Abstract
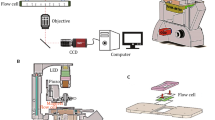
Single-molecule force spectroscopy is a powerful tool to analyze the architecture and interaction of large macromolecular assemblies that are refractory to high-resolution structural interrogations. Here, we describe an optical tweezers–based platform for extracting the mechanical fingerprints of individual nucleosome arrays bound with chromatin-associated complexes, such as the Polycomb repressive complex 2 (PRC2). This platform comprehensively characterizes the diverse binding modes of PRC2 on chromatin, measures their mechanical strengths, and is broadly applicable to the studies of other epigenetic machineries.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kornberg RD (1974) Chromatin structure: a repeating unit of histones and DNA. Science 184:868
Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ (1997) Crystal structure of the nucleosome core particle at 2.8A resolution. Nature 389:251–260
Fierz B, Poirier MG (2019) Biophysics of chromatin dynamics. Annu Rev Biophys 48:321–345
Zhou K, Gaullier G, Luger K (2019) Nucleosome structure and dynamics are coming of age. Nat Struct Mol Biol 26:3–13
Ordu O, Lusser A, Dekker NH (2016) Recent insights from in vitro single-molecule studies into nucleosome structure and dynamics. Biophys Rev 8:33–49
Cui Y, Bustamante C (2000) Pulling a single chromatin fiber reveals the forces that maintain its higher-order structure. Proc Natl Acad Sci U S A 97:127
Bennink ML et al (2001) Unfolding individual nucleosomes by stretching single chromatin fibers with optical tweezers. Nat Struct Biol 8:606–610
Poirier MG, Marko JF (2002) Mitotic chromosomes are chromatin networks without a mechanically contiguous protein scaffold. Proc Natl Acad Sci U S A 99:15393
Brower-Toland BD et al (2002) Mechanical disruption of individual nucleosomes reveals a reversible multistage release of DNA. Proc Natl Acad Sci U S A 99:1960
Pope LH et al (2005) Single chromatin fiber stretching reveals physically distinct populations of disassembly events. Biophys J 88:3572–3583
Mihardja S, Spakowitz AJ, Zhang Y, Bustamante C (2006) Effect of force on mononucleosomal dynamics. Proc Natl Acad Sci 103:15871
Hall MA et al (2009) High-resolution dynamic mapping of histone-DNA interactions in a nucleosome. Nat Struct Mol Biol 16:124–129
Kruithof M et al (2009) Single-molecule force spectroscopy reveals a highly compliant helical folding for the 30-nm chromatin fiber. Nat Struct Mol Biol 16:534–540
Vlijm R, Kim SH, De Zwart PL, Dalal Y, Dekker C (2017) The supercoiling state of DNA determines the handedness of both H3 and CENP-A nucleosomes. Nanoscale 9:1862–1870
Zaret KS, Lerner J, Iwafuchi-Doi M (2016) Chromatin scanning by dynamic binding of Pioneer factors. Mol Cell 62:665–667
Clapier CR, Iwasa J, Cairns BR, Peterson CL (2017) Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat Rev Mol Cell Biol 18:407–422
Allis CD, Jenuwein T (2016) The molecular hallmarks of epigenetic control. Nat Rev Genet 17:487–500
Hashemi Shabestari M, Meijering AEC, Roos WH, Wuite GJL, Peterman EJG (2017) In: Spies M, Chemla YR (eds) Methods in Enzymology, vol 582. Academic Press, Cambridge, Massachusetts, pp 85–119
Margueron R, Reinberg D (2011) The Polycomb complex PRC2 and its mark in life. Nature 469:343–349
Leicher R et al (2020) Single-molecule and in silico dissection of the interaction between Polycomb repressive complex 2 and chromatin. Proc Natl Acad Sci U S A 117:30465
Ngo TTM, Zhang Q, Zhou R, Yodh JG, Ha T (2015) Asymmetric unwrapping of nucleosomes under tension directed by DNA local flexibility. Cell 160:1135–1144
Wasserman MR, Schauer GD, O’Donnell ME, Liu S (2019) Replication fork activation is enabled by a single-stranded DNA gate in CMG helicase. Cell 178:600–611.e616
Ge EJ, Jani KS, Diehl KL, Müller MM, Muir TW (2019) Nucleation and propagation of heterochromatin by the histone methyltransferase PRC2: geometric constraints and impact of the regulatory subunit JARID2. J Am Chem Soc 141:15029–15039
Lowary PT, Widom J (1998) New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning11Edited by T. Richmond. J Mol Biol 276:19–42
Lusser A, Kadonaga JT (2004) Strategies for the reconstitution of chromatin. Nat Methods 1:19–26
Wang MD, Yin H, Landick R, Gelles J, Block SM (1997) Stretching DNA with optical tweezers. Biophys J 72:1335–1346
Wasserman MR, Liu S (2019) A tour de force on the double helix: exploiting DNA mechanics to study DNA-based molecular machines. Biochemistry 58:4667–4676
Müller MM, Muir TW (2015) Histones: at the crossroads of peptide and protein chemistry. Chem Rev 115:2296–2349
Marko JF, Siggia ED (1995) Stretching DNA. Macromolecules 28:8759–8770
Acknowledgments
We thank Matthew Reynolds for developing the Force-Extension Analyzer and the clustering analysis algorithm. This work is supported by the Robertson Foundation, the Pershing Square Sohn Cancer Research Alliance, and the National Institutes of Health (DP2HG010510).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Leicher, R., Liu, S. (2022). Probing the Interaction Between Chromatin and Chromatin-Associated Complexes with Optical Tweezers. In: Gennerich, A. (eds) Optical Tweezers. Methods in Molecular Biology, vol 2478. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2229-2_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2229-2_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2228-5
Online ISBN: 978-1-0716-2229-2
eBook Packages: Springer Protocols