Abstract
Malaria remains a major parasitic disease in the tropical and sub-tropical countries mainly due to dramatic increase in parasite lines resistant to commonly used anti-malarials. Characterization of novel metabolic pathways in the parasites and understanding their functional role is a prerequisite to design new anti-malarial strategies. Parasite proteases play key role in growth and differentiation of all the developmental stages across the parasite life cycle and present the most promising targets to develop new drugs against malaria. In Plasmodium falciparum genome database a total of 123 proteases are identified; these proteases belong to five different clans: Cysteine, Aspartic, Serine, Metallo-, and Threonine. Some of the most studied parasite proteases are those that are functional in the asexual blood stage cycle. Starting with the processing of key parasite ligand in merozoite, the invasive form of blood stage parasite, degradation of host hemoglobin in food-vacuole, regulation of levels of key metabolic pathways in cytosol and cellular organelles, degradation of misfolded and unused proteins, and rupture of host membrane for egress of daughter merozoites is mediated by these proteases. Here we discuss roles of some of the parasite proteases involved in various steps of the parasite intra-erythrocytic cycle.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Malaria
- Plasmodium falciparum
- Intra-erythrocytic cycle
- Merozoite invasion
- Merozoite egress
- Hemoglobin degradation
- Organelle proteases
1 Introduction
The phylum Apicomplexa includes various protozoan pathogens causing major parasite diseases in the developing world; the most important of these diseases is malaria which causes about 1 million deaths globally per year [1]. Malaria is caused by five different species of genus Plasmodium: P. falciparum, P. vivax, P. malariae, P. ovale, and P. knowlesi; among them P. falciparum is responsible for the most deadly form of malaria infections. Plasmodium has a complex life cycle which is completed in three major phases in two host systems. Infection in humans begins with a bite of infected female Anopheles mosquito that injects invasive form of the parasite, sporozoites, which reaches to liver hepatocytes. The sporozoite enter and exits several hepatocytes by ripping through the plasma-membrane before finally infecting one of the host cell. In the infected hepatocytes the parasite resides in a parasitophorous vacuole (PV), undergoes multiple rounds of mitotic nuclear division and organelle division and subsequently large number of merozoites are formed which are released into the blood stream. Merozoites are the blood stage invasive forms that initiate the asexual blood stage cycle. These merozoites invade the host erythrocyte and reside in the parasitophorous vacuole, wherein it develops into a ring stage form which then subsequently grows to develop into trophozoite and then divides many times to develop into schizont, which then ruptures releasing the newly formed merozoites into the blood stream to continue the cycle. During the asexual cycle some of the parasites differentiate into male and female gametocytes which are taken up by the mosquito during blood-meal. Within the mosquito mid-gut, the gametocytes develop into male and female gametes which undergo fertilization; the zygote formed subsequently develops into motile ookinete. The ookinete burrows itself into the mid-gut wall and encyst on the basal lamina. The oocysts undergoes meiosis and divide to form large number of sporozoites that then invade salivary glands from where they can be again injected into human host. The blood stage asexual cycle is responsible for all the clinical symptoms and pathogenicity in humans; therefore the blood stages of the parasites are the target of most of the drug/vaccine development programs.
In light of rapid increase in parasite populations that have multi drug resistance, there is a need to develop new drug targets against the malaria parasite. Recent developments in the fields of genomic, proteomics and metabolomics research have helped to identify new drug targets. Since it is possible to develop specific inhibitors for proteases that can target the defined active sites, the malarial proteases are among the leading potential targets for developing new modes of chemotherapy [2–4]. The detailed genomic studies have identified a total of 123 proteases in P. falciparum genome [5, 6]. A number of these proteases are shown to be involved in the mediation of processes within the erythrocytic cycle; these processes include: rupture of host erythrocyte and egress of merozoites; invasion of merozoites into host erythrocytes; degradation of host hemoglobin etc. Functional importance of parasite proteases have been highlighted by detailed studies, which supported their potential as drug targets. Indeed a number of inhibitors have been designed against cysteine and aspartic protease of the malaria parasite with aim to develop as new anti-malarials. These studies identified lead compounds that can block in vitro parasite development at nanomolar concentrations and have cured malaria in animal models [7–9]. In this chapter we summarize the role of proteases in the Plasmodium asexual life cycle and their potential scope to be developed as drug targets. Two major steps in the asexual life cycle of Plasmodium that are majorly dependent on proteases are merozoite invasion and their egress from host erythrocyte; in addition, degradation of host hemoglobin in food vacuole, an essential step in establishment and growth of parasite in the host erythrocyte, is also depends upon various classes of parasite proteases. Proteases in specific cellular organelle and in the cytosol of the parasite also play important role in regulating a number of metabolic pathways and cell cycle (Fig. 1).
2 Role of Proteases During Merozoite Invasion into Host Erythrocyte
Invasion of red blood cells by the malaria merozoite is the first and essential step in the asexual blood stage life cycle of the parasite. The molecular details of invasion of apicomplexan parasite into host cells are only recently becoming understood, and each of these steps is considered as targets of new drug and vaccine development. On coming in contact with the host erythrocyte, P. falciparum merozoite re-orients itself such that the apical pole of the parasite points towards and interacts with the host erythrocyte membrane. Two different sets of protein play important role during interaction of host erythrocyte and merozoites: proteins on the surface of the merozoite that are possibly involved in weak initial attachment with the RBCs; and proteins that are released from the apical secretory organelles of the merozoite, the rhoptries, micronemes, exonemes and dense granules, which are involved in secondary interactions [10]. After merozoite reorientation, the apical organelles release their protein content in sequential manner in response to a calcium-mediated signal [11, 12], most of these released proteins are mobilized to the parasite surface. On the surface, these parasite proteins make high affinity interactions between the parasite and host surfaces. A tight junction forms between the parasite and host, which translocates towards the rear of the parasite via interactions between the microneme proteins’ cytoplasmic tails and a cortical parasite actin-myosin system [13, 14]. The role of parasite proteolytic enzymes in these critical steps in the life cycle of Plasmodium has been studied extensively. The bulk of the evidence indicates a prime role for serine proteases of the subtilisin and rhomboid families in these steps; these proteases act primarily as maturases and ‘sheddases’, which are required to process, activate and ultimately remove ligands involved in interactions with the host cell. Processing of some of the major merozoite surface and apical proteins, which play key role in invasion, is mediate by different proteases which points towards the importance of proteases in invasion process. Processing and maturation of some of these important proteins is described here (Fig. 2) (Table 1).
2.1 Merozoite Surface Proteins
The Merozoite Surface Protein-1 (MSP-l), is a GPI anchored protein present in a large protein complex on the surface of Plasmodium merozoites [15]. It is suggested to be involved in initial low-affinity binding of the parasite to the host cell, and has been long considered to be a good vaccine candidate. MSP-l is initially expressed as a protein precursor of ~195 kDa, and is subjected to primary processing which is thought to take place whilst the parasites are developing within the host cell rather than during invasion itself. After signal peptide removal and GPI anchor modification, primary processing in P. falciparum results in the full length gene product being cleaved into four subunits known as MSP-133, MSP-l30, MSP-l35, and MSP-l42 (in order from the N- to C-terminus of the original gene product and so named based on their molecular weights) [16]. These fragments are bound together non-covalently in a complex, the MSP-l42 fragment remains attached on the surface of the merozoite anchoring the complex in the membrane via its GPI anchor. During invasion MSP-l42 is proteolytically cleaved into two fragments (called MSP-l33 and MSP-l19) in what is known as secondary processing. This processing result in the release of the MSP-l complex from the parasite surface-an event which appears to be important, as only the post-processing stub (MSP-l19) appears to be able to penetrate the moving junction and still be localized to the parasite surface after invasion is complete. The role of proteases in MSP-1 processing as well as shedding has been the subject of intense studies. The first step towards identification of the MSP-1 shedding protease was the observation that this activity is calcium dependent; sensitive to the serine protease inhibitors PMSF and DFP, and also that the protease responsible is bound to the parasite plasma membrane when the processing event occurs [17]. Two other important merozoite surface proteins, MSP-6 and MSP-7, also get processed in the parasite. The precursor MSP-6 protein is N-terminal processed to generate MSP-636. Similarly the precursor MSP-7 protein is N-terminal processed to generate MSP-733. The MSP-733 gets further cleaved to generate MSP-722 and MSP-711 fragments [18, 19].
2.2 Merozoite Apical Proteins
The P. falciparum apical membrane antigen-1 (AMA-l), another long-time vaccine candidate in Plasmodium is also shed during invasion and, as for MSPl, anti-AMA-1 antibodies and small peptide based inhibitors that block this processing impede merozoite invasion [20, 21]. While analyzing the activity responsible for PfAMAl shedding, Howell et al. discovered that PfAMAl is shed by a protease with the same characteristics and inhibition profile as that responsible for MSP-l shedding [22, 23]. They concluded that the same protease, named Merozoite Surface Sheddase (MESH,) (which was later defined as Subtilisins) is responsible for the shedding of the two proteins (Fig. 3).
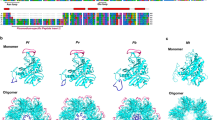
Primary Structure and processing of P. falciparum merozoite surface/apical proteins: MSP-1, MSP-6, MSP-7 and AMA-1. The primary precursor MSP-1 protein contains a number of variable, conserved and semi-conserved regions. Primary processing of this protein generate fragments of different sizes labelled as MSP-183, MSP1-30, MSP1-38 and MSP1-42; during invasion the MSP1-42 gets further cleaved into MSP1-33 and MSP1-19 [16]. The precursor MSP-6 protein is N-terminal processed to generate MSP-636. Similarly the precursor MSP-7 protein is N-terminal processed to generate MSP-733. The MSP-733 gets further cleaved to generate MSP-722 and MSP-711 fragments [18, 19]. AMA-1 is expressed as 83 kDa precursor protein consisting of N-terminal pro-sequence and a C-terminal trans-membrane region. In the micronemes the pro-sequence is cleaved off leaving 66 kDa protein containing three domains (I, II and III) attached to the membrane and is released on the merozoite surface. The 66 kDa is shed by juxtamembrane cleavage releasing 48 kDa fragment; further processing of this 48 kDa within the Domain III generate 44 kDa fragment which remains attached to the small polypeptide comprising the remainder of domain III via a intra-molecular disulfide bond [22, 23]
2.3 Role of Subtilisin-Like Serine Proteases During Invasion
The vital role of MESH during invasion of RBCs by the merozoites led investigators to search for candidate proteases in Plasmodium. The subtilisin-like family of proteases emerged as primary candidates owing to their calcium dependent serine protease activity and late stage expression pattern; characteristics’ similar to that of a putative MESH. Three P. falciparum genes encoding products belonging to the superfamily of subtilisin-like serine proteases, or subtilases, have been identified. Two of these genes, pfsub-1 and pfsub-2, were discovered and their gene products partially characterized some time ago [24, 25], whereas the presence of a third gene, pfsub-3, was revealed only by the P. falciparum genome project [26]. Both PfSUB-1 and PfSUB-2 are expressed in asexual blood stages and the mature enzymes accumulate in the apical regions of the merozoite. PfSUB1 is localized in special apical secretory organelles, the exonemes, and gets released in response to a calcium dependent signal into the parasitophorous vacuole just prior to schizont rupture and merozoite release [27, 28]. Selective inhibitors of PfSUB-1 do not inhibit shedding of MSP-1 or AMA-1, formally ruling out any involvement of PfSUB-1 in this process [29]. However, this inhibits egress of blood-stage P. falciparum, suggesting that PfSUB-1 is essential for parasite growth. The major role of PfSUB1 is processing of another protease PfSERA5 which is essential for parasite egress (as discussed later in this chapter). Nevertheless, it was later shown that PfSUB1 is required for pre-processing of MSP-1 along with MSP-6 and -7 prior to schizont rupture [30]. Global proteomic studies also identified several of merozoite surface and apical proteins [31] Overall, these and subsequent studies showed that the PfSUB1 thus plays an important role not in the actual invasion process but in priming the merozoites for invasion prior to their release from the schizont [29, 32, 33].
Another plausible candidate for a MESH, that is PfSUB2; a type I integral membrane protein, was identified by two research groups simultaneously [24, 25]. PfSUB2 represents a different sub class of eukaryotic pro-protein convertases as its deduced active site sequence resembles more with the bacterial subtilisins. Molecular modeling studies of PfSUB2 catalytic domain co-related with its proposed substrate specificity [24]. Further, PfSUB2 localization to the dense granules in merozoites made it an ideal candidate protein to function as MESH [23, 24]. Later it was shown that PfSUB2 localizes to the micronemes and is released just after schizont rupture to relocate to merozoite plasma membrane. In the same study it was shown that PfSUB2 specific peptide based inhibitor derived from its pro-domain can block MSP-1 and AMA-1 shedding [31]. Another study has shown that apart from MSP1and AMA-1, the PfSUB2 also mediate shedding of another invasion related protein, PTRAMP [34, 35]. Like PfSUB-1, PfSUB2 appears essential for blood-stages of the parasite as attempts to disrupt the sub2 gene in the rodent malaria P. berghei have been unsuccessful [36]. Recently Alam et al. have characterized PfSUB3 from Plasmodium falciparum [37].
2.4 Other Invasion Related Proteases: SERA-5, ABRA and Rhomboids
Another important protease family that plays important role in parasite invasion is the Serine repeat antigen (SERA) family. The human malarial parasite Plasmodium falciparum possesses nine SERA proteins, which belongs to cysteine protease family. Of these nine SERA proteins, six contains serine at active sites (serine-type) (SERA1 to SERA5 and SERA9) and three have cysteine at the active sites (cysteine-type) (SERA6 to SERA8) SERAs. Miller et al. tried knocking out eight of the nine SERAs located as a cluster on chromosome 2, the peripheral genes SERA-2, -3, -7 and -8 were dispensable the central genes, SERA-4, -5 and -6 remained refractory to deletion [38]. Joanne E. McCoubrie et al. then later tried knocking out four “serine type” SERA proteins; SERA1, SERA4, and SERA9 knockout lines were generated successfully, while SERA5, the most strongly expressed member of the SERA family, and SERA6 remained refractory to genetic deletion [39]. Serine repeat antigen-5 (SERA-5), also referred to simply as SERA, was initially identified as an abundant component of the PV that was shed in a soluble form at merozoite release. This and subsequent work [11] indicated that SERA5 was subjected to complex proteolytic processing, and that antibodies against it could interfere with merozoite release and erythrocyte invasion [40]. The central region of the molecule shared homology with the papain-like cysteine protease family, with the significant difference that the residue at the position of the active-site cysteine was replaced in SERA5 with a serine [41]. It is suggested that SERA5 can act as a protease despite its unusual active-site serine; recent studies showed that recombinant SERA-5 possesses autolytic activity, as well as chymotrypsin-like protease activity in trans against peptide substrates (Fig. 4) (Table 2) [42].
Primary Structure and processing of SERA5 during egress of P. falciparum merozoites from host erythrocytes. The precursor SERA5 (P126) is localized in the parasitophorous vacuole. Cleavage by PfSUB1 at two sites releases P56 which contains a central protease domain. The other two terminal fragments generated (P47 and P18), remain attached with each other due to disulfide-bond; this complex gets attached to the merozoite surface. The P56 fragment plays a proteolytic role during egress; later the P56 fragment is truncated by an unknown cysteine protease to modulate its function, perhaps by inactivation [11]
The probable role of SERA5 in invasion process is pointed out by processed proteolytic fragments derived from the N- and C-terminal regions of SERA-5, which associate with the merozoite surface [43, 44]. The significance of this is unclear, but it has been suggested that SERA-5 may play a role predominantly in merozoite release rather than invasion [42]. Later studies have linked the processing PfSERA5 in the parasitophorous vacuole and rupture of PVM during egress [33, 45]. Indeed, inhibition of processing of SERA5 shown to block the rupture of schizonts and release of merozoites [28]. Another member of this family, SERA-6 is also localized in the parasitophorous vacuole also gets processed by PfSUB1 to become active protein; SERA6 is also associated with egress and is essential for parasite survival [46].
Another P. falciparum merozoite peripheral surface protein proposed to mediate serine protease activity is known as acid basic repeat antigen, or ABRA. P. falciparum ABRA is a protein of about 100 kDa in size that accumulates during schizont maturation in the PV in a soluble form, but is also bound to the merozoite surface. Its name is derived from the presence within its sequence of two regions of highly charged tandem peptide repeats. The primary structure does not contain recognizable sequence motifs characteristic of major serine protease clans. The first suspicions that ABRA might be a protease came from the observation that the purified parasite protein consistently exhibited chymostatin-sensitive protease activity [47]. Recombinant ABRA produced in bacteria also appeared to possess proteolytic activity and the catalytic region was mapped to the N terminal domain of the protein that contains a serine residue, Ser317, previously proposed on the basis of sequence comparisons to be the active-site serine [47–49]. Possible role of ABRA has been suggested in erythrocyte binding during invasion [50]; however, importance of the predicted active-site Ser317 is not very clear. Clear orthologues of ABRA have been identified in P. vivax and two simian malarias [51] but an alignment of these sequences with that of ABRA shows that Ser317 is not conserved across species, being replaced by Glu in all the other sequences. It was found that disruption of the gene encoding MSP-3 resulted in the expression of truncated protein, which prevented trafficking of both MSP-3 and ABRA to the parasitophorous vacuole and merozoite surface [52]; however the resulting transgenic parasites lacking surface forms of both MSP-3 and ABRA were still capable of in vitro growth, which suggest that ABRA may not be playing a direct role in merozoite invasion.
As mentioned above, shedding of at least some Toxoplasma tachyzoite microneme proteins is mediated by a protease activity with the characteristics of rhomboids, which cleave within the TMD of integral membrane proteins [53]. Genes encoding rhomboid-like proteins are evident in the annotated P. falciparum genome [5, 6]. Earlier studies indicated that an activity of this nature may be present at the merozoite surface [54]. Later detailed studies using mammalian expression system showed that Plasmodium falciparum rhomboid protease PfROM-1 may be involved in cleavage of PfAMA-1 whereas PfROM-4 may be also involved in cleaving diverse adhesins including TRAP, CTRP, MTRAP, EBA-175, BAEBL, JESEBL, MAEBL, Rh1, Rh2a, Rh2b, and Rh4 [55, 56]; it was also shown that this cleavage relied on the adhesin transmembrane domains. However, later ROM-1 was shown to play role in sporozoite stage invasion and establishment of parasite into host hepatocyte [57, 58].
Another important parasite protease for which experimentally demonstrated link with invasion was shown is the Falcipain-1, an important cysteine protease of the parasite. Falcipain 1 was the first identified member of a small family of papain-like Plasmodium cysteine proteases; it was originally characterized as being primarily involved in haemoglobin catabolism during intra-erythrocytic growth [59]. However, studies with a radiolabelled cysteine protease chemical probe demonstrated, contrary to what would be predicted of a haemoglobinase, the Falcipain-1 expression peaks in merozoite and ring (the newly invaded parasite) stages of the erythrocytic cycle [60]. Indeed the enzyme was also localized at the apical end of the merozoite [60]. Treatment of cultures with falcipain 1 inhibitors derived from a positional scanning peptidyl epoxide library had no effect on intracellular growth of the parasite but appeared to very effectively prevent invasion by released merozoites, leading to the proposal that this protease has an important role in invasion. Some doubt was cast on this interpretation, however, by the recent demonstration that disruption of the P. falciparum falcipain-1 gene has no detectable effect on replication of asexual blood-stage parasites [61]. Although it is possible that up-regulation of other proteases may have compensated for the absence of Falcipain-1 in the knockout parasites, this work however proves that Falcipain-1 is not absolutely essential for replication of the asexual blood-stage parasite. As a result—and although its function remains obscure—the protease is unlikely to be considered a good target for anti-malarial drug development.
3 Role of Proteases During Rupture of Host Erythrocyte and Merozoite Egress
Rupture of host erythrocyte membrane and egress of merozoite into host milieu is a complicated process involving many steps; role of several and different classes of parasite proteases is suggested among these steps. In the early 1980s, the role of proteases in the mechanism of egress was pointed by Banyal et al. [62]. A number of serine and cysteine inhibitors have been studied for their effect on the egress of P. knowlesi merozoites. It was observed that mature schizonts accumulated upon treatment with a mixture of leupeptin, Chymostatin, antipain (a serine and cysteine protease inhibitor) and pepstatin (an aspartic protease inhibitor). Detailed studies showed that the process of egress is a two-step process, involving primary rupture of the parasitophorous vacuole membrane followed by a secondary rupture of the erythrocyte plasma membrane [63]. Using specific inhibitors and transgenic lines expressing GFP in different compartment of the infected erythrocyte, it was shown that the each step is mediated by distinct proteases; the primary vacuolar lysis step can be inhibited by cysteine protease inhibitors E-64 whereas the leupeptin and antipain can inhibit secondary erythrocyte rupture step [63]. Overall the egress may involve several proteases and their sequential processing by other enzymes. Proteases that have been implicated in parasite egress are: (1) aspartic proteases e.g. Plasmepsins and histo-aspartic proteases [64] (2) cysteine proteases e.g. falcipains [65, 66] (3) dipeptidyl peptidase 3 (PfDPAP3) [32] (4) Serine Repeat Antigens (SERAs)[38, 67]; and (5) serine protease subtilase 1 (PfSUB1) in the subtilisin S8 family [29].
The Plasmodium aspartic and cysteine proteases (plasmepsins and falcipains respectively) have been shown to function primarily as hemoglobinases in the parasite food vacuole as discussed later in this chapter. However, some evidence points towards their dual functionality as some members being also involved in the process of parasite egress from the host erythrocytes. The ability of Plasmepsin II to digest the host RBC cytoskeletal proteins like spectrin and actin and its localization in the host RBC cytosol outside the parasite provided the first indication towards this dual functionality and the possible involvement of food vacuole protease in the process of egress [64]. Similarly, Falcipain-2 was also demonstrated to be able to digest ankyrin and protein 4.1 at neutral pH [68, 69]. Further Dhawan et al. were able to inhibit the activity of recombinant Falcipain-2 using a peptide based on the cleavage site in ankyrin [65]. Transient silencing of Falcipain-2 also caused inhibition or merozoite egress in P. falciparum [70]. Subsequent gene disruption studies have shown that neither Plasmepsin II nor falcipain 2 is essential in asexual blood stages, and the knockout lines showed no defect in parasite egress from the host RBC [71–74]. However, the loss of falcipain2 was accompanied by an increased transcription of another similar gene falcipain 2′ [72]. The role of these proteases in egress cannot be completely ruled out but it is clear that there is some degree of redundancy involved which requires further attention. DPAP3 is a cathepsin-like cysteine protease identified as an important protease required for egress in an inhibitor based screening [32]. In the same study it was found that the inhibition of DPAP3 caused loss of PfSUB1indicating that it may be playing a role in PfSUB1 folding and activation and the inactive forms of PfSUB1 might be rapidly degraded. As described above, PfSUB1 is released from the exonemes just prior to schizont rupture in response to a calcium dependent signal into the PV where it carries out cleavage of SERA-5 along with other SERA proteins [29] which subsequently play key role in egress.
In addition to the parasite proteases, a host calcium-dependent protease, Calpain-1, is also required for efficient parasite egress of Plasmodium and Toxoplasma [75]. A cysteine protease inhibitor (DCG04) does not affect the parasite growth but prevents the release of parasite from the host cell. Selective extraction of treated cells identified host Calpain-1 as the target of this inhibitor. Calpain-1 is shown to be present in the cytoplasm of the infected host cell until the schizont stage of parasite growth, subsequently it shift to the membrane, indicating calcium binding and activation. Calpain-1 removal from erythrocytes prevented parasite egress and led to the growth arrest in the schizont stage, whereas reconstitution with recombinant calpain-1 could restore normal growth development.
4 Hemoglobin Degradation: Food Vacuole Proteases
Plasmodium parasite possesses a limited capacity for de novo synthesis of amino acids; the cellular amino acid pool in these parasites is thus derived from host cell hemoglobin after its degradation in a specialized form of lysosome called the ‘food vacuole.’ Apart from being a nutrient source degradation of hemoglobin is also important to maintain the osmotic integrity of the infected red blood cell. The food vacuole is an acidic compartment with pH around 5.2. Hemoglobin degradation is carried out by several vacuole-located proteases in a semi-ordered fashion. The process starts with an attack on the native hemoglobin. Enzymes capable of such attack include the aspartic proteases plasmepsin-I and plasmepsin-II, which cleave the alpha chain of hemoglobin, breaking the structure and exposing several other sites making it prone to other proteases’ attack. Further degradation process is carried out by the aspartic protease plasmepsin 4 (a histo-aspartic protease) and three falcipain proteins (falcipain 2, falcipain-2′ and falcipain-3), resulting in peptides that are larger than 20 amino acids in length. It has been suggested that falcipain 2 and 3 are also capable of attacking the native hemoglobin so may participate at the very first step [61, 70, 72, 76, 77]. These peptides in turn are degraded by other peptidases that breaks these peptides to smaller ones, around 5–8 amino acid in length. One of the candidate protease for this step is falcilysin, a zinc metalloprotease localized to multiple parasite compartments and proposed to play diverse function [78]. The final step within the food vacuole is catalyzed by dipeptidyl aminopeptidase 1, an enzyme that produces dipeptides.
4.1 Plasmepsins
The P. falciparum genome harbours ten aspartic protease genes (PM I, II, and IV–X and HAP) [71, 73, 79, 80] Out of these, three Plasmepsin (PM VI, VII and VIII) are not expressed in asexual blood stages. Rest all of the Plasmepsins are expressed in asexual stages at different locations. PM I and II are localized in the food vacuole and are considered to be the major players required at the very first step in the process of heamoglobin degradation [73, 81] Plasmepsin I and II carries out the first attack on the hemoglobin alpha chain opening up the structure for further protease cleavage It can then acted upon by other proteases including another plasmepsin, PM IV and Histo-aspartic protease (HAP) [73]. As in case of many of the parasite protease, Plasmepsins I and II are synthesized as pro-enzymes. Removal of the pro-domain is required to release the mature enzyme. Activation can be blocked with two tripeptide aldehyde compounds of low specificity, but the identity of the pro-plasmepsin processing enzyme has not been established yet [79]. The pro-plasmepsin convertase has been suggested as a promising target for new antimalarial drugs, since its blockage would inhibit the formation of all four food vacuole plasmepsins [82].
Malarial parasites, in vitro and in vivo, can be killed by specific inhibitors of Plasmepsins, indicating that these proteases are viable as drug targets. Analysis of substrate preferences and active site mutations has provided insight into the binding specificities of these different plasmepsins. Design of compounds able to inhibit several plasmepsins could be favorable, not only for efficient killing of the parasites, but also to impede the development of parasite resistance. HIV-1 protease inhibitors has provided a large pool of compounds, successfully utilized in the search for Plasmepsin inhibitors [83–86]. Some of the HIV-1 protease inhibitors currently on the market have demonstrated activity against Plasmepsin II as well as activity in P. falciparum infected erythrocytes [87, 88]. In addition these HIV-1 protease inhibitors have shown anti-parasitic activity in a murine malaria model [89].
4.2 Falcipains
There are three falcipains present in food vacuole; Falcipain-2 Falcipain-2′ and Falcipain-3. These cysteine proteases play a major role in degradation of hemoglobin and thus are most important protease present in food vacuole. Possible role of Falcipain-1 in merozoite invasion and gene deletion studies are discussed earlier in the chapter [60, 61]. FP2 and FP3 are the major hemoglobinases expressed in trophozoite stage, localize to the food vacuole, and degrade hemoglobin [66, 90, 91]. FP2′ is biochemically very similar to FP2, and share high sequence homology with FP2 [76]. Gene knockout and transient silencing analyses have revealed that Falcipain-2 is the major hemoglobinase as its disruption lead to low degradation rate of hemoglobin [70, 72]. Transient silencing of Falicpain-2 homologue, berghepain-2, in mouse malaria model caused inhibition of parasite growth [70, 92]. It is recently shown that FP-2 exists as a component of large protein complex consisting of several other proteases and heme-detoxification protein (HDP), and it is suggested that all these components work in a cooperated manner [77]. Falicpain-3 could not be disrupted which points towards essential role of FP3 in parasite [74]. Being most important of all Falicpains, FP-2 remained as first choice of all against which inhibitors were designed. Several studies are been done to develop lead compounds against FP-2. Different compounds ranging from peptide fluromethyl ketones, peptide vinyl sulfones, peptide aldehydes and a-ketoamides lot of chemical scaffolds have been used to develop inhibitors against FP-2 [93, 94]. In addition, A number of groups have been involved in developing falcipain-2 inhibitors using peptidomimetic approaches [95–98].
4.3 Aminopeptdiase
P. falciparum genome encodes nine exo-aminopeptidases, four of these enzymes are annotated as methionine aminopeptidases function in the catalytic removal of N-terminal initiator methionine during protein synthesis. The remaining five aminopeptidases are potential candidate enzymes for the release of free amino acids from hemoglobin-derived peptides [99]. The intra-erythrocytic stages of the human malaria parasite P. falciparum express two neutral metallo-aminopeptidases that are believed to be involved in the terminal stages of host hemoglobin digestion, an M1 alanyl aminopeptidase (PfM1AAP) and an M17 leucine aminopeptidase (PfM17LAP) [100, 101]. The M1 aminopeptidase harbors a trans-membrane domain and so thought to be a membrane protein. However, it was shown to be processed and thus being a soluble protein localized to parasite cytosol and around the food vacuole [102, 103]. Dalal and Kemba later showed using YFP-tagged transgenic line that PfM1AAP is localized to food vacuole and nucleus and not in cytosol [104]. In the same study it was also shown that PfM1AAP is essential as it cannot be knocked out in the parasite along with two other APs; PfM17LAP and aminopeptidase P (PfAPP). PfM18AAP, with highest expression levels in rings, is another member of aminopeptidase family. Functionally active recombinant enzyme, rPfM18AAP, and native enzyme in cytosolic extracts of malaria parasites are 560-kDa octomers that exhibit optimal activity at neutral pH and require the presence of metal ions to maintain enzymatic activity and stability [105]. As in case of human aspartyl aminopeptidase, the exopeptidase activity of PfM18AAP is exclusive to N-terminal acidic amino acids, glutamate and aspartate, making this enzyme of particular interest and suggesting that it may function alongside the malaria cytosolic neutral aminopeptidases in the release of amino acids from host hemoglobin-derived peptides. Whereas immune-cytochemical studies using transgenic P. falciparum parasites show that PfM18AAP is expressed in the cytosol, immunoblotting experiments revealed that the enzyme is also trafficked out of the parasite into the surrounding parasitophorous vacuole. Antisense-mediated knockdown of PfM18AAP results in a lethal phenotype as a result of significant intracellular damage and validates this enzyme as a target at which novel antimalarial drugs could be directed. The importance of parasite aminopeptidase in hemoglobin degradation has made these proteases as potent drug targets against Parasite. A number of structural and bioinformatic studies have been carried out to develop new antimalarial targeting aminopeptidases of P. falciparum [106–110].
5 Organelle Proteases and Cell Cycle Regulation
The malaria parasite Plasmodium possesses two essential organelles, which have prokaryotic origin, the mitochondrion and the relict plastid apicoplast. Both the organelles are essential for parasite survival and plays essential role in maintenance of parasite cell cycle. The metabolic pathways in the mitochondrion and the apicoplast may represent suitable drug targets in the parasite. Selected antibiotics such as doxycycline and clindamycin which target some of these prokaryotic metabolic pathways have already been shown to possess antiparasitic efficacies and are used in malaria treatments [111–115].
5.1 Mitochondrial Proteases
Plasmodium harbours a single mitochondrion during its asexual cycle which divides just before cytokinesis and is distributed equally as single organelle per progeny. Mitochondrion in Plasmodium is a validated drug target as the known antimalarial drug atovaquone acts on the respiratory chain complex III in mitochondrion. The only known proteases to be localized in P. falciparum mitochondrion are ClpQ [also called HslV (Heat shock loci V)] [116, 117]. In addition, Falcilysin is shown to have multiple localization in the parasite and only partially localized to mitochondrion while it is majorly involved in haemoglobin degradation in food vacuole and also in transit peptide degradation in the apicoplast [80]. The ClpQ protease is a mitochondrial resident protease machinery having ClpY [also called HslU (Heat shock loci U)] as the ATPase partner. ClpQY is the prokaryotic predecessor of the eukaryotic proteasomal machinery; in the ClpQY machinery the ClpY is the ATPase partner forming a hexameric head over the two hexameric core assemblies of ClpQ protease either on one or both sides [118]. The ClpQY machinery seems to play an essential role in the growth and survival of the parasite at least during the asexual blood stage as the disruption of the machinery by blocking ClpQ and ClpY interaction leads to parasite death with the death phenotype resembling apoptosis in eukaryotic cells [119]. The ClpQY machinery is essential in regulation of replication of mitochondrial genome in Trypanosoma brucei [120] knockdown of ClpQY results in over replication of minicircle DNA and abnormal segregation of kinetoplast leading to formation of large kDNA networks which ultimately blocks cell division. Further, the absence of a homolog in the human host leads to the conclusion that ClpQY protease machinery could be a promising drug target in all apicomplexan parasites. Indeed functional importance of ClpQY in P. falciparum is clearly shown. Disruption of ClpQY machinery in P. falciparum by using small peptide based inhibitors caused dysfunctioning of mitochondria and inhibited parasite growth [119] further, this initiated a cascade of protease and nuclease activation that caused apoptosis like cell death of the treated parasites [119]. Further, trans-expression of mutant inactive ClpQ protein caused dominant negative effect in the parasite which disrupted mitochondria development and caused parasite death [121]; these studies thus support the essentiality of the ClpQY protease machinery for parasite survival and candidature of ClpQ as a promising target for developing new anti-malarial.
5.2 Apicoplast Proteases
The discovery of the apicoplast in the Plasmodium spp. in 1996 instantly made it a key target for the development of new therapies against these pathogens owing to its prokaryotic origin and hence availability of various organelle pathways absent from the human host that could be targeted. The apicoplast is a reduced cyanobacterial plastid in the parasite and was acquired by the apicomplexan protozoans by secondary endosymbiosis. It plays an important role in biosynthesis of haem, isopentenyl diphophate and fatty acids [122], thus the apicoplast is considered to be crucial for parasite survival. Antibacterial agents such as ciprofloxacin, rifampicin and thiostrepton that target DNA replication, transcription and translation of the apicoplast, respectively, have been also shown to kill the parasite [123–126]. It is recently showed that the critical and essential function of the apicoplast is in isoprenoid precursors synthesis, and it possible to generate apicoplast minus parasites in vitro by chemically rescue using isoprenoid precursors supplemented media [127].
Apicoplast is a four membrane bound structure having a 35 kb genome; however about 95 % of its proteins are nuclear encoded which are imported via a complex protein translocation pathway which is yet to be fully elucidated. Since, majority of the proteins of the organelle depend on this pathway to be correctly delivered to their respective site of action to carry out their required function it is understandable that any perturbation in this pathway will lead to chaotic situation inside the apicoplast and hence may be lethal for the parasite. The protein import to the apicoplast makes use of a bipartite N-terminal extension of the protein; the first part of this sequence targets the protein into the secretory pathway while the second part called the transit peptide (TP) region takes the protein through the four surrounding membranes of the organelle. Two proteases have been proposed to take part in this process; the stromal processing peptidase (SPP) is considered to take part in the import process directly by being responsible for cleavage of the transit peptide to yield the mature protein [128] while the second protease falcilysin is indirectly linked with the process and is proposed to be the enzyme responsible for degradation of transit peptide [80].
Another protease localized to the apicoplast is ClpP, a serine type protease functioning in conjunction with an ATPase to form the complete protease machinery as in case of ClpQY. The parasite genome codes for four Clp ATPases termed as ClpB1, ClpB2, ClpC and ClpM and also an inactive version of the ClpP protease termed ClpR [129]. Three of these ATPases, ClpB1, ClpC and ClpM are localized to the apicoplast as did ClpR while ClpB2 localizes to the parasitophorus vacuole [129]. The ClpP protease is localized to the apicoplast and is proteolytically active; a specific inhibitor of ClpP can block apicoplast development caused significant inhibition of parasite growth in-vitro [130]. ClpP is thus a promising drug target and its inhibitor can be developed as lead anti-malarials.
6 Other Cellular Proteases
A number of cellular pathways employ proteases that do not directly distinguish their substrates but instead utilize a post-translational modification of the protein as the recognition signal. This includes complex multi-meric ATP dependent protease system, Proteasome.
6.1 Proteasome
The 20S proteasome is a multimeric self-compartmentalising protease machinery essential for the survival of every eukaryotic cell. The machinery consists of two central rings formed by the β subunits capped on both sides by α subunit rings. The diversity of the individual subunits varies among different organisms, ranging from a single subtype for each in archae-bacteria to seven subtypes of each in human. Apart from the basic proteolytic core complex, proteasome also has the 19S regulatory subunit which helps in substrate recognition and unfolding of the substrate driving it through the central proteolytic chamber. The functions of the proteasome range from simply degradation of misfolded proteins destined so by polyubiquitination, to regulation of the cell cycle by maintaining the levels of respective proteins such as cyclins and various transcription factors.
Protista, particularly the pathogenic protists are by far the only known eukaryotes to possess both 26S proteasome in addition to its prokaryotic predecessor ClpQY as described earlier [131]. In-silico studies have identified all the 14 subunits of 20S proteasome along with the subunits of the 19S regulatory particle [5, 132, 133]. However, given the degree of conservation and essentiality of the ubiquitin proteasome pathway it is surprising that despite the sequencing of several parasite genomes so little is known about its specific functional role in any of the parasites. Increasing evidence suggest essentiality of the proteasome machinery for the malaria parasites and hence it being a plausible drug target [134]. The irreversible inhibitor of the proteasome, lactacystin, could stall the growth of P. bergei parasite in-vitro as well as in-vivo. Also, the inhibitory effect of lactacystin on P. falciparum parasite was found to be cell cycle specific, the drug being able to kill parasite only if applied prior to initiation of DNA replication and not afterwards. A range of proteasome inhibitors have thus far been tested for their efficacy against the malarial parasite. These inhibitors including, lactacystin, salinosporamide A, MG132, epoxomicin, Thiostrepton and bortezomib were reported to inhibit parasite growth in vitro at low nanomolar concentrations [133–137].
6.2 Ubiquitination and Deubiquitinations
Ubiquitination is by far the best characterised post translational modification targeting the proteins for degradation, i.e., by the Proteasome. However, evidence that it also plays a key role in regulation of several other cellular pathways is mounting, wherein it serves as any other protein modification (such as phosphorylation) required for proper functioning or targeting of the protein. Several other ubiquitin like protein (Ubl) have been identified in eukaryotes, homologs for six of which have been found in Plasmodium including, Nedd8 (neural precursor cell expressed developmentally down-regulated 8) [138], small ubiquitin-related modifier (SUMO) [139], Hub1, ubiquitin-related modifier 1 (Urm1) [138], and autophagy-8 (Atg8) [140]. In P. falciparum, in silico studies identified four predicted sources of ubiquitin moieties. The polyubiquitin gene PFL0585w (contains five conserved ubiquitin repeats), two ubiquitin fusion proteins PfUBS27a and PfUBL40 (contain a ubiquitin moiety at their N-terminus) and PfsUb which is targeted to apicoplast [140–142]. Ubiquitin is removed by selective proteases known as Deubiquitinating proteases (DUBs) that selectively hydrolyse the isopeptide linkage. Two independent studies identified 18 and 29 Deubiquitinating enzymes (DUBs), respectively, in Plasmodium [139, 141]. DUBs are involved in removal of ubiquitin chain from proteins. Homologs of human UCH37, PfUCH54; and of UCHL3, PfUCHL3, are the only DUBs to be characterised so far and shown to exhibit deubiquitinating and deNeddylating activities [138, 143]. DUBs have been classified into at least five distinct subfamilies based on their sequence similarity and likely mechanisms of action including: (1) UBP (Ubiquitin specific processing protease). (2) OTU (Ovarian TUmour) related proteases. (3) UCH (Ubiquitin C-terminal hydrolases) and (4) Machado-Joseph disease protease (MJD). Of these, the first three are cysteine proteases, while the last one is a novel group of zinc-dependent metalloprotease. A bioinformatics approach to identify components of the ubiquitin mediated pathway in apicomplexan parasites identified three OTU family proteins in P. falciparum [144]. Additionally, the presence of a functional Deubiquitinating enzyme PfUCH54 has been shown by [145]. The deNeddylating activities of PfUCH54 and PfUCHL3 might be of particular interest for drug design, since such activity is not known for the mammalian homologs of the two DUBs.
Protein modification by SUMO is also found in P. falciparum; however, its role in the regulation of the parasite life cycle is poorly understood. SUMOlyated proteins are widely distributed in parasite and PfSir2 is one of the targets along with several other putative targets [146]. Functional studies of a SUMO-specific protease (SENP) of P. falciparum, PfSENP1 demonstrated that this protease has unique cleavage sequence preference relative to the human SENPs [139]. In addition, a small molecule inhibitors of this protease can inhibit P. falciparum replication in infected human blood.
6.3 Signal and Transit Peptide Cleaving Proteases
To survive in the host erythrocyte the parasite has evolved a powerful protein secretion system responsible for trafficking of protein to sub-cellular organelles, host erythrocyte cytosol, host erythrocyte membrane, and secretion into the host milieu. The key player in secretion and trafficking of parasite proteins are the signal peptidase that are involved in cleaving the signal sequence from the target proteins, after this processing the proteins are released from the membrane and then routed to their destination. A total of five signal peptidases have been identified in P. falciparum that may be involved in formation of a signal peptidase complex (SPC) [5, 147, 148]. Another processing protease is Signal Peptide Peptidase that is an aspartyl family protease and is involved in cleavage of remnant signal peptides after their release by signal peptidase. The P. falciparum SPP was earlier suggested to be involved in invasion of merozoite by targeting a possible substrate, Band-3 [149, 150]; however later studies established its localization in parasite ER its role in growth of the asexual stage parasite [151, 152]. As mentioned above a number of nuclear encoded proteins are trafficked to mitochondria, apicoplast and other organelles in the parasite depending upon the presence of specific targeting sequences. Upon reaching the organelle membrane these transit peptide sequences are cleaved off from the proteins. Not many proteases are identified to be involved in processing of these transit sequences in different parasite organelles, only Falcilycin is implicated in processing of transit peptide in these organelles [78, 80].
To modify the host erythrocyte and to evade the host immune response, the parasite exports large number of proteins beyond the parasitophorous membrane. Most of these proteins contain an N-terminal signal sequence responsible for entry into ER as in case of other secreted proteins; in addition, these proteins also contain a motif termed PEXEL (Plasmodium EXport ELement) which is responsible for trafficking of these proteins beyond parasitophorous membrane. The PEXEL motif is a pentameric sequence RxLxE/Q/D that is processed after the conserved leucine in the ER and then gets N-aceylated, this cleavage is suggested to be essential for trafficking of target proteins beyond parasite boundaries [153, 154]. An aspartic protease Plasmepsin V which resides in ER is shown to be responsible for cleavage of PEXEL and facilitating trafficking of the proteins [155].
7 Conclusion
In view of the development of resistance of the parasite to frontline antimalarials, evolution of insecticide-resistant mosquitoes and unavailability of a vaccine, there is an urgent need to identify new drugs targets in the malaria parasite and develop novel anti-malarials, In the future, antimalarial therapy must combine several features that are still far from being ideal, including: minimal toxic side effects, high efficacy against resistant strains of P. falciparum, activity against several Plasmodium species, optimal pharmacokinetic profile, and low cost of therapy. Protease seems to be important players in lot of mechanism of survival for the parasite and thus seems to be wonderful target for developing new anti-malarials. Protease like Subtilisins and SERAs are good drug targets are they play essential roles in egress pathway of parasite; food vacuole protease will remain the best choice as drug targets as they are involved in hemoglobin degradation, one of most important pathways for parasite survival. Recent work on organelle proteases has also shown them to other plausible targets for drug development. There is need of combined effort from protein structural studies, computational biology and synthetic chemistry to develop more potent antimalarials.
References
Snow RW, Guerra CA, Noor AM et al (2005) The global distribution of clinical episodes of Plasmodium falciparum malaria. Nature 434:214–217
Teixeira C, Gomes JR, Gomes P (2011) Falcipains, Plasmodium falciparum cysteine proteases as key drug targets against malaria. Curr Med Chem 18:1555–1572
Tschan S, Mordmuller B, Kun JF (2011) Threonine peptidases as drug targets against malaria. Expert Opin Ther Targets 15:365–378
McKerrow JH, Rosenthal PJ, Swenerton R et al (2008) Development of protease inhibitors for protozoan infections. Curr Opin Infect Dis 21:668–672
Wu Y, Wang X, Liu X et al (2003) Data-mining approaches reveal hidden families of proteases in the genome of malaria parasite. Genome Res 13:601–616
Cai H, Gu J, Wang Y (2010) Core genome components and lineage specific expansions in malaria parasites plasmodium. BMC Genomics 11(Suppl 3):S13
Rosenthal PJ (2011) Falcipains and other cysteine proteases of malaria parasites. Adv Exp Med Biol 712:30–48
Brady RL, Cameron A (2004) Structure-based approaches to the development of novel anti-malarials. Curr Drug Targets 5:137–149
Werbovetz KA (2000) Target-based drug discovery for malaria, leishmaniasis, and trypanosomiasis. Curr Med Chem 7:835–860
Cowman AF, Crabb BS (2006) Invasion of red blood cells by malaria parasites. Cell 124:755–766
Blackman MJ (2000) Proteases involved in erythrocyte invasion by the malaria parasite: function and potential as chemotherapeutic targets. Curr Drug Targets 1:59–83
Singh S, Alam MM, Pal-Bhowmick I et al (2010) Distinct external signals trigger sequential release of apical organelles during erythrocyte invasion by malaria parasites. PLoS Pathog 6:e1000746
Alexander DL, Mital J, Ward GE et al (2005) Identification of the moving junction complex of Toxoplasma gondii: a collaboration between distinct secretory organelles. PLoS Pathog 1:e17
Besteiro S, Dubremetz JF, Lebrun M (2011) The moving junction of apicomplexan parasites: a key structure for invasion. Cell Microbiol 13:797–805
Holder AA (1988) The precursor to major merozoite surface antigens: structure and role in immunity. Prog Allergy 41:72–97
Stafford WH, Blackman MJ, Harris A et al (1994) N-terminal amino acid sequence of the Plasmodium falciparum merozoite surface protein-1 polypeptides. Mol Biochem Parasitol 66:157–160
Blackman MJ, Holder AA (1992) Secondary processing of the Plasmodium falciparum merozoite surface protein-1 (MSP1) by a calcium-dependent membrane-bound serine protease: shedding of MSP133 as a noncovalently associated complex with other fragments of the MSP1. Mol Biochem Parasitol 50:307–315
Pachebat JA, Ling IT, Grainger M et al (2001) The 22 kDa component of the protein complex on the surface of Plasmodium falciparum merozoites is derived from a larger precursor, merozoite surface protein 7. Mol Biochem Parasitol 117:83–89
Pachebat JA, Kadekoppala M, Grainger M et al (2007) Extensive proteolytic processing of the malaria parasite merozoite surface protein 7 during biosynthesis and parasite release from erythrocytes. Mol Biochem Parasitol 151:59–69
Dutta S, Haynes JD, Barbosa A et al (2005) Mode of action of invasion-inhibitory antibodies directed against apical membrane antigen 1 of Plasmodium falciparum. Infect Immun 73:2116–2122
Treeck M, Zacherl S, Herrmann S et al (2009) Functional analysis of the leading malaria vaccine candidate AMA-1 reveals an essential role for the cytoplasmic domain in the invasion process. PLoS Pathog 5:e1000322
Howell SA, Withers-Martinez C, Kocken CH et al (2001) Proteolytic processing and primary structure of Plasmodium falciparum apical membrane antigen-1. J Biol Chem 276: 31311–31320
Howell S, Well I, Fleck S et al (2003) A single malaria merozoite serine protease mediates shedding of multiple surface proteins by juxtamembrane cleavage. J Biol Chem 278: 23890–23898
Barale JC, Blisnick T, Fujioka H et al (1999) Plasmodium falciparum subtilisin-like protease 2, a merozoite candidate for the merozoite surface protein 1-42 maturase. Proc Natl Acad Sci USA 96:6445–6450
Hackett F, Sajid M, Withers-Martinez C et al (1999) PfSUB-2: a second subtilisin-like protein in Plasmodium falciparum merozoites. Mol Biochem Parasitol 103:183–195
Gardner MJ, Hall N, Fung E et al (2002) Genome sequence of the human malaria parasite Plasmodium falciparum. Nature 419:498–511
Blackman MJ, Fujioka H, Stafford WH et al (1998) A subtilisin-like protein in secretory organelles of Plasmodium falciparum merozoites. J Biol Chem 273:23398–23409
Agarwal S, Singh MK, Garg S et al (2012) Ca(2+) -mediated exocytosis of subtilisin-like protease 1: a key step in egress of Plasmodium falciparum merozoites. Cell Microbiol, DOI: 10.1111/cmi.12086
Yeoh S, O’Donnell RA, Koussis K et al (2007) Subcellular discharge of a serine protease mediates release of invasive malaria parasites from host erythrocytes. Cell 131:1072–1083
Koussis K, Withers-Martinez C, Yeoh S et al (2009) A multifunctional serine protease primes the malaria parasite for red blood cell invasion. EMBO J 28:725–735
Silmon de Monerri NC, Flynn HR, Campos MG et al (2011) Global identification of multiple substrates for Plasmodium falciparum SUB1, an essential malarial processing protease. Infect Immun 79:1086–1097
Arastu-Kapur S, Ponder EL, Fonovic UP et al (2008) Identification of proteases that regulate erythrocyte rupture by the malaria parasite Plasmodium falciparum. Nat Chem Biol 4:203–213
Blackman MJ (2008) Malarial proteases and host cell egress: an ‘emerging’ cascade. Cell Microbiol 10:1925–1934
Harris PK, Yeoh S, Dluzewski AR et al (2005) Molecular identification of a malaria merozoite surface sheddase. PLoS Pathog 1:241–251
Green JL, Hinds L, Grainger M et al (2006) Plasmodium thrombospondin related apical merozoite protein (PTRAMP) is shed from the surface of merozoites by PfSUB2 upon invasion of erythrocytes. Mol Biochem Parasitol 150:114–117
Uzureau P, Barale JC, Janse CJ et al (2004) Gene targeting demonstrates that the Plasmodium berghei subtilisin PbSUB2 is essential for red cell invasion and reveals spontaneous genetic recombination events. Cell Microbiol 6:65–78
Alam A, Bhatnagar RK, Chauhan VS (2012) Expression and characterization of catalytic domain of Plasmodium falciparum subtilisin-like protease 3. Mol Biochem Parasitol 183:84–89
Miller SK, Good RT, Drew DR et al (2002) A subset of Plasmodium falciparum SERA genes are expressed and appear to play an important role in the erythrocytic cycle. J Biol Chem 277:47524–47532
McCoubrie JE, Miller SK, Sargeant T et al (2007) Evidence for a common role for the serine-type Plasmodium falciparum serine repeat antigen proteases: implications for vaccine and drug design. Infect Immun 75:5565–5574
Chulay JD, Lyon JA, Haynes JD et al (1987) Monoclonal antibody characterization of Plasmodium falciparum antigens in immune complexes formed when schizonts rupture in the presence of immune serum. J Immunol 139:2768–2774
Higgins DG, McConnell DJ, Sharp PM (1989) Malarial proteinase? Nature 340:604
Hodder AN, Drew DR, Epa VC et al (2003) Enzymic, phylogenetic, and structural characterization of the unusual papain-like protease domain of Plasmodium falciparum SERA5. J Biol Chem 278:48169–48177
Pang XL, Mitamura T, Horii T (1999) Antibodies reactive with the N-terminal domain of Plasmodium falciparum serine repeat antigen inhibit cell proliferation by agglutinating merozoites and schizonts. Infect Immun 67:1821–1827
Li J, Mitamura T, Fox BA et al (2002) Differential localization of processed fragments of Plasmodium falciparum serine repeat antigen and further processing of its N-terminal 47 kDa fragment. Parasitol Int 51:343–352
Delplace P, Bhatia A, Cagnard M et al (1988) Protein p126: a parasitophorous vacuole antigen associated with the release of Plasmodium falciparum merozoites. Biol Cell 64:215–221
Ruecker A, Shea M, Hackett F et al (2012) Proteolytic activation of the essential parasitophorous vacuole cysteine protease SERA6 accompanies malaria parasite egress from its host erythrocyte. J Biol Chem 287:37949–37963
Nwagwu M, Haynes JD, Orlandi PA et al (1992) Plasmodium falciparum: chymotryptic-like proteolysis associated with a 101-kDa acidic-basic repeat antigen. Exp Parasitol 75:399–414
Kushwaha A, Rao PP, Duttu VS et al (2000) Expression and characterisation of Plasmodium falciparum acidic basic repeat antigen expressed in Escherichia coli. Mol Biochem Parasitol 106:213–224
Lopera TM, Restrepo M, Blair S et al (1998) Humoral immune response to the anti-malaria vaccine SPf66 in the Colombian Atrato River region. Mem Inst Oswaldo Cruz 93:495–500
Curtidor H, Urquiza M, Suarez J et al (2001) Plasmodium falciparum acid basic repeat antigen (ABRA) peptides: erythrocyte binding and biological activity. Vaccine 19:4496–4504
Vargas-Serrato E, Barnwell JW, Ingravallo P et al (2002) Merozoite surface protein-9 of Plasmodium vivax and related simian malaria parasites is orthologous to p101/ABRA of P. falciparum. Mol Biochem Parasitol 120:41–52
Mills KE, Pearce JA, Crabb BS et al (2002) Truncation of merozoite surface protein 3 disrupts its trafficking and that of acidic-basic repeat protein to the surface of Plasmodium falciparum merozoites. Mol Microbiol 43:1401–1411
Zhou XW, Blackman MJ, Howell SA et al (2004) Proteomic analysis of cleavage events reveals a dynamic two-step mechanism for proteolysis of a key parasite adhesive complex. Mol Cell Proteomics 3:565–576
Howell SA, Hackett F, Jongco AM et al (2005) Distinct mechanisms govern proteolytic shedding of a key invasion protein in apicomplexan pathogens. Mol Microbiol 57:1342–1356
Baker RP, Wijetilaka R, Urban S (2006) Two Plasmodium rhomboid proteases preferentially cleave different adhesins implicated in all invasive stages of malaria. PLoS Pathog 2:e113
O’Donnell RA, Hackett F, Howell SA et al (2006) Intramembrane proteolysis mediates shedding of a key adhesin during erythrocyte invasion by the malaria parasite. J Cell Biol 174:1023–1033
Vera IM, Beatty WL, Sinnis P et al (2011) Plasmodium protease ROM1 is important for proper formation of the parasitophorous vacuole. PLoS Pathog 7:e1002197
Srinivasan P, Coppens I, Jacobs-Lorena M (2009) Distinct roles of Plasmodium rhomboid 1 in parasite development and malaria pathogenesis. PLoS Pathog 5:e1000262
Rosenthal PJ, Sijwali PS, Singh A et al (2002) Cysteine proteases of malaria parasites: targets for chemotherapy. Curr Pharm Des 8:1659–1672
Greenbaum DC, Baruch A, Grainger M et al (2002) A role for the protease falcipain 1 in host cell invasion by the human malaria parasite. Science 298:2002–2006
Sijwali PS, Kato K, Seydel KB et al (2004) Plasmodium falciparum cysteine protease falcipain-1 is not essential in erythrocytic stage malaria parasites. Proc Natl Acad Sci USA 101:8721–8726
Banyal HS, Misra GC, Gupta CM et al (1981) Involvement of malarial proteases in the interaction between the parasite and host erythrocyte in Plasmodium knowlesi infections. J Parasitol 67:623–626
Wickham ME, Culvenor JG, Cowman AF (2003) Selective inhibition of a two-step egress of malaria parasites from the host erythrocyte. J Biol Chem 278:37658–37663
Le Bonniec S, Deregnaucourt C, Redeker V et al (1999) Plasmepsin II, an acidic hemoglobinase from the Plasmodium falciparum food vacuole, is active at neutral pH on the host erythrocyte membrane skeleton. J Biol Chem 274:14218–14223
Dhawan S, Dua M, Chishti AH et al (2003) Ankyrin peptide blocks falcipain-2-mediated malaria parasite release from red blood cells. J Biol Chem 278:30180–30186
Dasaradhi PV, Korde R, Thompson JK et al (2007) Food vacuole targeting and trafficking of falcipain-2, an important cysteine protease of human malaria parasite Plasmodium falciparum. Mol Biochem Parasitol 156:12–23
Aoki S, Li J, Itagaki S et al (2002) Serine repeat antigen (SERA5) is predominantly expressed among the SERA multigene family of Plasmodium falciparum, and the acquired antibody titers correlate with serum inhibition of the parasite growth. J Biol Chem 277:47533–47540
Dua M, Raphael P, Sijwali PS et al (2001) Recombinant falcipain-2 cleaves erythrocyte membrane ankyrin and protein 4.1. Mol Biochem Parasitol 116:95–99
Hanspal M, Dua M, Takakuwa Y et al (2002) Plasmodium falciparum cysteine protease falcipain-2 cleaves erythrocyte membrane skeletal proteins at late stages of parasite development. Blood 100:1048–1054
Dasaradhi PV, Mohmmed A, Kumar A et al (2005) A role of falcipain-2, principal cysteine proteases of Plasmodium falciparum in merozoite egression. Biochem Biophys Res Commun 336:1062–1068
Omara-Opyene AL, Moura PA, Sulsona CR et al (2004) Genetic disruption of the Plasmodium falciparum digestive vacuole plasmepsins demonstrates their functional redundancy. J Biol Chem 279:54088–54096
Sijwali PS, Rosenthal PJ (2004) Gene disruption confirms a critical role for the cysteine protease falcipain-2 in hemoglobin hydrolysis by Plasmodium falciparum. Proc Natl Acad Sci USA 101:4384–4389
Liu J, Gluzman IY, Drew ME et al (2005) The role of Plasmodium falciparum food vacuole plasmepsins. J Biol Chem 280:1432–1437
Sijwali PS, Koo J, Singh N et al (2006) Gene disruptions demonstrate independent roles for the four falcipain cysteine proteases of Plasmodium falciparum. Mol Biochem Parasitol 150:96–106
Chandramohanadas R, Davis PH, Beiting DP et al (2009) Apicomplexan parasites co-opt host calpains to facilitate their escape from infected cells. Science 324:794–797
Singh N, Sijwali PS, Pandey KC et al (2006) Plasmodium falciparum: biochemical characterization of the cysteine protease falcipain-2′. Exp Parasitol 112:187–192
Chugh M, Sundararaman V, Kumar S et al (2013) Protein complex directs hemoglobin-to-hemozoin formation in Plasmodium falciparum. Proc Natl Acad Sci USA 110:5392–5397
Murata CE, Goldberg DE (2003) Plasmodium falciparum falcilysin: a metalloprotease with dual specificity. J Biol Chem 278:38022–38028
Francis SE, Banerjee R, Goldberg DE (1997) Biosynthesis and maturation of the malaria aspartic hemoglobinases plasmepsins I and II. J Biol Chem 272:14961–14968
Ponpuak M, Klemba M, Park M et al (2007) A role for falcilysin in transit peptide degradation in the Plasmodium falciparum apicoplast. Mol Microbiol 63:314–334
Francis SE, Gluzman IY, Oksman A et al (1994) Molecular characterization and inhibition of a Plasmodium falciparum aspartic hemoglobinase. EMBO J 13:306–317
Banerjee R, Liu J, Beatty W et al (2002) Four plasmepsins are active in the Plasmodium falciparum food vacuole, including a protease with an active-site histidine. Proc Natl Acad Sci USA 99(2):990–995
Silva AM, Lee AY, Gulnik SV et al (1996) Structure and inhibition of plasmepsin II, a hemoglobin-degrading enzyme from Plasmodium falciparum. Proc Natl Acad Sci USA 93:10034–10039
Moon RP, Tyas L, Certa U et al (1997) Expression and characterisation of plasmepsin I from Plasmodium falciparum. Eur J Biochem 244:552–560
Haque TS, Skillman AG, Lee CE et al (1999) Potent, low-molecular-weight non-peptide inhibitors of malarial aspartyl protease plasmepsin II. J Med Chem 42:1428–1440
Nezami A, Luque I, Kimura T et al (2002) Identification and characterization of allophenylnorstatine-based inhibitors of plasmepsin II, an antimalarial target. Biochemistry 41:2273–2280
Skinner-Adams TS, McCarthy JS, Gardiner DL et al (2004) Antiretrovirals as antimalarial agents. J Infect Dis 190:1998–2000
Parikh S, Gut J, Istvan E et al (2005) Antimalarial activity of human immunodeficiency virus type 1 protease inhibitors. Antimicrob Agents Chemother 49:2983–2985
Savarino A, Cauda R, Cassone A (2005) Aspartic proteases of Plasmodium falciparum as the target of HIV-1 protease inhibitors. J Infect Dis 191:1381–1382, author reply 1382–1383
Shenai BR, Sijwali PS, Singh A et al (2000) Characterization of native and recombinant falcipain-2, a principal trophozoite cysteine protease and essential hemoglobinase of Plasmodium falciparum. J Biol Chem 275:29000–29010
Dahl EL, Rosenthal PJ (2005) Biosynthesis, localization, and processing of falcipain cysteine proteases of Plasmodium falciparum. Mol Biochem Parasitol 139:205–212
Mohmmed A, Dasaradhi PV, Bhatnagar RK et al (2003) In vivo gene silencing in Plasmodium berghei - a mouse malaria model. Biochem Biophys Res Commun 309:506–511
Powers JC, Asgian JL, Ekici OD et al (2002) Irreversible inhibitors of serine, cysteine, and threonine proteases. Chem Rev 102:4639–4750
Rosenthal PJ, Wollish WS, Palmer JT et al (1991) Antimalarial effects of peptide inhibitors of a Plasmodium falciparum cysteine proteinase. J Clin Invest 88:1467–1472
Gibbons P, Verissimo E, Araujo NC et al (2010) Endoperoxide carbonyl falcipain 2/3 inhibitor hybrids: toward combination chemotherapy of malaria through a single chemical entity. J Med Chem 53:8202–8206
Liu Y, Lu WQ, Cui KQ et al (2012) Synthesis and biological activities of novel artemisinin derivatives as cysteine protease falcipain-2 inhibitors. Arch Pharm Res 35:1525–1531
Mane UR, Gupta RC, Nadkarni SS et al (2013) Falcipain inhibitors as potential therapeutics for resistant strains of malaria: a patent review. Expert Opin Ther Pat 23:165–187
Rizzi L, Sundararaman S, Cendic K et al (2011) Design and synthesis of protein-protein interaction mimics as Plasmodium falciparum cysteine protease, falcipain-2 inhibitors. Eur J Med Chem 46:2083–2090
Curley GP, O’Donovan SM, McNally J et al (1994) Aminopeptidases from Plasmodium falciparum, Plasmodium chabaudi chabaudi and Plasmodium berghei. J Eukaryot Microbiol 41:119–123
Gavigan CS, Dalton JP, Bell A (2001) The role of aminopeptidases in haemoglobin degradation in Plasmodium falciparum-infected erythrocytes. Mol Biochem Parasitol 117:37–48
Stack CM, Lowther J, Cunningham E et al (2007) Characterization of the Plasmodium falciparum M17 leucyl aminopeptidase. A protease involved in amino acid regulation with potential for antimalarial drug development. J Biol Chem 282:2069–2080
Florent I, Derhy Z, Allary M et al (1998) A Plasmodium falciparum aminopeptidase gene belonging to the M1 family of zinc-metallopeptidases is expressed in erythrocytic stages. Mol Biochem Parasitol 97:149–160
Allary M, Schrevel J, Florent I (2002) Properties, stage-dependent expression and localization of Plasmodium falciparum M1 family zinc-aminopeptidase. Parasitology 125:1–10
Dalal S, Klemba M (2007) Roles for two aminopeptidases in vacuolar hemoglobin catabolism in Plasmodium falciparum. J Biol Chem 282:35978–35987
Teuscher F, Lowther J, Skinner-Adams TS et al (2007) The M18 aspartyl aminopeptidase of the human malaria parasite Plasmodium falciparum. J Biol Chem 282:30817–30826
Poreba M, McGowan S, Skinner-Adams TS et al (2012) Fingerprinting the substrate specificity of M1 and M17 aminopeptidases of human malaria, Plasmodium falciparum. PLoS One 7:e31938
Dalal S, Ragheb DR, Klemba M (2012) Engagement of the S1, S1′ and S2′ subsites drives efficient catalysis of peptide bond hydrolysis by the M1-family aminopeptidase from Plasmodium falciparum. Mol Biochem Parasitol 183:70–77
Jones PM, Robinson MW, Dalton JP et al (2011) The Plasmodium falciparum malaria M1 alanyl aminopeptidase (PfA-M1): insights of catalytic mechanism and function from MD simulations. PLoS One 6:e28589
Skinner-Adams TS, Lowther J, Teuscher F et al (2007) Identification of phosphinate dipeptide analog inhibitors directed against the Plasmodium falciparum M17 leucine aminopeptidase as lead antimalarial compounds. J Med Chem 50:6024–6031
Harbut MB, Velmourougane G, Dalal S et al (2011) Bestatin-based chemical biology strategy reveals distinct roles for malaria M1- and M17-family aminopeptidases. Proc Natl Acad Sci USA 108:E526–E534
Waller RF, McFadden GI (2005) The apicoplast: a review of the derived plastid of apicomplexan parasites. Curr Issues Mol Biol 7:57–79
Goodman CD, Su V, McFadden GI (2007) The effects of anti-bacterials on the malaria parasite Plasmodium falciparum. Mol Biochem Parasitol 152:181–191
Schlitzer M (2007) Malaria chemotherapeutics part I: history of antimalarial drug development, currently used therapeutics, and drugs in clinical development. Chem Med Chem 2:944–986
Schlitzer M (2006) Selective enzyme inhibitor instead of an “iron-triggered cluster bomb”. Pharm Unserer Zeit 35:8–9
Dahl EL, Rosenthal PJ (2008) Apicoplast translation, transcription and genome replication: targets for antimalarial antibiotics. Trends Parasitol 24:279–284
Tschan S, Kreidenweiss A, Stierhof YD et al (2010) Mitochondrial localization of the threonine peptidase PfHslV, a ClpQ ortholog in Plasmodium falciparum. Int J Parasitol 40:1517–1523
Ramasamy G, Gupta D, Mohmmed A et al (2007) Characterization and localization of Plasmodium falciparum homolog of prokaryotic ClpQ/HslV protease. Mol Biochem Parasitol 152:139–148
Sousa MC, Trame CB, Tsuruta H et al (2000) Crystal and solution structures of an HslUV protease-chaperone complex. Cell 103:633–643
Rathore S, Jain S, Sinha D et al (2011) Disruption of a mitochondrial protease machinery in Plasmodium falciparum is an intrinsic signal for parasite cell death. Cell Death Dis 2:e231
Li Z, Lindsay ME, Motyka SA et al (2008) Identification of a bacterial-like HslVU protease in the mitochondria of Trypanosoma brucei and its role in mitochondrial DNA replication. PLoS Pathog 4:e1000048
Jain S, Rathore S, Asad M et al (2013) The prokaryotic ClpQ protease plays a key role in growth and development of mitochondria in Plasmodium falciparum. Cell Microbiol, DOI: 10.1111/cmi.12142
Ralph SA, van Dooren GG, Waller RF et al (2004) Tropical infectious diseases: metabolic maps and functions of the Plasmodium falciparum apicoplast. Nat Rev Microbiol 2:203–216
McConkey GA, Rogers MJ, McCutchan TF (1997) Inhibition of Plasmodium falciparum protein synthesis. Targeting the plastid-like organelle with thiostrepton. J Biol Chem 272:2046–2049
Lin Q, Katakura K, Suzuki M (2002) Inhibition of mitochondrial and plastid activity of Plasmodium falciparum by minocycline. FEBS Lett 515:71–74
Williamson DH, Preiser PR, Moore PW et al (2002) The plastid DNA of the malaria parasite Plasmodium falciparum is replicated by two mechanisms. Mol Microbiol 45:533–542
Chaubey S, Kumar A, Singh D et al (2005) The apicoplast of Plasmodium falciparum is translationally active. Mol Microbiol 56:81–89
Yeh E, DeRisi JL (2011) Chemical rescue of malaria parasites lacking an apicoplast defines organelle function in blood-stage Plasmodium falciparum. PLoS Biol 9:e1001138
van Dooren GG, Su V, D’Ombrain MC et al (2002) Processing of an apicoplast leader sequence in Plasmodium falciparum and the identification of a putative leader cleavage enzyme. J Biol Chem 277:23612–23619
El Bakkouri M, Rathore S, Calmettes C et al (2013) Structural insights into the inactive subunit of the apicoplast-localized caseinolytic protease complex of Plasmodium falciparum. J Biol Chem 288:1022–1031
Rathore S, Sinha D, Asad M et al (2010) A cyanobacterial serine protease of Plasmodium falciparum is targeted to the apicoplast and plays an important role in its growth and development. Mol Microbiol 77:873–890
Gille C, Goede A, Schloetelburg C et al (2003) A comprehensive view on proteasomal sequences: implications for the evolution of the proteasome. J Mol Biol 326:1437–1448
Mordmuller B, Fendel R, Kreidenweiss A et al (2006) Plasmodia express two threonine-peptidase complexes during asexual development. Mol Biochem Parasitol 148:79–85
Aminake MN, Mahajan A, Kumar V et al (2012) Synthesis and evaluation of hybrid drugs for a potential HIV/AIDS-malaria combination therapy. Bioorg Med Chem 20:5277–5289
Gantt SM, Myung JM, Briones MR et al (1998) Proteasome inhibitors block development of Plasmodium spp. Antimicrob Agents Chemother 42:2731–2738
Reynolds JM, El Bissati K, Brandenburg J et al (2007) Antimalarial activity of the anticancer and proteasome inhibitor bortezomib and its analog ZL3B. BMC Clin Pharmacol 7:13
Kreidenweiss A, Kremsner PG, Mordmuller B (2008) Comprehensive study of proteasome inhibitors against Plasmodium falciparum laboratory strains and field isolates from Gabon. Malar J 7:187
Prudhomme J, McDaniel E, Ponts N et al (2008) Marine actinomycetes: a new source of compounds against the human malaria parasite. PLoS One 3:e2335
Frickel EM, Quesada V, Muething L et al (2007) Apicomplexan UCHL3 retains dual specificity for ubiquitin and Nedd8 throughout evolution. Cell Microbiol 9:1601–1610
Ponder EL, Albrow VE, Leader BA et al (2011) Functional characterization of a SUMO deconjugating protease of Plasmodium falciparum using newly identified small molecule inhibitors. Chem Biol 18:711–721
Ponder EL, Bogyo M (2007) Ubiquitin-like modifiers and their deconjugating enzymes in medically important parasitic protozoa. Eukaryot Cell 6:1943–1952
Ponts N, Yang J, Chung DW et al (2008) Deciphering the ubiquitin-mediated pathway in apicomplexan parasites: a potential strategy to interfere with parasite virulence. PLoS One 3:e2386
Spork S, Hiss JA, Mandel K et al (2009) An unusual ERAD-like complex is targeted to the apicoplast of Plasmodium falciparum. Eukaryot Cell 8:1134–1145
Artavanis-Tsakonas K, Misaghi S, Comeaux CA et al (2006) Identification by functional proteomics of a deubiquitinating/deNeddylating enzyme in Plasmodium falciparum. Mol Microbiol 61:1187–1195
Le Roch KG, Johnson JR, Florens L et al (2004) Global analysis of transcript and protein levels across the Plasmodium falciparum life cycle. Genome Res 14:2308–2318
Artavanis-Tsakonas K, Weihofen WA, Antos JM et al (2010) Characterization and structural studies of the Plasmodium falciparum ubiquitin and Nedd8 hydrolase UCHL3. J Biol Chem 285:6857–6866
Issar N, Roux E, Mattei D et al (2008) Identification of a novel post-translational modification in Plasmodium falciparum: protein sumoylation in different cellular compartments. Cell Microbiol 10:1999–2011
Sharma S, Pradhan A, Chauhan VS et al (2005) Isolation and characterization of type I signal peptidase of different malaria parasites. J Biomed Biotechnol 2005:301–309
Tuteja R, Pradhan A, Sharma S (2008) Plasmodium falciparum signal peptidase is regulated by phosphorylation and required for intra-erythrocytic growth. Mol Biochem Parasitol 157:137–147
McPherson RA, Donald DR, Sawyer WH et al (1993) Proteolytic digestion of band 3 at an external site alters the erythrocyte membrane organisation and may facilitate malarial invasion. Mol Biochem Parasitol 62:233–242
Li X, Chen H, Oh SS et al (2008) A Presenilin-like protease associated with Plasmodium falciparum micronemes is involved in erythrocyte invasion. Mol Biochem Parasitol 158:22–31
Li X, Chen H, Bahamontes-Rosa N et al (2009) Plasmodium falciparum signal peptide peptidase is a promising drug target against blood stage malaria. Biochem Biophys Res Commun 380:454–459
Marapana DS, Wilson DW, Zuccala ES et al (2012) Malaria parasite signal peptide peptidase is an ER-resident protease required for growth but not for invasion. Traffic 13:1457–1465
Chang HH, Falick AM, Carlton PM et al (2008) N-terminal processing of proteins exported by malaria parasites. Mol Biochem Parasitol 160:107–115
Boddey JA, Moritz RL, Simpson RJ et al (2009) Role of the Plasmodium export element in trafficking parasite proteins to the infected erythrocyte. Traffic 10:285–299
Boddey JA, Hodder AN, Gunther S et al (2010) An aspartyl protease directs malaria effector proteins to the host cell. Nature 463:627–631
Klemba M, Gluzman I, Goldberg DE (2004) A Plasmodium falciparum dipeptidyl aminopeptidase I participates in vacuolar hemoglobin degradation. J Biol Chem 279:43000–43007
Acknowledgements
The Malaria Group at ICGEB is supported by Program Support Grant from Department of Biotechnology, Govt. of India, and research grant to AM under Indo-Swiss Joint Research Programme from Department of Science & Technology, Govt. of India. AM is a recipient of National Bioscience Award for Career Development from Department of Biotechnology, Govt. of India.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Rathore, S., Jain, S., Asad, M., Datta, G., Malhotra, P., Mohmmed, A. (2013). Role of Proteases During Intra-erythrocytic Developmental Cycle of Human Malaria Parasite Plasmodium falciparum . In: Chakraborti, S., Dhalla, N. (eds) Proteases in Health and Disease. Advances in Biochemistry in Health and Disease, vol 7. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-9233-7_13
Download citation
DOI: https://doi.org/10.1007/978-1-4614-9233-7_13
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-9232-0
Online ISBN: 978-1-4614-9233-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)