Abstract
The acute inflammatory response is a complex defense mechanism that has evolved to respond rapidly to injury, infection, and other disruptions in homeostasis. This robust responsiveness to biological stress likely endows the host with increased fitness, but over-robust or inadequate inflammation predisposes the host to various diseases. Importantly, well-compartmentalized inflammation is generally beneficial, but spillover of inflammation into the blood is a hallmark—and likely also a driver—of self-maintaining inflammation. The blood is also a key entry point and immunological interface for vectors of parasitic diseases, diseases that themselves incite systemic inflammation. The complex role of inflammation in health and disease has made this biological system difficult to understand comprehensively and modulate rationally for therapeutic purposes. Consequently, systems approaches have been applied in order to characterize dynamical properties and identify key control points in inflammation. This process begins with the collection of high-dimensional, experimental, and clinical data, followed by data reduction and data-driven modeling that finally informs mechanistic computational models for analysis, prediction, and rational modulation. These studies have suggested that the overall architecture of the inflammatory response includes a multiscale positive feedback from inflammation → tissue damage → inflammation, which is often inadequately controlled by negative feedback from anti-inflammatory mediators. Given the importance of the blood interface for the inflammatory response, and the accessibility of this compartment both as an immunological sampling reservoir for vectors as well as for diagnosis and therapy, we suggest that any rational efforts at modulating inflammation via the blood compartment must involve computational modeling.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Inflammation is an essential process in maintaining health and responding to disease. Acute inflammation is driven largely by the innate immune system, which not only serves as the first line of defense against invading pathogens but also functions to resolve tissue damage and restore homeostasis upon a variety of inflammatory conditions including sepsis, trauma, wound healing, and many more. A large aspect of the acute inflammatory response plays out in the blood, but usually only when inflammation is dysregulated. Dysregulated systemic inflammation also plays a significant role in the pathophysiology of other diseases that are not primarily attributed to innate immunity, such as cancer and diabetes. Although the list of diseases is broad and the processes important to each setting may differ in certain respects, the core architecture of the inflammatory response to biological stress is highly conserved [1]. An infection or a tissue injury/damage triggers an initially local cascade of events mediated by an array of cells (e.g., macrophages, neutrophils, dendritic cells, lymphocytes, etc.) and molecules (cytokines, free radicals, and damage-associated molecular pattern molecules (DAMPs)) that locate invading pathogens or stressed/damaged tissue, alert and recruit other cells and molecules, eliminate the offending agents, and finally restore the body to equilibrium [2]. When dysregulated or overexuberant, inflammation can be discerned in the systemic circulation in the form of altered levels of inflammatory cells and molecular mediators.
In sepsis and trauma, this response is concomitant with physiologic manifestations including changes in heart rate and body temperature, responses that act in a concerted fashion in order to help optimize host defense while minimizing tissue damage. Indeed, although a well-regulated inflammatory response is essential for proper healing and host defense, an overly exuberant response can become self-perpetuating and lead to organ dysfunction and death [3, 4]. These vastly different outcomes can be explained, at least in part, by the high-level architecture of the immune response, which includes a positive feedback loop from inflammation →damage/dysfunction → inflammation that can drive pathophysiology in inflammatory diseases (Fig. 11.1).
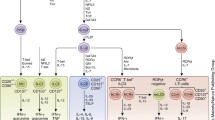
Complex structure of the innate immune response to biological stress. Following an initiating event (e.g., trauma, hemorrhage, infection), both pro- and anti-inflammatory influences (e.g., chemokines, cytokines, lipid products, and free radicals) are elaborated, leading to tissue damage or dysfunction. These stressed tissues elaborate damage-associated molecular patterns (DAMPs), which further propagate innate immune mechanisms. When the pro-inflammatory mediators exceed defined thresholds, both pro- and anti-inflammatory mediators spillover into the blood and may cause inflammation to feedback and spread systemically to other organs as well; we refer to this process as an inflammatory tipping point. Inflammatory mediators are also transferred to blood-feeding vectors and can serve to communicate the infection status of the host, as well as modulating anti-parasite immunity in the vector. Black curved arrows represent the pro-inflammatory feedback and gray curved arrows represent feedback from anti-inflammatory mediators
The detrimental effects of self-sustaining inflammation are likely responsible for the general perception of inflammation as an inherently harmful process [5, 6]. However, in addition to the aforementioned beneficial roles of inflammation in the resolution of tissue injury, recent studies suggest that morbidity and mortality are worse in animals and humans with low levels of early pro-inflammatory signals [7]. The emerging view of inflammation is indeed more nuanced, casting inflammation as a highly coordinated communication network that allows the body to sense and respond to challenges and subsequently restore homeostasis [8, 9]. One may consider the complexity resulting from this coordination to be an indicator of a well-regulated and properly orchestrated response, and consequently a less complex response would be indicative of a pathological dis- or misconnectivity of the network. Guided by insights from studies on the dysregulated physiology characteristic of sepsis and trauma/hemorrhage, which have reported that a decrease invariability/complexity of heart rate can presage increased morbidity and mortality, we have suggested that well-organized dynamic networks of mediators are crucial to an appropriate inflammatory response [10, 11]. Indeed, such networks are induced early in the response to experimental surgical trauma in mice, and these networks become disorganized and less complex with the addition of hemorrhagic shock to this minor trauma [10].How ever, emerging studies from our group also suggest that overly-robust, and possibly self-sustaining, inflammation manifests as networks that are highly complex.
The current paradigm for acute inflammation, based in large part on studies in response to trauma, hemorrhage, or infection, involves a dynamic cascade of cellular and molecular events. Innate immune cells such as mast cells, neutrophils, and macrophages are activated directly by bacterial endotoxin or indirectly by various stimuli elicited systemically upon trauma and hemorrhage [12–15], including the release of DAMPs (Fig. 11.1) [16–18]. These stimuli enter the systemic circulation and activate circulating monocytes and neutrophils [19], which subsequently migrate to compromised tissue by following along a chemoattractant gradient induced at the site of injury/infection [20]. Activated macrophages and neutrophils produce and secrete effectors that activate a variety of immune cells (including further activating themselves) as well as nonimmune cells such as endothelial cells. Both DAMPs and pro-inflammatory cytokines—primary among them tumor necrosis factor-α (TNF-α) [21–27]—promote immune cell activation and affect important physiological functions that feedback positively to promote further production of inflammatory mediators. This behavior may lead to inflammatory tipping points—and concomitant spillover of inflammatory mediators into the blood—indicative of cascading system failure that occurs at multiple scales and across multiple compartments [28] (Fig. 11.1). In turn, dysregulated inflammation in the blood may itself become a driver of further inflammation in other tissues (Fig. 11.1).
Inflammation Is a Complex System
As evidenced by the preceding description, inflammation, like most biological systems, is a highly nonlinear system with multiple feedback loops that may be discerned even when viewed in a coarse-grained, relatively abstract fashion (Fig. 11.1). Positive feedback loops allow rapid ramping up of a response to biological stress, while the negative feedback works to suppress inflammation and restore homeostasis once the threat (infection, damaged tissue, etc.) has been eliminated. We suggest that, as has likely occurred in many other complex biological systems [29], inflammation has evolved to be robust to a broad range of perturbations but at a cost of fragility in key control nodes that may account for the tipping point behavior described above [16, 29] (Fig. 11.1). Failure at these points can lead to disease; therefore, characterizing these failure modes, and especially the tipping point phenotype, is paramount for the development of effective therapeutic interventions [28]. Another property of a complex nonlinear system is the ability to exhibit vastly different behaviors that depend on initial conditions and parameters (i.e., strengths and rates of interactions of components) [30, 31]. This heterogeneity, which recapitulates the clinical observation of patient-to-patient variability, complicates the prediction of individual patient outcomes using the current suite of statistically based tools [17, 28]. As described below, a systems approach to inflammation can be useful, indeed necessary, to explain the behavior of the innate immune response in an individual patient to various biological conditions and ultimately allow for the modulation of this response in pathological conditions.
Modeling Inflammation
Modeling Methods for A Systems Biology of Inflammation
Systems biology approaches span a broad range of techniques, and can be categorized roughly into correlative or causative approaches, with focus on either learning basic principles of system organization and function [32–34] or building predictive computational models [32, 35]. Although there is overlap between these areas, most efforts at elucidating biological mechanisms from high-dimensional data have traditionally focused on particular points along this spectrum of computational approaches. We suggest that gleaning translationally relevant insights into the inflammatory response and its interconnected (patho) physiology will require the successful navigation of this spectrum, in a logical progression from data to models to understanding and prediction [17, 28] (Fig. 11.2).
Taming the data deluge: from high-dimensional data to data-driven and mechanistic models. Data-driven methods are used to reduce dimensionality and infer correlative and causal relationships among genes/proteins in the system. Quasi-mechanistic insights from these models, together expert knowledge and the network structure inferred by Dynamic Bayesian Networks inform mechanistic models that may be encoded as ODEs, ABMs, RBMs, etc. Predictions from simulation of these models are compared with experimental data under new conditions and the model is refined based on the discrepancies. Finally, the mechanistic model can be analyzed by a variety of methods to understand its dynamics and identify key control points. ODE ordinary differential equation, ABM agent-based model, RBM rule-based model
Correlative approaches, with which most biologists and clinicians are familiar, include regression techniques that build models predictive within the conditions of the data they were trained on [36]. Although these methods do not provide detailed mechanistic insight, these approaches can be used to understand abstract features of the response, such as the presence of nonlinearities and the order of the response. The main drawback of this class of models is that they are almost completely devoid of mechanistic insight, and can be very over-fit to the data on which they were trained. A less-utilized data-driven method is principal component analysis (PCA), which reduces a high-dimensional dataset into a few principal components that account for much of the observed variance in the data. When applied to time-series data, the variables (genes/proteins/etc.) that constitute these principal components may be interpreted as the principal drivers of the observed response and can give some mechanistic insights into the underlying process [10, 37]. In the setting of inflammation, correlative approaches such as PCA may facilitate the development of diagnostics by analyzing the cytokine milieu in the blood resulting from inflammatory spillover, in order to identify the health state of individuals and possibly inform patient-specific interventions [38]. While these methods correlate gene/protein levels to phenotype and can suggest relevant molecular players involved in a given inflammatory process, these methods do not provide much information about how the genes/proteins interact with each other [17, 33].
In order to better discern organizational aspects of interacting networks of mediators, such as co-regulation or auto-induction, a variety of methods have been developed. Hierarchical clustering and Bayesian methods use high-throughput genomic or proteomic data of several time points and/or conditions to correlate gene expression patterns with function and infer regulatory networks of correlated genes. Several developments in these methods over the past 15 years have yielded more informative networks that can be more easily translated into mechanistic models. Among these methods, Dynamic Bayesian Networks (DBNs) are particularly suited for inferring directed (causative) networks of interactions based on the probabilistic measure of how well the network can explain observed data. DBNs can be supplemented by additional experimental evidence and expert knowledge to hypothesize mechanistic models (Fig. 11.2).
Mechanistic models are derived from more detailed biological and physical descriptions of a system have a rich set of tools for both analysis and simulation. These models, based on causative interactions, can be constructed as ordinary differential equations (ODEs), rule-based models (RBMs), and agent-based models (ABMs) among other methods (including hybrid methods), and have the advantage of potentially being predictive outside the range of conditions/time points that they were calibrated on. Although it is often difficult to parameterize such models, they can unveil emergent phenomena that are not immediately obvious from the interactions that are encoded in the model. There are several analytic tools for ODE models especially that have been developed and used to decipher the organizational principles of networks (or subnetworks), the properties that explain the dynamics and robustness/sensitivity of a given complex system, and, perhaps most importantly, the critical points of control in the system [34] (Fig. 11.2). These tools are particularly important in order to help define the complex interplay between the inflammatory mediators in the blood and other compartments both within the host (organs/tissue) and without (e.g., in the case of interactions with blood-feeding vectors). Tools from dynamical systems theory allow identification of the possible steady state(s) of a system as well as the kinetics of the system’s time evolution. These tools have been used extensively to explain (or predict, depending on the context) diverse behaviors such as bistability, hysteresis, and oscillations in a variety of biological systems [39]. Bifurcation diagrams, in particular, can be used to map out the effects of a particular parameter on the possible steady state behaviors of a system, and to indicate the transition from a healthy steady state to a pathological one [14, 40–42]. The relative importance of parameters can also be quantified by calculating the change in the model output in response to changes in the parameter values using sensitivity analysis [34, 43]. These methods work in a complementary fashion to identify the key points that can be modulated to change the behavior of a system (Fig. 11.2).
The analysis of ODE models of biological systems can be approached from a control theory perspective as well. Achieving robustness and efficiency are core principles of both evolution as well as engineering. Indeed, feedback, a pervasive biological phenomenon, is also a fundamental component of control strategies [29]. An ODE model is the equivalent of a state-space representation of a control system. Thus, it is possible to decompose the biological system into a control structure and analyze the role of each component using control theoretic tools that characterize their robustness and identify the key mediators that modulate the performance of such a control system [44]. These analyses are especially relevant given that the tipping point phenomenon in the inflammatory response is likely the result of a failure of the body’s control structure to handle stress (Fig. 11.1).
Although ODE models are associated with a wide range of analytical tools, they are inappropriate descriptions for settings in which there are low numbers of molecules or in settings in which molecules are not well mixed and thus stochastic effects are at play. RBMs and ABMs (among others) are superior methods for such conditions, as they are able to handle stochastic simulations. Agent-based modeling software packages such as NetLogo [45] and SPARK [46] are especially useful because of their ability to encode and visualize spatially realistic effects as well. Hybrid models can be constructed to merge the advantages of both ODEs and ABMs, and are especially useful for describing phenomena that occur on different timescales. Moreover, the inflammatory response is a quintessential example of a system whose parts operate not only at various timescales but also across various compartments. Thus, a hybrid modeling approach that melds processes best modeled by ODE with processes best modeled by ABM/RBM is essential for a more complete description of inflammation. As a step in this direction, a hybrid model of pressure ulcer formation in spinal cord injury (SCI) patients was developed, abstracting the microscopic details of blood flow and oxygen availability as a series of resistors using differential equations, while encoding an abstracted cascade of inflammation and wound healing in response to simulated cycles of pressure on the peripheral tissue using ABM. Based on data on blood flow in noninjured human subjects versus SCI patients, the parameters gained from this hybrid model predicted the higher likelihood of pressure ulcer development in SCI patients (Solovyev et al., unpublished observations).
While we wish to navigate through process of data→data-driven model→ mechanistic model →prediction and understanding of the innate immune response, we seek to put it in the perspective of translational applications with a focus on clinical and preclinical settings. Much of the work in systems biology has understandably been in simpler, well-studied model organisms, but even among studies focused on preclinical science, there has been an overall lack of translation to the clinical arena. Translational systems biology is a framework with a focus on translational insights for novel diagnostic or therapeutic purposes and predictive mathematical models that inform in silico clinical trials [9, 47, 48]. Initially formulated to deal with the clinical challenge of integrating acute inflammation and organ dysfunction in critical illness, this work expanded to include healing of acute and chronic wounds and infections in various diseases, rational dynamic modulation of inflammation, and cross-species host–pathogen interactions.
Modeling and Rational Modulation of Inflammation in Sepsis and Trauma/Hemorrhage
Traumatic injury is often accompanied by hemorrhage and is a significant cause of morbidity and mortality in patients, especially among young people [49, 50]. These patients are particularly susceptible to multiple organ dysfunction syndrome (MODS), a poorly understood syndrome that may be partly attributed to excessive and dysregulated inflammation [4]. The complexity of the interactions between inflammation and organ physiology has likely stymied the development of therapies for MODS, and was the motivation for the development of both data-driven and mechanistic computational models [10, 12–14, 42, 51–54].
Several models of the acute inflammatory response to sepsis, trauma, and hemorrhage have been developed, models that provide insight into the mechanisms of inflammation at varying degrees of abstraction. Based on the typical progression of the inflammatory pathway described in the preceding section, an ODE model of acute inflammation consisting of pathogen, a single population of inflammatory cells, and a measure of global tissue damage/dysfunction interconnected the actions of pro-inflammatory cytokines and DAMPs for the first time, and described both recoverable infection and septic shock, as well as suggesting different therapeutic avenues for the diverse manifestations of sepsis [55]. With the addition of more interactions, including the positive feedback loop between inflammation and damage, a more complex model was used for simulating populations of patients in sepsis and anthrax [12, 52], and to test effects of probiotic treatment of necrotizing enterocolitis [56]. Another equation-based model was calibrated in various inflammatory scenarios in mice [12] and was calibrated on easily accessible circulating levels of inflammatory cytokines and nitric oxide (NO) reaction products, and explicitly included measurable physiological parameters such as blood pressure along with the more abstract global damage (a surrogate for both DAMPs and the health status of the individual; Fig. 11.1) in mice [12, 13, 51]. This calibrated model was capable of predicting, outside of its calibration set, dose ranges of endotoxin at which death is known to occur [12] in addition to predicting responses to combinations of insults on which it was not trained [12, 57, 58].
While investigating the role of initial trauma in the murine response to trauma/hemorrhagic shock, both correlative (transcriptomic analysis, PCA, regression) and causative (ODE) models were used in a complementary fashion, and suggested that the role of initial trauma is central in driving the inflammatory response, both systemically and in the liver [13]. Transcriptomic data indicated an overlap between the genes and pathways induced in trauma alone or trauma with hemorrhagic shock with differences in only the magnitude of expression. In agreement with this observation, a mechanistic mathematical model showed that using the same model with different initial conditions could differentiate the inflammatory responses to trauma versus hemorrhagic shock. Later, multivariate regression, PCA, and dynamic network analysis all suggested major mechanistic differences between sham cannulation and hemorrhagic shock and predicted that the majority of the inflammatory response to survivable trauma/hemorrhage was due mostly to the underlying tissue trauma induced by cannulation surgery [13]. The model was extended to include details of experimental trauma/hemorrhage in mice (e.g. bleeding rate and target blood pressure), and further validated using a unique, computerized platform for automated hemorrhage that was constructed specifically to test the behavior of this mathematical model [53].
The natural extension from understanding and predicting the inflammatory response is to modulate it in a rational fashion to reduce its detrimental effects. Whereas the modeling work described earlier can help identify targets for therapeutic intervention, and predictive models can be calibrated to account for individual variability while making therapeutic suggestions, synthetic biology can help drive further, clinically-useful developments. Indeed, recent advances have begun to lay the foundations for clinical applications of synthetic biology. These advances have focused on the engineering of synthetic biological circuits in bacterial cells that are introduced into the human host to sense and respond appropriately to transition the host from a diseased to a healthy state [59]. As noted in the review by Warren et al., advances need to be made in the use of mammalian synthetic biology in order to facilitate clinical translation [60–62] (see also Chap. 14).
In the setting of inflammatory diseases, key stumbling blocks to effective therapy involve the variability of individual responses to pro-inflammatory stimuli as well as the detrimental effects of an overly suppressed inflammatory response. Importantly, the blood compartment is both a component of the multiscale positive feedback loop of inflammation → damage → inflammation as well as being easily accessible for therapy [28]. Accordingly, we have envisioned a synthetic biological device using a human cell line to detect the circulating levels of pro-inflammatory mediators in the blood of an individual patient, and respond appropriately by producing an appropriate counter-stimulus—usually a neutralizing protein or receptor antagonist—for a given mediator. We have successfully created stably transfected human hepatocyte (HepG2-derived) cell lines expressing the mouse soluble TNF-α receptor (sTNFR) [63, 64], under control of the mouse variant of the central, TNF-α-responsive transcription factor NF-kB enhancer coupled to a reduced thymidine kinase promoter. These cells are housed in a bioreactor optimized for the growth and differentiation of hepatocytes, that directly connects the with host’s circulatory system. Initial proof-of-concept studies using this bioreactor that produces sTNFR constitutively in a rat bacterial endotoxin infusion model (a quantitative paradigm of acute inflammation that mimics many of the features of sepsis) show promising results for the dynamic modulation of TNF and other pro-inflammatory mediators, as well as ameliorating organ pathophysiology [65]. We suggest that the combination of mechanistic mathematical modeling—of both a given inflammatory disease as well as the effect of this type of biohybrid device on the disease—could be combined to engineer the optimal use of this type of synthetic biohybrid device in order to modulate inflammation systemically.
Moving Beyond the Host: Cross-Species Immune Signaling
The aforementioned examples have focused on the host’s inflammatory response to infection or injury. In the case of infectious diseases, however, the host is not an isolated system and instead part of an entire ecosystem involving the infectious agent/parasite as well as possible vectors. Infectious organisms have evolved alongside the host immune system and developed strategies for evasion and modulation of immunity in the host [66, 67]. In the case of diseases such as dengue and malaria, the addition of an invertebrate vector agent introduces a further layer of complexity in the disease process. Blood plays an expanded role in such diseases, serving not only as the site of immune system coordination within the host but also as an interface for communication and interaction among the parasite, vector, and host [66]. This complex ecology is being reassessed in light of the modern view of the vector as an organism that mounts an immune/inflammatory response in an attempt to control parasite growth, rather than as a willing partner in parasite transmission [66].
Studies show that in addition to the parasite Plasmodium falciparum, proteins and other biomolecules from the host are ingested and can persist in the mosquito vector Anopheles stephensi upon taking a blood meal [66]. Thus, the mosquito vector is likely to be sampling the current immune/inflammatory state of the vertebrate host, in essence getting a snapshot as well as early warning regarding the state of the host’s inflammatory equilibrium (Fig. 11.1). Several such blood-derived factors have been identified, including insulin, insulin-like peptides, and the cytokine transforming growth factor (TGF)-β1[66]. Moreover, these host molecules induce signaling in the mosquito midgut cells and modulate protein expression [68, 69]. For example, mammalian TGF-β1 induces mosquito responses including mitogen-activated protein (MAP) kinase signaling [68]. More recently, we found using a Luminex™ assay for multiple cytokines and chemokines that interleukin (IL)-10 was selectively retained in the mosquito midgut for up to five hours post-blood meal (unpublished observations). In vitro studies also showed that administration of human IL-10 can alter MAPK signaling in mosquito cells (Luckhart et al. unpublished observations). The full gamut of interspecies signaling factors is likely to include DAMPs and other inflammation-related molecules as well (Fig. 11.1). Below, we discuss these findings in greater detail.
TGF-β1 has been identified as a central player in the immune response to parasite infection within the host [70]. However, much less is known about the converse, namely the possible role of TGF-β1 on mosquito immunity and physiology. Human TGF-β1 ingested by the Anopheles stephensi mosquito via a blood meal was shown to induce expression of the mosquito homolog of the inducible nitric oxide synthase, AsNOS [71]. Inducible NOS is often associated with mammalian host defense responses to malaria, and studies have shown that the mosquito also regulates parasite development through complex, multiphasic expression of AsNOS [72]. Several additional observations including evidence of feedback regulation by the MAPK MEK (MAP kinase kinase)/extracellular signal-regulated kinase (ERK), along with dichotomous dose-dependent effects of mammalian TGF-β1 on AsNOS induction and parasite growth suggested that computational modeling might be beneficial in clarifying the underlying mechanisms [71]. An initial Boolean model of the system predicted oscillations in AsNOS as well as MEK/ERK, which was one possible mechanistic model consistent with experimental data. An ODE model of the same system gave quantitative predictions that fit reasonably well with the data. This model also highlighted the necessity of a persistent presence of TGF-β1 to drive the multiphasic response. However, experimental data had previously suggested that the half-life of TGF-β1 may be much shorter than the observed multiphasic time course of AsNOS. This discrepancy was reconciled with the model-generated hypothesis of an endogenous mosquito TGF-β1-like molecule that is induced by exogenous mammalian TGF-β1 and can drive the long-term AsNOS response. Indeed, the mosquito homolog of TGF-β1, As60A [73] was shown to have the same multiphasic dynamics that the model predicted the hypothesized TGF-β1-like molecule must have in order to maintain the observed AsNOS response [74]. These studies begin to provide insight into some of the conserved, cross-species mechanisms of immune modulation between the mammalian host and mosquito vector. Notably, they highlight a new role for the blood as a medium for the interface and biological communication between species, with particular implications for vector-borne diseases.
In keeping with the goal of translational systems biology, we wish to use computational modeling and analysis for the development of new therapeutic strategies. In the decade since the publication of the genomes of the malaria parasite(s) and of Anopheles gambiae, another major malaria mosquito vector, there have been several functional and comparative genomic analyses that have helped uncover regulatory networks through correlative studies [75–80]. Some of these studies have focused on the interface between vector and parasite, identifying gene clusters/networks responsible for the mosquito’s control of parasite growth [81]. However, most modeling work in malaria has been focused mainly on epidemiological aspects of the disease or very coarse-grained mechanistic modeling of host–pathogen–vector interactions [30], rather than models on the intra- and intercellular scale that build directly from the genomic studies or other quantitative experimental data. As outlined in Sect. 11.2.1, and partially illustrated in the preceding paragraph, a systematic approach starting with data-driven modeling and correlative studies that inform mechanistic models and analyses can help build a comprehensive understanding of the molecular and cellular mechanisms underlying the interspecies immune control of malaria parasite. This approach is essential for identifying master regulators in the mosquito vector that can point to therapeutic targets for disease control via genetic modification.
Just as we described the dynamic modulation of inflammation in the host via a bioreactor, we seek to modulate the interspecies immune response to infection via the use of transgenic mosquitoes, ideally at the blood-feeding interface. Genetically modified mosquito (GMM) vectors have become an attractive option for disease control in the past decade as efforts to eradicate mosquitoes or modulate human immunity to malaria infection have been met with reduced efficacy and other challenges. Key to the success of a strategy involving GMMs is ensuring that the modification remains dominant and spreads throughout the population while maintaining the fitness of the mosquito. Recent studies have generated mosquitoes with increased parasite killing but with detrimental effects on fitness [82, 83]. These studies are more descriptive of the phenotype than the underlying mechanism driving it, and much remains to be learned about the pathways driving the observed response. Thus, a systems-level understanding of blood factor-modulated immune response of the mosquito is needed to account for the trade-offs between parasite killing and mosquito fitness in potential interventions.
Conclusions and Future Prospects
The study of the inflammatory response dates back to Roman times, when it was first characterized by its physical manifestations of increased temperature, redness, pain, and swelling. Centuries of research have increased our understanding of inflammation beyond description of its symptoms, and unveiled an ever-increasing complexity underlying this primordial defense mechanism. The modern view of inflammation is that of a multifaceted communication process that manifests across multiple compartments of the body and multiple biological scales [28]. The blood is an important compartment among these, and serves at least three different functions in innate immunity. It is the medium through which inflammation progresses in its early stages as circulating monocytes and other inflammatory cells are recruited to sites of injury/infection. In settings of dysregulated or overexuberant inflammation, spillover of inflammatory mediators from the site of injury to the blood can contribute to a positive feedback, increasing systemic inflammation. Finally, inflammatory mediators in the host’s blood can transfer to blood-feeding vectors and directly modulate the immune response of the vector in a complex host–vector–parasite interaction. The highly complex nature of the immune response to biological stress and the multifaceted role of the circulatory system in this response are perhaps to blame for the lack of efficient and/or successful therapies for diseases such as sepsis, trauma, and MODS. A systems approach to inflammation can be helpful, perhaps even necessary, for the identification of better therapeutic strategies by taking advantage of both data-driven and mechanistic modeling. The methods highlighted in this chapter can provide novel insights into the innate immune system, increasing our understanding to suggest targets for rational modulation of inflammation as well as providing predictive simulations on which to base further basic research, drug discovery, and clinical trials.
Future possibilities include design of synthetic biological circuits for dynamic, individualized modulation of inflammation in the host, as well as control of the host–pathogen–vector interface to eliminate parasite transmission in vector-borne disease. While much progress remains to be made in order to realize these far-reaching goals, recent advances offer a promising outlook for the future of translational systems biology of inflammation.
Abbreviations
- ABM:
-
Agent-based model
- AsNOS:
-
Anopheles stephensi nitric oxide synthase
- DAMP:
-
Damage-associated molecular pattern molecule
- DBN:
-
Dynamic Bayesian Networks
- GMM:
-
Genetically modified mosquito
- MODS:
-
Multiple organ dysfunction syndrome
- ODE:
-
Ordinary differential equations
- PCA:
-
Principal component analysis
- RBM:
-
Rule-based model
- sTNFR:
-
Soluble tumor necrosis factor-α receptor
- TNF-α:
-
Tumor necrosis factor-α
References
Medzhitov R. Origin and physiological roles of inflammation. Nature. 2008;454(7203):428–35.
Brown KL, Cosseau C, Gardy JL, Hancock REW. Complexities of targeting innate immunity to treat infection. Trends Immunol. 2007;28(6):260–6.
Marshall JC. Inflammation, coagulopathy, and the pathogenesis of multiple organ dysfunction syndrome. Crit Care Med. 2001;29(7 Suppl):S99–S106.
Jarrar D, Chaudry IH, Wang P. Organ dysfunction following hemorrhage and sepsis: mechanisms and therapeutic approaches (Review). Int J Mol Med. 1999;4(6):575–83.
Waxman K. Shock: ischemia, reperfusion, and inflammation. New Horiz. 1996;4(2):153–60.
Peitzman AB, Billiar TR, Harbrecht BG, Kelly E, Udekwu AO, Simmons RL. Hemorrhagic shock. Curr Probl Surg. 1995;32(11):925–1002.
Namas R, Ghuma A, Torres A, Polanco P, Gomez H, Barclay D, et al. An adequately robust early TNF-a response is a hallmark of survival following trauma/hemorrhage. PLoS ONE. 2009;4(12):e8406.
Nathan C. Points of control in inflammation. Nature. 2002;420(6917):846–52.
Vodovotz Y, Csete M, Bartels J, Chang S, An G. Translational systems biology of inflammation. PLoS Comput Biol. 2008;4:1–6.
Mi Q, Constantine G, Ziraldo C, Solovyev A, Torres A, Namas R, et al. A dynamic view of trauma/hemorrhage-induced inflammation in mice: principal drivers and networks. PLoS ONE. 2011;6:e19424.
Namas R, Zamora R, Namas R, An G, Doyle J, Dick TE, et al. Sepsis: something old, something new, and a systems view. J Crit Care. 2012;27(3):314e1–11.
Chow CC, Clermont G, Kumar R, Lagoa C, Tawadrous Z, Gallo D, et al. The acute inflammatory response in diverse shock states. Shock. 2005;24:74–84.
Lagoa CE, Bartels J, Baratt A, Tseng G, Clermont G, Fink MP, et al. The role of initial trauma in the host’s response to injury and hemorrhage: insights from a comparison of mathematical simulations and hepatic transcriptomic analysis. Shock. 2006;26:592–600.
Reynolds A, Rubin J, Clermont G, Day J, Vodovotz Y, Ermentrout BG. A reduced mathematical model of the acute inflammatory response: I. Derivation of model and analysis of anti-inflammation. J Theor Biol. 2006;242(1):220–36.
Torres A, Bentley T, Bartels J, Sarkar J, Barclay D, Namas R, et al. Mathematical modeling of post-hemorrhage inflammation in mice: studies using a novel, computer-controlled, closed-loop hemorrhage apparatus. Shock. 2008;32(2):172–8.
Vodovotz Y, An G. Systems biology and inflammation. In: Yan Q, editor. Systems biology in drug discovery and development: methods and protocols. Totowa:Springer; 2009. 181–201.
Mi Q, Li NYK, Ziraldo C, Ghuma A, Mikheev M, Squires R, et al. Translational systems biology of inflammation: potential applications to personalized medicine. Personal Med. 2010;7:549–59.
Chen GY, Nuñez G. Sterile inflammation: sensing and reacting to damage. Nat Rev Immunol. 2010;10(12):826–37.
Parker SJ, Watkins PE. Experimental models of gram-negative sepsis. Br J Surg. 2001;88(1):22–30.
Bellingan G. Inflammatory cell activation in sepsis. Br Med Bull. 1999;55(1):12–29.
Jones AL, Selby P. Tumour necrosis factor: clinical relevance. Cancer Surv. 1989;8(4):817–36.
Cavaillon JM. Cytokines and macrophages. Biomed Pharmacother. 1994;48(10):445–53.
Kox WJ, Volk T, Kox SN, Volk HD. Immunomodulatory therapies in sepsis. Intensive Care Med. 2000;26 (Suppl 1):S124–8.
Dinarello CA. Proinflammatory cytokines. Chest. 2000;118(2):503–8.
Pinsky MR. Sepsis: a pro- and anti-inflammatory disequilibrium syndrome. Contrib Nephrol. 2001;(132):354–66.
Baugh JA, Bucala R. Mechanisms for modulating TNF alpha in immune and inflammatory disease. Curr Opin Drug Discov Dev. 2001;4(5):635–50.
Chen G, Goeddel DV. TNF-R1 signaling: a beautiful pathway. Science. 2002;296(5573):1634–5.
An G, Nieman G, Vodovotz Y. Computational and systems biology in trauma and sepsis: current state and future perspectives. Int J Burns Trauma. 2012;2:1–10.
Csete ME, Doyle JC. Reverse engineering of biological complexity. Science. 2002;295(5560):1664–9.
Mideo N, Day T, Read AF. Modelling malaria pathogenesis. Cell Microbiol. 2008;10(10):1947–55.
Vodovotz Y, Constantine G, Rubin J, Csete M, Voit EO, An G. Mechanistic simulations of inflammation: current state and future prospects. Math Biosci. 2009;217(1):1–10.
Mesarovic MD, Sreenath SN, Keene JD. Search for organising principles: understanding in systems biology. Syst Biol (Stevenage). 2004;1(1):19–27.
Janes KA, Yaffe MB. Data-driven modelling of signal-transduction networks. Nat Rev Mol Cell Biol. 2006;7(11):820–8.
Kitano H. Systems biology: a brief overview. Science. 2002;295(5560):1662–4.
Arkin A, Schaffer D. Network news: innovations in 21st century systems biology. Cell. 2011;144(6):844–9.
Mac Nally R. Regression and model-building in conservation biology, biogeography and ecology: the distinction between-and reconciliation of-“predictive” and “explanatory” models. Biodivers Conserv. 2000;9(5):655–71.
Janes KA, Gaudet S, Albeck JG, Nielsen UB, Lauffenburger DA, Sorger PK. The response of human epithelial cells to TNF involves an inducible autocrine cascade. Cell. 2006;124(6):1225–39.
Vodovotz Y, Constantine G, Faeder J, Mi Q, Rubin J, Bartels J, et al. Translational systems approaches to the biology of inflammation and healing. Immunopharmacol Immunotoxicol. 2010;32(2):181–95.
Angeli D, Ferrell JE, Sontag ED. Detection of multistability, bifurcations, and hysteresis in a large class of biological positive-feedback systems. Proc Natl Acad Sci U S A. 2004;101(7):1822–7.
Clermont G, Chow CC, Kumar R, Vodovotz Y. Mathematical simulation of the innate immune response. Crit Care Med. 2001;29(12Suppl):A111.
Bagci EZ, Vodovotz Y, Billiar TR, Ermentrout GB, Bahar I. Bistability in apoptosis: roles of Bax, Bcl−2, and mitochondrial permeability transition pores. Biophys J. 2006;90(5):1546–59.
Day J, Rubin J, Vodovotz Y, Chow CC, Reynolds A, Clermont G. A reduced mathematical model of the acute inflammatory response II. Capturing scenarios of repeated endotoxin administration. J Theor Biol. 2006;242(1):237–56.
Marino S, Hogue IB, Ray CJ, Kirschner DE. A methodology for performing global uncertainty and sensitivity analysis in systems biology. J Theor Biol. 2008;254(1):178–96.
Kurata H, El-Samad H, Iwasaki R, Ohtake H, Doyle JC, Grigorova I, et al. Module-based analysis of robustness tradeoffs in the heat shock response system. PLoS Comput Biol. 2006;2(7):e59.
Wilensky U. NetLogo. Center for connected learning and computer-based modeling, Northwestern University. Evanston, IL. 1999. http://ccl.northwestern.edu/netlogo/.
Solovyev A, Mikheev M, Zhou L, Dutta-Moscato J, Ziraldo C, An G, et al. SPARK: a framework for multi-scale agent-based biomedical modeling. Int J Agent Technol Syst. 2010;2:18–30.
An G, Hunt CA, Clermont G, Neugebauer E, Vodovotz Y. Challenges and rewards on the road to translational systems biology in acute illness: four case reports from interdisciplinary teams. J Crit Care. 2007;22:169–75.
An G, Faeder J, Vodovotz Y. Translational systems biology: introduction of an engineering approach to the pathophysiology of the burn patient. J Burn Care Res. 2008;29:277–85.
Kauvar DS, Wade CE. The epidemiology and modern management of traumatic hemorrhage: US and international perspectives. Crit Care. 2005;9 Suppl 5:S1–S9.
Kauvar DS, Lefering R, Wade CE. Impact of hemorrhage on trauma outcome: an overview of epidemiology, clinical presentations, and therapeutic considerations. J Trauma. 2006;60(6 Suppl):S3–11.
Prince JM, Levy RM, Bartels J, Baratt A, Kane JM III, Lagoa C, et al. In silico and in vivo approach to elucidate the inflammatory complexity of CD14-deficient mice. Mol Med. 2006;12(4–6):88–96.
Kumar R, Chow CC, Bartels JD, Clermont G, Vodovotz Y. A mathematical simulation of the inflammatory response to anthrax infection. Shock. 2008;29(1):104–11.
Torres A, Bentley T, Bartels J, Sarkar J, Barclay D, Namas R, et al. Mathematical modeling of posthemorrhage inflammation in mice: studies using a novel, computer-controlled, closed-loop hemorrhage apparatus. Shock. 2009;32(2):172–8.
Constantine G, Buliga M, Vodovotz Y, Bohnen N, Clermont G. Time varying patterns of organ failure. Int J Contemp Math Sci. 2010;5:2263–72.
Kumar R, Clermont G, Vodovotz Y, Chow CC. The dynamics of acute inflammation. J Theor Biol. 2004;230:145–55.
Arciero J, Rubin J, Upperman J, Vodovotz Y, Ermentrout GB. Using a mathematical model to analyze the role of probiotics and inflammation in necrotizing enterocolitis. PLoS ONE. 2010;5:e10066.
Vodovotz Y, Chow C, Bartels J, Lagoa C, Kumar R, Day J, Rubin J, Ermentrout B, Riviere B, Yotov I, Constantine G, Billiar T, Fink M, Clermont G. Mathematical simulations of sepsis and trauma. Proceedings of the 11th Congress of the European Shock Society. 2005.
Vodovotz Y, Chow CC, Bartels J, Lagoa C, Prince J, Levy R, et al. In silico models of acute inflammation in animals. Shock. 2006;26:235–44.
Ruder WC, Lu T, Collins JJ. Synthetic biology moving into the clinic. Science. 2011;333(6047):1248–52.
Arkin A, Fletcher D. Fast, cheap and somewhat in control. Genome Biol. 2006;7(8):114.
Karlsson M, Weber W, Fussenegger M. Design and construction of synthetic gene networks in mammalian cells. In: Weber W, Fussenegger M, editors. Synthetic gene networks, vol. 813. Humana Press (New York, NY).; 2012. pp. 359–76.
Weber W, Fussenegger M. Emerging biomedical applications of synthetic biology. Nat Rev Genet. 2012;13(1):21–35.
Fernandez-Botran R, Sun X, Crespo FA. Soluble cytokine receptors in biological therapy. Expert Opin Biol Ther. 2002;2(6):585–605.
Larrick JW, Wright SC. Native cytokine antagonists. Baillieres Clin Haematol. 1992;5(3):681–702.
Namas R, Mikheev M, Yin J, Over P, Young M, Constantine G, et al. Biohybrid device for the systemic control of acute inflammation. Disrupt Sci Technol. 2012;1(1).
Akman-Anderson L, Vodovotz Y, Zamora R, Luckhart S. Bloodfeeding as an Interface of mammalian and arthropod immunity. In: Beckage N, edtior. Insect Immunology. San Diego: Elsevier; 2007. pp. 149–177.
Gooding LR. Virus proteins that counteract host immune defenses. Cell. 1992;71(1):5–7.
Surachetpong W, Singh N, Cheung KW, Luckhart SMAPK. ERK signaling regulates the TGF-β1-dependent mosquito response to Plasmodium falciparum. PLoS Pathog. 2009;5(4):e1000366.
Surachetpong W, Pakpour N, Cheung KW, Luckhart S. Reactive oxygen species-dependent cell signaling regulates the mosquito immune response to Plasmodium falciparum. Antioxid Redox Signal. 2011;14(6):943–55.
Omer FM, Kurtzhals JAL, Riley EM. Maintaining the immunological balance in parasitic infections: a role for TGF-β. Parasitol Today. 2000;16(1):18–23.
Luckhart S, Lieber MJ, Singh N, Zamora R, Vodovotz Y. Low levels of mammalian TGF-β1 are protective against malaria parasite infection, a paradox clarified in the mosquito host. Exp Parasitol. 2008;118:290–6.
Luckhart S, Vodovotz Y, Cui L, Rosenberg R. The mosquito Anopheles stephensi limits malaria parasite development with inducible synthesis of nitric oxide. Proc Natl Acad Sci U S A. 1998;95:5700–5.
Crampton AL, Luckhart S. Isolation and characterization of As60 A, a transforming growth factor- β gene, from the malaria vector Anopheles stephensi. Cytokine. 2001;13(2):65–74.
Price I, Ermentrout B, Zamora R, Wang B, Azhar N, Mi Q, et al. In vivo, in vitro, and in silico studies suggest a conserved immune module that regulates malaria parasite transmission from mammals to mosquitoes. J Theor Biol. 2013;334:172–86.
Date SV, Stoeckert CJ. Computational modeling of the Plasmodium falciparum interactome reveals protein function on a genome-wide scale. Genome Res. 2006;16(4):542–9.
LaCount DJ, Vignali M, Chettier R, Phansalkar A, Bell R, Hesselberth JR, et al. A protein interaction network of the malaria parasite Plasmodium falciparum. Nature. 2005;438(7064):103–7.
Hu G, Cabrera A, Kono M, Mok S, Chaal BK, Haase S, et al. Transcriptional profiling of growth perturbations of the human malaria parasite Plasmodium falciparum. Nat Biotech. 2010;28(1):91–8.
Osta MA, Christophides GK, Vlachou D, Kafatos FC. Innate immunity in the malaria vector Anopheles gambiae: comparative and functional genomics. J Exp Biol. 2004;207(15):2551–63.
Zdobnov EM, von Mering C, Letunic I, Torrents D, Suyama M, Copley RR, et al. Comparative genome and proteome analysis of Anopheles gambiae and Drosophila melanogaster. Science. 2002;298(5591):149–59.
Winzeler EA. Applied systems biology and malaria. Nat Rev Micro. 2006;4(2):145–51.
Osta MA, Christophides GK, Kafatos FC. Effects of mosquito genes on Plasmodium development. Science. 2004;303(5666):2030–2.
Corby-Harris V, Drexler A, Watkins de Jong L, Antonova Y, Pakpour N, Ziegler R, et al. Activation of Akt signaling reduces the prevalence and intensity of malaria parasite infection and lifespan in Anopheles stephensi mosquitoes. PLoS Pathog. 2010;6(7):e1001003.
Dong Y, Das S, Cirimotich C, Souza-Neto JA, McLean KJ, Dimopoulos G. Engineered Anopheles immunity to Plasmodium infection. PLoS Pathog. 2011;7(12):e1002458.
Acknowledgments
This work was supported in part by the National Institutes of Health grants R01GM67240, P50GM53789, R33HL089082, R01HL080926, R01AI080799, R01HL76157, R01DC008290, and UO1 DK072146; National Institute on Disability and Rehabilitation Research grant H133E070024; a Shared University Research Award from IBM, Inc.; and grants from the Commonwealth of Pennsylvania, the Pittsburgh Life Sciences Greenhouse, and the Pittsburgh Tissue Engineering Initiative/Department of Defense.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this chapter
Cite this chapter
Azhar, N., Vodovotz, Y. (2014). Innate Immunity in Disease: Insights from Mathematical Modeling and Analysis. In: Corey, S., Kimmel, M., Leonard, J. (eds) A Systems Biology Approach to Blood. Advances in Experimental Medicine and Biology, vol 844. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-2095-2_11
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2095-2_11
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-2094-5
Online ISBN: 978-1-4939-2095-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)