Abstract
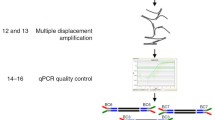
Multiple analyses such as DNA profiling, sequencing, or comparative genome hybridization (CGH) done on the single-cell level long for pre-amplification due to the diploid human genome. Isothermal whole genome amplification allows amplification of long DNA templates from single cells. When analysis needs to be performed under rare cell conditions additional care needs to be taken due to the fact that, even after pre-enrichment, few candidate target cells are still dispersed among an overwhelming number of non-target background cells. Here, we describe a protocol where we define a population of candidate target cells based on specific staining. Candidate cells are then isolated by laser microdissection and pressure catapulting (LMPC) and transferred onto a microliter reaction slide. This slide allows monitoring the single-cell isolation process and isothermal whole genome amplification in less than 2 μL. The amplification products obtained from single cells can be forwarded to multiple analyses.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Alix-Panabieres C, Pantel K (2014) Technologies for detection of circulating tumor cells: facts and vision. Lab Chip 14(1):57–62. doi:10.1039/c3lc50644d
Vona G, Beroud C, Benachi A, Quenette A, Bonnefont JP, Romana S, Dumez Y, Lacour B, Paterlini-Brechot P (2002) Enrichment, immunomorphological, and genetic characterization of fetal cells circulating in maternal blood. Am J Pathol 160(1):51–58
Vona G, Sabile A, Louha M, Sitruk V, Romana S, Schutze K, Capron F, Franco D, Pazzagli M, Vekemans M, Lacour B, Brechot C, Paterlini-Brechot P (2000) Isolation by size of epithelial tumor cells: a new method for the immunomorphological and molecular characterization of circulatingtumor cells. Am J Pathol 156(1):57–63
Nagrath S, Sequist LV, Maheswaran S, Bell DW, Irimia D, Ulkus L, Smith MR, Kwak EL, Digumarthy S, Muzikansky A, Ryan P, Balis UJ, Tompkins RG, Haber DA, Toner M (2007) Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450(7173):1235–1239. doi:10.1038/nature06385
Liu Z, Fusi A, Klopocki E, Schmittel A, Tinhofer I, Nonnenmacher A, Keilholz U (2011) Negative enrichment by immunomagnetic nanobeads for unbiased characterization of circulating tumor cells from peripheral blood of cancer patients. J Transl Med 9:70. doi:10.1186/1479-5876-9-70
Kroneis T, Gutstein-Abo L, Kofler K, Hartmann M, Hartmann P, Alunni-Fabbroni M, Walcher W, Dohr G, Petek E, Guetta E, Sedlmayr P (2010) Automatic retrieval of single microchimeric cells and verification of identity by on-chip multiplex PCR. J Cell Mol Med 14(4):954–969
Kroneis T, Geigl JB, El-Heliebi A, Auer M, Ulz P, Schwarzbraun T, Dohr G, Sedlmayr P (2011) Combined molecular genetic and cytogenetic analysis from single cells after isothermal whole-genome amplification. Clin Chem 57(7):1032–1041
Hoshino K, Huang YY, Lane N, Huebschman M, Uhr JW, Frenkel EP, Zhang X (2011) Microchip-based immunomagnetic detection of circulating tumor cells. Lab Chip 11(20):3449–3457. doi:10.1039/c1lc20270g
Stott SL, Hsu CH, Tsukrov DI, Yu M, Miyamoto DT, Waltman BA, Rothenberg SM, Shah AM, Smas ME, Korir GK, Floyd FP Jr, Gilman AJ, Lord JB, Winokur D, Springer S, Irimia D, Nagrath S, Sequist LV, Lee RJ, Isselbacher KJ, Maheswaran S, Haber DA, Toner M (2010) Isolation of circulating tumor cells using a microvortex-generating herringbone-chip. Proc Natl Acad Sci U S A 107(43):18392–18397. doi:10.1073/pnas.1012539107
Andreopoulou E, Yang LY, Rangel KM, Reuben JM, Hsu L, Krishnamurthy S, Valero V, Fritsche HA, Cristofanilli M (2012) Comparison of assay methods for detection of circulating tumor cells in metastatic breast cancer: AdnaGen AdnaTest BreastCancer Select/Detect versus Veridex Cell Search system. Int J Cancer 130(7):1590–1597. doi:10.1002/ijc.26111
Zhang L, Riethdorf S, Wu G, Wang T, Yang K, Peng G, Liu J, Pantel K (2012) Meta-analysis of the prognostic value of circulating tumor cells in breast cancer. Clin Cancer Res 18(20):5701–5710. doi:10.1158/1078-0432.ccr-12-1587
Alix-Panabieres C, Pantel K (2014) Challenges in circulating tumour cell research. Nat Rev Cancer 14(9):623–631. doi:10.1038/nrc3820
El-Heliebi A, Kroneis T, Zohrer E, Haybaeck J, Fischereder K, Kampel-Kettner K, Zigeuner R, Pock H, Riedl R, Stauber R, Geigl JB, Huppertz B, Sedlmayr P, Lackner C (2013) Are morphological criteria sufficient for the identification of circulating tumor cells in renal cancer? J Transl Med 11:214. doi:10.1186/1479-5876-11-214
El-Heliebi A, Kroneis T, Wagner K, Meditz K, Kolb D, Feichtinger J, Thallinger GG, Quehenberger F, Liegl-Atzwanger B, Rinner B (2014) Resolving tumor heterogeneity: genes involved in chordoma cell development identified by low-template analysis of morphologically distinct cells. PLoS One 9(2), e87663. doi:10.1371/journal.pone.0087663
Acknowledgement
This work was supported by the EU SAFE Network of Excellence (LSHB-CT-2004-503243, EU 6th Framework Package) and the County of Styria, Austria.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Kroneis, T., Chen, S., El-Heliebi, A. (2015). Low-Volume On-Chip Single-Cell Whole Genome Amplification for Multiple Subsequent Analyses. In: Kroneis, T. (eds) Whole Genome Amplification. Methods in Molecular Biology, vol 1347. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2990-0_17
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2990-0_17
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2989-4
Online ISBN: 978-1-4939-2990-0
eBook Packages: Springer Protocols