Abstract
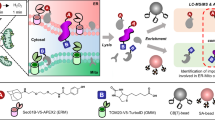
The endoplasmic reticulum (ER) is an essential organelle responsible for many cellular functions, including protein synthesis and folding, lipid synthesis, membrane trafficking, and storage of Ca2+. Therefore, global profiling of ER-associated proteins should be invaluable for understanding these biological processes. However, the difficulty of isolating the intact ER hampered proteome-wide analysis of ER proteins. This chapter describes a chemoproteomic approach for ER proteome analysis using ER-localizable reactive molecules (ERMs), which need neither ER fractionation nor genetic transformation. ERMs spontaneously accumulate in the ER of live cells, and the resultant high concentration of ERMs facilitates spatially limited chemical modification of ER-localized proteins with a detection/purification tag via simple intermolecular reactions. This enables the tag-mediated enrichment and quantitative analysis of the ER-associated proteins using liquid chromatography-tandem mass spectrometry (LC-MS/MS) coupled with SILAC technology.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Schwarz DS, Blower MD (2016) The endoplasmic reticulum: structure, function and response to cellular signaling. Cell Mol Life Sci 73:79–94
Lippincott-Schwartz J, Roberts TH, Hirschberg K (2000) Secretory protein trafficking and organelle dynamics in living cells. Annu Rev Cell Dev Biol 16:557–589
Friedman JR, Voeltz GK (2011) The ER in 3D: a multifunctional dynamic membrane network. Trends Cell Biol 21:709–717
Wu H, Carvalho P, Voeltz GK (2018) Here, there, and everywhere: the importance of ER membrane contact sites. Science 361:eann5835
Zhao L, Ackerman SL (2006) Endoplasmic reticulum stress in health and disease. Curr Opin Cell Biol 18:444–452
Chen X, Karnovsky A, Sans MD, Andrews PC, Williams JA (2010) Molecular characterization of the endoplasmic reticulum: insights from proteomic studies. Proteomics 10:4040–4052
Lee JS, Wu Y, Schnepp P, Fang J, Zhang X, Karnovsky A, Woods J, Stemmer PM, Liu M, Zhang K, Chen X (2015) Proteomics analysis of rough endoplasmic reticulum in pancreatic beta cells. Proteomics 15:1508–1511
Fujisawa A, Tamura T, Yasueda Y, Kuwata K, Hamachi I (2018) Chemical profiling of the endoplasmic reticulum proteome using designer labeling reagents. J Am Chem Soc 140:17060–17070
Yasueda Y, Tamura T, Fujisawa A, Kuwata K, Tsukiji S, Kiyonaka S, Hamachi I (2016) A set of organelle-localizable reactive molecules for mitochondrial chemical proteomics in living cells and brain tissues. J Am Chem Soc 138:7592–7602
Yasueda Y, Tamura T, Hamachi I (2016) Nucleus-selective chemical proteomics using Hoechst-tagged reactive molecules. Chem Lett 45:265–267
Meinig JM, Fu L, Peterson BR (2015) Synthesis of fluorophores that target small molecules to the endoplasmic reticulum of living mammalian cells. Angew Chem Int Ed 54:9696–9699
Bull VH, Thiede B (2012) Proteome analysis of tunicamycin induced ER stress. Electrophoresis 33:1814–1823
Kozutsumi Y, Segal M, Normington K, Gething MJ, Sambrook J (1988) The presence of malfolded proteins in the endoplasmic reticulum signals the induction of glucose-regulated proteins. Nature 332:462–464
Hayoun D, Kapp T, Edri-Brami M, Ventura T, Cohen M, Avidan A, Lichtenstein RG (2012) HSP60 is transported through the secretory pathway of 3-MCA-induced fibrosarcoma tumor cells and undergoes N-glycosylation. FEBS J 279:2083–2095
Mandal A, Mandal S, Park MH (2016) Global quantitative proteomics reveal up-regulation of endoplasmic reticulum stress response proteins upon depletion of eIF5A in HeLa cells. Sci Rep 6:25795
Zhao Q, Wang J, Levichkin IV, Stasinopoulos S, Ryan MT, Hoogenraad NJ (2002) A mitochondrial specific stress response in mammalian cells. EMBO J 21:4411–4419
Stout JM, Gousset MU, Drummond HA, Gray WIII, Pruett BE, Stec DE (2013) Sex-specific effects of heme oxygenase-2 deficiency on renovascular hypertension. J Am Soc Hypertens 7:328–335
Acknowledgments
We thank Dr. Alma Fujisawa and Dr. Yuki Yasueda for their great contribution to the development of ERMs and for helpful advice on the manuscript. This work was funded by a Grant-in-Aid for Scientific Research on Innovative Areas “Integrated Bio-metal Science” (19H05764) to T.T., and Japan Science and Technology Agency (JST) ERATO Grant JPMJER1802 to I.H. This work was also supported by a Grant-in-Aid for Scientific Research on Innovative Areas “Chemistry for Multimolecular Crowding Biosystems” (17H06348).
Conflicts of Interest
The authors declare that this work was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Tamura, T., Hamachi, I. (2023). Quantitative Analysis of the Endoplasmic Reticulum-Associated Proteins Using ER-Localizable Reactive Molecules. In: Luque-Garcia, J.L. (eds) SILAC. Methods in Molecular Biology, vol 2603. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2863-8_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2863-8_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2862-1
Online ISBN: 978-1-0716-2863-8
eBook Packages: Springer Protocols