Abstract
Next-generation sequencing (NGS) is driving our current approaches to molecular diagnosis and the characterization of genetic disorders. The arrival of NGS platforms has made whole-exome sequencing (WES) possible, leading to tremendous improvements in the technology, throughput, cost, and efficiency of variant detection and disease diagnosis. This in turn has brought a paradigm shift in clinical and medical genetics. WES and whole-genome sequencing (WGS) are fast becoming affordable, one-for-all tests and entering the clinical genetics arena. In this chapter, we summarize the evolving roles of WES, which range from its use as a disease diagnostic tool for rare single-gene Mendelian disorders, to a routine screening tool for monitoring disease progression and treatment response in cancer patients, as well as a tool for novel gene discovery in a research setting. This chapter highlights the achievements and limitations of the technology and emphasizes the challenges involved in the implementation of WES assays for clinical care.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Introduction
The paradigm shift from genetics to genomics was put in motion by a revolutionary study that described sequencing of the entire genome of Mycoplasma genitalium in a single run on a Roche 454 instrument [58]. The study revealed a highly parallel sequencing-by-synthesis method capable of sequencing 25 million bases, at 99 % or more accuracy, in a single 4 h run. Subsequently, several high-throughput flow cell-based sequencing methods became commercially available from Illumina (San Diego, California), Roche (454 Life Sciences Corporation, Branford, Connecticut), and Life Technologies (Carlsbad, California). These developments marked the beginning of a new era based on next-generation sequencing (NGS). Simultaneously, several sequence capture or target enrichment methods were evolving to improve the throughput and specificity of sequencing technology [1, 6, 30, 37, 55, 74]. With the rapid development of these advanced sequencing technologies, per-base sequencing costs are declining drastically, to a level at which almost complete resequencing of the human genome is becoming affordable, even in clinical settings [3, 57, 64, 87]. Nevertheless, the infrastructure requirements, analysis burden, and turnaround time requirements involved in clinically interpreting entire patient genomes for mutation detection bear significant issues. Whole-exome sequencing (WES), in contrast, which interrogates the roughly 1 % of the human genome that represents the entire coding region and harbors 85 % or more of causative mutations, is quite feasible and much more affordable in a clinical setting.
The successful implementation of NGS technology in clinical laboratories for diagnostic purposes began with gene panels designed to specifically target and sequence multiple genes related to a particular disorder. Soon several disease specific or phenotype specific gene panels became clinically available [31, 40, 48, 51, 70, 98, 99, 101]. These included highly heterogeneous disorders, such as congenital disorders of glycosylation (CDG), congenital muscular dystrophies (CMD), limb girdle muscular dystrophies, dilated cardiomyopathy, and mitochondrial disorders, each with several subtypes of overlapping phenotypes and associated with a large number of causative genes [2, 98]. Traditional molecular diagnostic approaches for such diseases followed a sequential, Sanger sequencing-based gene-by-gene analysis of known disease-associated genes. However, with the advent of NGS technologies and the decline in per-base sequencing cost, the NGS panel approach has become a significantly cheaper and quicker option, available as a single test. Subsequently, with the availability of better sequence chemistries and easier workflows, NGS technology moved into other clinical arenas, including cancer diagnosis [66], human leukocyte antigen locus characterization [39, 81], and pathogen genome sequencing for the purpose of evaluating resistance [85]. Rapid identification of novel disease genes and the revealing locus and allelic heterogeneity of inherited genetic disorders, both Mendelian and complex, has established WES as a comprehensive clinical test.
In this chapter, we discuss the various roles of WES in clinical medicine and provide an overview of how WES has transformed the diagnostic outlook on genetic disorders. We highlight the major successes and challenges of implementing WES assays in clinical genetics, concluding with a note on the future of whole-exome assays.
Whole-Exome Sequencing: Methodology
Exome Capture and Next-Generation Sequencing
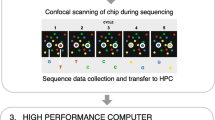
WES refers to sequencing of the entire protein-coding region of the human genome. This is achieved by parallel sequencing of all targeted regions (exons) using NGS technologies. Irrespective of the manufacturer and sequencing platform, the basic methodology or principles involved in WES are similar (Fig. 16.1). First, genomic DNA is fragmented either by optimized sonication or by restriction digestion to generate uniform libraries of DNA strands. This fragmented DNA is then enriched for protein-coding regions of the genome (exons), using unique adapter ligation chemistry that is proprietary to each individual commercial manufacturer [18]. Adapter-ligated DNA fragments are captured and amplified either on a solid surface (bridge amplification on a glass slide) or in solution (emulsion PCR on micro-beads). Finally, different massively parallel sequencing technologies are used to sequence all target DNA regions and produce what are called sequence reads, of different lengths, depending on the technology used. Sequence reads are computationally aligned to a reference exome and analyzed for sequence variations. The experimental design allows for each nucleotide to be represented in a large number of reads, which is referred to as “read depth” or “coverage.” Variant annotation using analytical pipelines helps filter false positives and non-contributive calls to identify causal mutations. WES therefore serves as a comprehensive method for rapid identification of exonic mutations, such as missense, nonsense, splice site, and small deletion and insertion mutations (indels); however, detection of copy number variations (CNVs) and structural variations (SVs) is still an issue.
Basic methodology of exome capture or target enrichment for whole-exome sequencing. The various steps involved are indicated by numbers. In step 1, genomic DNA is randomly fragmented into more or less uniform shorter segments, either by ultrasonication or restriction digestion with enzymes. In step 2, adapters with sequencing motifs and indices are ligated to the fragments. In step 3, biotinylated probes that are specific for target regions (exons) are added and allowed to hybridize. Step 4 involves addition of streptavidin beads to selectively capture all target regions by binding biotin. While the streptavidin beads (with bound target regions) are held by a magnet, unbound nonspecific DNA fragments are separated and washed away. Finally, in step 5, target regions are eluted by denaturation from the biotinylated probes. Although alternative methods for adapter ligation may be available, the basic concept for target (exon) capture is similar
Sequence Analysis and Variant Detection
Massively parallel sequencing of the entire exome generates terabytes of information. Sorting through and making sense of such massive volumes of data to identify causative genes and mutations requires multistep bioinformatics analysis. Upon initial generation of sequence base call files, they are converted into the more commonly used FASTQ file format for storage and later analysis [18]. Several open-source and in-house-developed software programs can be used to align sequence reads to a best-match location of a reference sequence and stored in what is called the BAM (binary alignment) file format [49]. These aligned reads are then processed to call out sequence variants depending on the presence and zygosity of variants. Information from this analysis, which includes inferred single nucleotide polymorphisms (SNPs) and insertions and deletions (indels) along with base coverage, quality, and score, is stored in a different file format termed variant call format (VCF) [22, 50]. Finally, each single call in the entire set of variants is annotated with a variety of customizable information, including gene name, genomic and cDNA coordinates, amino acid change, and functional classification, to help with the interpretation of causative variants [104].
Variant Analysis and Molecular Diagnosis
Analysis of the variants and identification of the disease causative gene and mutations in WES are daunting tasks compared to the traditional single-gene sequencing approach. Several predictive algorithms are being developed and made commercially available, but their reliability and interpretative ability is not well established. Most of the clinical laboratories that offer WES assays currently include various parameters, such as the functional effect of the observed variant, relevance of the gene to the clinical presentation, and mode of inheritance, to filter variant calls through in-house-validated pipelines and algorithms (Fig. 16.2). Finally, short-listed candidate variants are confirmed by the gold standard Sanger sequencing. Confirmed variants may fall into different categories based on previous association and functional effects of the variant (Table 16.1). In the event a new disease gene is identified, disease association requires further evidence. In silico analysis by prediction algorithms based on evolutionary conservation of the amino acid or nucleotide may increase confidence in an association, but is not definitive [18]. Segregation of mutations in the gene with presence of disease among family members may also provide additional evidence, but does not necessarily or fully associate the gene with disease. Functional studies are best, when available, because they may not only establish disease association but also provide insight into disease pathogenesis and treatment options. Alternatively, identification of mutations in the novel gene in unrelated individuals with similar phenotypes by rapid targeted single-gene testing may establish disease association, as well. While diagnostic laboratories focus on finding a pathogenic change in a known disease-associated gene, research testing of exomes is driven by the additional goal of new gene discovery and may include extensive functional analysis to establish disease association with the gene.
Basic pipeline for variant filtration in whole-exome sequencing analysis. Various parameters are included in WES algorithms to filter and remove nonpathogenic and false-positive variants from whole-exome variant data to create a manageable dataset (150–250 variants) that includes the candidate causative mutations. As indicated in the data filtration funnel, variants that do not meet QC metrics, such as those with poor coverage (<20×), are considered less likely to be real, treated as false positives, and therefore filtered. Variants with a minor allele frequency of >0.01 are polymorphisms by definition and less likely to be pathogenic. Silent changes and intronic variants beyond the consensus splice donor/acceptor sequences are less likely to be pathogenic and are often filtered in initial rounds of analysis. Familial variants may also be carefully filtered based on zygosity and segregation pattern. Though the basic parameters followed are common to all commercial and laboratory-developed algorithms, the thresholds and ranges for acceptability may vary. EVS = Exome Variant Server (NHLBI Exome Sequencing Project)
Exome Sequencing: A Transformative Technology
NGS approaches, and especially WES, have created hope for patients who may have already undergone a diagnostic odyssey of invasive approaches and clinical tests, and yet remain in the dark as to the underlying genetic cause of their condition. The potential of WES to provide molecular diagnoses by screening nearly all human exons for mutations was recognized early on, and attempts to explore its diagnostic potential were soon underway, heralding a new era in clinical and medical genetics.
WES as a Diagnostic Assay - Proven Potential
WES has facilitated characterization of several recessive as well as dominant diseases, revealing associations with new disease genes. Recessive traits, which are more commonly highlighted in consanguineous families, are comparatively easier to diagnose and implicate through WES because affected individuals within the family carry causative mutations in segments that are homozygous by descent. For example, in the case of first cousin mating, these regions account for approximately 10 % of the entire exome, thereby restricting the search to this small region. For dominant traits, however, the process is less straightforward. Molecular characterization of dominant traits is complicated by several factors, including reduced penetrance for certain genes, locus heterogeneity, and alleles that affect reproductive fitness. In such scenarios, the finding of independent de novo variants in the same gene among multiple unrelated affected individuals provides considerable evidence for disease association irrespective of allelic heterogeneity. The first successful demonstration of the potency of WES for rare variant identification and disease diagnosis came from an unexpected diagnosis of a patient referred for possible Bartter syndrome [16]. Due to an inconclusive clinical presentation, WES was performed for this individual, and informed variant analysis led to the identification of a homozygous mutation in the SLC26A3 gene. This study provided the first proof of concept of the application of WES for genetic disease diagnosis. Even though the gene was previously known to be disease causing (congenital chloride-losing diarrhea, CLD), the clinical overlap of the patient’s phenotype with that of Bartter syndrome [36] obviated suspicion of the gene. Substantial family information, including that of consanguinity, inheritance mode of the disease, and regions of excessive homozygosity due to identity by descent, helped with the molecular characterization. Moreover, reevaluation of additional study subjects with a presumptive diagnosis of Bartter syndrome identified mutations in SLC26A3. These findings not only established the diagnostic ability of WES but also expanded the phenotypic variability of SLC26A3-associated CLD.
Whole-Exome Sequencing Facilitates Gene Discovery
Traditional gene mapping tools, such as homozygosity mapping, linkage analysis, karyotyping, and copy number variation (CNV) analysis, have led to the identification of new disease genes [41, 44, 45, 102]; however, these methods require analysis of a cohort of multiple unrelated affected individuals to narrow down genomic regions of interest, before finally zeroing in on the candidate gene. In contrast, WES of a single family or a parent–proband trio can result in rapid gene identification. This was first reported approximately 5 years after the launch of the technology in 2005 [72]. Using WES, two potentially pathogenic variants were identified in a novel candidate gene, DHODH, thus implicating the gene in the autosomal recessive Miller syndrome. This condition is characterized by severe micrognathia, cleft lip or palate, limb defects, coloboma, and supernumerary nipples [65]. Even though the disease had been described several decades ago, not much about the causal gene or mode of inheritance was known until this study. Despite little understanding of how DHODH mutations cause Miller syndrome, the subsequent identification of mutations in additional patients by targeted gene sequencing confirmed disease association without functional analysis. Shortly thereafter, another novel disease gene association was reported by the same group, which identified MLL2 (KMT2D) to be the causative gene for Kabuki syndrome [71]. These findings strongly suggested that exome sequencing of a small number of affected individuals from unrelated kindred, or of multiple individuals from a single affected family, could be a powerful and efficient strategy for the identification of rare disease genes.
From Medical Genetics to Medical Genomics: A Shift in Paradigm
Beginning in early 2008, the NIH’s Undiagnosed Diseases Program (UDP) began offering clinical WES as a pilot program, with initial funds totaling $280,000 [60]. UDP’s explicit objectives were to provide molecular diagnosis to patients who remained undiagnosed despite thorough workup and to discover novel disease genes and disorders to gain insight into the pathogenesis of the clinical manifestations. After receiving several thousand applications from prospective participants, 160 individuals were enrolled, and the huge task of deciphering the underlying genetic causes began. Included was a healthy Colombian couple with two sons affected with an uncharacterized neurological illness, presenting with seizures, tremors, and several other complications. When one of the sons succumbed to the disease, the second son of the family was enrolled in the above-mentioned multi-institute initiative in hopes of identifying the underlying cause. After collaborative efforts for more than a year, a definitive diagnosis came from WES analysis. Furthermore, the molecular diagnosis was also established for almost 25 % (39/160) of the enrolled individuals overall. Novel disease genes, including NT5E, associated with arterial calcification disorder [90], and HINT3, an aprataxin-related gene causative of a familial distal myopathy [28], were identified, as well. Most of the diagnoses made, however, included known rare (≤1 in 10,000) or ultra-rare (<60 cases reported) diseases in individuals who had previously undergone multiple molecular and/or biochemical genetic tests. UDP’s experience suggested that, with comprehensive phenotypic information, accurate bioinformatics tools, and a methodological approach, WES can be an economical single test for disease diagnosis.
Implementation of Exome Sequencing in Clinical Medicine
Whereas the suitability of WES for clinical medicine was initially debated, the emerging consensus is that the future of diagnostic exome sequencing has already begun [54, 63]. As new genes and diseases are identified through clinical WES, the test is gaining popularity. Expected reductions in cost and improved reimbursement are also likely to mean wider implementation of WES in clinical medicine.
Mendelian Disorders and Exome Sequencing
The conventional approach, still widely in practice, for molecular diagnosis of single-gene Mendelian disorders follows serial interrogation of all exons and exon–intron boundaries of known disease-specific genes via traditional polymerase chain reaction (PCR) amplification and the gold standard Sanger sequencing. Unlike complex traits and disorders such as autism and intellectual disability, which can involve several causative genes and variants, Mendelian disorders are generally associated with mutations in a single gene. With the utilization of clinical genetics and molecular diagnosis, however, locus heterogeneity and overlapping disease phenotypes have shown that, even for Mendelian disorders, making a molecular diagnosis is less straightforward than previously thought. This notion favored the application of multi-gene panels in which all common disease-related genes are interrogated simultaneously through NGS. Consequently, there are now several individual disease gene panels available [2, 40, 98]. Even though the panel approach has reduced the diagnostic odyssey for patients and boosted diagnostic capacity, a substantial fraction of patients still remain without a molecular diagnosis. This can be attributed, in part, to the inability to detect mutations in regulatory and intronic regions. Nevertheless, most such cases are believed to be due to the involvement of previously unknown disease genes. One important feature in support of this is the occurrence of more than 85 % of causative mutations for Mendelian disorders in exonic regions of the genome [12]. This percentage, together with the growing potential of WES as a diagnostic tool, makes it a preferred approach for rare Mendelian disorders with genetic and phenotypic heterogeneity. Notably, however, causative variants detectable by a combination of conventional methodologies, including homozygosity mapping and candidate gene selection, may be missed by WES [10, 69]. Bloch-Zupan et al. [10] report a case of homozygous mutations in the SMOC2 gene, responsible for dental developmental defects, which were initially missed by WES due to poor coverage [10]. Overall, however, whereas homozygosity mapping or linkage analysis may be preferred for consanguineous and large pedigrees, WES is proving to be the most informative of these diagnostic tests [13, 14, 26, 59]. In some cases, WES has provided an accurate molecular diagnosis in patients previously diagnosed with a different disease, further cementing the value of this assay in clinically heterogeneous Mendelian disorders [47]. Besides establishing a molecular diagnosis in patients and providing carrier testing opportunities for family members, the identification of causative mutations in Mendelian diseases also guides patient management and family counseling [4], and opens up opportunities for therapeutic intervention and participation in clinical studies [75]. Finally, the identification of new disease genes and causative mutations contributes to our understanding of disease phenotype, pathogenesis, and gene function [77].
Complex Disorders and Exome Sequencing
Common complex diseases constitute a major part of overall disease burden in the general population. Most common diseases are complex, with extensive genetic heterogeneity resulting in clinically indistinguishable phenotypes. This includes conditions such as autism, intellectual disability, cardiac disease, and diabetes. X-chromosome-linked intellectual disability alone has been associated with more than 100 different genes. Similarly, autism spectrum disorders are linked to multiple genes, with no single gene accounting for more than 1 % of cases [9]. It is obvious that, even more so than for single-gene Mendelian disorders, the WES approach is advantageous for multifactorial and multigenic complex disease characterization. Recently, one single WES study investigating the genetic etiology of autosomal recessive forms of intellectual disability identified 50 novel candidate genes [68]. These include genes encoding proteins involved in transcription, translation, cell-cycle control, and fatty acid and energy metabolism critical for normal brain development and function. The discovery of such novel disease-associated genes not only improves our understanding of the underlying cause of disease manifestations but can also suggest novel targets for therapy and management.
Unlike most Mendelian disorders, diseases with complex genetic etiologies involve coding variants that present as risk factors rather than direct causes of disease. Such risk factors found by traditional methods to date include an APOE genotype that plays a role in late-onset alzheimer’s disease, complement factor H polymorphism in age-related macular degeneration, and an LRRK2 risk variant in Parkinson’s disease [19, 43, 92]. The application of WES to complex disease diagnosis will enable the identification of similar common protein-coding risk alleles, as well as rare risk alleles. Genome-wide association studies (GWAS) have been revolutionary in terms of uncovering common variants associated with complex disorders, but have not satisfactorily explained the heritability of these traits [17, 56, 62, 83]. With the advent of WES, the focus of complex trait genetics has shifted towards low-frequency and rare variants [79, 97], and the link between variants and complex traits is on its way to becoming clearer [11, 23, 27, 73, 80]. The routine use of WES in clinical laboratories will most likely identify more and more rare variants that have a strong causative effect on phenotype, unlike the common variants that, individually, contribute only minimally [24, 42].
Application of WES to Neoplastic Diseases
Historically, pathologists have relied on histomorphology to classify and diagnose neoplasms [8, 21]. Recent progress in cancer genomics, however, has pointed towards the utility of a more granular approach through the identification of genetic alterations common to morphologically diverse tumor types and through the discrimination of subgroups within what was thought to be a single tumor type [7]. Consequently, WES has been applied to tumor diagnostics to obtain a comprehensive picture of copy number alterations (CNAs) and of pathogenic mutations [52]. The potential of WES to detect somatic CNAs in cancer syndromes has been explored, as well [52, 82]. In a study involving 17 matched tumor and normal tissues from patients with metastatic castrate-resistant prostate cancer, targeted WES analysis successfully identified various common CNAs, such as androgen receptor (AR) gain and PTEN loss [52]. This study and others suggest that somatic CNAs that involve the amplification of oncogenes or deletion of tumor suppressors and are significant contributors to cancer etiology can now be monitored more comprehensively using WES than array-based technologies [15]. Unlike germ-line mutations, somatic mutation and CNA detection in cancer are performed by simultaneous exome sequencing of normal and tumor tissue from the same individual, followed by a comparison of copy number ratios of exonic regions in the two sample types [52]. This approach of analyzing the relative coverage (of tumor versus normal sample) distinguishes a true chromosomal deletion from a lack of coverage due to technical limitations. WES thus offers the combined efficiency of both array comparative genomic hybridization (aCGH), which detects CNAs by relative probe frequency [78], and single nucleotide polymorphism (SNP) array, which detects loss of heterozygosity (LOH) and absence of heterozygosity (AOH) by zygosity changes at known SNP loci [61]. Whereas the prohibitive cost and analysis burden of whole-genome sequencing (WGS) have limited its clinical application thus far, successful detection of somatic DNMT3A mutations in acute monocytic leukemia [105], PBRM1 mutations in renal carcinoma [100], BAP1 mutations in metastasizing uveal melanomas [34], and AR, NCOA2, PTEN, RB1, and TP53 CNAs in prostate cancer [94] by WES are confirming it as a cancer diagnostic and monitoring assay option.
There are several advantages to using WES for cancer genomics. First, it provides an exon-level resolution of CNAs. Second, the vast data available through comprehensive sequencing projects such as The Cancer Genome Atlas (TCGA) can be leveraged because whole-exome data for thousands of cancer cases from multiple studies are publicly available [95]. This makes integrative cancer detection strategies possible and drives personalized medicine approaches. Genotype-directed therapies are transforming cancer care, as seen with several drugs and target inhibitors in various cancer types, including chronic myeloid leukemia, colorectal adenocarcinoma, and melanoma [25, 53, 76]. The role of coexisting or co-occurring passenger mutations, separate from the driver mutations that actually cause the clonal expansion of cancer cells, is also being investigated so the two can be distinguished [5, 33]. Comparison of WES data across multiple patients is expected to contribute to the teasing out of the two, which could in turn translate into new drug targets. Despite these advantages, WES still has some limitations. These are primarily pertaining to coverage of certain exons and of genes with complex sequence context, as a result of which some mutations and CNAs may be missed. Additionally, CNAs involving gene-poor regions may not be detected due to assay design. Gene fusion events or chimeric gene products unique to cancer etiology and the more frequent large chromosomal aberration events, such as translocations, large deletions, or inversions, are not detected by WES. A comprehensive approach of various NGS technologies including WES, WGS, and transcriptome analysis is being explored, but clinical applicability is still rudimentary [67, 86, 93, 94].
From Diagnosis to Therapy: Advances in Clinical Care
Despite the proven potential of WES for clinical diagnostic purposes, one common criticism of the technology is the lack of evidence for its clinical usefulness. Pharmacogenomics is one area in which WES is expected to play a major role, especially by identifying variants that contribute to genotype-specific responses to drugs. One such example is related to the substitution of glutamic acid for valine at position 600 (p.V600E) in the BRAF gene in individuals with malignant melanoma [20]. This specific mutation acts by conferring a constant flux through the mitogen-activated protein kinase (MAPK) pathway, thereby promoting malignancy. The genotype-specific drug vemurafenib (PLX4032), recently approved by the FDA, is used for targeted intervention of metastatic melanoma [46, 106]. Eventually, however, tumor cells were found to develop resistance to the drug over time, but in a cohort of 20 melanoma patients treated with vemurafenib, WES identified the underlying cause for the development of drug resistance: a gain in copy number (by 2–13 times) of the mutant p.V600E BRAF allele [88].
Several other targeted therapies, such as imatinib for chronic myeloid leukemia, trastuzumab for breast cancer, irinotecan and panitumab for colorectal cancer, and erlotinib for lung cancer, may all be monitored for their treatment effect and resistance development using WES. Implementation of WES in the context of personalized medicine is highlighted by a recent study reporting a novel genetic risk factor linked to the VACTERL association [89]. A heterozygous mutation in the CPSI gene, identified by WES in monozygotic twins, is suspected of being the risk factor associated with the severe pulmonary artery hypertension observed post-surgery in the twin who underwent surgery. Generally, homozygous or compound heterozygous mutations in CPSI are associated with a rare urea cycle disorder; however, through WES analysis the authors clarified that there were no discordant de novo mutations between the two twins and that the observed complication must have been due to the combination of the observed heterozygous variant and an environmental trigger: in this case, surgery.
Limitations and Challenges of Implementing Exome Sequencing Assays
Despite being quite comprehensive, WES has yet to overcome several technical and analytic challenges before it can replace the current gold standard of Sanger sequencing, or even targeted NGS panels. These challenges are summarized here. The first and foremost technical challenge is the inefficiency to capture and sequence all target exons. Contrary to what is suggested by its name, WES currently misses around 5–8 % of the human exome because of low or no coverage [16]. Most of this is explained by sequence context, such as with high or low GC content or the presence of highly homologous pseudogenes [38]. Capture of all target exons is, of course, essential to avoid false-negative interpretations due to the presence of potentially causative mutations in missed exons. Highly repetitive sequences, which include interspersed repeats and tandem repeats, constitute more than half of the human genome. These highly homologous regions are co-enriched and co-sequenced along with the target regions [96]. This challenge may be countered by increasing the sequence read size, which is still limited with current NGS technologies. However, several alternative approaches, such as paired end sequencing and correlation of average read depth differences to detect repeat regions, are being explored [96]. A second challenge is storage and management of the vast amount of sequencing data generated by the technology. This demands a large investment in infrastructure and technology, which is a major strain for diagnostic laboratories. A third limitation is the variant detection capability of WES. With high coverage and read depth, point mutations and small indels in exonic regions can be detected with high efficiency, but those in regulatory regions are not. In addition, larger multi-exon or multi-gene deletions and duplications, which contribute to a significant proportion of the mutation spectrum for several genes, as well as gene-fusion or chimeric events common in cancer, are not efficiently detected. Besides variant detection capability, another major challenge of the test involves assessment of the clinical implications of variants identified. Most of the observed variants may not be clinically predictable or actionable due to lack of sufficient evidence. However, with the routine practice of WES and accumulation of relevant information, this concern would gradually be reduced. The fifth challenge to implementing WES assays in clinical care is the requirement of additional training for physicians to help them interpret test results and reports. With a more comprehensive set of variants available for consideration in the patient’s clinical context, clinicians who see the patient, if trained in this area, would be able to make the optimal interpretation as to the causative gene. Alternatively or ideally simultaneously, extensive phenotypic information may be collected beforehand and made available to the pathologists and laboratorians interpreting the data. Finally, a considerable challenge facing the clinics and laboratories that offer these tests is the constantly changing technology. Recently, members of the Standardization of Clinical Testing workgroup (Nex-StoCT) have laid out guidelines for the validation and implementation of NGS-based tests [29]. With NGS technology changing all the time, however, these aspects also change and can become a hurdle to implementation.
Despite the challenges and limitations, WES and WGS have stirred tremendous interest, with the future of clinical care promising expedited diagnosis and more personalized medicine. Moreover, implementation of WES in medical practice will potentially aid the advancement of our understanding of human biology and pathogenesis.
A Look to the Future of Whole-Exome Assays
Current commercially available NGS technologies have already revolutionized the diagnostic capacity of modern clinical genetics. Nevertheless, advanced so-called “third-generation” sequencing technologies, such as Helicos Heliscope (Helicos Biosciences Corporation, Cambridge, MA), PacBio SMRT (Pacific Biosciences, California), and Nanopore sequencers (Oxford Nanopore Technologies, Oxford, UK), are being actively developed to further improve genomic sequencing applications [32]. These third-generation sequencing platforms differ from the current technologies in that the initial target capture and enrichment step, which involves DNA amplification, is no longer required. The input patient DNA is sequenced and analyzed at the single-molecule level with the help of engineered protein polymerases [32]. This will not only cut cost and turnaround time but also have the added advantage of avoiding any in vitro amplification bias. Upon thorough validation and optimization of their diagnostic ability, these future technologies promise to move today’s medical practice to the anticipated next level of care.
Currently, even more so than the sequencing technology and needed coverage improvements, the progress in data analysis tools and candidate variant filtration is of major concern. WES alone, which interrogates about 1 % of the human genome, returns a list of about 20,000 variant calls [91]. Family information, such as the mode of inheritance within a family, linkage analysis or variant data, i.e., the WES profile of unaffected family members, helps eliminate familial normal variations and track down disease-causing mutations [60], but performing additional tests including WES on multiple family members increases diagnostic costs and is not ideal for a variety of reasons. As more and more exomes are analyzed and sequence variants reported in publicly available databases, however, variant analysis and disease diagnosis by WES will certainly become easier and faster.
Meanwhile, with the implementation of WES and NGS technologies in clinical pathology becoming more common, the need for trained pathologists capable of interpreting the data and assessing the potential impact on an individual’s health is growing. The training of future pathologists is now under discussion, and teaching curricula in genomics and personalized medicine are being actively developed for residents [35, 84]. A national committee of Pathology Program Directors and other experts has also recently formed to develop model curricula and promote their widespread implementation [35, 103]. The implementation of WES and WGS in clinical practice has, therefore, added a new dimension to the already multifaceted roles of pathologists.
Conclusions
With more than 85 % of causative mutations harbored in as little as 1 % of the entire human genome, the use of WES as the most efficient strategy for disease diagnosis seems well justified. Even though WGS has the potential to identify CNVs and point mutations in exons, as well as in regulatory regions of the introns, the cost, time, and the analysis burden currently involved has meant WGS is on hold for clinical implementation, at least for now. Substantial proof-of-principle studies and evidence of diagnostic capability, affordability, and feasibility in the clinical setting have supported the use of WES. Currently, it is offered for clinical diagnosis by multiple major clinical laboratories across the USA, and as the technology improves and becomes less expensive, more laboratories are beginning to develop the test.
Clinicians who contemplate ordering a WES assay should first consider other available tests, such as relatively comprehensive gene panels. Gene panels, which interrogate only a limited number of genes, each more or less associated with the patient’s clinical presentation, more completely retain the integrity of the individual’s genetic information. Appropriate ethical guidelines and data-masking features during data analysis will likely overcome this difference eventually and make WES widely acceptable for rare diseases, cancer, and prenatal and infectious disease diagnosis. Finally, reductions in cost, more robust technologies, and improved data storage processes will soon make clinical WGS feasible, as well. The future of medical care can be envisioned as an integrated approach, with pathologists, geneticists, and other physicians all contributing to make informed decisions about patient management and treatment.
References
Albert TJ, Molla MN, Muzny DM, Nazareth L, Wheeler D, Song X, Richmond TA, Middle CM, Rodesch MJ, Packard CJ, Weinstock GM, Gibbs RA. Direct selection of human genomic loci by microarray hybridization. Nat Methods. 2007;4:903–5.
Ankala A, Kohn JN, Dastur R, Gaitonde P, Khadilkar SV, Hegde M. Ancestral founder mutations in Calpain-3 in indian agarwal community—historical, clinical and molecular perspective. Muscle Nerve. 2013;47:931–7.
Ansorge WJ. Next-generation DNA sequencing techniques. N Biotechnol. 2009;25:195–203.
Antonarakis SE, Beckmann JS. Mendelian disorders deserve more attention. Nat Rev Genet. 2006;7:277–82.
Barton NH. Genetic hitchhiking. Philos Trans R Soc Lond B Biol Sci. 2000;355:1553–62.
Bashiardes S, Veile R, Helms C, Mardis ER, Bowcock AM, Lovett M. Direct genomic selection. Nat Methods. 2005;2:63–9.
Berman J. Modern classification of neoplasms: reconciling differences between morphologic and molecular approaches. BMC Cancer. 2005;5:100.
Berman JJ. Tumor taxonomy for the developmental lineage classification of neoplasms. BMC Cancer. 2004;4:88.
Betancur C. Etiological heterogeneity in autism spectrum disorders: more than 100 genetic and genomic disorders and still counting. Brain Res. 2011;1380:42–77.
Bloch-Zupan A, Jamet X, Etard C, Laugel V, Muller J, Geoffroy V, Strauss JP, Pelletier V, Marion V, Poch O, Strahle U, Stoetzel C, Dollfus H. Homozygosity mapping and candidate prioritization identify mutations, missed by whole-exome sequencing, in SMOC2, causing major dental developmental defects. Am J Hum Genet. 2011;89:773–81.
Bogardus C. Missing heritability and GWAS utility. Obesity (Silver Spring). 2009;17:209.
Botstein D, Risch N. Discovering genotypes underlying human phenotypes: past successes for mendelian disease, future approaches for complex disease. Nat Genet. 2003;33(Suppl):228–37.
Cabral RM, Kurban M, Wajid M, Shimomura Y, Petukhova L, Christiano AM. Whole-exome sequencing in a single proband reveals a mutation in the CHST8 gene in autosomal recessive peeling skin syndrome. Genomics. 2012;99:202–8.
Campeau PM, Lu JT, Sule G, Jiang MM, Bae Y, Madan S, Hogler W, Shaw NJ, Mumm S, Gibbs RA, Whyte MP, Lee BH. Whole-exome sequencing identifies mutations in the nucleoside transporter gene SLC29A3 in dysosteosclerosis, a form of osteopetrosis. Hum Mol Genet. 2012;21:4904–9.
Chang H, Jackson DG, Kayne PS, Ross-Macdonald PB, Ryseck RP, Siemers NO. Exome sequencing reveals comprehensive genomic alterations across eight cancer cell lines. PLoS One. 2011;6:e21097.
Choi M, Scholl UI, Ji W, Liu T, Tikhonova IR, Zumbo P, Nayir A, Bakkaloglu A, Ozen S, Sanjad S, Nelson-Williams C, Farhi A, Mane S, Lifton RP. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc Natl Acad Sci U S A. 2009;106:19096–101.
Cirulli ET, Goldstein DB. Uncovering the roles of rare variants in common disease through whole-genome sequencing. Nat Rev Genet. 2010;11:415–25.
Coonrod EM, Durtschi JD, Margraf RL, Voelkerding KV. Developing genome and exome sequencing for candidate gene identification in inherited disorders. Arch Pathol Lab Med. 2012;137:415–33.
Corder EH, Saunders AM, Strittmatter WJ, Schmechel DE, Gaskell PC, Small GW, Roses AD, Haines JL, Pericak-Vance MA. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer’s disease in late onset families. Science. 1993;261:921–3.
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, Teague J, Woffendin H, Garnett MJ, Bottomley W, Davis N, Dicks E, Ewing R, Floyd Y, Gray K, Hall S, Hawes R, Hughes J, Kosmidou V, Menzies A, Mould C, Parker A, Stevens C, Watt S, Hooper S, Wilson R, Jayatilake H, Gusterson BA, Cooper C, Shipley J, Hargrave D, Pritchard-Jones K, Maitland N, Chenevix-Trench G, Riggins GJ, Bigner DD, Palmieri G, Cossu A, Flanagan A, Nicholson A, Ho JW, Leung SY, Yuen ST, Weber BL, Seigler HF, Darrow TL, Paterson H, Marais R, Marshall CJ, Wooster R, Stratton MR, Futreal PA. Mutations of the BRAF gene in human cancer. Nature. 2002;417:949–54.
de Coronado S, Haber MW, Sioutos N, Tuttle MS, Wright LW. NCI thesaurus: using science-based terminology to integrate cancer research results. Stud Health Technol Inform. 2004;107:33–7.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, Philippakis AA, del Angel G, Rivas MA, Hanna M, McKenna A, Fennell TJ, Kernytsky AM, Sivachenko AY, Cibulskis K, Gabriel SB, Altshuler D, Daly MJ. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491–8.
Dickson SP, Wang K, Krantz I, Hakonarson H, Goldstein DB. Rare variants create synthetic genome-wide associations. PLoS Biol. 2010;8:e1000294.
Do R, Kathiresan S, Abecasis GR. Exome sequencing and complex disease: practical aspects of rare variant association studies. Hum Mol Genet. 2012;21:R1–9.
Druker BJ, Talpaz M, Resta DJ, Peng B, Buchdunger E, Ford JM, Lydon NB, Kantarjian H, Capdeville R, Ohno-Jones S, Sawyers CL. Efficacy and safety of a specific inhibitor of the BCR-ABL tyrosine kinase in chronic myeloid leukemia. N Engl J Med. 2001;344:1031–7.
Dundar H, Ozgul RK, Yalnizoglu D, Erdem S, Oguz KK, Tuncel D, Temucin CM, Dursun A. Identification of a novel twinkle mutation in a family with infantile onset spinocerebellar ataxia by whole exome sequencing. Pediatr Neurol. 2012;46:172–7.
Eichler EE, Flint J, Gibson G, Kong A, Leal SM, Moore JH, Nadeau JH. Missing heritability and strategies for finding the underlying causes of complex disease. Nat Rev Genet. 2010;11:446–50.
Gahl WA, Markello TC, Toro C, Fajardo KF, Sincan M, Gill F, Carlson-Donohoe H, Gropman A, Pierson TM, Golas G, Wolfe L, Groden C, Godfrey R, Nehrebecky M, Wahl C, Landis DM, Yang S, Madeo A, Mullikin JC, Boerkoel CF, Tifft CJ, Adams D. The National Institutes of Health Undiagnosed Diseases Program: insights into rare diseases. Genet Med. 2012;14:51–9.
Gargis AS, Kalman L, Berry MW, Bick DP, Dimmock DP, Hambuch T, Lu F, Lyon E, Voelkerding KV, Zehnbauer BA, Agarwala R, Bennett SF, Chen B, Chin EL, Compton JG, Das S, Farkas DH, Ferber MJ, Funke BH, Furtado MR, Ganova-Raeva LM, Geigenmuller U, Gunselman SJ, Hegde MR, Johnson PL, Kasarskis A, Kulkarni S, Lenk T, Liu CS, Manion M, Manolio TA, Mardis ER, Merker JD, Rajeevan MS, Reese MG, Rehm HL, Simen BB, Yeakley JM, Zook JM, Lubin IM. Assuring the quality of next-generation sequencing in clinical laboratory practice. Nat Biotechnol. 2012;30:1033–6.
Gnirke A, Melnikov A, Maguire J, Rogov P, LeProust EM, Brockman W, Fennell T, Giannoukos G, Fisher S, Russ C, Gabriel S, Jaffe DB, Lander ES, Nusbaum C. Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nat Biotechnol. 2009;27:182–9.
Gowrisankar S, Lerner-Ellis JP, Cox S, White ET, Manion M, LeVan K, Liu J, Farwell LM, Iartchouk O, Rehm HL, Funke BH. Evaluation of second-generation sequencing of 19 dilated cardiomyopathy genes for clinical applications. J Mol Diagn. 2010;12:818–27.
Gullapalli RR, Desai KV, Santana-Santos L, Kant JA, Becich MJ. Next generation sequencing in clinical medicine: challenges and lessons for pathology and biomedical informatics. J Pathol Inform. 2012;3:40.
Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100:57–70.
Harbour JW, Onken MD, Roberson ED, Duan S, Cao L, Worley LA, Council ML, Matatall KA, Helms C, Bowcock AM. Frequent mutation of BAP1 in metastasizing uveal melanomas. Science. 2010;330:1410–3.
Haspel RL, Arnaout R, Briere L, Kantarci S, Marchand K, Tonellato P, Connolly J, Boguski MS, Saffitz JE. A call to action: training pathology residents in genomics and personalized medicine. Am J Clin Pathol. 2010;133:832–4.
Hebert SC. Bartter syndrome. Curr Opin Nephrol Hypertens. 2003;12:527–32.
Hodges E, Xuan Z, Balija V, Kramer M, Molla MN, Smith SW, Middle CM, Rodesch MJ, Albert TJ, Hannon GJ, McCombie WR. Genome-wide in situ exon capture for selective resequencing. Nat Genet. 2007;39:1522–7.
Hoischen A, Gilissen C, Arts P, Wieskamp N, van der Vliet W, Vermeer S, Steehouwer M, de Vries P, Meijer R, Seiqueros J, Knoers NV, Buckley MF, Scheffer H, Veltman JA. Massively parallel sequencing of ataxia genes after array-based enrichment. Hum Mutat. 2010;31:494–9.
Holcomb CL, Hoglund B, Anderson MW, Blake LA, Bohme I, Egholm M, Ferriola D, Gabriel C, Gelber SE, Goodridge D, Hawbecker S, Klein R, Ladner M, Lind C, Monos D, Pando MJ, Proll J, Sayer DC, Schmitz-Agheguian G, Simen BB, Thiele B, Trachtenberg EA, Tyan DB, Wassmuth R, White S, Erlich HA. A multi-site study using high-resolution HLA genotyping by next generation sequencing. Tissue Antigens. 2011;77:206–17.
Jones MA, Bhide S, Chin E, Ng BG, Rhodenizer D, Zhang VW, Sun JJ, Tanner A, Freeze HH, Hegde MR. Targeted polymerase chain reaction-based enrichment and next generation sequencing for diagnostic testing of congenital disorders of glycosylation. Genet Med. 2011;13:921–32.
Kerem B, Rommens JM, Buchanan JA, Markiewicz D, Cox TK, Chakravarti A, Buchwald M, Tsui LC. Identification of the cystic fibrosis gene: genetic analysis. Science. 1989;245:1073–80.
Kiezun A, Garimella K, Do R, Stitziel NO, Neale BM, McLaren PJ, Gupta N, Sklar P, Sullivan PF, Moran JL, Hultman CM, Lichtenstein P, Magnusson P, Lehner T, Shugart YY, Price AL, de Bakker PI, Purcell SM, Sunyaev SR. Exome sequencing and the genetic basis of complex traits. Nat Genet. 2012;44:623–30.
Klein RJ, Zeiss C, Chew EY, Tsai JY, Sackler RS, Haynes C, Henning AK, SanGiovanni JP, Mane SM, Mayne ST, Bracken MB, Ferris FL, Ott J, Barnstable C, Hoh J. Complement factor H polymorphism in age-related macular degeneration. Science. 2005;308:385–9.
Kurotaki N, Imaizumi K, Harada N, Masuno M, Kondoh T, Nagai T, Ohashi H, Naritomi K, Tsukahara M, Makita Y, Sugimoto T, Sonoda T, Hasegawa T, Chinen Y, Tomita Ha HA, Kinoshita A, Mizuguchi T, Yoshiura Ki K, Ohta T, Kishino T, Fukushima Y, Niikawa N, Matsumoto N. Haploinsufficiency of NSD1 causes Sotos syndrome. Nat Genet. 2002;30:365–6.
Lander ES, Botstein D. Homozygosity mapping: a way to map human recessive traits with the DNA of inbred children. Science. 1987;236:1567–70.
Lee JT, Li L, Brafford PA, Van Den Eijnden M, Halloran MB, Sproesser K, Haass NK, Smalley KSM, Tsai J, Bollag G. PLX4032, a potent inhibitor of the B‐Raf V600E oncogene, selectively inhibits V600E‐positive melanomas. Pigment Cell Melanoma Res. 2010;23:820–7.
Leidenroth A, Sorte HS, Gilfillan G, Ehrlich M, Lyle R, Hewitt JE. Diagnosis by sequencing: correction of misdiagnosis from FSHD2 to LGMD2A by whole-exome analysis. Eur J Hum Genet. 2012;20:999–1003.
Lemke JR, Riesch E, Scheurenbrand T, Schubach M, Wilhelm C, Steiner I, Hansen J, Courage C, Gallati S, Bürki S. Targeted next generation sequencing as a diagnostic tool in epileptic disorders. Epilepsia. 2012;53:1387–98.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Lin X, Tang W, Ahmad S, Lu J, Colby CC, Zhu J, Yu Q. Applications of targeted gene capture and next-generation sequencing technologies in studies of human deafness and other genetic disabilities. Hear Res. 2012;288:67–76.
Lonigro RJ, Grasso CS, Robinson DR, Jing X, Wu YM, Cao X, Quist MJ, Tomlins SA, Pienta KJ, Chinnaiyan AM. Detection of somatic copy number alterations in cancer using targeted exome capture sequencing. Neoplasia. 2011;13:1019–25.
Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat SM, Supko JG, Haluska FG, Louis DN, Christiani DC, Settleman J, Haber DA. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med. 2004;350:2129–39.
Majewski J, Rosenblatt DS. Exome and whole-genome sequencing for gene discovery: the future is now! Hum Mutat. 2012;33:591–2.
Mamanova L, Coffey AJ, Scott CE, Kozarewa I, Turner EH, Kumar A, Howard E, Shendure J, Turner DJ. Target-enrichment strategies for next-generation sequencing. Nat Methods. 2010;7:111–8.
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ, McCarthy MI, Ramos EM, Cardon LR, Chakravarti A, Cho JH, Guttmacher AE, Kong A, Kruglyak L, Mardis E, Rotimi CN, Slatkin M, Valle D, Whittemore AS, Boehnke M, Clark AG, Eichler EE, Gibson G, Haines JL, Mackay TF, McCarroll SA, Visscher PM. Finding the missing heritability of complex diseases. Nature. 2009;461:747–53.
Mardis ER. Next-generation DNA sequencing methods. Annu Rev Genomics Hum Genet. 2008;9:387–402.
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z, Dewell SB, Du L, Fierro JM, Gomes XV, Godwin BC, He W, Helgesen S, Ho CH, Irzyk GP, Jando SC, Alenquer ML, Jarvie TP, Jirage KB, Kim JB, Knight JR, Lanza JR, Leamon JH, Lefkowitz SM, Lei M, Li J, Lohman KL, Lu H, Makhijani VB, McDade KE, McKenna MP, Myers EW, Nickerson E, Nobile JR, Plant R, Puc BP, Ronan MT, Roth GT, Sarkis GJ, Simons JF, Simpson JW, Srinivasan M, Tartaro KR, Tomasz A, Vogt KA, Volkmer GA, Wang SH, Wang Y, Weiner MP, Yu P, Begley RF, Rothberg JM. Genome sequencing in microfabricated high-density picolitre reactors. Nature. 2005;437:376–80.
Martinez FJ, Lee JH, Lee JE, Blanco S, Nickerson E, Gabriel S, Frye M, Al-Gazali L, Gleeson JG. Whole exome sequencing identifies a splicing mutation in NSUN2 as a cause of a Dubowitz-like syndrome. J Med Genet. 2012;49:380–5.
Maxmen A. Exome sequencing deciphers rare diseases. Cell. 2011;144:635–7.
McCarroll SA, Kuruvilla FG, Korn JM, Cawley S, Nemesh J, Wysoker A, Shapero MH, de Bakker PI, Maller JB, Kirby A, Elliott AL, Parkin M, Hubbell E, Webster T, Mei R, Veitch J, Collins PJ, Handsaker R, Lincoln S, Nizzari M, Blume J, Jones KW, Rava R, Daly MJ, Gabriel SB, Altshuler D. Integrated detection and population-genetic analysis of SNPs and copy number variation. Nat Genet. 2008;40:1166–74.
McClellan J, King MC. Genetic heterogeneity in human disease. Cell. 2010;141:210–7.
Mefford HC. Diagnostic exome sequencing: are we there yet? N Engl J Med. 2012;367:1951–3.
Metzker ML. Sequencing technologies: the next generation. Nat Rev Genet. 2009;11:31–46.
Miller M, Fineman R, Smith DW. Postaxial acrofacial dysostosis syndrome. J Pediatr. 1979;95:970–5.
Mitra AP, Pagliarulo V, Yang D, Waldman FM, Datar RH, Skinner DG, Groshen S, Cote RJ. Generation of a concise gene panel for outcome prediction in urinary bladder cancer. J Clin Oncol. 2009;27:3929–37.
Morin RD, Johnson NA, Severson TM, Mungall AJ, An J, Goya R, Paul JE, Boyle M, Woolcock BW, Kuchenbauer F, Yap D, Humphries RK, Griffith OL, Shah S, Zhu H, Kimbara M, Shashkin P, Charlot JF, Tcherpakov M, Corbett R, Tam A, Varhol R, Smailus D, Moksa M, Zhao Y, Delaney A, Qian H, Birol I, Schein J, Moore R, Holt R, Horsman DE, Connors JM, Jones S, Aparicio S, Hirst M, Gascoyne RD, Marra MA. Somatic mutations altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas of germinal-center origin. Nat Genet. 2010;42:181–5.
Najmabadi H, Hu H, Garshasbi M, Zemojtel T, Abedini SS, Chen W, Hosseini M, Behjati F, Haas S, Jamali P, Zecha A, Mohseni M, Puttmann L, Vahid LN, Jensen C, Moheb LA, Bienek M, Larti F, Mueller I, Weissmann R, Darvish H, Wrogemann K, Hadavi V, Lipkowitz B, Esmaeeli-Nieh S, Wieczorek D, Kariminejad R, Firouzabadi SG, Cohen M, Fattahi Z, Rost I, Mojahedi F, Hertzberg C, Dehghan A, Rajab A, Banavandi MJ, Hoffer J, Falah M, Musante L, Kalscheuer V, Ullmann R, Kuss AW, Tzschach A, Kahrizi K, Ropers HH. Deep sequencing reveals 50 novel genes for recessive cognitive disorders. Nature. 2011;478:57–63.
Need AC, Shashi V, Hitomi Y, Schoch K, Shianna KV, McDonald MT, Meisler MH, Goldstein DB. Clinical application of exome sequencing in undiagnosed genetic conditions. J Med Genet. 2012;49:353–61.
Neveling K, Collin RW, Gilissen C, van Huet RA, Visser L, Kwint MP, Gijsen SJ, Zonneveld MN, Wieskamp N, de Ligt J. Next‐generation genetic testing for retinitis pigmentosa. Hum Mutat. 2012;33:963–72.
Ng SB, Bigham AW, Buckingham KJ, Hannibal MC, McMillin MJ, Gildersleeve HI, Beck AE, Tabor HK, Cooper GM, Mefford HC, Lee C, Turner EH, Smith JD, Rieder MJ, Yoshiura K, Matsumoto N, Ohta T, Niikawa N, Nickerson DA, Bamshad MJ, Shendure J. Exome sequencing identifies MLL2 mutations as a cause of Kabuki syndrome. Nat Genet. 2010;42:790–3.
Ng SB, Buckingham KJ, Lee C, Bigham AW, Tabor HK, Dent KM, Huff CD, Shannon PT, Jabs EW, Nickerson DA, Shendure J, Bamshad MJ. Exome sequencing identifies the cause of a mendelian disorder. Nat Genet. 2010;42:30–5.
Nielsen R. Genomics: in search of rare human variants. Nature. 2010;467:1050–1.
Okou DT, Steinberg KM, Middle C, Cutler DJ, Albert TJ, Zwick ME. Microarray-based genomic selection for high-throughput resequencing. Nat Methods. 2007;4:907–9.
Oti M, Brunner HG. The modular nature of genetic diseases. Clin Genet. 2007;71:1–11.
Paez JG, Janne PA, Lee JC, Tracy S, Greulich H, Gabriel S, Herman P, Kaye FJ, Lindeman N, Boggon TJ, Naoki K, Sasaki H, Fujii Y, Eck MJ, Sellers WR, Johnson BE, Meyerson M. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science. 2004;304:1497–500.
Peltonen L, Perola M, Naukkarinen J, Palotie A. Lessons from studying monogenic disease for common disease. Hum Mol Genet. 2006;15 Spec No 1, R67–74.
Pinkel D, Segraves R, Sudar D, Clark S, Poole I, Kowbel D, Collins C, Kuo WL, Chen C, Zhai Y, Dairkee SH, Ljung BM, Gray JW, Albertson DG. High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet. 1998;20:207–11.
Prasad A, Merico D, Thiruvahindrapuram B, Wei J, Lionel AC, Sato D, Rickaby J, Lu C, Szatmari P, Roberts W, Fernandez BA, Marshall CR, Hatchwell E, Eis PS, Scherer SW. A discovery resource of rare copy number variations in individuals with autism spectrum disorder. G3 (Bethesda). 2012;2:1665–85.
Pritchard JK. Are rare variants responsible for susceptibility to complex diseases? Am J Hum Genet. 2001;69:124–37.
Proll J, Danzer M, Stabentheiner S, Niklas N, Hackl C, Hofer K, Atzmuller S, Hufnagl P, Gully C, Hauser H, Krieger O, Gabriel C. Sequence capture and next generation resequencing of the MHC region highlights potential transplantation determinants in HLA identical haematopoietic stem cell transplantation. DNA Res. 2011;18:201–10.
Sathirapongsasuti JF, Lee H, Horst BAJ, Brunner G, Cochran AJ, Binder S, Quackenbush J, Nelson SF. Exome sequencing-based copy-number variation and loss of heterozygosity detection: ExomeCNV. Bioinformatics. 2011;27:2648–54.
Schork NJ, Murray SS, Frazer KA, Topol EJ. Common vs. rare allele hypotheses for complex diseases. Curr Opin Genet Dev. 2009;19:212–9.
Schrijver I, Natkunam Y, Galli S, Boyd SD. Integration of genomic medicine into pathology residency training: the stanford open curriculum. J Mol Diagn. 2013;15:141–8.
Serizawa M, Sekizuka T, Okutani A, Banno S, Sata T, Inoue S, Kuroda M. Genomewide screening for novel genetic variations associated with ciprofloxacin resistance in Bacillus anthracis. Antimicrob Agents Chemother. 2010;54:2787–92.
Shah SP, Morin RD, Khattra J, Prentice L, Pugh T, Burleigh A, Delaney A, Gelmon K, Guliany R, Senz J, Steidl C, Holt RA, Jones S, Sun M, Leung G, Moore R, Severson T, Taylor GA, Teschendorff AE, Tse K, Turashvili G, Varhol R, Warren RL, Watson P, Zhao Y, Caldas C, Huntsman D, Hirst M, Marra MA, Aparicio S. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature. 2009;461:809–13.
Shendure J, Ji H. Next-generation DNA sequencing. Nat Biotechnol. 2008;26:1135–45.
Shi H, Moriceau G, Kong X, Lee MK, Lee H, Koya RC, Ng C, Chodon T, Scolyer RA, Dahlman KB, Sosman JA, Kefford RF, Long GV, Nelson SF, Ribas A, Lo RS. Melanoma whole-exome sequencing identifies (V600E)B-RAF amplification-mediated acquired B-RAF inhibitor resistance. Nat Commun. 2012;3:724.
Solomon BD, Pineda-Alvarez DE, Hadley DW, Program NCS, Teer JK, Cherukuri PF, Hansen NF, Cruz P, Young AC, Blakesley RW, Lanpher B, Mayfield Gibson S, Sincan M, Chandrasekharappa SC, Mullikin JC. Personalized genomic medicine: lessons from the exome. Mol Genet Metab. 2011;104:189–91.
St. Hilaire C, Ziegler SG, Markello TC, Brusco A, Groden C, Gill F, Carlson-Donohoe H, Lederman RJ, Chen MY, Yang D. NT5E mutations and arterial calcifications. N Engl J Med. 2011;364:432–42.
Stower H. The exome factor. Genome Biol. 2011;12:407.
Tan EK. Identification of a common genetic risk variant (LRRK2 Gly2385Arg) in Parkinson’s disease. Ann Acad Med Singapore. 2006;35:840–2.
Taylor BS, Ladanyi M. Clinical cancer genomics: how soon is now? J Pathol. 2011;223:318–26.
Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS, Arora VK, Kaushik P, Cerami E, Reva B, Antipin Y, Mitsiades N, Landers T, Dolgalev I, Major JE, Wilson M, Socci ND, Lash AE, Heguy A, Eastham JA, Scher HI, Reuter VE, Scardino PT, Sander C, Sawyers CL, Gerald WL. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18:11–22.
TCGA. Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474:609–15.
Treangen TJ, Salzberg SL. Repetitive DNA and next-generation sequencing: computational challenges and solutions. Nat Rev Genet. 2012;13:36–46.
Vaags AK, Lionel AC, Sato D, Goodenberger M, Stein QP, Curran S, Ogilvie C, Ahn JW, Drmic I, Senman L, Chrysler C, Thompson A, Russell C, Prasad A, Walker S, Pinto D, Marshall CR, Stavropoulos DJ, Zwaigenbaum L, Fernandez BA, Fombonne E, Bolton PF, Collier DA, Hodge JC, Roberts W, Szatmari P, Scherer SW. Rare deletions at the neurexin 3 locus in autism spectrum disorder. Am J Hum Genet. 2012;90:133–41.
Valencia CA, Ankala A, Rhodenizer D, Bhide S, Littlejohn MR, Keong LM, Rutkowski A, Sparks S, Bonnemann C, Hegde M. Comprehensive mutation analysis for congenital muscular dystrophy: a clinical PCR-based enrichment and next-generation sequencing panel. PLoS One. 2013;8:e53083.
Valencia CA, Rhodenizer D, Bhide S, Chin E, Littlejohn MR, Keong LM, Rutkowski A, Bonnemann C, Hegde M. Assessment of target enrichment platforms using massively parallel sequencing for the mutation detection for congenital muscular dystrophy. J Mol Diagn. 2012;14:233–46.
Varela I, Tarpey P, Raine K, Huang D, Ong CK, Stephens P, Davies H, Jones D, Lin ML, Teague J, Bignell G, Butler A, Cho J, Dalgliesh GL, Galappaththige D, Greenman C, Hardy C, Jia M, Latimer C, Lau KW, Marshall J, McLaren S, Menzies A, Mudie L, Stebbings L, Largaespada DA, Wessels LF, Richard S, Kahnoski RJ, Anema J, Tuveson DA, Perez-Mancera PA, Mustonen V, Fischer A, Adams DJ, Rust A, Chan-on W, Subimerb C, Dykema K, Furge K, Campbell PJ, Teh BT, Stratton MR, Futreal PA. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature. 2011;469:539–42.
Vasta V, Ng SB, Turner EH, Shendure J, Hahn SH. Next generation sequence analysis for mitochondrial disorders. Genome Med. 2009;1:100.
Vissers LE, van Ravenswaaij CM, Admiraal R, Hurst JA, de Vries BB, Janssen IM, van der Vliet WA, Huys EH, de Jong PJ, Hamel BC, Schoenmakers EF, Brunner HG, Veltman JA, van Kessel AG. Mutations in a new member of the chromodomain gene family cause CHARGE syndrome. Nat Genet. 2004;36:955–7.
Wall DP, Tonellato PJ. The future of genomics in pathology. F1000 Med Rep. 2012;4:14.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Yan XJ, Xu J, Gu ZH, Pan CM, Lu G, Shen Y, Shi JY, Zhu YM, Tang L, Zhang XW, Liang WX, Mi JQ, Song HD, Li KQ, Chen Z, Chen SJ. Exome sequencing identifies somatic mutations of DNA methyltransferase gene DNMT3A in acute monocytic leukemia. Nat Genet. 2011;43:309–15.
Yang H, Higgins B, Kolinsky K, Packman K, Go Z, Iyer R, Kolis S, Zhao S, Lee R, Grippo JF, Schostack K, Simcox ME, Heimbrook D, Bollag G, Su F. RG7204 (PLX4032), a selective BRAFV600E inhibitor, displays potent antitumor activity in preclinical melanoma models. Cancer Res. 2010;70:5518–27.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this chapter
Cite this chapter
Ankala, A., Hegde, M.R. (2015). Implementation of Exome Sequencing Assay. In: Netto, G., Schrijver, I. (eds) Genomic Applications in Pathology. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-0727-4_16
Download citation
DOI: https://doi.org/10.1007/978-1-4939-0727-4_16
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-0726-7
Online ISBN: 978-1-4939-0727-4
eBook Packages: MedicineMedicine (R0)