A, Gortler M, Altstock RT, Shtevi A, Weisshaus O, Fire E, Dotan N, Schnaar RL (2004) Intact cell adhesion to glycan microarrays. Glycobiology 14:197–203
Hovis JS, Boxer SG (2001) Patterning and composition arrays of supported lipid bilayers by microcontact printing. Langmuir 17:3400–3405
Groves JT, Mahal LK, Bertozzi CR (2001) Control of cell adhesion and growth with micropatterned supported lipid membranes. Langmuir 17:5129–5133
Anderson DG, Peng W, Akinc A, Hossain N, Kohn A, Padera R, Langer R, Sawicki JA (2004) A polymer library approach to suicide gene therapy for cancer. Proc Natl Acad Sci U S A 101:16028–16033
Yang J, Mei Y, Hook AL, Taylor M, Urquhart AJ, Bogatyrev SR, Langer R, Anderson DG, Davies MC, Alexander MR (2010) Polymer surface functionalities that control human embryoid body cell adhesion revealed by high throughput surface characterization of combinatorial material microarrays. Biomaterials 31:8827–8838
Anderson DG, Levenberg S, Langer R (2004) Nanoliter-scale synthesis of arrayed biomaterials and application to human embryonic stem cells. Nat Biotechnol 22:863–866
Flaim CJ, Chien S, Bhatia SN (2005) An extracellular matrix microarray for probing cellular differentiation. Nat Methods 2:119–125
Ankam S, Teo BKK, Kukumberg M, Yim EKF (2013) High throughput screening to investigate the interaction of stem cells with their extracellular microenvironment. Organogenesis 9:128–142
Mei Y, Saha K, Bogatyrev SR, Yang J, Hook AL, Kalcioglu ZI, Cho SW, Mitalipova M, Pyzocha N, Rojas F, Van Vliet KJ, Davies MC, Alexander MR, Langer R, Jaenisch R, Anderson DG (2010) Combinatorial development of biomaterials for clonal growth of human pluripotent stem cells. Nat Mater 9:768–778
Zhang R, Mjoseng HK, Hoeve MA, Bauer NG, Pells S, Besseling R, Velugotla S, Tourniaire G, Kishen REB, Tsenkina Y, Armit C, Duffy CRE, Helfen M, Edenhofer F, de Sousa PA, Bradley M (2013) A thermoresponsive and chemically defined hydrogel for long-term culture of human embryonic stem cells. Nat Commun 4(1335):1–10
Cheng XH, Wang YB, Hanein Y, Bohringer KF, Ratner BD (2004) Novel cell patterning using microheater-controlled thermoresponsive plasma films. J Biomed Mater Res A 70A:159–168
Hook AL, Chang CY, Yang J, Atkinson S, Langer R, Anderson DG, Davies MC, Williams P, Alexander MR (2013) Discovery of novel materials with broad resistance to bacterial attachment using combinatorial polymer microarrays. Adv Mater 25:2542–2547
Anglin E, Davey R, Herrid M, Hope S, Kurkuri M, Pasic P, Hor M, Fenech M, Thissen H, Voelcker NH (2010) Cell microarrays for the screening of factors that allow the enrichment of bovine testicular cells. Cytometry Part A: the journal of the International Society for Analytical Cytology 77:881–889
Pernagallo S, Wu M, Gallagher MP, Bradley M (2011) Colonising new frontiers-microarrays reveal biofilm modulating polymers. J Mater Chem 21:96–101
Unadkat HV, Hulsman M, Cornelissen K, Papenburg BJ, Truckenmuller RK, Carpenter AE, Wessling M, Post GF, Uetz M, Reinders MJ, Stamatialis D, van Blitterswijk CA, de Boer J (2011) An algorithm-based topographical biomaterials library to instruct cell fate. Proc Natl Acad Sci U S A 108:16565–16570
Moe AA, Suryana M, Marcy G, Lim SK, Ankam S, Goh JZ, Jin J, Teo BK, Law JB, Low HY, Goh EL, Sheetz MP, Yim EK (2012) Microarray with micro- and nano-topographies enables identification of the optimal topography for directing the differentiation of primary murine neural progenitor cells. Small 8:3050–3061
Ohnaga T, Shimada Y, Moriyama M, Kishi H, Obata T, Takata K, Okumura T, Nagata T, Muraguchi A, Tsukada K (2013) Polymeric microfluidic devices exhibiting sufficient capture of cancer cell line for isolation of circulating tumor cells. Biomed Microdevices 15:611–616
Li N, Ho CM (2008) Photolithographic patterning of organosilane monolayer for generating large area two-dimensional B lymphocyte arrays. Lab Chip 8:2105–2112
Ellmark P, Hogerkorp CM, Ek S, Belov L, Berglund M, Rosenquist R, Christopherson RI, Borrebaeck CAK (2008) Phenotypic protein profiling of different B cell sub-populations using antibody CD-microarrays. Cancer Lett 265:98–106
Sekine K, Revzin A, Tompkins RG, Toner M (2006) Panning of multiple subsets of leukocytes on antibody-decorated poly(ethylene) glycol-coated glass slides. J Immunol Methods 313:96–109
Wu JQ, Wang B, Belov L, Chrisp J, Learmont J, Dyer WB, Zaunders J, Cunningham AL, Dwyer DE, Saksena NK (2007) Antibody microarray analysis of cell surface antigens on CD4+ and CD8+ T cells from HIV+ individuals correlates with disease stages. Retrovirology 4:83
Gao J, Liu C, Liu D, Wang Z, Dong S (2010) Antibody microarray-based strategies for detection of bacteria by lectin-conjugated gold nanoparticle probes. Talanta 81:1816–1820
Marimon JM, Monasterio A, Ercibengoa M, Pascual J, Prieto I, Simon L, Perez-Trallero E (2010) Antibody microarray typing, a novel technique for Streptococcus pneumoniae serotyping. J Microbiol Methods 80:274–280
Niu W, Narayanaswamy R, Scouras A, Hart GT, Davies J, Ellington AD, Iyer VR, Marcotte EM (2006) Systematic profiling of cellular phenotypes with spotted cell microarrays reveals new mating pheromone response genes. Faseb J 20:A928–A928
Bochner BR, Gadzinski P, Panomitros E (2001) Phenotype MicroArrays for high-throughput phenotypic testing and assay of gene function. Genome Res 11:1246–1255
Schwenk JM, Stoll D, Templin MF, Joos TO (2002) Cell microarrays: an emerging technology for the characterization of antibodies. Biotechniques 33:S54–S61
Chin VI, Taupin P, Sanga S, Scheel J, Gage FH, Bhatia SN (2004) Microfabricated platform for studying stem cell fates. Biotechnol Bioeng 88:399–415
Woodruff K, Fidalgo LM, Gobaa S, Lutolf MP, Maerkl SJ (2013) Live mammalian cell arrays. Nat Methods 10:550–552
Hong BJ, Sunkara V, Park JW (2005) DNA microarrays on nanoscale-controlled surface. Nucleic Acids Res 33:e106
Sunkara V, Hong BJ, Park JW (2007) Sensitivity enhancement of DNA microarray on nano-scale controlled surface by using a streptavidin-fluorophore conjugate. Biosens Bioelectron 22:1532–1537
Wang WJ, Itaka K, Ohba S, Nishiyama N, Chung UI, Yamasaki Y, Kataoka K (2009) 3D spheroid culture system on micropatterned substrates for improved differentiation efficiency of multipotent mesenchymal stem cells. Biomaterials 30:2705–2715
Berdondini L, Chippalone M, van der Wal PD, Imfeld K, de Rooij NF, Koudelka-Hep M, Tedesco M, Martinoia S, van Pelt J, Le Masson G, Garenne A (2006) A microelectrode array (MEA) integrated with clustering structures for investigating in vitro neurodynamics in confined interconnected sub-populations of neurons. Sensor Actuat B-Chem 114:530–541
Maccione A, Gandolfo M, Tedesco M, Nieus T, Imfeld K, Martinoia S, Berdondini L (2010) Experimental investigation on spontaneously active hippocampal cultures recorded by means of high-density MEAs: analysis of the spatial resolution effects. Frontiers in Neuroengineering 3(4):1–12
Maccione A, Garofalo M, Nieus T, Tedesco M, Berdondini L, Martinoia S (2012) Multiscale functional connectivity estimation on low-density neuronal cultures recorded by high-density CMOS Micro Electrode Arrays. J Neurosci Methods 207:161–171
Marconi E, Nieus T, Maccione A, Valente P, Simi A, Messa M, Dante S, Baldelli P, Berdondini L, Benfenati F (2012) Emergent functional properties of neuronal networks with controlled topology. PLoS One 7:e34648
Bakkum DJ, Frey U, Radivojevic M, Russell TL, Muller J, Fiscella M, Takahashi H, Hierlemann A (2013) Tracking axonal action potential propagation on a high-density microelectrode array across hundreds of sites. Nat Commun 4(2181):1–12
Xia YN, Whitesides GM (1998) Soft lithography. Angew Chem Int Ed 37:551–575
Khademhosseini A, Yeh J, Jon S, Eng G, Suh KY, Burdick JA, Langer R (2004) Molded polyethylene glycol microstructures for capturing cells within microfluidic channels. Lab Chip 4:425–430
Chiu DT, Jeon NL, Huang S, Kane RS, Wargo CJ, Choi IS, Ingber DE, Whitesides GM (2000) Patterned deposition of cells and proteins onto surfaces by using three-dimensional microfluidic systems. Proc Natl Acad Sci U S A 97:2408–2413
Khademhosseini A, Suh KY, Jon S, Eng G, Yeh J, Chen GJ, Langer R (2004) A soft lithographic approach to fabricate patterned microfluidic channels. Anal Chem 76:3675–3681
Whitesides GM, Ostuni E, Takayama S, Jiang X, Ingber DE (2001) Soft lithography in biology and biochemistry. Annu Rev Biomed Eng 3:335–373
Narsinh KH, Sun N, Sanchez-Freire V, Lee AS, Almeida P, Hu SJ, Jan T, Wilson KD, Leong D, Rosenberg J, Yao M, Robbins RC, Wu JC (2011) Single cell transcriptional profiling reveals heterogeneity of human induced pluripotent stem cells. J Clin Invest 121:1217–1221
Kang JH, Krause S, Tobin H, Mammoto A, Kanapathipillai M, Ingber DE (2012) A combined micromagnetic-microfluidic device for rapid capture and culture of rare circulating tumor cells. Lab Chip 12:2175–2181
Nagrath S, Sequist LV, Maheswaran S, Bell DW, Irimia D, Ulkus L, Smith MR, Kwak EL, Digumarthy S, Muzikansky A, Ryan P, Balis UJ, Tompkins RG, Haber DA, Toner M (2007) Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450:1235–1239
Karabacak NM, Spuhler PS, Fachin F, Lim EJ, Pai V, Ozkumur E, Martel JM, Kojic N, Smith K, Chen PI, Yang J, Hwang H, Morgan B, Trautwein J, Barber TA, Stott SL, Maheswaran S, Kapur R, Haber DA, Toner M (2014) Microfluidic, marker-free isolation of circulating tumor cells from blood samples. Nat Protoc 9:694–710
Hur SC, Mach AJ, Di Carlo D (2011) High-throughput size-based rare cell enrichment using microscale vortices. Biomicrofluidics 5(022206):1–10
Mach AJ, Kim JH, Arshi A, Hur SC, Di Carlo D (2011) Automated cellular sample preparation using a Centrifuge-on-a-Chip. Lab Chip 11:2827–2834
Dalerba P, Kalisky T, Sahoo D, Rajendran PS, Rothenberg ME, Leyrat AA, Sim S, Okamoto J, Johnston DM, Qian DL, Zabala M, Bueno J, Neff NF, Wang JB, Shelton AA, Visser B, Hisamori S, Shimono Y, van de Wetering M, Clevers H, Clarke MF, Quake SR (2011) Single-cell dissection of transcriptional heterogeneity in human colon tumors. Nat Biotechnol 29:1120–1127
Gossett DR, Tse HTK, Lee SA, Ying Y, Lindgren AG, Yang OO, Rao JY, Clark AT, Di Carlo D (2012) Hydrodynamic stretching of single cells for large population mechanical phenotyping. Proc Natl Acad Sci U S A 109:7630–7635
Tse HTK, Gossett DR, Moon YS, Masaeli M, Sohsman M, Ying Y, Mislick K, Adams RP, Rao JY, Di Carlo D (2013) Quantitative diagnosis of malignant pleural effusions by single-cell mechanophenotyping. Sci Transl Med 5(212):212ra163, 1–9
Wlodkowic D, Faley S, Zagnoni M, Wikswo JP, Cooper JM (2009) Microfluidic single-cell array cytometry for the analysis of tumor apoptosis. Anal Chem 81:5517–5523
Powell AA, Talasaz AH, Zhang H, Coram MA, Reddy A, Deng G, Telli ML, Advani RH, Carlson RW, Mollick JA, Sheth S, Kurian AW, Ford JM, Stockdale FE, Quake SR, Pease RF, Mindrinos MN, Bhanot G, Dairkee SH, Davis RW, Jeffrey SS (2012) Single cell profiling of circulating tumor cells: transcriptional heterogeneity and diversity from breast cancer cell lines. PLoS One 7:e33788
Faley SL, Copland M, Wlodkowic D, Kolch W, Seale KT, Wikswo JP, Cooper JM (2009) Microfluidic single cell arrays to interrogate signalling dynamics of individual, patient-derived hematopoietic stem cells. Lab Chip 9:2659–2664
Lecault V, VanInsberghe M, Sekulovic S, Knapp DJHF, Wohrer S, Bowden W, Viel F, McLaughlin T, Jarandehei A, Miller M, Falconnet D, White AK, Kent DG, Copley MR, Taghipour F, Eaves CJ, Humphries RK, Piret JM, Hansen CL (2011) High-throughput analysis of single hematopoietic stem cell proliferation in microfluidic cell culture arrays. Nat Methods 8:581–586
Boedicker JQ, Li L, Kline TR, Ismagilov RF (2008) Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics. Lab Chip 8:1265–1272
Kastrup CJ, Boedicker JQ, Pomerantsev AP, Moayeri M, Bian Y, Pompano RR, Kline TR, Sylvestre P, Shen F, Leppla SH, Tang WJ, Ismagilov RF (2008) Spatial localization of bacteria controls coagulation of human blood by ‘quorum acting’. Nat Chem Biol 4:742–750
Han Q, Bagheri N, Bradshaw EM, Hafler DA, Lauffenburger DA, Love JC (2012) Polyfunctional responses by human T cells result from sequential release of cytokines. Proc Natl Acad Sci U S A 109:1607–1612
Yamanaka YJ, Szeto GL, Gierahn TM, Forcier TL, Benedict KF, Brefo MSN, Lauffenburger DA, Irvine DJ, Love JC (2012) Cellular barcodes for efficiently profiling single-cell secretory responses by microengraving. Anal Chem 84:10531–10536
Jin A, Ozawa T, Tajiri K, Obata T, Kondo S, Kinoshita K, Kadowaki S, Takahashi K, Sugiyama T, Kishi H, Muraguchi A (2009) A rapid and efficient single-cell manipulation method for screening antigen-specific antibody-secreting cells from human peripheral blood. Nat Med 15:1088–1092
Berdondini L, Imfeld K, Maccione A, Tedesco M, Neukom S, Koudelka-Hep M, Martinoia S (2009) Active pixel sensor array for high spatio-temporal resolution electrophysiological recordings from single cell to large scale neuronal networks. Lab Chip 9:2644–2651
Park K, Jang J, Irimia D, Sturgis J, Lee J, Robinson JP, Toner M, Bashir R (2008) ‘Living cantilever arrays’ for characterization of mass of single live cells in fluids. Lab Chip 8:1034–1041
Bierwolf J, Lutgehetmann M, Feng K, Erbes J, Deichmann S, Toronyi E, Stieglitz C, Nashan B, Ma PX, Pollok JM (2011) Primary rat hepatocyte culture on 3D nanofibrous polymer scaffolds for toxicology and pharmaceutical research. Biotechnol Bioeng 108:141–150
Li J, Tao R, Wu W, Cao HC, Xin JJ, Li J, Guo J, Jiang LY, Gao CY, Demetriou AA, Farkas DL, Li LJ (2010) 3D PLGA scaffolds improve differentiation and function of bone marrow mesenchymal stem cell-derived hepatocytes. Stem Cells Dev 19:1427–1436
Salmenpera P, Kankuri E, Bizik J, Siren V, Virtanen I, Takahashi S, Leiss M, Fassler R, Vaheri A (2008) Formation and activation of fibroblast spheroids depend on fibronectin-integrin interaction. Exp Cell Res 314:3444–3452
Bao BA, Lai CP, Naus CC, Morgan JR (2012) Pannexin1 drives multicellular aggregate compaction via a signaling cascade that remodels the actin cytoskeleton. J Biol Chem 287:8407–8416
Jukes JM, Both SK, Leusink A, Sterk LMT, Van Blitterswijk CA, De Boer J (2008) Endochondral bone tissue engineering using embryonic stem cells. Proc Natl Acad Sci U S A 105:6840–6845
Lumelsky N, Blondel O, Laeng P, Velasco I, Ravin R, McKay R (2001) Differentiation of embryonic stem cells to insulin-secreting structures similar to pancreatic islets. Science 292:1389–1394
Kehat I, Kenyagin-Karsenti D, Snir M, Segev H, Amit M, Gepstein A, Livne E, Binah O, Itskovitz-Eldor J, Gepstein L (2001) Human embryonic stem cells can differentiate into myocytes with structural and functional properties of cardiomyocytes. J Clin Invest 108:407–414
Carraro A, Hsu WM, Kulig KM, Cheung WS, Miller ML, Weinberg EJ, Swart EF, Kaazempur-Mofrad M, Borenstein JT, Vacanti JP, Neville C (2008) In vitro analysis of a hepatic device with intrinsic microvascular-based channels. Biomed Microdevices 10:795–805
Jang KJ, Cho HS, Kang do H, Bae WG, Kwon TH, Suy KY (2011) Fluid-shear-stress-induced translocation of aquaporin-2 and reorganization of actin cytoskeleton in renal tubular epithelial cells. Integrative biology : quantitative biosciences from nano to macro 3:134–141
Kimura H, Yamamoto T, Sakai H, Sakai Y, Fujii T (2008) An integrated microfluidic system for long-term perfusion culture and on-line monitoring of intestinal tissue models. Lab Chip 8:741–746
Park J, Koito H, Li J, Han A (2009) Microfluidic compartmentalized co-culture platform for CNS axon myelination research. Biomed Microdevices 11:1145–1153
Gu A, Shively JE (2011) Angiopoietins-1 and -2 play opposing roles in endothelial sprouting of embryoid bodies in 3D culture and their receptor Tie-2 associates with the cell-cell adhesion molecule PECAM1. Exp Cell Res 317:2171–2182
Rouwkema J, de Boer J, Van Blitterswijk CA (2006) Endothelial cells assemble into a 3-dimensional prevascular network in a bone tissue engineering construct. Tissue Eng 12:2685–2693
Rivron NC, Vrij EJ, Rouwkema J, Le Gac S, van den Berg A, Truckenmuller RK, van Blitterswijk CA (2012) Tissue deformation spatially modulates VEGF signaling and angiogenesis. Proc Natl Acad Sci U S A 109:6886–6891
Khademhosseini A, Eng G, Yeh J, Kucharczyk PA, Langer R, Vunjak-Novakovic G, Radisic M (2007) Microfluidic patterning for fabrication of contractile cardiac organoids. Biomed Microdevices 9:149–157
Fu CY, Tseng SY, Yang SM, Hsu L, Liu CH, Chang HY (2014) A microfluidic chip with a U-shaped microstructure array for multicellular spheroid formation, culturing and analysis. Biofabrication 6(1):015009. doi:10.1088/1758-5082/6/1/015009
Dong Y, Tan OL, Loessner D, Stephens C, Walpole C, Boyle GM, Parsons PG, Clements JA (2010) Kallikrein-related peptidase 7 promotes multicellular aggregation via the alpha(5)beta(1) integrin pathway and paclitaxel chemoresistance in serous epithelial ovarian carcinoma. Cancer Res 70:2624–2633
Frey O, Misun PM, Fluri DA, Hengstler JG, Hierlemann A (2014) Reconfigurable microfluidic hanging drop network for multi-tissue interaction and analysis. Nat Commun 5:4250. doi:10.1038/ncomms5250
Torisawa YS, Takagi A, Nashimoto Y, Yasukawa T, Shiku H, Matsue T (2007) A multicellular spheroid array to and viability realize spheroid formation, culture, assay on a chip. Biomaterials 28:559–566
Hsiao AY, Torisawa YS, Tung YC, Sud S, Taichman RS, Pienta KJ, Takayama S (2009) Microfluidic system for formation of PC-3 prostate cancer co-culture spheroids. Biomaterials 30:3020–3027
Xu Z, Gao Y, Hao Y, Li E, Wang Y, Zhang J, Wang W, Gao Z, Wang Q (2013) Application of a microfluidic chip-based 3D co-culture to test drug sensitivity for individualized treatment of lung cancer. Biomaterials 34:4109–4117
Torisawa YS, Shiku H, Yasukawa T, Nishizawa M, Matsue T (2005) Multi-channel 3-D cell culture device integrated on a silicon chip for anticancer drug sensitivity test. Biomaterials 26:2165–2172
Wagner I, Materne EM, Brincker S, Sussbier U, Fradrich C, Busek M, Sonntag F, Sakharov DA, Trushkin EV, Tonevitsky AG, Lauster R, Marx U (2013) A dynamic multi-organ-chip for long-term cultivation and substance testing proven by 3D human liver and skin tissue co-culture. Lab Chip 13:3538–3547
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Rothbauer, M., Charwat, V., Ertl, P. (2016). Cell Microarrays for Biomedical Applications. In: Li, P., Sedighi, A., Wang, L. (eds) Microarray Technology. Methods in Molecular Biology, vol 1368. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-3136-1_19
Download citation
DOI: https://doi.org/10.1007/978-1-4939-3136-1_19
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3135-4
Online ISBN: 978-1-4939-3136-1
eBook Packages: Springer Protocols