Abstract
Numerous interphase molecular cytogenetic approaches are useful for the analysis of chromosomes in normal and abnormal human cells. Interphase fluorescence in situ hybridization techniques offer unique possibilities to visualize individual chromosomes or chromosomal regions in single nondividing cells isolated from any given tissue. Despite technological difficulties encountered during studying human interphase chromosomes in health and disease, molecular cytogenetics or cytogenomics (“chromosomics”) does provide solutions for high-resolution single-cell analysis of genome organization, structure, and behavior at all stages of the cell cycle. However, usually relatively little attention is paid to interphase molecular cytogenetics in current biomedical literature. Looking through the voluminous amount of original research papers and reviews dedicated to human interphase chromosomes, one can conclude that the technological aspects of studying human interphase chromosomes applied to basic and clinical research are rarely addressed. In an attempt to fill this gap, the present chapter provides a description of technological solutions in human interphase cytogenetics.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Preimplantation Genetic Diagnosis
- Interphase Nucleus
- Interphase Chromosome
- Molecular Cytogenetic Technique
- Asynchronous Replication
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Introduction
It is generally accepted that almost all fluorescence in situ hybridization (FISH) protocols are applicable for developing an interphase FISH (I-FISH) method. To learn more about numerous approaches and applications of useful FISH-based techniques and detailed protocols, readers can refer to recent FISH application guides edited by the authors with the contribution of leading international experts in the field of molecular cytogenetics and cytogenomics, who also contributed to this book (Fluorescence In Situ Hybridization (FISH)—Application Guide, edited by Thomas Liehr, Springer, 2009, and Fluorescence In Situ Hybridization (FISH): Protocols and Applications, edited by Joanna M. Bridger and Emanuela V. Volpi, Humana Press, 2010).
Recently we have attempted to give an overview of currently applied molecular cytogenetic techniques with a special emphasis on their technological abilities for studying human interphase chromosomes (Vorsanova et al. 2010a, freely available at Molecular Cytogenetics (BioMed Central), an open access journal dedicated to different aspects of chromosome and genome biology, http://www.molecularcytogenetics.org/content/3/1/1). Here we present an updated review dedicated to technological achievements in human interphase cytogenetics.
According to Gersen and Keagle (2005), it is estimated that more than one million cytogenetic and molecular cytogenetic analyses are performed each year. Taken together, these analyses represent the standard of care in medical genetics and routine clinical workups for numerous patients suffering from congenital malformations, mental diseases, cancers, or reproductive problems (Carter 2007; Liehr 2009; Vorsanova et al. 2010b). The significance of molecular cytogenetic techniques for molecular diagnosis has been repeatedly shown, and these techniques are recognized as a valuable addition or even alternative to conventional cytogenetics (Liehr and Claussen 2002; Iourov et al. 2008a; Bejjani and Shaffer 2008). In addition, molecular cytogenetic technologies are widely used in basic biomedical research (Liehr et al. 2004). For instance, the thousands of articles mentioning at least one molecular cytogenetic technique are indexed in browsable scientific databases (for more details, see Iourov et al. 2008a; Chap. 12, and the web page about multicolor FISH at http://www.med.uni-jena.de/fish/mFISH/mFISHlit.htm website, managed by Dr. Thomas Liehr, Jena, Germany). Thus, it is certain that the role of molecular cytogenetics in current biomedicine is appreciable.
Two essential advantages of molecular cytogenetics can be noted: (1) the ability to provide either a high-resolution on-chip scan of the whole genome or to visualize single specific genomic loci (Bejjani and Shaffer 2008), and (2) the capability to analyze DNA (RNA)-based genome organization, structure, and behavior in single cells (Levsky and Singer 2003; Iourov et al. 2006a). The first advantage is appreciable when analyzing mixed DNA isolated from a large amount of cells and is rarely appreciated in single-cell genomic studies (Iourov et al. 2012; Vanneste et al. 2012). The second advantage of molecular cytogenetic techniques is consistently emphasized but is usually applied to studying metaphase chromosomes of mitotic cells. However, eukaryotic cells are more likely to be in interphase. Therefore, surveys of genome organization, structure, and behavior do not evaluate an essential part of cellular life. In molecular diagnosis, interphase analysis is also not commonly applied. One might suggest a lack of reproducibility and the low resolution of interphase cytogenetic techniques. However, a brief look through molecular cytogenetic studies of somatic genomic variations and genome behavior in interphase nuclei (Walter et al. 2006; Goetze et al. 2007; Iourov et al. 2008b, 2012; Rouquette et al. 2010) and developments in interphase cytogenetics (Iourov et al. 2006c; Liehr 2009; Vorsanova et al. 2010a, b) demonstrates that this idea is unsupported. Furthermore, numerous laboratories elaborating such techniques are able to solve important practical and research tasks without notable difficulties. Evidently, interphase molecular cytogenetics requires additional attention, which is the intention of the present chapter.
Molecular Cytogenetic Techniques and Interphase Cytogenetics
There are currently two essential platforms available for developments in molecular cytogenetics: FISH, including comparative genomic hybridization (CGH), and peptide nucleic acid (PNA) probing for analysis of chromosomal DNA (Liehr and Claussen 2002; Iourov et al. 2008a; Liehr 2009; Vorsanova et al. 2010a). The resolution of such techniques is usually established against cytogenetic banding analysis (the gold standard of resolution for genetic analyses). Single-cell molecular cytogenetics analyzes either metaphase plates or interphase nuclei. Study of metaphase plates is traditionally made by means of several detection technologies [spectral karyotyping (SKY) or multicolor FISH (MFISH)] (Schrock et al. 1996; Speicher et al. 1996) or specific DNA probe sets (chromosome-enumeration/centromeric, site-specific, whole-painting, microdissected) (Yurov et al. 1996; Soloviev et al. 1998a, b; Liehr et al. 2002; Nietzel et al. 2001). If modified, these techniques can be applied to interphase chromosomal analysis, but this “translation” (transfer of technology) requires major efforts (Vorsanova et al. 2010a; Iourov et al. 2010). Table 11.1 provides an overview of the molecular cytogenetic techniques used for metaphase and interphase analysis of single cells. As one can see, molecular cytogenetics allows us to perform high-resolution analysis of chromosomal structure and behavior at all stages of the cell cycle. Nonetheless, molecular cytogenetic methods are preferentially used for detecting metaphase chromosome imbalances and rearrangements or for whole-genome scans by CGH (Liehr et al. 2004; Vorsanova et al. 2010b).
Visualization is the key stage of studying interphase chromosomes. Without direct (microscopic) visualization of DNA-base chromosomal structures, related research is certainly incomplete. Thus, FISH-based techniques offer the unique possibility to depict either whole chromosomes or specific genomic loci in single cells. In other words, if one wishes to obtain valid data on human interphase chromosomes, one will undertake an I-FISH study. Further, we attempt to review the technique in context of applications to single-cell chromosomal analysis, which is the basis of interphase molecular cytogenetics.
I-FISH
FISH represents a general-purpose technique for studies of both the whole genome and specific genomic loci. The resolution of molecular cytogenetic is essentially determined according to the DNA sequence size of probes hybridizing in situ. DNA probes are designated as centromeric and telomeric (hybridizing to repetitive-sequence DNA), site-specific (hybridizing to euchromatic DNA, i.e., DNA sequences of a gene), or whole chromosome painting (wcp; hybridizing to DNA of whole chromosomes) (Table 11.2).
FISH with chromosome-specific DNA probes: FISH painting of repetitive genomic sequences is performed with centromeric (chromosome-enumeration or chromosome-specific) or telomeric DNA probes. Analysis of telomeres is an important area of biomedical research (Aubert et al. 2012). In such approaches, PNA/DNA probes possessing TTAGGG repetitive motifs are used. Representing the technological basis of telomere biology (cancer and aging research), telomere FISH and related techniques are poorly applicable for diagnosis. On the other hand, I-FISH with telomeric probes is applicable for analysis of nuclear organization (Klewes et al. 2011).
I-FISH using centromeric DNA probes has become an integral part of molecular diagnosis in medical genetics, oncology, and reproductive medicine (Cremer et al. 1986; Vorsanova et al. 1986, 1991, 2005b; Baumgartner et al. 2006; Yurov et al. 2007; Iourov et al. 2008b). Additionally, it is repeatedly demonstrated that I-FISH with these probes is highly applicable in chromosome biology studies encompassing genome research (chromosomal and nuclear), evolution, behavior, and variation in health and disease. These DNA probes feature nearly 100% hybridization efficiency and chromosome specificity. As a result, analysis of an individual homologous chromosome pair in interphase becomes possible. Moreover, extreme interindividual variations of pericentromeric heterochromatic DNA has led to developing quantitative FISH (QFISH) solving numerous problems encountered during metaphase and interphase analysis of chromosomes (Iourov et al. 2005; Vorsanova et al. 2005a). It is noteworthy that the resolution of related assays is poorly determined by DNA sequence size of loci assessed (Table 11.1), inasmuch as centromeric I-FISH is used for the analysis of phenomena involving large chromosomal regions or even whole chromosomes. For instance, I-FISH with chromosome-enumeration probes allows the detection of numerical chromosome imbalances (aneuploidy and polyploidy) in interphase. The latter is the most frequent application of the method (Fig. 11.1) and is required for pre-/postnatal diagnosis, cancer diagnosis/prognosis, and somatic genomic variation surveys. The nearly 100 % hybridization efficiency of centromeric DNA probes and chromosome-specific DNA sequences forming pericentromeric (heterochromatic) chromosomal regions (apart from shared alphoid DNA of chromosomes 5 and 19, 13 and 21, and 14 and 22) (Yurov et al. 1996; Lee et al. 1997; Vorsanova et al. 2002, 2005a) is the essential advantage that this technique possesses. However, heteromorphisms of pericentromeric DNAs can produce the lack of a signal leading, thereby, to impossibility of the I-FISH assay application. Fortunately, such extreme heteromorphisms (centromeric DNA variations) are rare in the general population (Verma and Luke 1992; Liehr et al. 1998; Vorsanova et al. 2002, 2005a).
Two- and three-color interphase fluorescence in situ hybridization (I-FISH) with centromeric DNA probes: (a) normal diploid nucleus with two signals for chromosome 1 and chromosome 15; (b) monosomic nucleus with two signals for chromosome 1 and one signal for chromosome 15; (c) trisomic nucleus with two signals for chromosome 1 and three signals for chromosome 15; (d) normal diploid nucleus with two signals for chromosome 1, chromosome 9, and chromosome 16; (e) monosomic nucleus with two signals for chromosome 1 and chromosome 9 and one signal for chromosome 16; (f) trisomic nucleus with two signals for chromosome 1 and chromosome 16 and three signals for chromosome 9; (g) triploid nucleus with three signals for chromosome 16 and chromosome 18; (h) tetraploid nucleus with two signals for chromosome X and chromosome Y; (i) tetraploid nucleus with two signals for chromosome X and chromosome Y and four signals for chromosome 1. (Copyright © Vorsanova et al. 2010a; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0)
I-FISH with site-specific probes: Site-specific DNA probes (YACs, BACs, PACs, cosmids) are used either to map chromosomal regions within which a breakpoint is located or to evaluate chromosomal imbalances by a targeted FISH assay (diagnosis of known microdeletion and microduplication syndromes) (Iourov et al. 2008a; Liehr 2009), aneuploidy and/or recurrent chromosome abnormalities during preimplantation genetic diagnosis (Fung et al. 2001; Stumm et al. 2006; Lu et al. 2009), prenatal diagnosis (Soloviev et al. 1995; Vorsanova et al. 2005b; Liehr 2009), oncocytogenetic analysis (Liehr and Claussen 2002; Mitelman et al. 2007), and copy number variation precision (Carter 2007). Probing small genomic loci (<1 Mb), site-specific probes are applied to studying gene-specific nuclear organization and its relevance to genome behavior (Goetze et al. 2007; Strickfaden et al. 2010). However, relatively moderate hybridization efficiency (<70 %) hinders using the approaches in numerous areas of biomedical research and diagnosis. Alternatively, a number of FISH procedures with these types of probes (i.e., hematological and tumor diagnosis) are found effective for molecular cytogenetic diagnosis and have cutoffs varying between 90 and 95 % (Liehr 2009). As a result, interphase molecular cytogenetic studies by I-FISH with site-specific probes are commonly applied in preimplantation, prenatal, and postnatal diagnosis as well as in cancer cytogenetics (Fig. 11.2). Although repeatedly noted to be of significant importance for detecting gene fusions resulting from interchromosomal translocations (cancer biomarkers) (Mitelman et al. 2007) and to be useful for preimplantation diagnosis (Stumm et al. 2006), such I-FISH modifications have considerable disadvantages. To be more precise, the hybridization efficiency of site-specific probes is usually between 40 % and 70 %. This irregularity of hybridization efficiency can produce false-positive or false-negative data. Moreover, one has to use probes hybridizing to well-characterized chromosomal/genomic DNA loci (i.e., oncogenes or genes/genomic loci within deletion or duplication regions). Few well-characterized approaches using these DNA probes may be of importance for detecting continuously reported chromosomal rearrangements in cancer cells (Virgili et al. 2008; Nicholson and Duesberg 2009; Sen and Hopwood 2010) and deletions/duplications in a clinical population (Halder et al. 2008; Weise et al. 2012). Nonetheless, application of site-specific probes is the best way to visualize interphase chromosomal DNAs less than 1 Mb. Simultaneous hybridization of centromeric and site-specific probes (mFISH) (Fig. 11.3) is applicable for diagnostics and survey of somatic genome variations.
I-FISH with site-specific DNA probes: (a) normal diploid nucleus with two signals for chromosome 21; (b) trisomic nucleus with three signals for chromosome 21; (c) interphase nucleus exhibiting colocalization of ABL and BCR genes, probably caused by t(9;22)/Philadelphia chromosome. (Copyright © Vorsanova et al. 2010a; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0)
Five-color I-FISH (mFISH) with DNA probes for chromosomes 18, X, and Y (centromeric probes) as well as 13 and 21 (site-specific probes). A presumably normal (diploid) male nucleus isolated from the adult human brain. (Copyright © Vorsanova et al. 2010a; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0)
FISH with wcp probe and MFISH/SKY: Wcp probe combinations allow performing 24-color MFISH or SKY for metaphase analysis of interchromosomal chromosome rearrangements in cancers and individuals with constitutional chromosomal abnormities (Liehr and Claussen 2002; Liehr et al. 2004). In interphase chromosomal analysis, MFISH and SKY are hardly applicable. Occasional studies applied MFISH-based approaches for visualizing all chromosomes in fibroblast interphase nuclei and prometaphase rosettes (Walter et al. 2006). Similar assays with 2–5 wcp probes are frequently encountered in molecular cytogenetic diagnosis of structural alterations to metaphase chromosomes, as well (Liehr et al. 2004), and in investigation of genome organization in interphase nuclei (Rouquette et al. 2010). Nevertheless, I-FISH with wcp probes is sophisticated and too poorly reproducible to become competitive with other interphase molecular cytogenetic techniques. It is therefore unsurprising that FISH chromosomal painting by wcp is generally recognized as completely useless for identification of interphase chromosome numbers and structure (Fig. 11.4). Alternatively, basic research of chromosome architecture in interphase is usually performed using I-FISH with wcp providing for visualization of chromosome territories and their positioning relative to nuclear structures. Additionally, I-FISH-wcp approaches were almost the only way to study genomic organization in interphase until more effective techniques have been elaborated (Walter et al. 2006; Rouquette et al. 2010). Finally, these techniques are all limited in their abilities to paint chromosome territories (volumes) only (Table 11.2).
I-FISH with two-whole chromosome painting (wcp) for chromosomes 7 and 21. (a) Ambiguous chromosome territories provide information neither about number of chromosomes nor about structure of chromosomes (chromosome 7, green signal; chromosome 21, red signals), whereas this individual presented with regular unbalanced t(7;21). Details of this case are described in Vorsanova et al. (2008). (b) Chromosome territories in an interphase nucleus of a cell isolated from the ataxia-telangiectasia brain (chromosome 7, green signals; chromosome 14, red signal). Note the impossibility to identify number of chromosomes 14. (Copyright © Vorsanova et al. 2010a; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0)
Interphase chromosome-specific MCB: By microdissection of chromosomal loci for obtaining a set of probes that produce multicolor pseudo-G-banding, a high-resolution molecular cytogenetic technique for analysis of metaphase chromosomes termed MCB (multicolor banding) was proposed (Liehr et al. 2002). Further, this idea has been adapted to interphase chromosomal analysis and has provided for elaboration of interphase chromosome-specific MCB (ICS-MCB). To visualize a homologous pair of interphase chromosomes in their integrity, one has to generate MCB (Iourov et al. 2007, 2009a, b; Manvelyan et al. 2008; Iourov 2012). Figure 11.5 gives an example of aneuploidy detection in an interphase nucleus isolated from the Alzheimer’s disease brain (Iourov et al. 2009a). ICS-MCB can be widely applied for basic research of somatic genomic variations, chromosome structural and functional organization in interphase, and supramolecular disease mechanisms. Apparently, the sole disadvantage of this technique is the impossibility to analyze more than one homologous chromosome pair at a time (Iourov et al. 2007; Iourov 2012). This state-of-the-art molecular cytogenetic technology is discussed in detail in Chap. 9 of this book.
Interphase chromosome-specific multicolor banding (ICS-MCB) with chromosome 21-specific probe. Monosomy (loss) of chromosome 21 in a nucleus isolated from the Alzheimer’s disease brain. (Copyright © Vorsanova et al. 2010a; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0)
Fiber FISH: Probably the highest molecular cytogenetic resolution is achieved by DNA fiber FISH. This technique provides a mapping resolution of 1–3 Mb (metaphase analysis). Applied to interphase nuclei, it achieves a resolution of 50 kb or even more. The high resolution is attributed to a higher degree of chromatin decondensation than other FISH techniques. Applied to naked DNA fibers, fiber FISH show that chromatin fully decondenses, leading to a resolution ranging from 1 to 400 kb. Furthermore, DNA fiber FISH provides a mapping tool supplementary to restriction mapping permitting accurate gap and overlap sizing (Raap et al. 1996; Weier 2001). The latter, however, is currently out of the scope of human genome research, inasmuch as genomic loci are supposed to be all mapped in a definitive manner by the Human Genome Project.
Heng et al. (1992) were able to release the chromatin fibers from cells arrested at G1 and G2 by different chemicals and alkaline lysis procedure. They have also demonstrated fluorescence-labeled probes to hybridize specifically to single-copy genomic DNA sequences of the free chromatin. FISH signals have been detected for sequences separated by 21 kb (the closest position) and 350 kb (the far position), with exact correspondence between the observed and expected distances. The resolution of this technique is likely to approach 10 kb, and the coverage should span millions of base pairs. According to these data, authors have concluded that free chromatin mapping can be generally used to study the structure and organization of mammalian interphase genomes.
To improve the DNA resolution of FISH, Wiegant and his collaborators have adapted a nuclear extraction technique, resulting in highly extended DNA loops arranged around the nuclear matrix in a halo-like structure (Wiegant et al. 1992). In situ hybridization signals depicting alphoid and cosmid DNAs appear as beads-on-a-string, which, according to preliminary experiments, results from the association of individual probe fragments. By multicolor hybridization the authors were able to determine relative map positions and to detect easily a 10-kb overlap between individual cosmid clones, each of which shows linear beaded signals, suggesting that the DNA was essentially linearized in these experiments. The resolution range was defined as 10–200 kb, and probably as little as a few kilobases, thus greatly extending the abilities of fiber FISH. Fiber FISH was also found useful for investigation of genomic organization and mapping, stalled transcription, and genomic rearrangements (including large deletions within gene sequences) (Weier 2001). Although this technique is based on obtaining DNA fibers from interphase nuclei, it cannot be directly attributed to I-FISH. Single-cell fiber FISH (especially when large cell populations are analyzed) is highly complicated.
Immuno-FISH: Immuno-FISH combines immunohistochemical detection of proteins with FISH to specific DNA (RNA) targets (Dundas et al. 2001; Yang et al. 2004). A simple protocol of immuno-FISH using cytospin centrifuge and fixation without acetic acid in 80 % methanol is effective for detecting colocalization of entromeric alpha-satellite DNA sequences with the kinetochore CENP-B proteins (Marcais et al. 1999). Such FISH analyses of chromosome 21-specific alphoid DNA and immunostaining of kinetochores on extended interphase chromatin fibers and interphase nuclei indicated that centromeric kinetochore-specific proteins bind to restricted areas of centromeric DNA arrays. In general application, this approach allows prevention of protein and DNA loss during processing cell suspensions for cytogenetic and immunochemical evaluation. Immuno-FISH is found to be applicable in cancer research/diagnosis (immunophenotyping during single-cell genetic analysis), studies of chromosome structure and organization, transplantation research, and identification of supramolecular disease mechanisms (Meaburn et al. 2009; Strickfaden et al. 2010). Figure 11.6 shows immuno-FISH on interphase neuronal cells of the adult human brain (more details in Iourov et al. 2009b,c).
Immuno-FISH (I-FISH) using centromeric probe for chromosome Y (DYZ3) with immunostaining by NeuN (neuron-specific antibody) performed for the analysis of cells isolated from the human brain. (Copyright © Vorsanova et al. 2010a; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0)
Fast-FISH with microwave activation for I-FISH: Usually FISH using chromosome-specific or site-specific DNA probes is performed during 1 or 2 days. Several fast-FISH protocols were developed using microwave activation for rapid hybridization and detection. Microwave activation for FISH has been proposed by Dr. Ilia Soloviev in 1994. In contrast to standard FISH protocols, this method offers an opportunity to detect hybridization signals within a few minutes, granting 10- to 15-fold detection time reduction. No signal amplification is used to minimize the overlapping and nonspecific background of hybridization signals during chromosomal analysis of interphase nuclei. Microwave activation makes FISH applicable to cells containing cytoplasm (Fig. 11.7). This technique was highly reproducible and applicable for many different chromosome-specific DNA probes. For instance, we have tested alpha-satellite DNA probes specific for chromosomes 1, 3–19, 21, 22, X, and Y. The procedure has allowed rapid chromosome detection in interphase nuclei and metaphase plates of peripheral blood and amniotic fluid (cell suspensions were older than 2 years). Chromosome-specific repetitive DNA probes with FISH microwave activation are to be used for rapid diagnosis of common chromosomal syndromes including chromosome aneuploidies, fast sex determination in prenatal screening, and routine chromosome identification (Soloviev et al. 1994). For more specific purposes, it seems that laboratory microwave ovens are required. However, a common commercially available microwave oven is a handy alternative to a thermal cycler for fast-FISH. Comparable results have been obtained for chromosome 1-/X-specific satellite DNA probes. In addition, the complete fast-FISH procedure was accelerated. An optimized condition for a commercially available X-specific alpha-satellite probe by fast-FISH technique has been also developed for quantitative microscopy (Durm et al. 1997). For highly repetitive DNA probes, the hybridization (renaturation) time and the number of subsequent washing steps can be reduced considerably by omitting denaturing chemical agents (formamide). The appropriate hybridization temperature and time allow a clear discrimination between major and minor binding sites by quantitative fluorescence microscopy. The well-defined physical conditions for hybridization permit automatization of the procedure (Iourov et al. 2008c). Highly fluorescent major binding sites are obtained when denaturation is performed at 74 °C and hybridization is performed during 60 min. These conditions have shown the best microwave activation for denaturation and hybridization to accelerate the procedure. It is to be noted that slides with the target material and the hybridization buffer are placed in a standard microwave oven. After denaturation for 20 s at 900 W, hybridization is performed for 4 min at 90 W. The suitability of a microwave oven for fast-FISH was confirmed with a chromosome 1-specific alpha-satellite probe. In this series of tests, denaturation was performed at 630 W for 60 s and hybridization at 90 W for 5 min. The results were analyzed quantitatively and compared to the results obtained by fast-FISH. The major binding sites were clearly discriminated by their brightness (Durm et al. 1997).
Another method for FISH signals enhancing by microwave pulses during DNA–DNA hybridization using a single- or low-copy probe has shown application of microwaves to be effective in diagnostic or research practice because of the enhancement of weak signals. Microwave FISH has been compared systematically with simple FISH protocols, and it was possible to demonstrate that microwave irradiation leads to better FISH results, especially during the first 100 min of hybridization (Weise et al. 2005).
General advantages and limitations: All FISH-based methods require (1) obtaining cells suspensions or performing other biopsy preparations for the analysis, (2) denaturation and hybridization, and (3) microscopic visual/digital analysis of hybridization results (Iourov et al. 2006b; Iourov 2009). The first stage does not cause any complication, when I-FISH is used, because any cell type of a human organism can be processed for such analyses (Iourov et al. 2006b; Vorsanova et al. 2010a). This is the essential advantage of interphase molecular cytogenetics in contrast to classical cytogenetics (metaphase analysis), that is, the ability to analyze chromosomes in all cell types. Regardless of I-FISH limitations (Liehr and Claussen 2002), some modifications such as ICS-MCB allow a view of interphase chromosomes in their integrity. As mentioned earlier and in Chap. 9, ICS-MCB still has some limitations, being, however, the unique way to visualize the whole banded chromosome in a nucleus (Iourov et al. 2007). I-FISH denaturation and hybridization are performed identically to classical FISH-based approaches (Liehr 2009), and no additional drawbacks can be attributed to these procedures. I-FISH microscopic or digital analysis is not associated with any special problem (Iourov et al. 2008c, 2009c). There are also possibilities to apply digital analysis for studying interphase chromosomes: QFISH analysis of signal colocalization (gene fusions: chromosomal translocations in interphase nuclei), ICS-MCB (visualization of chromosomal structures), increasing “signal visibility,” and automatic signal detection. Digital analysis is required for multicolor FISH-based assays (SKY, MFISH, multiprobe interphase FISH, or mFISH), which are usually applied to increase the potential of FISH-based assays through simultaneous analysis of multiple targets (Iourov et al. 2009a, b; Lu et al. 2009). A combination of FISH techniques (i.e., mFISH with 2–5 probes, QFISH, and ICS-MCB) has become the basis for an integrated approach toward molecular diagnosis and genome (chromosome) research at supramolecular level in interphase. Usually, the way that FISH results are evaluated (i.e., visual or digital) is determined by features of DNA probes (amount of probes per reaction and DNA sequence affinity) and detection. Consequently, it is better to subdivide I-FISH techniques this way (Table 11.2).
Several general problems of I-FISH application do exist. Differences of hybridization efficiency complicate simultaneous applications of different probe sets (Iourov et al. 2006a). Site-specific probes signals can be overlooked when wcp or centromeric probes are used (because of intensity differences). Probably the simplest solution is the ICS-MCB. However, some interphase FISH protocols with established probe combinations are proven to be effective for diagnostic purposes (Gersen and Keagle 2005; Liehr 2009). DNA replication during the S-phase of the cell cycle is another major source of unusual I-FISH signal appearance. There are recommendations concerning this type of I-FISH artifacts in the available literature, but the analysis can still be hindered by replicative signals. The latter mainly concerns site-specific probes, being, however, observed during I-FISH with centromeric probes, as well (Fig. 11.8). An additional source of artifacts that can be misinterpreted (i.e., considered as false-positive chromosome abnormalities) is the specificity of nuclear genome organization or interphase chromosome architecture. Here, the problem is related to chromosomal associations (Leitch 2000; Iourov et al. 2005; Krueger and Osborne 2006), significantly affecting I-FISH results and becoming even more important when taking into account that numerous cell types are prone to exhibit chromosomal associations/pairing (Fig. 11.8). Such problems are easily managed by QFISH (Iourov et al. 2005) (Fig. 11.8).
Problems of I-FISH with centromeric/site-specific DNA probes. (a, b) Replication of specific genomic loci (LSI21 probe): some nuclei exhibit replicated signals, whereas in some nuclei it is not apparent; note the distance between signals can be more than a diameter of a signal. (c) Asynchronous replication of a signal (DXZ1) in case of tetrasomy of chromosome X; note difficulty to make a definitive conclusion about number of signals in the right nucleus. (d) Two-color FISH with centromeric/site-specific DNA probes for chromosome 1 shows chromosomal associations in a nucleus isolated from the adult human brain; note impossibility to identify number of chromosomes. (e) Quantitive FISH (QFISH) demonstrates an association of centromeric regions of homologous chromosomes 9, but not a monosomy or chromosome loss. (Copyright © Vorsanova et al. 2010a; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0)
I-FISH for Analysis of DNA Replication
I-FISH allows the visualization of replicating genomic DNA sequences in interphase nuclei. FISH has been shown to help discriminate between nonreplicated and replicated regions of the genome in interphase nuclei, based on the number of specific fluorescent signals (Selig et al. 1992). In normal diploid cells, FISH results on nonreplicated DNA are seen as a single signal whereas replicated loci are characterized by doublets (doubling of a signal). The distribution of these two patterns in unsynchronized cell populations can be used to determine the replication time (S phase) of a DNA sequence. The availability of well-mapped genomic probes and the possibility to compare results from different cell lines make this a convenient approach, by which domains of replication timing control mapped at any chromosomal position can be addressed and the relationship to various gene expression patterns can be deduced. Because there appear to be important but poorly understood correlations among replication timing, chromatin structure, and transcriptional competence in mammalian cells, this technique seems to be valuable for understanding related molecular interrelationships.
I-FISH studies have established that monoallelically expressed genes display the unusual property of asynchronous replication, and in those genes that exhibit transcription randomly monoallelic, the asynchronous replication is also random (Ensminger and Chess 2004). By examining the replication timing of genes in a number of human trisomies, authors consistently find one allele replicating early and the other two alleles replicating late, similar to previous observations in X-chromosome trisomies.
I-FISH with chromosome 21-specific cosmid probes was also previously used to identify trisomy 21 in cultured and uncultured amniotic cells. Proper identification of chromosome 21 numbers was made in 65–75 % of trisomic cells and in 70–75 % of normal disomic cells by using all the tested probes. The efficiency of FISH analysis for the total population of interphase cells and cells in the postreplication periods (late S, G2) of the cell cycle was assessed (Fig. 11.9). Selective scoring of cells in the postreplicative period (a pair of FISH signals on replicated interphase chromosomes) increased the amount of informative nuclei by as much as 95–97 %. The approach was found to determine overlapping chromosomes, artificial doubling of FISH signals on each chromatid of interphase chromosomes, background, and polyploidy. Cosmid probes and integral analysis of hybridization-positive nuclei in pre- and postreplication periods may, therefore, be applicable for improving prenatal diagnosis of trisomy 21. Interestingly, that I-FISH showed that additional chromosomes 21 can induce changing in the replication pattern of an allelic pair: from a synchronous pattern mimicking concomitantly expressed alleles to unsynchronized ones appearing as signals displaying an allele-specific mode of expression (Amiel et al. 1998). A similar phenomenon of asynchronous replication of alleles in genomes carrying an extra chromosome was found in autosomal aneuploidy (trisomy of chromosomes 18 and 13) and sex chromosome aneuploidy (47,XXX and 47,XXY) (Amiel et al. 1999). These data suggest that gross phenotypic abnormalities associated with chromosomal aneuploidy result not only from overexpression of extra gene copies (increased gene dosage) but also from altered expression of genes located on the remaining two homologous chromosomes.
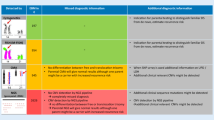
FISH hybridization with the cosmid probe (21q22.2) on cultured amniotic fluid trisomic 21 cells. (a) Cell with asynchronously replicating loci (unreplicated locus with singlet signal and replicated locus with doublet signal). (b) Replicated cell shows two closely paired hybridization signals corresponding to each chromatid of chromosome 21. (c) Cell with trisomy 21 in the postreplicative stage of the cell cycle. (From Soloviev et al. 1995. Reproduced with permission of John Wiley &Sons, Ltd., in the format reuse in a book/textbook vis Copyright Clearance Center). (d) Examples of FISH on interphase nuclei with chromosome X-specific centromeric and region-specific probes (locus Xq28) show different types of signals (SD and SD) in a girl with Rett syndrome (RTT). Cy3-labeled centromeric alphoid DNA probe was used. Two single red signals indicate simultaneously replicating centromeric DNA from both X chromosomes. PAC clone 671D9 (MeCP2 gene) was labeled by biotin and detected with FITC-avidin. Two asynchronously replicating loci could be seen: one single green signal represents late-replicating X chromosome and one double green signal represents early-replicating X chromosome. Interphase nuclei were counterstained with fluorescent dye Hoechst 33258 (blue color). (From Vorsanova et al. 2001a Brain &development by Nihon Shoni Shinkeigaku. Reproduced with permission of Elsevier BV in the format reuse in a book/textbook via Copyright Clearance Center)
A study of replication timing by I-FISH using chromosome X-specific DNA probes was used to determine the loci with altered replication and transcription in Rett syndrome (RTT), a epigenetic disease caused by mutations in MECP2. It was detected that a feature of RTT patients is the MECP2 locus escaping inactivation in late-replicating chromosome X (Fig. 11.9). Therefore, region Xq28 could contain genes, including MECP2, escaping X-inactivation and featured by biallelic expression from the active as from inactive chromosomes X (Vorsanova et al. 2001a). These results support the hypothesis proposing the disturbances in dosage compensation effect caused by aberrant activation of the inactive X-chromosome genes in RTT (biallelic expression in contrast to monoallelic) (Vorsanova et al. 2001a, b) and indicate that normal MЕCP2 allele can escape X-inactivation and, in contrast, reduce the pathogenic effect of a mutated allele in RTT.
In the light of the tight relationship between replication timing and expression of a given DNA sequence, the replication timing of FMR1 alleles on active and inactive X chromosomes was analyzed by I-FISH (Yeshaya et al. 1999). The authors concluded that the FMR1 locus is subjected to X-inactivation and the delaying effect of the trinucleotide expansion (causing fragile X syndrome) is superimposed on the delay in replication associated with X-inactivation. Thus, a significant epigenetic marker of the interphase chromosome replicative activity is asynchronous replication of monoallelically expressed genes and the synchronous replication of biallelically expressed genes.
Testing a similar hypothesis in microdeletion syndromes (i.e., a microdeletion can affect epigenetic profiling of genes located outside the missing segment), Yeshaya et al. (2009) analyzed the replication patterns of two genes: SNRPN, a normally monoallelically expressed gene (assigned to 15q11.13) and RB1, a biallelically expressed gene (assigned to 13.q14) in the genomes of patients carrying the 22q11.2 deletion (DiGeorge/velocardiofacial syndrome) and those carrying the 7q11.23 deletion (Williams syndrome). In each affected individual, an aberrant and reversed pattern of replication was shown. In other words, a monoallelic gene replicated more synchronously than a biallelic gene. This inverted pattern, which appears to be nonspecific for those deletions, clearly distinguishes cells of deletion carriers from unaffected individuals. As a result, a potential epigenetic marker for suspecting a hidden microdeletion that is too small to be detected by conventional karyotyping methods was proposed (Fig. 11.10).
FISH signals in PHA-stimulated lymphocytes at interphase, following FISH with RB1. Cells with two singlets (SS cells) in which neither allele has replicated (a–c); cells with two doublets (DD cells) in which both alleles have replicated (d–f); and cells with one singlet and one doublet (SD cells) (g–i), which are S-phase cells in which one allele has replicated while its partner has not. (Copyright © Yeshaya et al. 2009; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0)
Litmanovitch et al. (1998) have used I-FISH for studying replication patterns of alpha-satellite DNA sequences in the light of the human centromere structure and function. They showed an association between replication timing of alpha-satellite DNA sequences and centromere function. Chromosomes having homologous alpha-satellite loci, which replicated synchronously, were revealed to be associated with a lower occurrence of chromosome-specific aneuploidy, whereas chromosomes exhibiting asynchrony with long intervals between early- and late-replicating loci showed the highest occurrence of chromosome-specific aneuploidy. The latter supports the hypothesis suggesting that loss of replication control within loc. composed of human centromeric DNAs affects essential centromere functions, such as ensuring proper sister chromatid separation and proper chromosomal segregation during cell division.
Chromosome Architecture and Behavior in Interphase
Chromosome architecture in interphase is consistently shown to be a driving force for crucial intranuclear processes. Specific arrangement of interphase chromosomes is likely to play a role in the regulation of genome activity and cell division as well as formation of chromosome rearrangements occurring during meiosis and mitosis (Leitch 2000; Iourov 2012). To analyze genome organization in interphase, numerous approaches are to be applied, among which I-FISH appears to play the leading role. Several applications of I-FISH approaches for interphase chromosome analysis can be proposed: (1) identification of chromosome positioning and its relation to other nuclear compartments/structures (I-FISH with wcp, interphase MFISH, or ICS-MCB); (2) analyzing positioning of specific genomic loci in relation to each other (associations of whole chromosomes or chromosomal loci) and their behavior (transcriptional/replicative activity) to get a view of functional nuclear genome/chromosome organization and its driving forces (I-FISH with centromeric, site-specific, and wcp, mFISH/QIFSH, or ICS-MCB); and (3) assessment of chromosome arrangement or behavior and its relationship to genome, epigenome, and proteome profiling for delineation of possible consequences of specific interphase chromosome architecture (somatic chromosomal mutations) (I-FISH with centromeric, site-specific, and wcp, mFISH/QIFSH, ICS-MCB, and immuno-FISH). I-FISH analysis of spatial chromosome organization is also influenced by specificity of methods used for structural preservation of nuclei. There are some reports about dependence of fixation on I-FISH results, whereas other studies have not provided similar data. Suspension FISH (S-FISH) is likely to be an alternative for I-FISH spatial genome analysis using standard fixation protocols and is able to leave aside related problems (Steinhaeuser et al. 2002). This technique is discussed in detail in Chap. 10 of this book. In brief, advantages of this approach are referred to the possibility of studying three-dimensionally (3D) preserved nuclei from any human tissue, whereas other 3D preservation techniques require specific conditions of cell cultivation. The latter makes I-FISH lose its main advantage—the opportunity to analyze nondividing cells.
Molecular Cytogenetic Diagnosis
Molecular cytogenetic identification of chromosomal aberrations by I-FISH has been already mentioned in this chapter as well as in a number of comprehensive reviews (Leitch 2010; Iourov et al. 2006a, 2009c; Yurov et al. 2009; Sen and Hopwood 2010; Vorsanova et al. 2010a, b). However, some additional comments about more specific problems of medical cytogenetics seem to be required. Because studying chromosomes in interphase nuclei has undoubtedly profound effects on molecular cancer and prenatal diagnosis, it is obvious that it is impossible to refer all the I-FISH diagnostic studies. To list some technical solutions in molecular cytogenetic diagnosis by I-FISH, we have preferred to focus on difficulties encountered during the introduction and usage for diagnostic purposes. Newly introduced interphase techniques are primarily used for research purposes and are rarely tested for diagnostic validity. Limiting practical application of such I-FISH protocols requires reevaluating the drawbacks. However, the majority of these can be eliminated by application of additional FISH-based approaches (i.e., QFISH). Another problem comes from the diagnosis of somatic chromosomal mosaicism. Regardless of some attempts to propose guidelines or criteria for mosaicism definition (for details, see Iourov et al. 2009c; Vorsanova et al. 2010b), additional studies of somatic mosaicism seem to be strongly required. For instance, a large-scale study aimed to uncover somatic genomic variations in several unaffected human tissues might lead the way. Finally, it is still poorly understood whether data obtained through interphase analysis can be more valid than those obtained by metaphase analysis. From the “structural point of view” (analyzing structural chromosome imbalances), metaphase chromosomal analysis is likely to be more precise. On the other hand, mosaics require large cell populations to be analyzed, and this problem is even more notable when cases of complex, hidden (cryptic), or dynamic mosaicism are evaluated. Metaphase analysis in these cases can be applied for thorough definition of all cell lines, because simple I-FISH analyses (apart from ICS-MCB) are hardly able to show precisely the structure of rearranged chromosomes in a given cell line. More sophisticated studies can require additional data to obtain. For instance, parental origin of chromosomes or epigenetic features addressed by either QFISH (Iourov et al. 2005) or pod-FISH (Weise et al. 2010) could be useful for more thorough confirmational or exclusive diagnosis.
Molecular cytogenetic diagnosis should be performed using a panel of FISH-based techniques (Liehr 2009; Bridger and Volpi 2010; Vorsanova et al. 2010b). To achieve the highest resolution, one can combine molecular cytogenetic techniques based on different platforms (array CGH with I-FISH; metaphase FISH-based techniques with I-FISH, etc.). Cases of complex mosaics or balanced structural chromosome abnormalities seem to especially require such a complex diagnostic procedure. Consequently, regardless of significant developments in molecular interphase cytogenetics, I-FISH techniques remain an addition to whole-genome screening approaches based on array CGH (array CHG) and/or metaphase cytogenetic analysis used for the diagnosis. Only a few targeted I-FISH assays for identification of known caner-associated translocations in interphase and preimplantation genetic diagnosis seem to be applicable in routine molecular cytogenetic diagnosis. To this end, the diagnostic potential of I-FISH is to be more thoroughly analyzed for becoming a routine testing procedure in molecular diagnosis.
Conclusion
According to the present overview of molecular cytogenetic techniques for visualizing chromosomes in interphase, we conclude that a firm technological basis does exist for high-resolution analyses of chromosomes in almost all human tissues. I-FISH advanced by developments in interphase molecular cytogenetics is almost the unique technological issue for studying functional consequences of spatiotemporal chromosome arrangement (architecture) in the interphase nuclei, elucidating the role of such immense intercellular genomic diversity or somatic genomic variations (somatic mosaicism), and proposing new diagnostic solutions for medical genetics, reproductive medicine, and oncology. I-FISH provides for assessment of genome variations and behavior (including DNA replication) in all the cell types of the human organism (all stages of the cell cycle) at molecular resolutions. The combinations of interphase molecular cytogenetic techniques (i.e., mFISH, QFISH, ICS-MCB, S-FISH, pod-FISH, immuno-FISH, etc.) have already given rise to several biomedical discoveries or even new biomedical directions (i.e., molecular neurocytogenetics; for details, see Chap. 3). Therefore, one can insist that developments in interphase molecular cytogenetics are promising for basic and diagnostic research in genetics, cellular and molecular biology, and molecular (genome) medicine. In summary, describing the technological solutions for studying human interphase chromosomes allows us to conclude that interphase molecular cytogenetics opens new opportunities for genetics and cell biology.
References
Amiel A, Avivi L, Gaber E, Fejgin MD (1998) Asynchronous replication of allelic loci in Down syndrome. Eur J Hum Genet 6(4):359–364
Amiel A, Korenstein A, Gaber E, Avivi L (1999) Asynchronous replication of alleles in genomes carrying an extra autosome. Eur J Hum Genet 7(2):223–230
Aubert G, Hills M, Lansdorp PM (2012) Telomere length measurement-caveats and a critical assessment of the available technologies and tools. Mutat Res 730(1–2):59–67
Baumgartner A, Weier JF, Weier H-UG (2006) Chromosome-specific DNA repeat probes. J Histochem Cytochem 54:1363–1370
Bejjani BA, Shaffer LG (2008) Clinical utility of contemporary molecular cytogenetics. Annu Rev Genomics Hum Genet 9:71–86
Bridger JM, Volpi EV (eds) (2010) Fluorescence in situ hybridization (FISH): protocols and applications. Humana Press, New York
Carter NP (2007) Methods and strategies for analyzing copy number variation using DNA microarrays. Nat Genet 39:S16–S21
Cremer T, Landegent J, Brückner A, Scholl HP, Schardin M, Hager HD et al (1986) Detection of chromosome aberrations in the human interphase nucleus by visualization of specific target DNAs with radioactive and non-radioactive in situ hybridization techniques: diagnosis of trisomy 18 with probe L1.84. Hum Genet 74(4):346–352
Dundas SR, Boyle S, Bellamy CO, Hawkins W, Garden OJ, Ross JA, Bickmore W (2001) Dual Y-chromosome painting and immunofluorescence staining of archival human liver transplant biopsies. J Histochem Cytochem 49:1321–1322
Durm M, Haar F-M, Hausmann M, Ludwig H, Cremer C (1997) Optimized Fast-FISH with α-satellite probes: acceleration by microwave activation. Braz J Med Biol Res 30(1):15–22
Ensminger AW, Chess A (2004) Coordinated replication timing of monoallelically expressed genes along human autosomes. Hum Mol Genet 13:651–658
Fung J, Weier H-UG, Pedersen RA (2001) Detection of structural and numerical chromosome abnormalities in interphase cells using spectral imaging. J Histochem Cytochem 49:797–798
Gersen SL, Keagle MB (2005) The principles of clinical cytogenetics, 2nd edn. Humana Press, Totowa, NJ
Goetze S, Mateos-Langerak J, van Driel R (2007) Three-dimensional genome organization in interphase and its relation to genome function. Semin Cell Dev Biol 18(5):707–714
Halder A, Jain M, Kabra M, Gupta N (2008) Mosaic 22q11.2 microdeletion syndrome: diagnosis and clinical manifestations of two cases. Mol Cytogenet 1:18
Heng HH, Squire J, Tsui LC (1992) High-resolution mapping of mammalian genes by in situ hybridization to free chromatin. Proc Natl Acad Sci U S A 89(20):9509–9513
Iourov IY (2009) Microscopy and imaging systems. In: Liehr T (ed) Fluorescence in situ hybridization (FISH)—application guide. Springer Verlag, Berlin, pp 75–84
Iourov IY (2012) To see an interphase chromosome or: how a disease can be associated with specific nuclear genome organization. BioDiscovery 4:5. doi:10.7750/BioDiscovery.2012.4.5
Iourov IY, Soloviev IV, Vorsanova SG, Monakhov VV, Yurov YB (2005) An approach for quantitative assessment of fluorescence in situ hybridization (FISH) signals for applied human molecular cytogenetics. J Histochem Cytochem 53:401–408
Iourov IY, Vorsanova SG, Yurov YB (2006a) Chromosomal variations in mammalian neuronal cells: known facts and attractive hypotheses. Int Rev Cytol 249:143–191
Iourov IY, Vorsanova SG, Pellestor F, Yurov YB (2006b) Brain tissue preparations for chromosomal PRINS labeling. Methods Mol Biol 334:123–132
Iourov IY, Vorsanova SG, Yurov YB (2006c) Intercellular genomic (chromosomal) variations resulting in somatic mosaicism: mechanisms and consequences. Curr Genomics 7:435–446
Iourov IY, Liehr T, Vorsanova SG, Yurov YB (2007) Interphase chromosome-specific multicolor banding (ICS-MCB): a new tool for analysis of interphase chromosomes in their integrity. Biomol Eng 24:415–417
Iourov IY, Vorsanova SG, Yurov YB (2008a) Recent patents on molecular cytogenetics. Recent Pat DNA Gene Seq 2:6–15
Iourov IY, Vorsanova SG, Yurov YB (2008b) Molecular cytogenetics and cytogenomics of brain diseases. Curr Genomics 9:452–465
Iourov IY, Vorsanova SG, Yurov YB (2008c) Fluorescence intensity profiles of in situ hybridization signals depict genome architecture within human interphase nuclei. Tsitol Genet 42(5):3–8
Iourov IY, Vorsanova SG, Liehr T, Yurov YB (2009a) Aneuploidy in the normal, Alzheimer’s disease and ataxia-telangiectasia brain: differential expression and pathological meaning. Neurobiol Dis 34:212–220
Iourov IY, Vorsanova SG, Liehr T, Kolotii AD, Yurov YB (2009b) Increased chromosome instability dramatically disrupts neural genome integrity and mediates cerebellar degeneration in the ataxia-telangiectasia brain. Hum Mol Genet 18:2656–2669
Iourov IY, Vorsanova SG, Soloviev IV, Yurov YB (2009c) Interphase FISH: detection of intercellular genomic variations and somatic chromosomal mosaicism. In: Liehr T (ed) Fluorescence in situ hybridization (FISH)—application guide. Springer, Berlin, pp 301–311
Iourov IY, Vorsanova SG, Yurov YB (2010) Somatic genome variations in health and disease. Curr Genomics 11(6):387–396
Iourov IY, Vorsanova SG, Yurov YB (2012) Single cell genomics of the brain: focus on neuronal diversity and neuropsychiatric diseases. Curr Genomics 13(6):477–488
Klewes L, Höbsch C, Katzir N, Rourke D, Garini Y, Mai S (2011) Novel automated three-dimensional genome scanning based on the nuclear architecture of telomeres. Cytometry A 79(2):159–166
Krueger C, Osborne CS (2006) Raising the curtains on interchromosomal interactions. Trends Genet 22:637–639
Lee C, Wevrick R, Fisher RB, Ferguson-Smith MA, Lin CC (1997) Human centromeric DNAs. Hum Genet 100:291–304
Leitch AR (2000) Higher levels of organization in the interphase nucleus of cycling and differentiated cells. Microbiol Mol Biol Rev 64(1):138–152
Levsky JM, Singer RH (2003) Fluorescence in situ hybridization: past, present and future. J Cell Sci 116(pt 14):2833–2838
Liehr T (2009) Fluorescence in situ hybridization (FISH)—application guide. Springer, Berlin
Liehr T, Claussen U (2002) Multicolor-FISH approaches for the characterization of human chromosomes in clinical genetics and tumor cytogenetics. Curr Genomics 3:231–235
Liehr T, Pfeiffer RA, Trautmann U, Gebhart E (1998) Centromeric alphoid DNA heteromorphisms of chromosome 22 as revealed by FISH-technique. Clin Genet 53:231–232
Liehr T, Heller A, Starke H, Rubtsov N, Trifonov V, Mrasek K, Weise A, Kuechler A, Claussen U (2002) Microdissection based high resolution multicolor banding for all 24 human chromosomes. Int J Mol Med 9:335–339
Liehr T, Starke H, Weise A, Lehrer H, Claussen U (2004) Multicolor FISH probe sets and their applications. Histol Histopathol 19:229–237
Litmanovitch T, Altaras MM, Dotan A, Avivi L (1998) Asynchronous replication of homologous alpha-satellite DNA loci in man is associated with nondisjunction. Cytogenet Cell Genet 81(1):26–35
Lu CM, Kwan J, Baumgartner A, Weier JF, Wang M, Escudero T, Munne S, Zitzelsberger HF, Weier H-UG (2009) DNA probe pooling for rapid delineation of chromosomal breakpoints. J Histochem Cytochem 57:587–597
Manvelyan M, Hunstig F, Mrasek K, Bhatt S, Pellestor F, Weise A, Liehr T (2008) Position of chromosomes 18, 19, 21 and 22 in 3D-preserved interphase nuclei of human and gorilla and white hand gibbon. Mol Cytogenet 1:9
Marcais B, Vorsanova SG, Roizes G, Yurov YB (1999) Analysis of alphoid DNA variation and kinetochore size in human chromosome 21: evidence against pathological significance of alphoid satellite DNA diminutions. Tsitol Genet 33(1):25–31
Meaburn KJ, Gudla PR, Khan S, Lockett SJ, Misteli T (2009) Disease-specific gene repositioning in breast cancer. J Cell Biol 187:801–812
Mitelman F, Johanson B, Martens F (2007) The impact of translocations and gene fusions on cancer causation. Nat Rev Cancer 7:233–245
Nicholson JM, Duesberg P (2009) On the karyotypic origin and evolution of cancer cells. Cancer Genet Cytogenet 194:96–110
Nietzel A, Rocchi M, Starke H, Heller A, Fiedler W, Wlodarska I et al (2001) A new multicolor-FISH approach for the characterization of marker chromosomes: centromere-specific multicolor-FISH (cenM-FISH). Hum Genet 108:199–204
Raap A, Florijn RJ, Blonden LA, Wiegant J, Vaandrager J-W, Vrolijk H et al (1996) Fiber FISH as a DNA mapping tool. Methods 9(1):67–73
Rouquette J, Cremer C, Cremer T, Fakan S (2010) Functional nuclear architecture studied by microscopy: present and future. Int Rev Cell Mol Biol 282:1–90
Schrock E, du Manoir S, Veldman T, Schoell B, Weinberg J, Ferguson-Smith MA et al (1996) Multicolor spectral karyotyping of human chromosomes. Science 273:494–497
Selig S, Okumura K, Ward DC, Cedar H (1992) Delineation of DNA replication time zones by fluorescence in situ hybridization. EMBO J 11(3):1217–1225
Sen S, Hopwood V (2010) Molecular cytogenetic evidence for multistep tumorigenesis: implications for risk assessment and early detection. Cancer Biomark 9(1–6):113–132
Soloviev IV, Yuri B, Yurov YB, Vorsanova SG, Malet P (1994) Microwave activation of fluorescence in situ hybridization: a novel method for rapid chromosome detection and analysis. Focus 16(4):115–116
Soloviev IV, Yurov YB, Vorsanova SG, Fayet F, Roizes G, Malet P (1995) Prenatal diagnosis of trisomy 21 using interphase fluorescence in situ hybridization of postreplicated cells with site-specific cosmid and cosmid contig probes. Prenat Diagn 15:237–248
Soloviev IV, Yurov YB, Vorsanova SG, Malet P, Zerova TE, Buzhievskaya TI (1998a) Double color in situ hybridization of alpha-satellite chromosome 13, 21 specific cosmid clones for a rapid screening of their specificity. Tsitol Genet 32:60–64
Soloviev IV, Yurov YB, Vorsanova SG, Marcais B, Rogaev EI, Kapanadze BI et al (1998b) Fluorescent in situ hybridization analysis of α-satellite DNA in cosmid libraries specific for human chromosomes 13, 21 and 22. Russ J Genet 34:1247–1255
Speicher MR, Ballard GS, Ward DC (1996) Karyotyping human chromosomes by combinatorial multi-fluor FISH. Nat Genet 12:368–375
Steinhaeuser U, Starke H, Nietzel A, Lindenau J, Ullmann P, Claussen U et al (2002) Suspension (S)-FISH, a new technique for interphase nuclei. J Histochem Cytochem 50:1697–1698
Strickfaden H, Zunhammer A, van Koningsbruggen S, Köhler D, Cremer T (2010) 4D chromatin dynamics in cycling cells: Theodor Boveri’s hypotheses revisited. Nucleus 1(3):284–297
Stumm M, Wegner R-D, Bloechle M, Eckel H (2006) Interphase M-FISH applications using commercial probes in prenatal and PGD diagnostics. Cytogenet Genome Res 114:296–301
Vanneste E, Bittman L, Van der Aa N, Voet T, Vermeesch JR (2012) New array approaches to explore single cells genomes. Front Genet 3:44
Verma RS, Luke S (1992) Variation in alphoid DNA sequences escape detection of aneuploidy in interphase FISH technique. Genomics 14:113–116
Virgili A, Brazma D, Reid AG, Howard-Reeves J, Valgañón M, Chanalaris A et al (2008) FISH mapping of Philadelphia negative BCR/ABL1 positive CML. Mol Cytogenet 1:14
Vorsanova SG, Yurov YB, Alexandrov IA, Demidova IA, Mitkevich SP, Tirskaya AF (1986) 18p- syndrome: an unusual case and diagnosis by in situ hybridization with chromosome 18-specific alphoid DNA sequence. Hum Genet 72:185–187
Vorsanova SG, Yurov YB, Deryagin GV, Soloviev IV, Bytenskaya GA (1991) Diagnosis of aneuploidy by in situ hybridization: analysis of interphase nuclei. Bull Exp Biol Med 112:413–415
Vorsanova SG, Yurov YB, Kolotii AD, Soloviev IV (2001a) FISH analysis of replication and transcription of chromosome X loci: new approach for genetic analysis of Rett syndrome. Brain Dev 23:S191–S195
Vorsanova SG, Yurov YB, Ulas VY, Demidova IA, Kolotii AD, Gorbatchevskaia NL, Beresheva AK, Soloviev IV (2001b) Cytogenetic and molecular-cytogenetic studies of Rett syndrome (RTT): a retrospective analysis of a Russian cohort of RTT patients (the investigation of 57 girls and three boys). Brain Dev 23:S196–S201
Vorsanova SG, Yurov YB, Brusquant D, Carles E, Roizes G (2002) Two new cases of the christchurch (Ch1) chromosome 21: evidence for clinical consequences of de novo deletion 21p-. Tsitol Genet 36(1):46–49
Vorsanova SG, Iourov IY, Beresheva AK, Demidova IA, Monakhov VV, Kravets VS et al (2005a) Non-disjunction of chromosome 21, alphoid DNA variation, and sociogenetic features of Down syndrome. Tsitol Genet 39(6):30–36
Vorsanova SG, Kolotii AD, Iourov IY, Monakhov VV, Kirillova EA, Soloviev IV, Yurov YB (2005b) Evidence for high frequency of chromosomal mosaicism in spontaneous abortions revealed by interphase FISH analysis. J Histochem Cytochem 53:375–380
Vorsanova SG, Iourov IY, Voinova-Ulas VY, Weise A, Monakhov VV, Kolotii AD et al (2008) Partial monosomy 7q34-qter and 21pter-q22.13 due to cryptic unbalanced translocation t(7;21) but not monosomy of the whole chromosome 21: a case report plus review of the literature. Mol Cytogenet 1:13
Vorsanova SG, Yurov YB, Iourov IY (2010a) Human interphase chromosomes: a review of available molecular cytogenetic technologies. Mol Cytogenet 3:1
Vorsanova SG, Yurov YB, Soloviev IV, Iourov IY (2010b) Molecular cytogenetic diagnosis and somatic genome variations. Curr Genomics 11:440–446
Walter J, Joffe B, Bolzer A, Albiez H, Benedetti PA, Muller S et al (2006) Towards many colors in FISH on 3D-preserved interphase nuclei. Cytogenet Genome Res 114:367–378
Weier H-UG (2001) DNA Fiber mapping techniques for the assembly of high-resolution physical maps. J Histochem Cytochem 49(8):939–948
Weise A, Liehr T, Claussen U, Halbhuber K-J (2005) Increased efficiency of fluorescence in situ hybridization (FISH) using the microwave. J Histochem Cytochem 53(10):1301–1303
Weise A, Gross M, Hinreiner S, Witthuhn V, Mkrtchyan H, Liehr T (2010) POD-FISH: a new technique for parental origin determination based on copy number variation polymorphism. Methods Mol Biol 659:291–298
Weise A, Mrasek K, Klein E, Mulatinho M, Llerena JC Jr, Hardekopf D et al (2012) Microdeletion and microduplication syndromes. J Histochem Cytochem 60(5):346–358
Wiegant J, Kalle W, Mullenders L, Brookes S, Hoovers JM, Dauwerse JG et al (1992) High-resolution in situ hybridization using DNA halo preparations. Hum Mol Genet 1(8):587–591
Yang F, Shao C, Vedanarayanan V, Ehrlich M (2004) Cytogenetic and immuno-FISH analysis of the 4q subtelomeric region, which is associated with facioscapulohumeral muscular dystrophy. Chromosoma (Berl) 112:350–359
Yeshaya J, Shalgi R, Shohat M, Avivi L (1999) FISH-detected delay in replication timing of mutated FMR1 alleles on both active and inactive X-chromosomes. Hum Genet 105(1–2):86–97
Yeshaya J, Amir I, Rimon A, Freedman J, Shohat M, Avivi L (2009) Microdeletion syndromes disclose replication timing alterations of genes unrelated to the missing DNA. Mol Cytogenet 2:11
Yurov YB, Soloviev IV, Vorsanova SG, Marcais B, Roizes G, Lewis R (1996) High resolution fluorescence in situ hybridization using cyanine and fluorescein dyes: ultra-rapid chromosome detection by directly fluorescently labeled alphoid DNA probes. Hum Genet 97:390–398
Yurov YB, Vorsanova SG, Iourov IY, Demidova IA, Beresheva AK, Kravetz VS et al (2007) Unexplained autism is frequently associated with low-level mosaic aneuploidy. J Med Genet 44(8):521–525
Yurov YB, Vorsanova SG, Iourov IY (2009) GIN ‘n’ CIN hypothesis of brain aging: deciphering the role of somatic genetic instabilities and neural aneuploidy during ontogeny. Mol Cytogenet 2:23
Acknowledgments
The authors are supported by DLR/BMBF (RUS 2011–2013) and RFFI grant program 12-04-00215-а (Russian Federation, 2012–2014). We gratefully acknowledge the NICHD Brain and Tissue Bank for Developmental Disorders at the University of Maryland, Baltimore, MD, USA for providing the brain tissue samples for I-FISH experiments (partially shown in Figs. 11.1, 11.3, 11.4, 11.5, 11.6, and 11.8).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media, LLC
About this chapter
Cite this chapter
Vorsanova, S.G., Yurov, Y.B., Iourov, I.Y. (2013). Technological Solutions in Human Interphase Cytogenetics. In: Yurov, Y., Vorsanova, S., Iourov, I. (eds) Human Interphase Chromosomes. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-6558-4_11
Download citation
DOI: https://doi.org/10.1007/978-1-4614-6558-4_11
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-6557-7
Online ISBN: 978-1-4614-6558-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)