Abstract
The development of a facile genome engineering technology based on transcription activator-like effector nucleases (TALENs) has led to significant advances in diverse areas of science and medicine. In this review, we provide a broad overview of the development of TALENs and the use of this technology in basic science, biotechnology, and biomedical applications. This includes the discovery of DNA recognition by TALEs, engineering new TALE proteins to diverse targets, general advances in nuclease-based editing strategies, and challenges that are specific to various applications of the TALEN technology. We review examples of applying TALENs for studying gene function and regulation, generating disease models, and developing gene therapies. The current status of genome editing and future directions for other uses of these technologies are also discussed.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Key words
1 Introduction
The emergence of transcription activator-like effector nucleases (TALENs) has made genome-editing tools widely accessible to any laboratory with basic molecular biology expertise. The development of the TALEN technology and its use in various biotechnological applications build on the considerable progress in genome editing over the previous decade with other approaches. Accordingly, the availability of the TALEN technology over the past few years has led to numerous advances in genome editing in a diverse range of cell types and organisms. This facile genome editing approach has facilitated new strategies to model disease, develop novel genetic therapies, or create desired phenotypic properties through highly specific rewriting of the genome. In this chapter, the development and use of TALEN technologies are reviewed and discussed.
1.1 Genome-Editing Systems
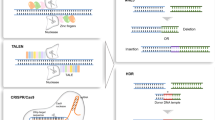
Genome editing with engineered site-specific endonucleases has emerged as a technology to selectively replace or correct disrupted genes, in contrast to conventional genetic engineering methods of gene addition [1, 2]. There are numerous platforms for generating site-specific gene modifications in the genome, but to date the most successful have been based on zinc finger nucleases [3, 4], TALENs [5, 6], and, more recently, the RNA-guided CRISPR/Cas9 system [7–9]. These systems are at present the most developed publicly available platforms for robust and efficient targeted gene editing. In particular, the recent development of TALENs and CRISPR/Cas9 has dramatically advanced genome editing due to their ease of engineering and efficient genetic modification [6–20]. Other systems in development include meganucleases [21, 22], triplex-forming oligonucleotide (TFO) complexes [23], and programmable recombinases based on zinc finger protein [24–27] or TALE DNA-binding domains [28]. Historically, meganucleases have been difficult to engineer due to interdependence of the DNA-binding and cleavage domains, although recent developments in directed evolution of meganucleases [29–31] and fusion of meganucleases to TALE DNA-binding proteins [32, 33] are providing promising new opportunities with this technology. TFO complexes have thus far been limited by relatively low levels of gene modification, but oligonucleotide-mediated gene editing can be improved with the incorporation of TALENs [34]. Programmable recombinases are a promising next-generation gene editing technology, but target site requirements, overall efficiency, and unknown off-target effects are still major challenges to the widespread adoption of this technology [35].
1.2 Nuclease-Mediated Genome Editing
Engineered nucleases generate targeted genome modifications by creating a targeted double-strand break in the genome that stimulates cellular DNA repair through either homology-directed repair (HDR) or nonhomologous end joining (NHEJ) [36, 37] (Fig. 1). Briefly, HDR uses a designed synthetic donor DNA template to guide repair and can be used to create specific sequence changes to genome, including the targeted addition of whole genes. HDR has enabled integration of gene cassettes of up to 8 kb in the absence of selection at high frequency (~6 %) in human cells [38]. Generally, gene correction strategies have been based solely on HDR, the efficiency of which is dependent on the genomic target, cell type, cell-cycle state, and efficient delivery of an exogenous DNA template [39–43]. In many cases, antibiotic selection is used in tandem with genome editing for gene correction in cell types with low levels of HDR repair [40–42]. In contrast to genome modification by HDR, the template-independent religation of DNA ends by NHEJ is a stochastic, error-prone repair process that introduces random small insertions and deletions at the DNA breakpoint (Fig. 1). Gene editing by NHEJ has been used in mammalian cells to disrupt genes [44, 45], delete chromosomal segments [46–48], or restore aberrant reading frames [49, 50]. This chapter reviews how TALENs have been used to exploit NHEJ and HDR DNA repair processes to create highly specific changes to a desired gene.
Mechanisms of DNA repair following nuclease-induced double-strand breaks. (a) In the absence of a DNA repair template, the break is repaired by nonhomologous end joining, which is an error-prone process and can lead to small insertions or deletions. Alternatively, two adjacent nuclease-induced breaks can be used to excise the intervening chromosomal DNA from the genome. (b) If a DNA repair template is provided with homology to the target site surrounding the break, it will be used to guide homology-directed repair. In this way, particular small changes to the DNA sequence or the insertion of whole-gene expression cassettes can be directed to specific genome target sites
2 Development of TALENs
2.1 TALE DNA Recognition
In 2009, two landmark studies described the simple and modular TALE DNA-binding domain [14, 15]. These novel DNA-binding proteins are naturally occurring transcriptional activators from the plant pathogen Xanthomonas. As reported in these studies, the TALE DNA-binding domain consists of numerous tandem repeats, with each repeat specifying recognition of a single base pair of DNA. Importantly, single-base-pair recognition by each repeat is determined by alteration of only two hypervariable amino acids, termed repeat variable diresidues (RVDs), and each repeat appears to recognize DNA in a modular manner. This simple mode of DNA recognition was confirmed in structural studies of a naturally occurring TALE bound to its cognate DNA target [51, 52]. These discoveries were quickly expanded upon to create novel TALE proteins by engineering a custom RVD array to recognize a user-specified DNA target [53–55]. The only sequence requirement for TALE binding is that each target site be immediately preceded by a 5′-thymine for efficient DNA recognition, although more recently modified proteins have been developed to accept other nucleotides at this position [56, 57]. These novel DNA-binding domains were then fused to transcriptional activator domains [53, 55], nuclease catalytic domains [11, 54, 55], epigenetic modifying domains [58, 59], and recombinases [28] to generate an array of programmable enzymes for manipulating genes in complex genomes.
Although naturally occurring TALEs have a modular RVD recognition code, several studies have shown that some RVDs, specifically those targeting guanosine, display unexpected recognition of degenerate bases in the context of engineered TALE DNA-binding domains [55, 60]. However, more specific RVDs, such as NH or NK for recognition of guanosine, can result in significantly reduced activity of the reengineered TALE protein [60–62]. Recently, a publicly available web server has been developed that generates TALE targets utilizing more specific RVDs predicted to have minimal impact on activity [63]. Other publicly available web servers are available to assist in generating RVD arrays that are predicted to have high activity and specificity [64–66]. Together, these studies demonstrate the overall robustness of TALE DNA recognition and its utility in generating highly active nucleases at novel targets of interest.
2.2 Assembly of RVD Arrays to Create Customized TALE DNA-Binding Domains
Synthesizing custom TALE DNA-binding domains requires contiguous assembly of many RVD repeats, each only differing by two amino acids, into a destination TALE array. The large number of repeats, typically 15–20, makes this process difficult with conventional recombinant DNA technology. To overcome this technical challenge, several approaches have been developed that iteratively assemble new TALE arrays in a highly efficient and rapid manner. Custom TALE arrays can be rapidly created from a relatively small library of plasmids using publicly available reagents utilizing “Golden Gate” molecular cloning techniques to assemble new arrays within a few days [13, 53]. These methods are simple and only require reagents and equipment commonly found in molecular biology labs, although the overall throughput of assembly is limited. Other protocols are well suited to high-throughput generation of TALE arrays using solid-phase assembly [12, 67] or ligation-independent cloning techniques [68]. Notably, with the proper equipment, these high-throughput assembly methods are able to generate dozens to hundreds of TALEN constructs in 1 day. Alternatively, TALE arrays can also be custom ordered and pre-validated through commercial sources such as Life Technologies, Cellectis Bioresearch, Transposagen Biopharmaceuticals, and System Biosciences.
2.3 TALE Nuclease Architectures
Conventionally, TALEN monomers are created as a fusion of the TALE DNA-binding domain to the nonspecific endonuclease catalytic domain of FokI. Site-specific double-strand breaks are created when two separate nuclease monomers bind to adjacent target DNA sequences on opposite strands in a tail-to-tail fashion, thereby permitting dimerization of FokI and cleavage of the target DNA (Fig. 2). Thus, since FokI acts as a dimer, TALENs are designed in pairs to guide two separate FokI monomers to a desired target site. Several TALEN architectures have been described that demonstrate improved nuclease activity by truncating the C-terminus of the TALE DNA-binding domain [11, 55, 69]. These studies also show that the translocation domain on the TALE N-terminus can be removed without impacting activity. Moreover, these truncations can be used to restrict the length of the sequence allowed between the TALEN monomers [55] and may be useful for restricting potential off-target mutagenesis. Directed evolution of the TALE DNA-binding domain has also yielded mutants that have higher observed gene editing activity against episomal and chromosomal targets [70]. Alternate nuclease catalytic domains are also possible; for example, fusions of TALEs to monomeric meganucleases have recently been shown to improve targeting of these enzymes [32].
TALEN architecture and structure. (a) The TALE DNA-binding domain consists of the array of RVDs engineered to recognize specific sequences, along with fixed N- and C-terminal domains (orange), fused to the catalytic domain of the FokI endonuclease (blue). (b) Schematic of the TALEN structure, with TALEs (orange, PDB 3UGM) fused to the FokI domain (blue, PDB 2FOK) on DNA (green)
2.4 Enhancement of Nuclease Activity
Several improvements have been made to enhance the specificity of the FokI chimeric nucleases. A major advance was the identification of mutations that require heterodimerization of the nuclease pairs [71–73], thereby preventing potential homodimerization of nuclease monomers at unintended target loci. Furthermore, introduction of distinct obligate heterodimer mutations can be used to create two independent TALENs by preventing unexpected interactions between monomers from either pair [48]. Introduction of inactivating mutations to the FokI domain on one of the two nuclease domains in each pair can be used to generate targeted nickases. The single-strand nicks generated by these enzymes facilitate high levels of HDR but do not stimulate error-prone NHEJ repair [74, 75]. TALE nickases therefore display significantly reduced mutagenesis at off-target loci. Finally, directed evolution was utilized to find mutations that enhance the activity of FokI in a target site-independent manner [76].
2.5 Relaxation of the 5′-Thymine Targeting Requirement
The range of DNA sequences that can be targeted by TALEs is constrained by a strict requirement of a thymine base at the zero base position (N0) [55]. The crystal structure of a natural TALE protein suggests that there is a cryptic repeat domain in the N-terminus of the protein that specifically recognizes thymine [51, 52]. Novel TALE architectures have been developed to overcome this requirement by engineering this region of the TALE N-terminus to recognize alternative bases at this position [56, 57] or by utilizing TALE-like domains from related plant pathogens [77, 78] that naturally recognize guanine at the N0 position. However, DNA-binding activity of these TALE architectures may be reduced, especially for targets with adenosine and cytosine bases at the N0 position. Further work in this area may yield TALE scaffolds that can readily target sequences with any base at the N0 position with high efficiency.
2.6 Targeting Methylated DNA
The methylation status of the target DNA locus is known to impact DNA binding of TALE proteins, particularly with chromosomal targets directly containing 5′-methylated cytosine (5mC) [79, 80]. As a result, DNA methylation can significantly reduce or completely eliminate TALE binding. Methylation analysis of a target loci can be used to generate TALENs targeted to open chromatin; however this further restricts the utility of TALENs for site-specific gene modification. Global demethylation of a target genome using chemical modifiers such as 5′-aza-2′-deoxycytidine can rescue TALE binding [79]; however these methods are commonly associated with undesirable toxicity. More attractive methods have been developed that substitute specific RVDs in TALE proteins to efficiently bind particular methylated and/or demethylated cytosines in the target sequence. Thus, TALE proteins can be reengineered either to be insensitive to cytosine methylation by using the N* RVD that binds to both cytosine and 5mC [81] or by utilizing RVDs that specifically recognize 5mC (NG) or cytosine (HD) [82]. By substituting these particular RVDs, TALENs can be engineered to target these sites with high efficiency. It is also noteworthy that TALEs have been shown to target regions that are insensitive to DNase I, indicating that these proteins are able to access sites located in heterochromatin [83]. These studies were performed in dividing cells, and future work is necessary to determine the role of DNA replication in facilitating access to these target sites.
2.7 Delivery of TALENs
TALEN monomers are readily delivered by DNA expression cassettes or directly as mRNA by conventional transfection methods. However, the size of TALEN monomers and the highly repetitive array of RVD sequences present a significant challenge to viral delivery of TALEN constructs, thereby potentially limiting their utility in some gene editing applications. Adenovirus presents an attractive delivery vehicle for delivering gene constructs encoding both TALEN monomers [84], although adenovirus has limited tropism in some cell types and is highly immunogenic. Interestingly, this study also demonstrated that lentivirus was unable to deliver intact TALEN gene cassettes, due to rearrangements in the TALEN-coding region caused by the repetitive structure of RVD arrays. This limitation was overcome by the development of recoded TALEN constructs, termed re-TALEs, that can be efficiently expressed by lentiviral delivery [20], although this method may require reoptimization and synthesis of each new TALE gene. In contrast to DNA or mRNA delivery, direct protein delivery of TALENs can be achieved by utilizing cell-penetrating peptides covalently bound to purified TALEN proteins [85]. This method enables efficient genome editing in cells without the risk of spontaneous integration of the TALEN DNA expression construct into the genome that can be caused by non-viral and viral gene delivery. Furthermore, previous evidence suggests that protein delivery of gene-editing nucleases may reduce off-target activity by limiting the duration of nuclease exposure [86].
3 Applications in Basic Science and Biotechnology
Conventional genetic engineering methods involve the addition of new genes to cellular genomes by random integration of foreign genetic material into the chromosomal DNA. In contrast, genome editing using engineered nucleases enables precise manipulation at nearly any desired locus with high efficiency. Importantly, genome editing can generate a variety of genetic mutations without leaving any exogenous DNA sequences in the target genome. The development of high-throughput TALE assembly methods, in combination with high success rates of engineering highly active TALEN pairs, has resulted in the unprecedented ability to manipulate any gene of interest in a diverse array of organisms (Table 1). As one example of the breadth of TALEN assembly and applicability, libraries of TALENs have been generated to target 18,740 human protein-coding genes [80].
A powerful application of the TALEN technology is to rapidly and efficiently generate cellular models of human disease or to interrogate disease-related mutations or genes. This approach has been exploited to create disease-associated genetic mutations in somatic and stem-cell models for a variety of human diseases [87]. Notably, in this study, few if any TALEN-associated off-target mutations were detectable in many of the modified cell populations. High-throughput TALEN assembly was also used to interrogate a large panel of genes related to epigenetic regulation or cancer, with successful modification of >85 % of targeted genes [12]. The ease of TALEN technologies has enabled researchers to rapidly generate large genomic deletions to quickly interrogate microRNA function [88, 89]. These notable examples demonstrate that TALENs are a versatile tool to interrogate and study small and large genetic elements in complex genomes.
TALENs have also enabled rapid gene modification to efficiently generate transgenic species or to knockout genes of interest. This has enabled the study of a variety of genes of interest in a diverse range of organisms, including mice [90, 91], rats [92], pigs [93], cows [93], zebrafish [47, 94, 95], C. elegans [96, 97], newts [98], silkworm [99], flies [100], mosquitos [101], and frogs [102]. In addition, genome engineering is an exciting method to address challenges in plant engineering [103, 104]. Many plant genes are arranged in tandem arrays, making it difficult to selectively alter single genes to study or impart new gene function. The ability of TALENs to discriminate between relatively few mismatches makes this technology particularly powerful for altering specific gene arrays. An example of this approach is the application of TALENs in rice to generate disease resistance, as well as the rapid modification of numerous other genes [105]. Other studies have demonstrated that TALENs are a powerful platform to rapidly modify plant genes, including Arabidopsis thaliana [106], barley [107], and Brachypodium [105].
4 Applications for Gene Therapies
Gene therapies using designer nucleases have shown promise to correct the genetic basis of human diseases [1, 2]. The significant advances made in the efficiency and precision of novel genome engineering technologies across the past decade have led to the development of TALENs targeted to numerous genes related to a range of human diseases (Table 1). In contrast to gene replacement therapies, genome editing can directly correct mutations associated with disease. For example, we developed TALENs to generate small insertions and deletions to restore the reading frame of the dystrophin gene as a novel method to correct the molecular basis of Duchenne muscular dystrophy [50]. TALENs have also been used to correct mutations associated with epidermolysis bullosa [108], sickle cell disease [109, 110], beta-thalassemia [111], xeroderma pigmentosum [112], and alpha-1 antitrypsin deficiency [113] by homologous recombination and to correct mitochondrial DNA disorders [114] by deletion of aberrant sequences.
Beyond correction of mutant genes, gene editing strategies have been developed to modify genes in order to modulate disease phenotypes. ZFNs targeted to the gene encoding the HIV-coreceptor CCR5 are currently in clinical trials and have laid the groundwork for genome editing as a novel treatment modality [44, 115]. Studies have demonstrated that TALENs can also introduce efficient mutations to CCR5 [11, 55, 116, 117] and present an alternative gene editing technology for this application. TALENs have also been designed to target and eliminate hepatitis B viral genomes from human cells [118, 119]. TALENs have been utilized to disrupt the myostatin gene [120], the loss of which leads to hypertrophy of skeletal muscle that could be used to treat a range of diseases, including muscular dystrophies. Collectively, these studies show that TALENs are a powerful technology to generate a variety of gene modifications to correct human diseases.
5 Discussion
Over the past 5 years, the rapid advancement of genome editing technologies has led to widespread adoption of various gene editing platforms for a diverse range of applications [1–6]. TALEN technologies have made effective gene-editing tools accessible to nearly any researcher at low cost. The robustness of this technology has enabled researchers to rapidly and efficiently interrogate a large number of genes in a range of organisms (Table 1). Importantly, TALENs have impressive observed specificity and several advances in this field have further improved the fidelity of this approach [56, 57, 60, 61, 63, 121]. The specificity and efficiency of these approaches may be further improved as second-generation technologies are developed, such as TALE recombinases [28] and single-chain TALE-meganuclease fusions [32, 33]. The easily programmable TALE DNA-binding domain has also been a boon to creating other synthetic enzymes to regulate gene expression [53, 83, 122] and the epigenome [59, 58]. Although the recent advent of CRISPR/Cas9-based genome-engineering tools has provided an alternative facile method for gene editing [7, 123, 124], there are many differences between the two technologies and various applications could benefit from the strengths of each approach. Collectively, TALENs and other TALE-based gene-modifying tools have introduced publicly available, low-cost, efficient, and rapid gene modification that is accessible to any lab and has enabled studies for a remarkable variety of applications.
References
Perez-Pinera P, Ousterout DG, Gersbach CA (2012) Advances in targeted genome editing. Curr Opin Chem Biol 16(3–4):268–277
Gaj T, Gersbach CA, Barbas CF III (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31(7):397–405
Urnov FD, Rebar EJ, Holmes MC et al (2010) Genome editing with engineered zinc finger nucleases. Nat Rev Genet 11(9):636–646
Gersbach CA, Gaj T, Barbas CF III (2014) Synthetic zinc finger proteins: the advent of targeted gene regulation and genome modification technologies. Acc Chem Res 47:2309–2318
Mussolino C, Cathomen T (2012) TALE nucleases: tailored genome engineering made easy. Curr Opin Biotechnol 23:644–650
Joung JK, Sander JD (2012) TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol 14(1):49–55
Sander JD, Joung JK (2014) CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol 32:347–355
Cong L, Ran FA, Cox D et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121):819–823
Mali P, Yang L, Esvelt KM et al (2013) RNA-guided human genome engineering via Cas9. Science 339(6121):823–826
Hockemeyer D, Wang H, Kiani S et al (2011) Genetic engineering of human pluripotent cells using TALE nucleases. Nat Biotechnol 29(8):731–734
Mussolino C, Morbitzer R, Lütge F et al (2011) A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic Acids Res 39(21):9283–9293
Reyon D, Tsai SQ, Khayter C et al (2012) FLASH assembly of TALENs for high-throughput genome editing. Nat Biotechnol 30:460–465
Cermak T, Doyle EL, Christian M et al (2011) Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res 39(12):e82
Moscou MJ, Bogdanove AJ (2009) A simple cipher governs DNA recognition by TAL effectors. Science 326(5959):1501
Boch J, Scholze H, Schornack S et al (2009) Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326(5959):1509–1512
Cho SW, Kim S, Kim JM et al (2013) Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol 31(3):230–232
Hou Z, Zhang Y, Propson NE et al (2013) Efficient genome engineering in human pluripotent stem cells using Cas9 from Neisseria meningitidis. Proc Natl Acad Sci U S A 110(39):15644–15649
Jinek M, East A, Cheng A et al (2013) RNA-programmed genome editing in human cells. eLife 2:e00471
Horii T, Tamura D, Morita S et al (2013) Generation of an ICF syndrome model by efficient genome editing of human induced pluripotent stem cells using the CRISPR system. Int J Mol Sci 14(10):19774–19781
Yang L, Guell M, Byrne S et al (2013) Optimization of scarless human stem cell genome editing. Nucleic Acids Res 41(19):9049–9061
Redondo P, Prieto J, Muñoz IG et al (2008) Molecular basis of xeroderma pigmentosum group C DNA recognition by engineered meganucleases. Nature 456(7218):107–111
Silva G, Poirot L, Galetto R et al (2011) Meganucleases and other tools for targeted genome engineering: perspectives and challenges for gene therapy. Curr Gene Ther 11(1):11–27
Parekh-Olmedo H, Kmiec EB (2007) Progress and prospects: targeted gene alteration (TGA). Gene Ther 14(24):1675–1680
Gersbach CA, Gaj T, Gordley RM et al (2011) Targeted plasmid integration into the human genome by an engineered zinc-finger recombinase. Nucleic Acids Res 39(17):7868–7878
Gaj T, Mercer AC, Gersbach CA et al (2011) Structure-guided reprogramming of serine recombinase DNA sequence specificity. Proc Natl Acad Sci U S A 108(2):498–503
Gordley RM, Gersbach CA, Barbas CF III (2009) Synthesis of programmable integrases. Proc Natl Acad Sci U S A 106(13):5053–5058
Gaj T, Mercer AC, Sirk SJ et al (2013) A comprehensive approach to zinc-finger recombinase customization enables genomic targeting in human cells. Nucleic Acids Res 41(6):3937–3946
Mercer AC, Gaj T, Fuller RP et al (2012) Chimeric TALE recombinases with programmable DNA sequence specificity. Nucleic Acids Res 40(21):11163–11172
Jarjour J, West-Foyle H, Certo MT et al (2009) High-resolution profiling of homing endonuclease binding and catalytic specificity using yeast surface display. Nucleic Acids Res 37(20):6871–6880
Takeuchi R, Lambert AR, Mak AN et al (2011) Tapping natural reservoirs of homing endonucleases for targeted gene modification. Proc Natl Acad Sci U S A 108(32):13077–13082
Takeuchi R, Choi M, Stoddard BL (2014) Redesign of extensive protein-DNA interfaces of meganucleases using iterative cycles of in vitro compartmentalization. Proc Natl Acad Sci U S A 111(11):4061–4066
Boissel S, Jarjour J, Astrakhan A et al (2014) megaTALs: a rare-cleaving nuclease architecture for therapeutic genome engineering. Nucleic Acids Res 42(4):2591–2601
Beurdeley M, Bietz F, Li J et al (2013) Compact designer TALENs for efficient genome engineering. Nat Commun 4:1762
Rivera-Torres N, Strouse B, Bialk P et al (2014) The position of DNA cleavage by TALENs and cell synchronization influences the frequency of gene editing directed by single-stranded oligonucleotides. PLoS One 9(5):e96483
Gaj T, Sirk SJ, Barbas CF III (2014) Expanding the scope of site-specific recombinases for genetic and metabolic engineering. Biotechnol Bioeng 111(1):1–15
Rouet P, Smih F, Jasin M (1994) Introduction of double-strand breaks into the genome of mouse cells by expression of a rare-cutting endonuclease. Mol Cell Biol 14(12):8096–8106
Rouet P, Smih F, Jasin M (1994) Expression of a site-specific endonuclease stimulates homologous recombination in mammalian cells. Proc Natl Acad Sci U S A 91(13):6064–6068
Moehle EA, Rock JM, Lee Y-L et al (2007) Targeted gene addition into a specified location in the human genome using designed zinc finger nucleases. Proc Natl Acad Sci 104(9):3055–3060
Urnov F, Miller J, Lee Y et al (2005) Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435:646–651
Sebastiano V, Maeder ML, Angstman JF et al (2011) In situ genetic correction of the sickle cell anemia mutation in human induced pluripotent stem cells using engineered zinc finger nucleases. Stem Cells 29(11):1717–1726
Zou J, Mali P, Huang X et al (2011) Site-specific gene correction of a point mutation in human iPS cells derived from an adult patient with sickle cell disease. Blood 118(17):4599–4608
Yusa K, Rashid ST, Strick-Marchand H et al (2011) Targeted gene correction of α1-antitrypsin deficiency in induced pluripotent stem cells. Nature 478(7369):391–394
Li H, Haurigot V, Doyon Y et al (2011) In vivo genome editing restores haemostasis in a mouse model of haemophilia. Nature 475(7355):217–221
Perez EE, Wang J, Miller JC et al (2008) Establishment of HIV-1 resistance in CD4+ T cells by genome editing using zinc-finger nucleases. Nat Biotechnol 26(7):808–816
Holt N, Wang J, Kim K et al (2010) Human hematopoietic stem/progenitor cells modified by zinc-finger nucleases targeted to CCR5 control HIV-1 in vivo. Nat Biotechnol 28(8):839–847
Lee HJ, Kim E, Kim JS (2010) Targeted chromosomal deletions in human cells using zinc finger nucleases. Genome Res 20(1):81–89
Gupta A, Hall VL, Kok FO et al (2013) Targeted chromosomal deletions and inversions in zebrafish. Genome Res 23(6):1008–1017
Sollu C, Pars K, Cornu TI et al (2010) Autonomous zinc-finger nuclease pairs for targeted chromosomal deletion. Nucleic Acids Res 38(22):8269–8276
Chapdelaine P, Pichavant C, Rousseau J et al (2010) Meganucleases can restore the reading frame of a mutated dystrophin. Gene Ther 17(7):846–858
Ousterout DG, Perez-Pinera P, Thakore PI et al (2013) Reading frame correction by targeted genome editing restores dystrophin expression in cells from Duchenne muscular dystrophy patients. Mol Ther 21(9):1718–1726
Mak AN, Bradley P, Cernadas RA et al (2012) The crystal structure of TAL effector PthXo1 bound to its DNA target. Science 335(6069):716–719
Deng D, Yan C, Pan X et al (2012) Structural basis for sequence-specific recognition of DNA by TAL effectors. Science 335(6069):720–723
Zhang F, Cong L, Lodato S et al (2011) Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nat Biotechnol 29(2):149–153
Christian M, Cermak T, Doyle EL et al (2010) Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186(2):757–761
Miller JC, Tan S, Qiao G et al (2011) A TALE nuclease architecture for efficient genome editing. Nat Biotechnol 29(2):143–148
Doyle EL, Hummel AW, Demorest ZL et al (2013) TAL effector specificity for base 0 of the DNA target is altered in a complex, effector- and assay-dependent manner by substitutions for the tryptophan in cryptic repeat −1. PLoS One 8(12):e82120
Lamb BM, Mercer AC, Barbas CF III (2013) Directed evolution of the TALE N-terminal domain for recognition of all 5′ bases. Nucleic Acids Res 41(21):9779–9785
Konermann S, Brigham MD, Trevino AE et al (2013) Optical control of mammalian endogenous transcription and epigenetic states. Nature 500(7463):472–476
Maeder ML, Angstman JF, Richardson ME et al (2013) Targeted DNA demethylation and activation of endogenous genes using programmable TALE-TET1 fusion proteins. Nat Biotechnol 31(12):1137–1142
Christian ML, Demorest ZL, Starker CG et al (2012) Targeting G with TAL effectors: a comparison of activities of TALENs constructed with NN and NK repeat variable di-residues. PLoS One 7(9):e45383
Streubel J, Blucher C, Landgraf A et al (2012) TAL effector RVD specificities and efficiencies. Nat Biotechnol 30(7):593–595
Cong L, Zhou R, Kuo YC et al (2012) Comprehensive interrogation of natural TALE DNA-binding modules and transcriptional repressor domains. Nat Commun 3:968
Lin Y, Fine EJ, Zheng Z et al (2014) SAPTA: a new design tool for improving TALE nuclease activity. Nucleic Acids Res 42:e47
Doyle EL, Booher NJ, Standage DS et al (2012) TAL effector-nucleotide targeter (TALE-NT) 2.0: tools for TAL effector design and target prediction. Nucleic Acids Res 40(Web Server issue):W117–W122
Sander JD, Maeder ML, Reyon D et al (2010) ZiFiT (Zinc Finger Targeter): an updated zinc finger engineering tool. Nucleic Acids Res 38(Suppl 2):W462–W468
Fine EJ, Cradick TJ, Zhao CL et al (2014) An online bioinformatics tool predicts zinc finger and TALE nuclease off-target cleavage. Nucleic Acids Res 42(6):e42
Briggs AW, Rios X, Chari R et al (2012) Iterative capped assembly: rapid and scalable synthesis of repeat-module DNA such as TAL effectors from individual monomers. Nucleic Acids Res 40(15):e117
Schmid-Burgk JL, Schmidt T, Kaiser V et al (2013) A ligation-independent cloning technique for high-throughput assembly of transcription activator-like effector genes. Nat Biotechnol 31(1):76–81
Bedell VM, Wang Y, Campbell JM et al (2012) In vivo genome editing using a high-efficiency TALEN system. Nature 491(7422):114–118
Sun N, Bao Z, Xiong X et al (2014) SunnyTALEN: a second-generation TALEN system for human genome editing. Biotechnol Bioeng 111:683–691
Miller JC, Holmes MC, Wang J et al (2007) An improved zinc-finger nuclease architecture for highly specific genome editing. Nat Biotechnol 25(7):778–785
Doyon Y, Vo TD, Mendel MC et al (2010) Enhancing zinc-finger-nuclease activity with improved obligate heterodimeric architectures. Nat Methods 8(1):74–79
Szczepek M, Brondani V, Buchel J et al (2007) Structure-based redesign of the dimerization interface reduces the toxicity of zinc-finger nucleases. Nat Biotechnol 25(7):786–793
Ramirez CL, Certo MT, Mussolino C et al (2012) Engineered zinc finger nickases induce homology-directed repair with reduced mutagenic effects. Nucleic Acids Res 40(12):5560–5568
Wang J, Friedman G, Doyon Y et al (2012) Targeted gene addition to a predetermined site in the human genome using a ZFN-based nicking enzyme. Genome Res 22(7):1316–1326
Guo J, Gaj T, Barbas C III (2010) Directed evolution of an enhanced and highly efficient FokI cleavage domain for zinc finger nucleases. J Mol Biol 400:96–107
de Lange O, Schreiber T, Schandry N et al (2013) Breaking the DNA-binding code of Ralstonia solanacearum TAL effectors provides new possibilities to generate plant resistance genes against bacterial wilt disease. New Phytol 199(3):773–786
Li L, Atef A, Piatek A et al (2013) Characterization and DNA-binding specificities of Ralstonia TAL-like effectors. Mol Plant 6(4):1318–1330
Bultmann S, Morbitzer R, Schmidt CS et al (2012) Targeted transcriptional activation of silent oct4 pluripotency gene by combining designer TALEs and inhibition of epigenetic modifiers. Nucleic Acids Res 40(12):5368–5377
Kim Y, Kweon J, Kim A et al (2013) A library of TAL effector nucleases spanning the human genome. Nat Biotechnol 31(3):251–258
Valton J, Dupuy A, Daboussi F et al (2012) Overcoming transcription activator-like effector (TALE) DNA binding domain sensitivity to cytosine methylation. J Biol Chem 287(46):38427–38432
Deng D, Yin P, Yan C et al (2012) Recognition of methylated DNA by TAL effectors. Cell Res 22(10):1502–1504
Perez-Pinera P, Ousterout DG, Brunger JM et al (2013) Synergistic and tunable human gene activation by combinations of synthetic transcription factors. Nat Methods 10(3):239–242
Holkers M, Maggio I, Liu J et al (2013) Differential integrity of TALE nuclease genes following adenoviral and lentiviral vector gene transfer into human cells. Nucleic Acids Res 41(5):e63
Liu J, Gaj T, Patterson JT et al (2014) Cell-penetrating peptide-mediated delivery of TALEN proteins via bioconjugation for genome engineering. PLoS One 9(1):e85755
Gaj T, Guo J, Kato Y et al (2012) Targeted gene knockout by direct delivery of zinc-finger nuclease proteins. Nat Methods 9(8):805–807
Ding Q, Lee YK, Schaefer EA et al (2013) A TALEN genome-editing system for generating human stem cell-based disease models. Cell Stem Cell 12(2):238–251
Kim YK, Wee G, Park J et al (2013) TALEN-based knockout library for human microRNAs. Nat Struct Mol Biol 20(12):1458–1464
Zhang Z, Xiang D, Heriyanto F et al (2013) Dissecting the roles of miR-302/367 cluster in cellular reprogramming using TALE-based repressor and TALEN. Stem Cell Reports 1(3):218–225
Wang H, Hu YC, Markoulaki S et al (2013) TALEN-mediated editing of the mouse Y chromosome. Nat Biotechnol 31(6):530–532
Takada S, Sato T, Ito Y et al (2013) Targeted gene deletion of miRNAs in mice by TALEN system. PLoS One 8(10):e76004
Tesson L, Usal C, Menoret S et al (2011) Knockout rats generated by embryo microinjection of TALENs. Nat Biotechnol 29(8):695–696
Carlson DF, Tan W, Lillico SG et al (2012) Efficient TALEN-mediated gene knockout in livestock. Proc Natl Acad Sci U S A 109(43):17382–17387
Sander JD, Cade L, Khayter C et al (2011) Targeted gene disruption in somatic zebrafish cells using engineered TALENs. Nat Biotechnol 29(8):697–698
Huang P, Xiao A, Zhou M et al (2011) Heritable gene targeting in zebrafish using customized TALENs. Nat Biotechnol 29(8):699–700
Lo TW, Pickle CS, Lin S et al (2013) Precise and heritable genome editing in evolutionarily diverse nematodes using TALENs and CRISPR/Cas9 to engineer insertions and deletions. Genetics 195(2):331–348
Wood AJ, Lo TW, Zeitler B et al (2011) Targeted genome editing across species using ZFNs and TALENs. Science 333(6040):307
Hayashi T, Sakamoto K, Sakuma T et al (2014) Transcription activator-like effector nucleases efficiently disrupt the target gene in Iberian ribbed newts (Pleurodeles waltl), an experimental model animal for regeneration. Dev Growth Differ 56(1):115–121
Ma S, Zhang S, Wang F et al (2012) Highly efficient and specific genome editing in silkworm using custom TALENs. PLoS One 7(9):e45035
Beumer KJ, Trautman JK, Christian M et al (2013) Comparing zinc finger nucleases and transcription activator-like effector nucleases for gene targeting in Drosophila. G3 (Bethesda) 3(10):1717–1725
Smidler AL, Terenzi O, Soichot J et al (2013) Targeted mutagenesis in the malaria mosquito using TALE nucleases. PLoS One 8(8):e74511
Lei Y, Guo X, Liu Y et al (2012) Efficient targeted gene disruption in Xenopus embryos using engineered transcription activator-like effector nucleases (TALENs). Proc Natl Acad Sci U S A 109(43):17484–17489
Voytas DF (2013) Plant genome engineering with sequence-specific nucleases. Annu Rev Plant Biol 64:327–350
Mahfouz MM, Li L, Shamimuzzaman M et al (2011) De novo-engineered transcription activator-like effector (TALE) hybrid nuclease with novel DNA binding specificity creates double-strand breaks. Proc Natl Acad Sci U S A 108(6):2623–2628
Shan Q, Wang Y, Chen K et al (2013) Rapid and efficient gene modification in rice and Brachypodium using TALENs. Mol Plant 6(4):1365–1368
Christian M, Qi Y, Zhang Y et al (2013) Targeted mutagenesis of Arabidopsis thaliana using engineered TAL effector nucleases. G3 (Bethesda) 3(10):1697–1705
Wendt T, Holm PB, Starker CG et al (2013) TAL effector nucleases induce mutations at a pre-selected location in the genome of primary barley transformants. Plant Mol Biol 83(3):279–285
Osborn MJ, Starker CG, McElroy AN et al (2013) TALEN-based gene correction for epidermolysis bullosa. Mol Ther 21(6):1151–1159
Sun N, Liang J, Abil Z et al (2012) Optimized TAL effector nucleases (TALENs) for use in treatment of sickle cell disease. Mol Biosyst 8(4):1255–1263
Voit RA, Hendel A, Pruett-Miller SM et al (2014) Nuclease-mediated gene editing by homologous recombination of the human globin locus. Nucleic Acids Res 42(2):1365–1378
Ma N, Liao B, Zhang H et al (2013) Transcription activator-like effector nuclease (TALEN)-mediated gene correction in integration-free beta-thalassemia induced pluripotent stem cells. J Biol Chem 288(48):34671–34679
Dupuy A, Valton J, Leduc S et al (2013) Targeted gene therapy of xeroderma pigmentosum cells using meganuclease and TALEN. PLoS One 8(11):e78678
Choi SM, Kim Y, Shim JS et al (2013) Efficient drug screening and gene correction for treating liver disease using patient-specific stem cells. Hepatology 57(6):2458–2468
Bacman SR, Williams SL, Pinto M et al (2013) Specific elimination of mutant mitochondrial genomes in patient-derived cells by mitoTALENs. Nat Med 19(9):1111–1113
Tebas P, Stein D, Tang WW et al (2014) Gene editing of CCR5 in autologous CD4 T cells of persons infected with HIV. N Engl J Med 370(10):901–910
Mussolino C, Alzubi J, Fine EJ et al (2014) TALENs facilitate targeted genome editing in human cells with high specificity and low cytotoxicity. Nucleic Acids Res 42(10):6762–6773
Ye L, Wang J, Beyer AI et al (2014) Seamless modification of wild-type induced pluripotent stem cells to the natural CCR5Delta32 mutation confers resistance to HIV infection. Proc Natl Acad Sci U S A 111:9591–9596
Bloom K, Ely A, Mussolino C et al (2013) Inactivation of hepatitis B virus replication in cultured cells and in vivo with engineered transcription activator-like effector nucleases. Mol Ther 21(10):1889–1897
Chen J, Zhang W, Lin J et al (2014) An efficient antiviral strategy for targeting hepatitis B virus genome using transcription activator-like effector nucleases. Mol Ther 22(2):303–311
Xu L, Zhao P, Mariano A et al (2013) Targeted myostatin gene editing in multiple mammalian species directed by a single pair of TALE nucleases. Mol Ther Nucleic Acids 2:e112
Guilinger JP, Pattanayak V, Reyon D et al (2014) Broad specificity profiling of TALENs results in engineered nucleases with improved DNA-cleavage specificity. Nat Methods 11(4):429–435
Maeder ML, Linder SJ, Reyon D et al (2013) Robust, synergistic regulation of human gene expression using TALE activators. Nat Methods 10(3):243–245
Mali P, Esvelt KM, Church GM (2013) Cas9 as a versatile tool for engineering biology. Nat Methods 10(10):957–963
Hsu PD, Lander ES, Zhang F (2014) Development and applications of CRISPR-Cas9 for genome engineering. Cell 157(6):1262–1278
Sung YH, Baek IJ, Kim DH et al (2013) Knockout mice created by TALEN-mediated gene targeting. Nat Biotechnol 31(1):23–24
Zu Y, Tong X, Wang Z et al (2013) TALEN-mediated precise genome modification by homologous recombination in zebrafish. Nat Methods 10(4):329–331
Acknowledgements
This work was supported by a US National Institutes of Health (NIH) Director’s New Innovator Award (DP2OD008586), National Science Foundation (NSF) Faculty Early Career Development (CAREER) Award (CBET-1151035), NIH R01DA036865, NIH R21AR065956, NIH UH3TR000505, NIH P30AR066527, the Duke Coulter Translational Partnership, and an American Heart Association Scientist Development Grant (10SDG3060033). D.G.O. was supported by an American Heart Association Mid-Atlantic Affiliate Predoctoral Fellowship.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Ousterout, D.G., Gersbach, C.A. (2016). The Development of TALE Nucleases for Biotechnology. In: Kühn, R., Wurst, W., Wefers, B. (eds) TALENs. Methods in Molecular Biology, vol 1338. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2932-0_3
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2932-0_3
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2931-3
Online ISBN: 978-1-4939-2932-0
eBook Packages: Springer Protocols