Abstract
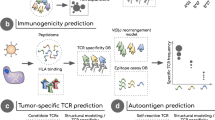
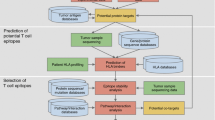
Cancer vaccines are a natural way of fighting the development and progression of cancer as they harness the power of immune system to tweak it into killing cancerous cells. One of the most important agents in an immune system, the cytotoxic T cells (CTL), play a major role and the CTL epitopes in the form of an immunotherapeutic product have been shown to help mount an immune response towards tumor cell destruction. Immunoinformatics and molecular modeling tools have proven powerful towards the prediction of plausible CTL epitopes as well as other epitopes, cutting short the time and cost. We focus on the sequential methodology using these tools as well as some databases to generate a succinct list of enterprising subtype-specific or promiscuous peptide epitopes.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Pronker ES, Weenen TC, Commandeur H, Claassen EHJHM, Osterhaus ADME (2013) Risk in vaccine research and development quantified. PLoS One 8(3):e57755
Tu SH, Huang HI, Lin SI, Liu HY, Sher YP, Chiang SK, Chong P, Roffler S, Tseng GC, Chen HW, Liu SJ (2012) A novel HLA-A2-restricted CTL epitope of tumor-associated antigen L6 can inhibit tumor growth in vivo. J Immunother 35(3):235–244
Bellone S, Anfossi S, O'Brien TJ, Cannon MJ, Silasi DA, Azodi M, Schwartz PE, Rutherford TJ, Pecorelli S, Santin AD (2009) Induction of human tumor-associated differentially expressed gene-12 (TADG-12/TMPRSS3)-specific cytotoxic T lymphocytes in human lymphocyte antigen-A2.1-positive healthy donors and patients with advanced ovarian cancer. Cancer 115(4):800–811
Neumann F, Kubuschok B, Ertan K, Schormann C, Stevanovic S, Preuss KD, Schmidt W, Pfreundschuh M (2011) A peptide epitope derived from the cancer testis antigen HOM-MEL-40/SSX2 capable of inducing CD4+ and CD8+ T-cell as well as B-cell responses. Cancer Immunol Immunother 60(9):1333–1346
Gritzapis AD, Fridman A, Perez SA, La Monica N, Papamichail M, Aurisicchio L, Baxevanis CN (2009) HER-2/neu (657-665) represents an immunogenic epitope of HER-2/neu oncoprotein with potent antitumor properties. Vaccine 28(1):162–170
Mishra S, Sinha S (2006) Prediction and molecular modeling of T cell epitopes derived from placental alkaline phosphatase for use in cancer immunotherapy. J Biomol Struct Dyn 24(2):109–121
Mishra S, Sinha S (2009) Immunoinformatics and modeling perspective of T cell epitope-based cancer immunotherapy: a holistic picture. J Biomol Struct Dyn 27(3):293–306
Jørgensen KW, Buus S, Nielsen M (2010) Structural properties of MHC class II ligands, implications for the prediction of MHC class II epitopes. PLoS One 5(12):e15877
van der Bruggen P, Stroobant V, Vigneron N, Van den Eynde B (2013) Peptide database: T cell-defined tumor antigens. Cancer Immun 13:15, http://cancerimmunity.org/peptide/
Parker KC, Bednarek MA, Coligan JE (1994) Scheme for ranking potential HLA-A2 binding peptides based on independent binding of individual peptide side-chains. J Immunol 152:163
Rammensee H-G, Friede T, Stevanovic S (1995) MHC ligands and peptide motifs: 1st listing. Immunogenetics 41:178–228
Rammensee, H-G. Bachmann, J., Stevanovic, S. (1997) MHC ligands and peptide motifs. Landes Bioscience (International distributor—except North America). Springer, Heidelberg
Singh H, Raghava GP (2003) ProPred1: prediction of promiscuous MHC class-I binding sites. Bioinformatics 19:1009–1014
Singh H, Raghava GPS (2001) ProPred: prediction of HLA-DR binding sites. Bioinformatics 17(12):1236–1237
Lundegaard C, Lamberth K, Harndahl M, Buus S, Lund O, Nielsen M (2008) NetMHC-3.0: accurate web accessible predictions of human, mouse and monkey MHC class I affinities for peptides of length 8–11. Nucleic Acids Res 36(Web Server issue):W509–W512
Lundegaard C, Lund O, Nielsen M (2008) Accurate approximation method for prediction of class I MHC affinities for peptides of length 8, 10 and 11 using prediction tools trained on 9mers. Bioinformatics 24(11):1397–1398
Nussbaum AK, Kuttler C, Hadeler KP, Rammensee H-G, Schild H (2001) PAProC: a prediction algorithm for proteasomal cleavages available on the WWW. Immunogenetics 53:87–94
Nielsen M, Lundegaard C, Lund O, Kesmir C (2005) The role of the proteasome in generating cytotoxic T cell epitopes: Insights obtained from improved predictions of proteasomal cleavage. Immunogenetics 57(1–2):33–41
Holzhütter HG, Kloetzel P-M (2000) A kinetic model of vertebrate 20S proteasome accounting for the generation of major proteolytic fragments from oligomeric peptide substrates. Biophys J 79:1196–1205
Bhasin M, Raghava GPS (2004) Analysis and prediction of affinity of TAP binding peptides using cascade SVM. Protein Sci 13(3):596–607
Hakenberg J, Nussbaum A, Schild H, Rammensee H-G, Kuttler C, Holzhütter H-G, Kloetzel P-M, Kaufmann SHE, Mollenkopf H-J (2003) MAPPP—MHC-I antigenic peptide processing prediction. Appl Bioinformatics 2(3):155–158
Larsen MV, Lundegaard C, Lamberth K, Buus S, Lund O, Nielsen M (2007) Large-scale validation of methods for cytotoxic T-lymphocyte epitope prediction. BMC Bioinformatics 8:424
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this protocol
Cite this protocol
Mishra, S., Sinha, S. (2014). Immunoinformatics, Molecular Modeling, and Cancer Vaccines. In: De, R., Tomar, N. (eds) Immunoinformatics. Methods in Molecular Biology, vol 1184. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-1115-8_28
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1115-8_28
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-1114-1
Online ISBN: 978-1-4939-1115-8
eBook Packages: Springer Protocols