Abstract
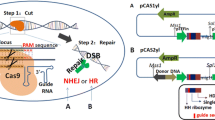
The oleaginous yeast Yarrowia lipolytica has emerged as an industrially relevant chassis to produce various valuable chemicals. Metabolic engineering of Y. lipolytica relies on the availability of genetic engineering tools. Existing engineering strategies for this yeast include homologous recombination, random integration, and episomal plasmid-based gene expression. CRISPR-Cas9 based genome-editing toolbox has also been developed to facilitate multiplexed gene disruption and regulation. Alternative to Cas9, the CRISPR effector Cas12a has also been adopted to perform genome engineering in multiple species. Due to its distinctive features such as short and simple crRNA structure, the ability to process its own crRNA and T-rich PAM sequence (TTTN), Cas12a holds promising potential to be developed as an efficient genome-editing tool. In this chapter, we describe the protocol to implement multiplexed genome editing in Y. lipolytica. The delivery of AsCas12a and crRNA expression via a single plasmid was described. CRISPR-Cas12a-based genome editing could expand the genetic toolbox of Y. lipolytica, whihc is complementary to the classical Cas9-based tools.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Yang Z, Zhang Z (2018) Engineering strategies for enhanced production of protein and bio-products in Pichia pastoris: a review. Biotechnol Adv 36(1):182–195
Spagnuolo M et al (2018) Alternative substrate metabolism in Yarrowia lipolytica. Front Microbiol 9:1077
Yaguchi A, Spagnuolo M, Blenner M (2018) Engineering yeast for utilization of alternative feedstocks. Curr Opin Biotechnol 53:122–129
Ma J et al (2020) Synthetic biology, systems biology, and metabolic engineering of Yarrowia lipolytica toward a sustainable biorefinery platform. J Ind Microbiol Biotechnol 47(9-10):845–862
Blazeck J et al (2014) Harnessing Yarrowia lipolytica lipogenesis to create a platform for lipid and biofuel production. Nat Commun 5:3131
Xu P et al (2016) Engineering Yarrowia lipolytica as a platform for synthesis of drop-in transportation fuels and oleochemicals. Proc Natl Acad Sci 113(39):10848–10853
Spagnuolo M, Yaguchi A, Blenner M (2019) Oleaginous yeast for biofuel and oleochemical production. Curr Opin Biotechnol 57:73–81
Markham KA et al (2018) Rewiring Yarrowia lipolytica toward triacetic acid lactone for materials generation. Proc Natl Acad Sci 115(9):2096–2101
Lv Y et al (2019) Optimizing oleaginous yeast cell factories for flavonoids and hydroxylated flavonoids biosynthesis. ACS Synth Biol 8(11):2514–2523
Lv Y et al (2020) Coupling metabolic addiction with negative autoregulation to improve strain stability and pathway yield. Metab Eng 61:79–88
Gu Y et al (2020) Refactoring Ehrlich pathway for high-yield 2-phenylethanol production in Yarrowia lipolytica. ACS Synth Biol 9(3):623–633
Gu Y et al (2020) Engineering Yarrowia lipolytica as a chassis for de novo synthesis of five aromatic-derived natural products and chemicals. ACS Synth Biol 9(8):2096–2106
Gao S et al (2017) Iterative integration of multiple-copy pathway genes in Yarrowia lipolytica for heterologous beta-carotene production. Metab Eng 41:192–201
Larroude M et al (2018) A synthetic biology approach to transform Yarrowia lipolytica into a competitive biotechnological producer of β-carotene. Biotechnol Bioeng 115(2):464–472
Liu H et al (2020) Genetic and bioprocess engineering to improve squalene production in Yarrowia lipolytica. Bioresour Technol 317:123991
Larroudé M et al (2018) Synthetic biology tools for engineering Yarrowia lipolytica. Biotechnol Adv 36(8):2150–2164
Wong L et al (2017) YaliBricks, a versatile genetic toolkit for streamlined and rapid pathway engineering in Yarrowia lipolytica. Metab Eng Commun 5(Supplement C):68–77
Schwartz CM et al (2016) Synthetic RNA polymerase III promoters facilitate high-efficiency CRISPR-Cas9-mediated genome editing in Yarrowia lipolytica. ACS Synth Biol 5(4):356–359
Schwartz C et al (2016) Standardized markerless gene integration for pathway engineering in Yarrowia lipolytica. ACS Synth Biol 6(3):402–409
Schwartz C et al (2018) Multiplexed CRISPR activation of cryptic sugar metabolism enables Yarrowia lipolytica growth on cellobiose. Biotechnol J 13(9):1700584
Schwartz C et al (2017) CRISPRi repression of nonhomologous end-joining for enhanced genome engineering via homologous recombination in Yarrowia lipolytica. Biotechnol Bioeng 114(12):2896–2906
Zetsche B et al (2017) Multiplex gene editing by CRISPR–Cpf1 using a single crRNA array. Nat Biotechnol 35(1):31–34
Yang Z, Edwards H, Xu P (2020) CRISPR-Cas12a/Cpf1-assisted precise, efficient and multiplexed genome-editing in Yarrowia lipolytica. Metab Eng Commun 10:e00112
Acknowledgments
This work was supported by Bill & Melinda Gates Foundation under grant NO. OPP1188443.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Yang, Z., Xu, P. (2021). Implementing CRISPR-Cas12a for Efficient Genome Editing in Yarrowia lipolytica. In: Wheeldon, I., Blenner, M. (eds) Yarrowia lipolytica. Methods in Molecular Biology, vol 2307. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1414-3_7
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1414-3_7
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1413-6
Online ISBN: 978-1-0716-1414-3
eBook Packages: Springer Protocols