Abstract
The evolution of Bordetella pertussis from a common ancestor similar to Bordetella bronchiseptica has occurred through large-scale gene loss, inactivation and rearrangements, largely driven by the spread of insertion sequence element repeats throughout the genome. B. pertussis is widely considered to be monomorphic, and recent evolution of the B. pertussis genome appears to, at least in part, be driven by vaccine-based selection. Given the recent global resurgence of whooping cough despite the wide-spread use of vaccination, a more thorough understanding of B. pertussis genomics could be highly informative. In this chapter we discuss the evolution of B. pertussis, including how vaccination is changing the circulating B. pertussis population at the gene-level, and how new sequencing technologies are revealing previously unknown levels of inter- and intra-strain variation at the genome-level.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 The Ongoing Problem of Bordetella pertussis
Whooping cough, the infectious respiratory disease caused by the bacterium Bordetella pertussis, has been resurgent in many countries for the past two decades. This resurgence comes in spite of a global vaccination programme, with 90% of the target population receiving a single dose of pertussis-containing vaccine, and 85% receiving three doses (WHO 2018). In addition, there has been a shift in the epidemiological profile of the disease: whereas once most cases were reported in infants and unvaccinated children, the resurgence is also affecting vaccinated children, adolescents and adults, Fig. 1 (Strebel et al. 2001; Clark 2014). Data from the Centers for Disease Control, for example, show that from 1990–1997, the mean incidence of whooping cough per year in 11–19 year-olds was 2.54 per 100,000 people, whereas from 2010–2017, it was 20.43 (Centers for Disease Control 2019).
Reported incidence of whooping cough per 100,000 in the USA, 1990–2017, showing increase in all age brackets. (Data source: CDC National notifiable diseases surveillance reports Centers for Disease Control (2019))
The original vaccination programme, introduced in the 1940s and 1950s, used a whole-cell vaccine (WCV). Initially, cases of the disease appeared to drop significantly. Due to perceived reactogenicity of the WCV (now largely discredited), it was replaced in many countries throughout the 1990s and early 2000s with an acellular vaccine (ACV) (for example: Cherry 1990, 1992, 1996; Cherry et al. 1993; Blumberg et al. 1993; Moore et al. 2004). The acellular vaccine contains one to five B. pertussis antigens, including Pertussis toxin (ptx), pertactin (prn), filamentous haemagglutinin (FHA) and the fimbrial proteins Fim2 and Fim3. These days, most developed countries use the ACV, although many developing countries continue to use the WCV. The recent use of the ACV has been strongly implicated in whooping cough’s resurgence. However, concerns were raised about waning immunity conveyed by the WCV beginning in the early 1990s and, in many countries, the resurgence does seem to pre-date the switch to the ACV (De Serres et al. 1995; Cherry 1996; de Melker et al. 1997).
Three main potential causes are thought to have contributed to the recent observed increase in whooping cough cases: increased awareness of the disease coupled with improved diagnosis due to a switch from culture-based to PCR- and serology-based techniques; waning immunity, particularly that conveyed by the ACV compared to the WCV; and genetic variations in circulating B. pertussis strains, away from the vaccine strains (Sealey et al. 2015; Clark 2014; Ausiello and Cassone 2014). Here we focus on the latter from two different but highly interrelated perspectives: variation at the gene-level, and variation at the genome-level, with particular consideration of how recent developments in genomic research have contributed to our understanding of evolution and variation in a species which traditionally has been described as highly monomorphic (for example: Bart et al. 2010; Mooi 2010).
2 The Speciation of Bordetella pertussis
The study of the bordetellae has focussed largely on the three classical, pathogenic, Bordetella species: B. bronchiseptica, B. pertussis and B. parapertussis. However, the Bordetella genus contains many additional species which have been isolated from extremely diverse environments, including marine sponges, bioreactors, nitrifying sludge and mural paintings in ancient tombs (Wang et al. 2007; Bianchi et al. 2005; Sun et al. 2019; Tazato et al. 2015). Using previously published 16S sequence data derived from many Bordetella species to create a phylogenetic tree, Hamidou Soumana et al. (2017) demonstrated that eight out of the ten clades contained soil-dwelling bordetellae; the permeation of the environmental phenotype throughout the phylogenetic tree hints at a soil-based origin for the Bordetella genus.
Key evolutionary milestones within the genus involve species that are capable of both environmental and pathogenic lifestyles, such as the key ancestor to the classical Bordetella species, a B. bronchiseptica-like bacterium. Multilocus sequence typing (MLST) studies of B. bronchiseptica have established two distinct complexes of isolates, complex I and IV. The majority of strains isolated from humans originate from Complex IV (Diavatopoulos et al. 2005; Park et al. 2012). Using the much higher discriminatory power of next generation sequencing (NGS) data, it has been demonstrated that B. pertussis and B. parapertussis evolved from different complexes, with B. parapertussis sharing a more recent ancestor with Complex I and B. pertussis with Complex IV (Linz et al. 2016).
Despite their apparently different evolutionary trajectories, B. pertussis and B. parapertussis cause remarkably similar pathologies in humans. In an example of convergent evolution, the genomes of both species have evolved primarily through genome reduction, mediated through homologous recombination of insertion sequences (IS). As seen in Table 1, the genome of B. bronchiseptica contains very few IS elements; in fact, the reference strain RB50 was originally believed to contain no IS elements at all (Parkhill et al. 2003; Preston et al. 2004). In contrast, B. parapertussis genomes contain around 30 IS elements, usually 22 copies of IS 1001 and 9 copies of IS 1002. The B. pertussis genome contains the most IS elements: up to ten copies of IS 1002, around 20 copies of IS 1663, and over 200 copies of IS 481. The appearance and expansion of these IS elements is thought to have led to the speciation of B. pertussis and, separately, B. parapertussis, from their B. bronchiseptica-like ancestors (Parkhill et al. 2003; Preston et al. 2004; Diavatopoulos et al. 2005).
Genome reduction was key in the speciation of B. pertussis. The significant, IS-mediated, reduction in genome size of the B. pertussis genome (around 4.1 Mb) compared to the B. bronchiseptica genome (around 5.3 Mb) has led to a streamlined genome, depleted of many metabolic, membrane transport, surface structure synthesis and gene expression regulatory genes when compared to B. bronchiseptica genomes (Parkhill et al. 2003). Comparative genomic studies between B. bronchiseptica and B. pertussis reveal that the latter has around 1,200 fewer genes (Parkhill et al. 2003; Linz et al. 2016). In addition, insertions of IS elements into genes which are functional in B. bronchiseptica has resulted in the existence of over 350 pseudogenes in B. pertussis, compared to only around 20 in B. bronchiseptica. This sculpting of the B. pertussis genome via IS-mediated homologous recombination has produced a highly specialised pathogenic bacterium which is niche-restricted to the human nasopharynx. Traditionally, B. pertussis has been described as a monomorphic species (for example: Mooi 2010; Bart et al. 2010); however, since the advent of whole genome sequencing, genomics has been revealing that the bacterium may be less clonal than previously thought, with the introduction of vaccination and continued homologous recombination between IS elements driving gene- and genome-level variations respectively.
3 Vaccination Has Accelerated B. pertussis Gene-Level Evolution
3.1 Changes to Circulating Alleles
Over the last several decades, allele-typing of selected genes has shown a number of similar trends, characterised largely by the drift of genes away from the vaccine alleles (for example: Mooi et al. 1998; van der Zee et al. 1996; Mooi et al. 2001). One well-reviewed example involves pertussis toxin. Prior to the 1990s, the predominant ptx promoter allele was ptxP1. A new allele, ptxP3, was first observed in 1988 (Bart et al. 2010). In ptxP3, a SNP in the binding site for ptx’s transcriptional regulator, BvgA, appears to increase the binding affinity between the promotor and regulator, thus increasing transcription and causing ptxP3-carrying strains to produce more ptx. The expression of other proteins involved in complement resistance is also altered and, together, these changes increase the transmissibility and severity of the disease caused by ptxP3-carrying strains (Mooi et al. 2009; Bart et al. 2010; King et al. 2013; de Gouw et al. 2014). This new allele spread rapidly throughout the 1990s, and ptxP3 is now present in greater than 90% of recent isolates (Lam et al. 2012; Bart et al. 2010). A thorough screen of 343 strains representing 19 countries and six continents, spanning 90 years of B. pertussis isolation, showed that similarly rapid selective sweeps have also occurred in other antigen-related genes, including ptxA, prn, and fim3 (Bart et al. 2014a).
In addition, analysis of 100 strains isolated during a 2012 whooping cough outbreak in the UK, after the introduction of the ACV, showed that the evolution of the antigens included in the ACV is occurring more rapidly than that of other B. pertussis surface proteins, which are presumably under similar levels of pressure from the human immune system (Sealey et al. 2015). Importantly, this analysis also showed that numerous different strains were circulating during the outbreak, rather than one particularly virulent strain or allelic profile being responsible. The same was also true for strains circulating during outbreaks in the USA, in California in 2010, and in Vermont and Washington in 2012 (Bowden et al. 2014; Bowden et al. 2016). This suggests that the strains circulating during outbreaks tend to be the same as those that circulate during non-outbreak periods, but that some unknown trigger causes an increase in whooping cough cases in regular four-yearly cycles.
Supporting the idea that the recent allelic changes we are seeing in B. pertussis are, in part, a response to the introduction of vaccination, Xu et al. (2015) and Du et al. (2016) showed that, in countries where vaccine uptake has been lower or delayed, the rate of change to the allelic profile of circulating strains has also been delayed. In the Philippines, for example, where the WCV is still in use, prn2 has yet to appear, despite being the allele most frequently seen in ACV-adopting countries (Galit et al. 2015).
3.2 The Proliferation of Antigen-Deficient Strains
Another gene-level phenomenon which has been observed recently is the emergence of strains deficient in one or more of the antigens used in the ACV. During the pre-ACV era, antigen-deficient strains were occasionally isolated, albeit at very low frequencies. Individual strains with non-functional pertactin genes, for example, were isolated in Europe, North America and Japan in the 1990s (Mastrantonio et al. 1999; Weigand et al. 2018; Miyaji et al. 2013). The landmark study by Bart et al. (2014a), in which all but around 20 strains were isolated prior to 2007, did not identify any strains which were prn-deficient (although one, BP310, has subsequently been resequenced by Zomer et al. (2018) and is likely to be deficient). Since the mid-2000s, however, the number of prn-deficient strains being isolated globally has increased rapidly. A study of Australian isolates from 2008–2012 showed an increase from 5% prn-deficient strains in 2008 to 78% prn-deficient strains in 2012 (Lam et al. 2014). Pertactin-deficiency appears to be polyclonal, affecting both dominant prn alleles, prn1 and prn2, and arising through several different mechanisms including insertions of IS 481, large deletions, and SNPs, with no single predominant causative mutation (Hegerle et al. 2012; Queenan et al. 2013; Barkoff et al. 2019).
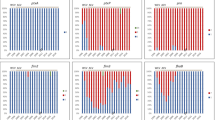
As with changes to allelic profiles, countries with different vaccination strategies appear to be differently affected by the proliferation of antigen-deficiency. A longitudinal study of prn-deficiency in European countries between 1998 and 2015, Fig. 2, showed that the number of prn-deficient strains is increasing in all screened countries but, the earlier a country introduced the ACV, the higher the percentage of strains currently found to be deficient (Barkoff et al. 2019). The rapid recent increase in prn-deficiency suggests that, although the deficiency may have always occurred in some strains by chance, it has been strongly selected for by the ACV compared to the WCV. Further supporting the idea of ACV-mediated selection pressure is the sustained decrease of prn-deficient strains circulating in Japan since pertactin was removed from the Japanese ACV in 2012 (Hiramatsu et al. 2017).
Correlation between the introduction of a primary acellular pertussis vaccine containing pertactin (PRN) in a European country and the proportion of PRN-deficient isolates found in the study, 2012–2015. (Reproduced with permission from Barkoff et al. (2019))
A smaller number of strains deficient in other antigens included in the ACV have also been identified. Bart et al. (2015) and Weigand et al. (2018) identified several geographically independent recent strains which were unable to produce FHA; the same strains were often also prn-deficient. In addition, a handful of strains have been isolated globally which are deficient in both prn and ptx (Bouchez et al. 2009; Williams et al. 2016; Weigand et al. 2018). However, both FHA and Pertussis toxin are thought to play more vital roles in whooping cough disease development than pertactin (Carbonetti 2010; Henderson et al. 2012; Serra et al. 2011). Hence, although occasional strains may develop deficiency in these antigens, the reduced ability of these deficient strains to cause disease may mean that this kind of antigen deficiency is unlikely to proliferate in the same way as prn-deficiency.
4 Recent Sequencing Advances Highlight Genome-Level Variation
4.1 The Limitations of Short-Read Sequencing
Whilst the wide availability of whole genome sequencing throughout the 2000s enabled a variety of high-throughput strain screens, the highly repetitive nature of the B. pertussis genome has made the assembly of closed, single-contig, B. pertussis genomes difficult. IS 481, together with the smaller number of copies of other repeated regions, such as IS 1002, IS 1663 and the rRNA operon, has confounded attempts to assemble closed genomes using short-read sequencing technologies, which produce sequencing reads shorter than the repeated section. The hundreds of B. pertussis genomes assembled using short-read sequencing alone have therefore tended to consist of several hundred contigs, ostensibly one per repeat in the genome. Thus, the majority of high-throughput screens throughout the 2000s were focussed on the gene-level differences between strains already discussed.
The presence of so many IS elements, however, means that assembly of closed B. pertussis genomes could be particularly informative: IS elements are able to move around the genome through homologous recombination, potentially causing genome-level structural changes which may be discernible only through single-contig assemblies (Bentley et al. 2008; Siguier et al. 2014). Despite the discovery that most whooping cough outbreaks tend to be polyclonal in nature, B. pertussis remains a relatively clonal species, with a low SNP rate compared to many other bacteria. In other species, in addition to gene-level variations, differences at a whole-genome level are known to contribute to altered gene expression and phenotypic diversity (Darch et al. 2014; Sousa et al. 1997). IS-mediated rearrangement may affect gene regulation and/or expression in B. pertussis by a number of mechanisms, including IS 481’s inwards and outwards-facing promoters, as well as changing the distance of genes from the origin of replication (Amman et al. 2018). Limited evidence has already shown that certain genome-level differences can affect phenotype in B. pertussis in this way (Brinig et al. 2006).
The speciation of B. pertussis from B. bronchiseptica via IS element-mediated homologous recombination resulted in a variety of genomic arrangement differences between the two species alongside the reduction in genome size, and it is likely that IS-mediated genomic rearrangement in B. pertussis is an ongoing process (Parkhill et al. 2003). Indeed, prior to the advent of long-read sequencing, pulsed-field gel electrophoresis (PFGE) was one of the few methods able to discriminate between highly clonal B. pertussis strains: isolate screens using PFGE indicated that strains which seemed otherwise alike could vary significantly in terms of PFGE type (van Gent et al. 2015; Bisgard et al. 2001; Advani et al. 2004; Advani et al. 2013). Although the existence of numerous PFGE types was widely seen, however, it could not be confirmed how different PFGE types arose; they could represent different genomic arrangements, but could also have arisen due to mutagenesis at PFGE restriction sites, for example.
Thus, closed B. pertussis genome sequences may validate and further reveal genome-level differences between strains which otherwise appear to have highly similar or identical DNA content. The recent availability of long-read sequencing techniques, which can produce sequencing reads longer than 1,000 bp, has therefore revolutionised our ability to discover and investigate genome-level variations in B. pertussis.
4.2 Long-Read Sequencing Shows Extensive Inter-strain Genome Rearrangement
The first study to take advantage of long reads utilised Pacific Biosciences (PacBio) sequencing to produce closed, fully annotated, genomes for two B. pertussis strains: BP1917 and BP1920 (Bart et al. 2014b). The arrangement of these two strains differed significantly, with three large inversions and a variety of deletion and/or insertion events between the pair. Having proven the ability of long reads to close the genomes of BP1917 and BP1920, Bart et al. (2015) next sequenced 11 B. pertussis strains which represented the pandemic ptxP3 lineage, again using PacBio sequencing to produce 10 kb-long reads. This cohort, which also included several strains deficient in prn and/or FHA, were characteristically similar in terms of SNPs but again showed significant differences in genome arrangement.
As is common for a developing technology, the cost of PacBio sequencing has rapidly decreased. Thus, higher-throughput strain screens have become increasingly feasible. Figure 3 shows the dramatic increase in closed genome sequences for the classical Bordetella species available from the NCBI’s RefSeq database since 2014. Bowden et al. (2016) conducted the first whooping cough outbreak screen to utilise long-read sequencing alongside short-read sequencing in hybrid, sequencing 31 strains which had circulated during US whooping cough outbreaks in 2010 and 2012. The hybrid approach has been shown to improve the accuracy of assemblies produced using long-read sequencing, which still have an intrinsically higher error rate than short-read-only assemblies, particularly in homopolymeric tracts (Au et al. 2012; Koren et al. 2012). In the 31 genomes studied, 21 different arrangement profiles were observed; most consisted of inversions around the origin of replication. Bowden et al. also validated the arrangements using whole genome optical mapping and found that, in all cases, the boundaries between rearranged sections were composed of a repeated element: an insertion sequence, or the rRNA operon. The vast majority of the boundaries were IS 481 (89%), whilst the rest were composed of an rRNA operon, IS 1002 or a combination of IS 1002 and IS 481 together.
The most thorough investigation of B. pertussis genomic rearrangement to date also used a hybrid assembly strategy, combining PacBio long reads with Illumina short reads to close the genomes of 257 strains, dating from 1939 to 2014 (Weigand et al. 2017). When clustered based on their arrangement profiles, most isolates clustered according to allelic profile; for example, most ptxP1 strains shared similar arrangements with other ptxP1 strains. This clustering indicates that most structures are relatively stable, as supported by a clinical isolate which showed the same structure before and after 11 serial passages. Furthermore, these findings suggest that lineages are conserved not just in terms of SNPs, but also in genomic arrangement. Interestingly, Weigand et al. note that, on average, only half of their predicted IS 481 target sites are occupied in any given genome, suggesting a potential for further IS-mediated structural changes in future generations, assuming these sites are not non-permissive.
5 How Else Might B. pertussis Generate Diversity Through Genome-Level Variation?
The primary metric commonly used to assess diversity of bacterial species is SNPs. However, B. pertussis is a textbook example of a clonal bacterial pathogen: variation, when judged by SNPs alone, is extremely limited, even taking into account the accelerated mutation of B. pertussis genes since the introduction of vaccination. Bart et al. (2010) estimated the mutation rate between B. pertussis isolates to be 1 SNP per 8,675 bases, compared to, for example, 1 SNP per 3,000 bases in Mycobacterium tuberculosis, and 1 SNP per 6,700 bases in Escherichia coli O157:H7 (Fleischmann et al. 2002; Gutacker et al. 2002; Zhang et al. 2006). Diversity within a species is vital for its survival, in order to drive adaptation; this is particularly true for pathogens, which are under pressure from the immune system (Mooi 2010). Therefore, a prominent question in B. pertussis genomics is: despite limited SNP diversity, how does B. pertussis generate diversity? The large numbers of closed genomes assembled using long-read sequencing have proven that rearrangements are a rich source of genome-level diversity, but can genomics also reveal other types of genome-level variation?
5.1 Harnessing Deletion as a Driver of Diversity
King et al. (2010) analysed the size of B. pertussis genomes of strains isolated over a 60-year period, demonstrating that genome streamlining has been an ongoing process, with recently isolated strains having smaller genomes and higher numbers of pseudogenes. Thus, B. pertussis is described as a species which is still undergoing genome reduction. Like genomic rearrangement, reduction is driven primarily by homologous recombination between insertion sequences. The large numbers of homologous IS elements in B. pertussis therefore produce a fertile mutational landscape capable of the generation of diversity.
Many bacterial species also generate diversity through the gain of genes, often by horizontal gene transfer (HGT), resulting in fluid gene content of the population, enabling the population to effectively respond to evolutionary bottlenecks that may arise over long or short timescales. In B. pertussis, however, HGT appears to occur very rarely (Linz et al. 2016). Gene content of a species or genus is often analysed using a “pangenome” approach, which consists of analysing which genes are consistently present (the core genome) in the population, and which genes are variably present (the accessory genome) (reviewed in Medini et al. 2005; Rouli et al. 2015). A number of studies have undertaken this analysis in B. pertussis using either comparative genomic hybridization (CGH) (for example: Zhang et al. 2006; Caro et al. 2006; Heikkinen et al. 2007; King et al. 2008) or NGS (for example: Park et al. 2012; Ding et al. 2017). These have shown that, despite extremely limited HGT and otherwise high levels of clonality between strains, B. pertussis maintains a moderate accessory genome, largely through gene loss rather than gene gain. For example, the most comprehensive pangenome study to date, using CGH on 171 B. pertussis strains, revealed that 15% of the genes present in the population appeared variably in the 171 strains studied. (King et al. 2010). By using a set of probes which included B. bronchiseptica and B. parapertussis, King et al. were able to avoid biasing their analysis towards genes that were present only in the B. pertussis reference strain, Tohama I, which has been shown to lack over 45 kb of the accessory genome of the population (Caro et al. 2008; Bouchez et al. 2008).
There is a lack of knowledge about the phenotypic impact of gene deletions in B. pertussis, however. As the cost of sequencing has plummeted, the frequency and ease with which genomes, and their constituent mutations, are published has far outpaced the publishing of their phenotypic impact. To cope with the deluge of data, ontology schemes strive to categorize genes into functional groups and estimate their function based on sequence homology. A variety of nucleotide polymorphisms in B. pertussis, such as those in ptxP3, have had their phenotypic impacts analysed, with some providing clear fitness advantages in the mouse model (Mooi et al. 2009). In contrast, many key gene deletions have yet to be experimentally characterized in B. pertussis. Using the ontology scheme Clusters of Orthologous Genes (COG), King et al. (2010) showed that, as expected, housekeeping genes were underrepresented in the deleted genes, whilst genes of unknown function were overrepresented by 25%. There therefore remains genetic “dark matter”, genes with unknown function which are overrepresented in gene deletions.
Whilst B. pertussis is undisputedly undergoing genome reduction, evolution acts on phenotypes rather than genotypes. Therefore, it is likely that the B. pertussis genome is undergoing streamlining as an effect of certain phenotypes being selected against. It has been theorised that the transcriptional and translational cost of superfluous genes far outstrips the mere cost of DNA replication of such regions (Adler et al. 2014). B. pertussis maintains many seemingly functionless pseudogenes, despite the vast majority being shown to be transcriptionally inactive in vitro and in the mouse model (Bart et al. 2010; King et al. 2008; de Gouw et al. 2014). This supports the idea that the deletion of some genes provides a greater fitness benefit than the deletion of some others. Nonetheless, there is also evidence that the DNA content of the species is under selection, as pseudogenes have been shown to be mildly enriched in gene deletions, suggesting that streamlining of the DNA is also favoured to some extent (Kuo and Ochman 2010). Thus, the process of B. pertussis genome streamlining is likely to be a balance between entirely passive and entirely directed.
There have been five deleted regions, totalling over 50 genes, that have been deleted in all recently isolated clinical strains in comparison to the reference strain Tohama I (King et al. 2010; Heikkinen et al. 2007; Bouchez et al. 2008). In addition to clinical strains, Bart et al. (2014a) also investigated two strains that were used to make WCVs. In one of the vaccine strains, the five deleted regions were present; if these regions impact cell surface antigens, the immunity conveyed by the WCV could therefore also be affected. This highlights the clinical importance of understanding the continual evolution of B. pertussis, which is in part driven by genome reduction.
5.2 Harnessing Duplication as a Driver of Diversity
Homologous recombination between IS elements has not only caused rearrangements and deletions in the B. pertussis genome, but also duplications ranging from single genes to large, multi-gene, regions. The general paradigm under which duplications occur is that a gene is duplicated, thus freeing the second copy from the purifying selection of the original copy of the gene, potentially allowing it over time to evolve a new function. However, the second copy of the gene may also maintain the same function of the first gene. These types of events are “canonical” duplications that are well documented in the bacterial kingdom (for example: Ohta 1989; Lynch 2002; Magadum et al. 2013). Taking the genes from Tohama I and clustering them based on 90% nucleotide homology using the tool CDHIT (available as a web server: http://weizhong-lab.ucsd.edu/cdhit_suite/cgi-bin/index.cgi?cmd=cd-hit-est), it can be seen that Tohama I maintains two copies of nine separate genes ranging from 97% to 100% homology (excluding IS elements and rRNA genes) The maintenance of these duplications provides further evidence that it is not the genome size of the bacterium itself that is the primary target of streamlining but certain phenotypic traits which are coded in the DNA.
Before the cost of NGS rapidly decreased in the late 2000s and early 2010s, duplications were primarily inferred by increased spot intensity in CGH or disturbances to southern blotting or PFGE patterns. Using these techniques, a number of multi-gene duplications were serendipitously discovered in the B. pertussis population. Using the power of long-read sequencing technologies, a number of studies had revealed further duplications, bringing the total to 13 serendipitously discovered mutations (Dalet et al. 2004; Caro et al. 2006; Heikkinen et al. 2007; Weigand et al. 2016; Dienstbier et al. 2018; Ring et al. 2018; Weigand et al. 2018). A recent study by Abrahams et al. (In preparation) systematically analysed the B. pertussis population in search of large multi-gene duplications. Previously published short-read sequencing data was utilised and read depth abnormalities were used to predict duplications. In the 473 strains analysed, over 400 duplications were found.
Abrahams et al. (In preparation) presents a deep description of duplications in B. pertussis. In addition to the quantity of duplications, over 90% of duplications were found at 11 “hotspot” loci but with varying gene contents, similar to a situation described previously in M. tuberculosis (Weiner et al. 2012). Interestingly, when the CNVs at each hotspot loci were mapped to a phylogenetic tree based on core genome SNPs, they appeared not to be vertically inherited and instead appeared to occur spontaneously many times at similar loci with subtly different gene content in each mutation, suggesting the existence of a potential phenotypic driver at those loci.
Large multi-gene duplications are known to be unstable in the bacterial kingdom, and this has also been demonstrated in B. pertussis. For instance, Dalet et al. (2004) noticed that subculturing a single isolate produced both high haemolytic and average haemolytic single colonies. Further analysis showed that colonies with high haemolysis had a duplication of the locus encoding adenylate cyclase, a key virulence factor. Dalet et al. further demonstrated that subculturing a strain with a duplication produced colonies with a single copy of the locus, thus indicating a mixed population, ostensibly caused by an unstable locus. This early study used PFGE and southern blotting to screen colonies for copy number of the locus. These “pre-genomics” tools provide high quality data, but largely answer very specific research hypotheses, in contrast to sequencing experiments, which shed light on a vast range of research questions. Abrahams et al. used ultra-long nanopore sequencing reads (over 3,000 reads longer than 50 kb) to confirm the presence of between 1 and 5 copies of a single locus within an otherwise clonal population. This tentatively supports the findings of Dalet et al., showing that in a single sample there exists a variety of genetic configurations of a single locus. This study demonstrates the potential of long-read sequencing to not only confirm long-predicted genomic structural variations in B. pertussis, but also to play a key role in the discovery and further investigation of a variety of entirely unpredicted genomic phenomena. The next steps in understanding these new genomic phenomena in B. pertussis should aim to elucidate the existence and extent of any phenotypic effects stemming from large duplicated regions, as well as any contribution they make to whooping cough virulence.
6 What Is the Future for B. pertussis Genomic Research?
Changes to the allelic profile of circulating B. pertussis strains have been recorded for many decades, in the pre-genomics era and beyond. The wide-spread availability of whole genome sequencing since the early 2000s, though, has enabled the screening of larger numbers of strains isolated over the last hundred years, thus allowing us to understand, longitudinally, the extent to which B. pertussis has been evolving on the gene-level. As seen above, there is evidence that many of these gene-level changes, in terms of both allelic profile and antigenic deficiency, have been influenced by the introduction of first the WCV and then the ACV. Since the 2010s, long-read sequencing has allowed us to investigate B. pertussis on a new level, that of the whole genome. The existence of a wide variety of inter-strain genomic rearrangements is now well-established, and more recent evidence has begun to show that other types of genome-level differences, such as large tandem duplications, also exist. However, the contribution of these observed gene- and genome-level variations to observed phenotypes is yet to be fully understood. In addition to informing our understanding of the continued evolution of the B. pertussis genome, a more thorough understanding of B. pertussis genomics could also contribute significantly to the future of whooping cough prevention strategies (Cherry 2019).
Long-read sequencing will have a major part to play in any future investigation of B. pertussis genomic variations. Until recently, high-throughput long-read sequencing was restricted to larger laboratories which could afford a PacBio sequencing system, thus all the early long-read studies described here took place at national health laboratories: the CDC in the US, and the Centre for Infectious Diseases Control in the Netherlands. Oxford Nanopore Technologies (ONT) sequencing may provide a more accessible alternative for smaller laboratories and, indeed, two studies from late 2018 have shown the feasibility of assembling multiple closed B. pertussis genomes using ONT sequencing in hybrid with Illumina sequencing (Ring et al. 2018; Bouchez et al. 2018). In addition, ultra-long ONT sequencing has recently revealed yet further B. pertussis structural complexity, in the form of highly mixed populations (Abrahams et al. In preparation). Thus, it is likely that future studies of B. pertussis genome structure will utilise both PacBio and ONT sequencing. For investigating the most complicated structural features, such as very long tandem duplications, there will likely be a preference for ONT sequencing, as there is theoretically no upper limit to the length of sequencing read which could be produced by nanopore sequencing (Schmid et al. 2018). It is also likely that any studies utilising long reads to investigate B. pertussis will use them in hybrid with a more accurate short-read technology, although improvements to both long-read technologies mean that highly accurate long-read-only assemblies are on the horizon, which could enable both base-level and genome-level interrogations using a single technology (Wenger et al. 2019; Oxford Nanopore Technologies 2018).
Alongside sequencing, there will still remain a place for other genomics tools, such as PFGE and optical mapping. The most recent survey of B. pertussis genomic diversity in the US, by Weigand et al. (2019), demonstrates the potential for such a wholistic approach. Using a combination of short-read sequencing, long-read sequencing, multilocus variable-number tandem-repeat analysis (MLVA), PFGE and optical mapping, Weigand et al. were able in a single study to characterise the gene- and genome-level profiles, including allelic-profile, antigen deficiency, genome arrangement and the existence of several large tandem duplications, in 170 strains isolated between 2000 and 2013. Such detailed analyses will likely provide a springboard for future studies, for both the continued surveillance of B. pertussis evolution, and the investigation of any correlation between genotypic, genomic and phenotypic differences.
In summary, genomics has shown, and continues to show, that B. pertussis is not necessarily the entirely monomorphic species it is traditionally believed to be. Although the allelic profile of B. pertussis changed in response to the introduction of the WCV, and more rapidly since the switch to the ACV, diversity at the gene-level remains very limited when compared to many other bacteria. However, the wide availability of WGS, and particularly the more recent long-read sequencing technologies, have revealed dynamic and substantial genome-level variations, both between and within strains. Future work may utilise a wholistic approach, focussing on the further elucidation and phenotypic characterisation of both gene- and genome-level phenomena together, ultimately informing our understanding of how diversity is generated in species with limited base-level inter-strain variation and, perhaps, the role this has played in the resurgence of whooping cough.
Abbreviations
- ACT:
-
adenylate cyclase toxin
- ACV:
-
acellular vaccine
- CGH:
-
comparative genomic hybridization
- COG:
-
Clusters of Orthologous Genes
- FHA:
-
Filamentous haemagglutinin
- Fim(2/3):
-
Fimbrial protein (2/3)
- HGT:
-
horizontal gene transfer
- INDEL:
-
insertions and deletions
- IS:
-
insertion sequence
- MLST:
-
multilocus sequence typing
- MLVA:
-
multilocus variable-number tandem-repeat analysis
- NGS:
-
next generation sequencing
- ONT:
-
Oxford Nanopore Technologies
- PacBio:
-
Pacific Biosciences
- PERISCOPE:
-
PERtussIS Correlates Of Protection Europe (consortium)
- PFGE:
-
pulsed field gel electrophoresis
- prn:
-
Pertactin
- ptx:
-
Pertussis toxin
- SMRT:
-
single-molecule real-time
- SNP:
-
single nucleotide polymorphism
- WCV:
-
whole-cell vaccine
- WGS:
-
whole genome sequencing
References
Abrahams JS, Ring N, Quick J, Loman N, Peng Y, Weigand MA, Tondella ML, Williams MM, Bagby S, Gorringe AR, Preston A (In preparation) Duplications drive diversity in the monomorphic pathogen Bordetella pertussis on an underestimated scale
Adler M, Anjum M, Berg OG, Andersson DI, Sandegren L (2014) High fitness costs and instability of gene duplications reduce rates of evolution of new genes by duplication-divergence mechanisms. Mol Biol Evol 31(6):1526–1535. https://doi.org/10.1093/molbev/msu111
Advani A, Donnelly D, Hallander H (2004) Reference system for characterization of Bordetella pertussis pulsed-field gel electrophoresis profiles. J Clin Microbiol 42(7):2890–2897. https://doi.org/10.1128/jcm.42.7.2890-2897.2004
Advani A, Hallander HO, Dalby T, Krogfelt KA, Guiso N, Njamkepo E, von Konnig CH, Riffelmann M, Mooi FR, Sandven P, Lutynska A, Fry NK, Mertsola J, He Q (2013) Pulsed-field gel electrophoresis analysis of Bordetella pertussis isolates circulating in Europe from 1998 to 2009. J Clin Microbiol 51(2):422–428. https://doi.org/10.1128/jcm.02036-12
Amman F, D’Halluin A, Antoine R, Huot L, Bibova I, Keidel K, Slupek S, Bouquet P, Coutte L, Caboche S, Locht C, Vecerek B, Hot D (2018) Primary transcriptome analysis reveals importance of IS elements for the shaping of the transcriptional landscape of Bordetella pertussis. RNA Biol 15(7):967–975. https://doi.org/10.1080/15476286.2018.1462655
Au KF, Underwood JG, Lee L, Wong WH (2012) Improving PacBio long read accuracy by Short read alignment. PLoS One 7(10):e46679. https://doi.org/10.1371/journal.pone.0046679
Ausiello CM, Cassone A (2014) Acellular pertussis vaccines and pertussis resurgence: revise or replace? MBio 5(3):e01339–e01314. https://doi.org/10.1128/mBio.01339-14
Barkoff AM, Mertsola J, Pierard D, Dalby T, Hoegh SV, Guillot S, Stefanelli P, van Gent M, Berbers G, Vestrheim D, Greve-Isdahl M, Wehlin L, Ljungman M, Fry NK, Markey K, He Q (2019) Pertactin-deficient Bordetella pertussis isolates: evidence of increased circulation in Europe, 1998 to 2015. Euro surveillance: bulletin Europeen sur les maladies transmissibles Eur Commun Dis Bull 24(7). https://doi.org/10.2807/1560-7917.es.2019.24.7.1700832
Bart MJ, van Gent M, van der Heide HG, Boekhorst J, Hermans P, Parkhill J, Mooi FR (2010) Comparative genomics of prevaccination and modern Bordetella pertussis strains. BMC Genomics 11:627. https://doi.org/10.1186/1471-2164-11-627
Bart MJ, Harris SR, Advani A, Arakawa Y, Bottero D, Bouchez V, Cassiday PK, Chiang CS, Dalby T, Fry NK, Gaillard ME, van Gent M, Guiso N, Hallander HO, Harvill ET, He Q, van der Heide HG, Heuvelman K, Hozbor DF, Kamachi K, Karataev GI, Lan R, Lutynska A, Maharjan RP, Mertsola J, Miyamura T, Octavia S, Preston A, Quail MA, Sintchenko V, Stefanelli P, Tondella ML, Tsang RS, Xu Y, Yao SM, Zhang S, Parkhill J, Mooi FR (2014a) Global population structure and evolution of Bordetella pertussis and their relationship with vaccination. MBio 5(2):e01074. https://doi.org/10.1128/mBio.01074-14
Bart MJ, Zeddeman A, van der Heide HGJ, Heuvelman K, van Gent M, Mooi FR (2014b) Complete genome sequences of Bordetella pertussis isolates B1917 and B1920, representing two predominant global lineages. Genome Announc 2(6). https://doi.org/10.1128/genomeA.01301-14
Bart MJ, van der Heide HGJ, Zeddeman A, Heuvelman K, van Gent M, Mooi FR (2015) Complete genome sequences of 11 Bordetella pertussis strains representing the pandemic ptxP3 lineage. Genome Announc 3(6). https://doi.org/10.1128/genomeA.01394-15
Bentley DR, Balasubramanian S, Swerdlow HP, Smith GP, Milton J, Brown CG, Hall KP, Evers DJ, Barnes CL, Bignell HR, Boutell JM, Bryant J, Carter RJ, Keira Cheetham R, Cox AJ, Ellis DJ, Flatbush MR, Gormley NA, Humphray SJ, Irving LJ, Karbelashvili MS, Kirk SM, Li H, Liu X, Maisinger KS, Murray LJ, Obradovic B, Ost T, Parkinson ML, Pratt MR, Rasolonjatovo IM, Reed MT, Rigatti R, Rodighiero C, Ross MT, Sabot A, Sankar SV, Scally A, Schroth GP, Smith ME, Smith VP, Spiridou A, Torrance PE, Tzonev SS, Vermaas EH, Walter K, Wu X, Zhang L, Alam MD, Anastasi C, Aniebo IC, Bailey DM, Bancarz IR, Banerjee S, Barbour SG, Baybayan PA, Benoit VA, Benson KF, Bevis C, Black PJ, Boodhun A, Brennan JS, Bridgham JA, Brown RC, Brown AA, Buermann DH, Bundu AA, Burrows JC, Carter NP, Castillo N, Chiara ECM, Chang S, Neil Cooley R, Crake NR, Dada OO, Diakoumakos KD, Dominguez-Fernandez B, Earnshaw DJ, Egbujor UC, Elmore DW, Etchin SS, Ewan MR, Fedurco M, Fraser LJ, Fuentes Fajardo KV, Scott Furey W, George D, Gietzen KJ, Goddard CP, Golda GS, Granieri PA, Green DE, Gustafson DL, Hansen NF, Harnish K, Haudenschild CD, Heyer NI, Hims MM, Ho JT, Horgan AM, Hoschler K, Hurwitz S, Ivanov DV, Johnson MQ, James T, Huw Jones TA, Kang GD, Kerelska TH, Kersey AD, Khrebtukova I, Kindwall AP, Kingsbury Z, Kokko-Gonzales PI, Kumar A, Laurent MA, Lawley CT, Lee SE, Lee X, Liao AK, Loch JA, Lok M, Luo S, Mammen RM, Martin JW, McCauley PG, McNitt P, Mehta P, Moon KW, Mullens JW, Newington T, Ning Z, Ling Ng B, Novo SM, O’Neill MJ, Osborne MA, Osnowski A, Ostadan O, Paraschos LL, Pickering L, Pike AC, Chris Pinkard D, Pliskin DP, Podhasky J, Quijano VJ, Raczy C, Rae VH, Rawlings SR, Chiva Rodriguez A, Roe PM, Rogers J, Rogert Bacigalupo MC, Romanov N, Romieu A, Roth RK, Rourke NJ, Ruediger ST, Rusman E, Sanches-Kuiper RM, Schenker MR, Seoane JM, Shaw RJ, Shiver MK, Short SW, Sizto NL, Sluis JP, Smith MA, Ernest Sohna Sohna J, Spence EJ, Stevens K, Sutton N, Szajkowski L, Tregidgo CL, Turcatti G, Vandevondele S, Verhovsky Y, Virk SM, Wakelin S, Walcott GC, Wang J, Worsley GJ, Yan J, Yau L, Zuerlein M, Mullikin JC, Hurles ME, McCooke NJ, West JS, Oaks FL, Lundberg PL, Klenerman D, Durbin R, Smith AJ (2008) Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456(7218):53–59. https://doi.org/10.1038/nature07517
Bianchi F, Careri M, Mustat L, Malcevschi A, Musci M (2005) Bioremediation of toluene and naphthalene: development and validation of a GC-FID method for their monitoring. Ann Chim 95(7–8):515–524
Bisgard KM, Christie CD, Reising SF, Sanden GN, Cassiday PK, Gomersall C, Wattigney WA, Roberts NE, Strebel PM (2001) Molecular epidemiology of Bordetella pertussis by pulsed-field gel electrophoresis profile: Cincinnati, 1989-1996. J Infect Dis 183(9):1360–1367. https://doi.org/10.1086/319858
Blumberg DA, Lewis K, Mink CM, Christenson PD, Chatfield P, Cherry JD (1993) Severe reactions associated with diphtheria-tetanus-pertussis vaccine: detailed study of children with seizures, hypotonic-hyporesponsive episodes, high fevers, and persistent crying. Pediatrics 91(6):1158–1165
Bouchez V, Caro V, Levillain E, Guigon G, Guiso N (2008) Genomic content of bordetella pertussis clinical isolates circulating in areas of intensive children vaccination. PLoS One 3(6):e2437. https://doi.org/10.1371/journal.pone.0002437
Bouchez V, Brun D, Cantinelli T, Dore G, Njamkepo E, Guiso N (2009) First report and detailed characterization of B. pertussis isolates not expressing pertussis toxin or pertactin. Vaccine 27(43):6034–6041. https://doi.org/10.1016/j.vaccine.2009.07.074
Bouchez V, Baines SL, Guillot S, Brisse S (2018) Complete genome sequences of Bordetella pertussis clinical isolate FR5810 and reference strain Tohama from combined Oxford Nanopore and Illumina sequencing. Microbiol Resour Announc 7(19). https://doi.org/10.1128/mra.01207-18
Bowden KE, Williams MM, Cassiday PK, Milton A, Pawloski L, Harrison M, Martin SW, Meyer S, Qin X, DeBolt C, Tasslimi A, Syed N, Sorrell R, Tran M, Hiatt B, Tondella ML (2014) Molecular epidemiology of the pertussis epidemic in Washington state in 2012. J Clin Microbiol 52(10):3549–3557. https://doi.org/10.1128/jcm.01189-14
Bowden KE, Weigand MR, Peng Y, Cassiday PK, Sammons S, Knipe K, Rowe LA, Loparev V, Sheth M, Weening K, Tondella ML, Williams MM, Blokesch M (2016) Genome structural diversity among 31 Bordetella pertussis isolates from two recent U.S. whooping cough statewide epidemics. Mol Biol Physiol 1(3). https://doi.org/10.1128/mSphere.00036-16
Brinig MM, Cummings CA, Sanden GN, Stefanelli P, Lawrence A, Relman DA (2006) Significant gene order and expression differences in Bordetella pertussis despite limited gene content variation. J Bacteriol 188(7):2375–2382. https://doi.org/10.1128/JB.188.7.2375-2382.2006
Carbonetti NH (2010) Pertussis toxin and adenylate cyclase toxin: key virulence factors of Bordetella pertussis and cell biology tools. Future Microbiol 5:455–469. https://doi.org/10.2217/fmb.09.133
Caro V, Hot D, Guigon G, Hubans C, Arrive M, Soubigou G, Renauld-Mongenie G, Antoine R, Locht C, Lemoine Y, Guiso N (2006) Temporal analysis of French Bordetella pertussis isolates by comparative whole-genome hybridization. Microbes Infect 8(8):2228–2235. https://doi.org/10.1016/j.micinf.2006.04.014
Caro V, Bouchez V, Guiso N (2008) Is the sequenced Bordetella pertussis strain Tohama I representative of the species? J Clin Microbiol 46(6):2125–2128. https://doi.org/10.1128/jcm.02484-07
Centers for Disease Control (2019) Reported pertussis incidence by age group and year | CDC. https://www.cdc.gov/pertussis/surv-reporting/cases-by-age-group-and-year.html#modalIdString_CDCTable_0. Accessed 20 Feb 2019
Cherry JD (1990) ‘Pertussis vaccine encephalopathy’: it is time to recognize it as the myth that it is. JAMA 263(12):1679–1680
Cherry JD (1992) Pertussis: the trials and tribulations of old and new pertussis vaccines. Vaccine 10(14):1033–1038
Cherry JD (1996) Historical review of pertussis and the classical vaccine. J Infect Dis 174(Suppl 3):S259–S263
Cherry JD (2019) The 112-year odyssey of pertussis and pertussis vaccines—mistakes made and implications for the future. J Pediatr Infect Dis Soc. https://doi.org/10.1093/jpids/piz005
Cherry JD, Holtzman AE, Shields WD, Buch D, Nielsen C, Jacobsen V, Christenson PD, Zachau-Christiansen B (1993) Pertussis immunization and characteristics related to first seizures in infants and children. J Pediatr 122(6):900–903
Clark TA (2014) Changing pertussis epidemiology: everything old is new again. J Infect Dis 209(7):978–981. https://doi.org/10.1093/infdis/jiu001
Dalet K, Weber C, Guillemot L, Njamkepo E, Guiso N (2004) Characterization of adenylate cyclase-hemolysin gene duplication in a Bordetella pertussis isolate. Infect Immun 72(8):4874–4877. https://doi.org/10.1128/iai.72.8.4874-4877.2004
Darch SE, McNally A, Harrison F, Corander J, Barr HL, Paszkiewicz K, Holden S, Fogarty A, Crusz SA, Diggle SP (2014) Recombination is a key driver of genomic and phenotypic diversity in a Pseudomonas aeruginosa population during cystic fibrosis infection. Scientific Reports, Published online: 12 January 2015. https://doi.org/10.1038/srep07649
de Gouw D, Hermans PW, Bootsma HJ, Zomer A, Heuvelman K, Diavatopoulos DA, Mooi FR (2014) Differentially expressed genes in Bordetella pertussis strains belonging to a lineage which recently spread globally. PLoS One 9(1):e84523. https://doi.org/10.1371/journal.pone.0084523
de Melker HE, Conyn-van Spaendonck MA, Rumke HC, van Wijngaarden JK, Mooi FR, Schellekens JF (1997) Pertussis in the Netherlands: an outbreak despite high levels of immunization with whole-cell vaccine. Emerg Infect Dis 3(2):175–178. https://doi.org/10.3201/eid0302.970211
De Serres G, Boulianne N, Douville Fradet M, Duval B (1995) Pertussis in Quebec: ongoing epidemic since the late 1980s. Can Commun Dis Rep Releve des maladies transmissibles au Canada 21(5):45–48
Diavatopoulos DA, Cummings CA, Schouls LM, Brinig MM, Relman DA, Mooi FR (2005) Bordetella pertussis, the causative agent of whooping cough, evolved from a distinct, human-associated lineage of B. bronchiseptica. PLoS Pathog 1(4):e45. https://doi.org/10.1371/journal.ppat.0010045
Dienstbier A, Pouchnik D, Wildung M, Amman F, Hofacker IL, Parkhill J, Holubova J, Sebo P, Vecerek B (2018) Comparative genomics of Czech vaccine strains of Bordetella pertussis. Pathog Dis 76(7). https://doi.org/10.1093/femspd/fty071
Ding W, Max Planck Institute for Developmental Biology T, Germany, Baumdicker F, Mathematisches Institut A-LUoF, 79104 Freiburg, Germany, Neher RA, Max Planck Institute for Developmental Biology T, Germany, Biozentrum and SIB Swiss Institute of Bioinformatics UoB, 4056 Basel, Switzerland (2017) panX: pan-genome analysis and exploration. Nucleic Acids Res 46(1). https://doi.org/10.1093/nar/gkx977
Du Q, Wang X, Liu Y, Luan Y, Zhang J, Li Y, Liu X, Ma C, Li H, Wang Z, He Q (2016) Direct molecular typing of Bordetella pertussis from nasopharyngeal specimens in China in 2012-2013. Eur J Clin Microbiol Infect Dis 35(7):1211–1214. https://doi.org/10.1007/s10096-016-2655-3
Fleischmann RD, Alland D, Eisen JA, Carpenter L, White O, Peterson J, DeBoy R, Dodson R, Gwinn M, Haft D, Hickey E, Kolonay JF, Nelson WC, Umayam LA, Ermolaeva M, Salzberg SL, Delcher A, Utterback T, Weidman J, Khouri H, Gill J, Mikula A, Bishai W, Jacobs WR Jr, Venter JC, Fraser CM (2002) Whole-genome comparison of Mycobacterium tuberculosis clinical and laboratory strains. J Bacteriol 184(19):5479–5490
Galit SR, Otsuka N, Furuse Y, Almonia DJ, Sombrero LT, Capeding RZ, Lupisan SP, Saito M, Oshitani H, Hiramatsu Y, Shibayama K, Kamachi K (2015) Molecular epidemiology of Bordetella pertussis in the Philippines in 2012-2014. Int J Infect Dis 35:24–26. https://doi.org/10.1016/j.ijid.2015.04.001
Gutacker MM, Smoot JC, Migliaccio CA, Ricklefs SM, Hua S, Cousins DV, Graviss EA, Shashkina E, Kreiswirth BN, Musser JM (2002) Genome-wide analysis of synonymous single nucleotide polymorphisms in Mycobacterium tuberculosis complex organisms: resolution of genetic relationships among closely related microbial strains. Genetics 162(4):1533–1543
Hamidou Soumana I, Linz B, Harvill ET (2017) Environmental origin of the genus Bordetella. Front Microbiol 8. https://doi.org/10.3389/fmicb.2017.00028
Hegerle N, Paris AS, Brun D, Dore G, Njamkepo E, Guillot S, Guiso N (2012) Evolution of French Bordetella pertussis and Bordetella parapertussis isolates: increase of Bordetellae not expressing pertactin. Clin Microbiol Infect 18(9):E340–E346. https://doi.org/10.1111/j.1469-0691.2012.03925.x
Heikkinen E, Kallonen T, Saarinen L, Sara R, King AJ, Mooi FR, Soini JT, Mertsola J, He Q (2007) Comparative genomics of Bordetella pertussis reveals progressive gene loss in Finnish strains. PLoS One 2(9):e904. https://doi.org/10.1371/journal.pone.0000904
Henderson MW, Inatsuka CS, Sheets AJ, Williams CL, Benaron DJ, Donato GM, Gray MC, Hewlett EL, Cotter PA (2012) Contribution of Bordetella filamentous hemagglutinin and adenylate cyclase toxin to suppression and evasion of interleukin-17-mediated inflammation. Infect Immun 80(6):2061–2075. https://doi.org/10.1128/iai.00148-12
Hiramatsu Y, Miyaji Y, Otsuka N, Arakawa Y, Shibayama K, Kamachi K (2017) Significant decrease in Pertactin-deficient Bordetella pertussis isolates, Japan. Emerg Infect Dis 23(4):699–701. https://doi.org/10.3201/eid2304.161575
King AJ, van Gorkom T, Pennings JL, van der Heide HG, He Q, Diavatopoulos D, Heuvelman K, van Gent M, van Leeuwen K, Mooi FR (2008) Comparative genomic profiling of Dutch clinical Bordetella pertussis isolates using DNA microarrays: identification of genes absent from epidemic strains. BMC Genomics 9:311. https://doi.org/10.1186/1471-2164-9-311
King AJ, van Gorkom T, van der Heide HG, Advani A, van der Lee S (2010) Changes in the genomic content of circulating Bordetella pertussis strains isolated from the Netherlands, Sweden, Japan and Australia: adaptive evolution or drift? BMC Genomics 11:64. https://doi.org/10.1186/1471-2164-11-64
King AJ, van der Lee S, Mohangoo A, van Gent M, van der Ark A, van de Waterbeemd B (2013) Genome-wide gene expression analysis of Bordetella pertussis isolates associated with a resurgence in pertussis: elucidation of factors involved in the increased fitness of epidemic strains. PLoS One 8(6):e66150. https://doi.org/10.1371/journal.pone.0066150
Koren S, Schatz MC, Walenz BP, Martin J, Howard J, Ganapathy G, Wang Z, Rasko DA, McCombie WR, Jarvis ED, Phillippy AM (2012) Hybrid error correction and de novo assembly of single-molecule sequencing reads. Nat Biotechnol 30(7):693–700. https://doi.org/10.1038/nbt.2280
Kuo CH, Ochman H (2010) The extinction dynamics of bacterial pseudogenes. PLoS Genet 6(8):e1001050. https://doi.org/10.1371/journal.pgen.1001050
Lam C, Octavia S, Bahrame Z, Sintchenko V, Gilbert GL, Lan R (2012) Selection and emergence of pertussis toxin promoter ptxP3 allele in the evolution of Bordetella pertussis. Infect Genet Evol 12(2):492–495. https://doi.org/10.1016/j.meegid.2012.01.001
Lam C, Octavia S, Ricafort L, Sintchenko V, Gilbert GL, Wood N, McIntyre P, Marshall H, Guiso N, Keil AD, Lawrence A, Robson J, Hogg G, Lan R (2014) Rapid increase in Pertactin-deficient Bordetella pertussis isolates, Australia. Emerg Infect Dis 20(4):626–633. https://doi.org/10.3201/eid2004.131478
Linz B, Ivanov YV, Preston A, Brinkac L, Parkhill J, Kim M, Harris SR, Goodfield LL, Fry NK, Gorringe AR, Nicholson TL, Register KB, Losada L, Harvill ET (2016) Acquisition and loss of virulence-associated factors during genome evolution and speciation in three clades of Bordetella species. BMC Genomics 17(1):767. https://doi.org/10.1186/s12864-016-3112-5
Lynch M (2002) Genomics. Gene duplication and evolution. Science 297(5583):945–947. https://doi.org/10.1126/science.1075472
Magadum S, Banerjee U, Murugan P, Gangapur D, Ravikesavan R (2013) Gene duplication as a major force in evolution. J Genet 92(1):155–161
Mastrantonio P, Spigaglia P, van Oirschot H, van der Heide HG, Heuvelman K, Stefanelli P, Mooi FR (1999) Antigenic variants in Bordetella pertussis strains isolated from vaccinated and unvaccinated children. Microbiology 145(Pt 8):2069–2075. https://doi.org/10.1099/13500872-145-8-2069
Medini D, Donati C, Tettelin H, Masignani V, Rappuoli R (2005) The microbial pan-genome. Curr Opin Genet Dev 15(6):589–594. https://doi.org/10.1016/j.gde.2005.09.006
Miyaji Y, Otsuka N, Toyoizumi-Ajisaka H, Shibayama K, Kamachi K (2013) Genetic analysis of Bordetella pertussis isolates from the 2008–2010 pertussis epidemic in Japan. PLoS One 8(10):e77165. https://doi.org/10.1371/journal.pone.0077165
Mooi FR (2010) Bordetella pertussis and vaccination: the persistence of a genetically monomorphic pathogen. Infect Genet Evol 10(1):36–49. https://doi.org/10.1016/j.meegid.2009.10.007
Mooi FR, van Oirschot H, Heuvelman K, van der Heide HG, Gaastra W, Willems RJ (1998) Polymorphism in the Bordetella pertussis virulence factors P.69/pertactin and pertussis toxin in The Netherlands: temporal trends and evidence for vaccine-driven evolution. Infect Immun 66(2):670–675
Mooi FR, van Loo IH, King AJ (2001) Adaptation of Bordetella pertussis to vaccination: a cause for its reemergence? Emerg Infect Dis 7(3 Suppl):526–528. https://doi.org/10.3201/eid0707.010708
Mooi FR, van Loo IH, van Gent M, He Q, Bart MJ, Heuvelman KJ, de Greeff SC, Diavatopoulos D, Teunis P, Nagelkerke N, Mertsola J (2009) Bordetella pertussis strains with increased toxin production associated with pertussis resurgence. Emerg Infect Dis 15(8):1206–1213. https://doi.org/10.3201/eid1508.081511
Moore DL, Le Saux N, Scheifele D, Halperin SA (2004) Lack of evidence of encephalopathy related to pertussis vaccine: active surveillance by IMPACT, Canada, 1993–2002. Pediatr Infect Dis J 23(6):568–571
Ohta T (1989) Role of gene duplication in evolution. Genome 31(1):304–310
Oxford Nanopore Technologies (2018) Clive G Brown: nanopore community meeting 2018 talk. https://nanoporetech.com/about-us/news/clive-g-brown-nanopore-community-meeting-2018-talk. Accessed 15 Jan 2019
Park J, Zhang Y, Buboltz AM, Zhang X, Schuster SC, Ahuja U, Liu M, Miller JF, Sebaihia M, Bentley SD, Parkhill J, Harvill ET (2012) Comparative genomics of the classical Bordetella subspecies: the evolution and exchange of virulence-associated diversity amongst closely related pathogens. BMC Genomics 13:545. https://doi.org/10.1186/1471-2164-13-545
Parkhill J, Sebaihia M, Preston A, Murphy LD, Thomson N, Harris DE, Holden MTG, Churcher CM, Bentley SD, Mungall KL, Cerdeño-Tárraga AM, Temple L, James K, Harris B, Quail MA, Achtman M, Atkin R, Baker S, Basham D, Bason N, Cherevach I, Chillingworth T, Collins M, Cronin A, Davis P, Doggett J, Feltwell T, Goble A, Hamlin N, Hauser H, Holroyd S, Jagels K, Leather S, Moule S, Norberczak H, O’Neil S, Ormond D, Price C, Rabbinowitsch E, Rutter S, Sanders M, Saunders D, Seeger K, Sharp S, Simmonds M, Skelton J, Squares R, Squares S, Stevens K, Unwin L, Whitehead S, Barrell BG, Maskell DJ (2003) Comparative analysis of the genome sequences of Bordetella pertussis, Bordetella parapertussis and Bordetella bronchiseptica. Nat Genet 35(1):32–40. https://doi.org/10.1038/ng1227
Preston A, Parkhill J, Maskell DJ (2004) The bordetellae: lessons from genomics. Nat Rev Microbiol 2(5):379–390. https://doi.org/10.1038/nrmicro886
Queenan AM, Cassiday PK, Evangelista A (2013) Pertactin-negative variants of Bordetella pertussis in the United States. N Engl J Med 368(6):583–584. https://doi.org/10.1056/NEJMc1209369
Ring N, Abrahams JS, Jain M, Olsen H, Preston A, Bagby S (2018) Resolving the complex Bordetella pertussis genome using barcoded nanopore sequencing. Microb Genom 4(11). https://doi.org/10.1099/mgen.0.000234
Rouli L, Merhej V, Fournier PE, Raoult D (2015) The bacterial pangenome as a new tool for analysing pathogenic bacteria. New Microbes New Infect 7:72–85. https://doi.org/10.1016/j.nmni.2015.06.005
Schmid M, Frei D, Patrignani A, Schlapbach R, Frey JE, Remus-Emsermann MNP, Ahrens CH (2018) Pushing the limits of de novo genome assembly for complex prokaryotic genomes harboring very long, near identical repeats. bioRxiv. https://doi.org/10.1101/300186
Sealey KL, Harris SR, Fry NK, Hurst LD, Gorringe AR, Parkhill J, Preston A (2015) Genomic analysis of isolates from the United Kingdom 2012 pertussis outbreak reveals that vaccine antigen genes are unusually fast evolving. J Infect Dis 212(2):294–301. https://doi.org/10.1093/infdis/jiu665
Serra DO, Conover MS, Arnal L, Sloan GP, Rodriguez ME, Yantorno OM, Deora R (2011) FHA-mediated cell-substrate and cell-cell adhesions are critical for Bordetella pertussis biofilm formation on abiotic surfaces and in the mouse nose and the trachea. PLoS One 6(12):e28811. https://doi.org/10.1371/journal.pone.0028811
Siguier P, Laboratoire de Microbiologie et Génétique Moléculaires UMdR, Centre National de Recherche Scientifique, Toulouse Cedex, France, Gourbeyre E, Laboratoire de Microbiologie et Génétique Moléculaires UMdR, Centre National de Recherche Scientifique, Toulouse Cedex, France, Chandler M, Laboratoire de Microbiologie et Génétique Moléculaires UMdR, Centre National de Recherche Scientifique, Toulouse Cedex, France (2014) Bacterial insertion sequences: their genomic impact and diversity. FEMS Microbiol Rev 38(5):865–891. https://doi.org/10.1111/1574-6976.12067
Sousa C, de Lorenzo V, Cebolla A (1997) Modulation of gene expression through chromosomal positioning in Escherichia coli. Microbiology 143(Pt 6):2071–2078. https://doi.org/10.1099/00221287-143-6-2071
Strebel P, Nordin J, Edwards K, Hunt J, Besser J, Burns S, Amundson G, Baughman A, Wattigney W (2001) Population-based incidence of pertussis among adolescents and adults, Minnesota, 1995-1996. J Infect Dis 183(9):1353–1359. https://doi.org/10.1086/319853
Sun H, Narihiro T, Ma X, Zhang XX, Ren H, Ye L (2019) Diverse aromatic-degrading bacteria present in a highly enriched autotrophic nitrifying sludge. Sci Total Environ 666:245–251. https://doi.org/10.1016/j.scitotenv.2019.02.172
Tazato N, Handa Y, Nishijima M, Kigawa R, Sano C, Sugiyama J (2015) Novel environmental species isolated from the plaster wall surface of mural paintings in the Takamatsuzuka tumulus: Bordetella muralis sp. nov., Bordetella tumulicola sp. nov. and Bordetella tumbae sp. nov. Int J Syst Evol Microbiol 65(12):4830–4838. https://doi.org/10.1099/ijsem.0.000655
van der Zee A, Vernooij S, Peeters M, van Embden J, Mooi FR (1996) Dynamics of the population structure of Bordetella pertussis as measured by IS1002-associated RFLP: comparison of pre- and post-vaccination strains and global distribution. Microbiology 142(Pt 12):3479–3485. https://doi.org/10.1099/13500872-142-12-3479
van Gent M, Heuvelman CJ, van der Heide HG, Hallander HO, Advani A, Guiso N, Wirsing von Konig CH, Vestrheim DF, Dalby T, Fry NK, Pierard D, Detemmerman L, Zavadilova J, Fabianova K, Logan C, Habington A, Byrne M, Lutynska A, Mosiej E, Pelaz C, Grondahl-Yli-Hannuksela K, Barkoff AM, Mertsola J, Economopoulou A, He Q, Mooi FR (2015) Analysis of Bordetella pertussis clinical isolates circulating in European countries during the period 1998-2012. Eur J Clin Microbiol Infect Dis 34(4):821–830. https://doi.org/10.1007/s10096-014-2297-2
Wang F, Grundmann S, Schmid M, Dorfler U, Roherer S, Charles Munch J, Hartmann A, Jiang X, Schroll R (2007) Isolation and characterization of 1,2,4-trichlorobenzene mineralizing Bordetella sp. and its bioremediation potential in soil. Chemosphere 67(5):896–902. https://doi.org/10.1016/j.chemosphere.2006.11.019
Weigand MR, Peng Y, Loparev V, Johnson T, Juieng P, Gairola S, Kumar R, Shaligram U, Gowrishankar R, Moura H, Rees J, Schieltz DM, Williamson Y, Woolfitt A, Barr J, Tondella ML, Williams MM (2016) Complete genome sequences of four Bordetella pertussis vaccine reference strains from serum Institute of India. Genome Announc 4(6). https://doi.org/10.1128/genomeA.01404-16
Weigand MR, Peng Y, Loparev V, Batra D, Bowden KE, Burroughs M, Cassiday PK, Davis JK, Johnson T, Juieng P, Knipe K, Mathis MH, Pruitt AM, Rowe L, Sheth M, Tondella ML, Williams MM (2017) The history of Bordetella pertussis genome evolution includes structural rearrangement. J Bacteriol 199(8). https://doi.org/10.1128/jb.00806-16
Weigand MR, Pawloski LC, Peng Y, Ju H, Burroughs M, Cassiday PK, Davis JK, DuVall M, Johnson T, Juieng P, Knipe K, Loparev VN, Mathis MH, Rowe LA, Sheth M, Williams MM, Tondella ML (2018) Screening and genomic characterization of filamentous hemagglutinin-deficient Bordetella pertussis. Infect Immun 86(4). https://doi.org/10.1128/iai.00869-17
Weigand MR, Williams MM, Peng Y, Kania D, Pawloski LC, Tondella ML (2019) Genomic survey of Bordetella pertussis diversity, United States, 2000-2013. Emerg Infect Dis 25(4):780–783. https://doi.org/10.3201/eid2504.180812
Weiner B, Gomez J, Victor TC, Warren RM, Sloutsky A, Plikaytis BB, Posey JE, van Helden PD, Gey van Pittius NC, Koehrsen M, Sisk P, Stolte C, White J, Gagneux S, Birren B, Hung D, Murray M, Galagan J (2012) Independent large scale duplications in multiple M. tuberculosis lineages overlapping the same genomic region. PLoS One 7(2). https://doi.org/10.1371/journal.pone.0026038
Wenger AM, Peluso P, Rowell WJ, Chang P-C, Hall RJ, Concepcion GT, Ebler J, Fungtammasan A, Kolesnikov A, Olson ND, Toepfer A, Chin C-S, Alonge M, Mahmoud M, Qian Y, Phillippy AM, Schatz MC, Myers G, DePristo MA, Ruan J, Marschall T, Sedlazeck FJ, Zook JM, Li H, Koren S, Carroll A, Rank DR, Hunkapiller MW (2019) Highly-accurate long-read sequencing improves variant detection and assembly of a human genome. bioRxiv, 519025
WHO (2018) Global and regional immunization profile. WHO. https://www.who.int/immunization/monitoring_surveillance/data/gs_gloprofile.pdf?ua=1. Accessed 9 Jan 2019
Williams MM, Sen K, Weigand MR, Skoff TH, Cunningham VA, Halse TA, Tondella ML (2016) Bordetella pertussis strain lacking Pertactin and pertussis toxin. Emerg Infect Dis 22(2):319–322. https://doi.org/10.3201/eid2202.151332
Xu Y, Liu B, Grondahl-Yli-Hannuksila K, Tan Y, Feng L, Kallonen T, Wang L, Peng D, He Q, Zhang S (2015) Whole-genome sequencing reveals the effect of vaccination on the evolution of Bordetella pertussis. Sci Rep 5:12888. https://doi.org/10.1038/srep12888
Zhang W, Qi W, Albert TJ, Motiwala AS, Alland D, Hyytia-Trees EK, Ribot EM, Fields PI, Whittam TS, Swaminathan B (2006) Probing genomic diversity and evolution of Escherichia coli O157 by single nucleotide polymorphisms. Genome Res 16(6):757–767. https://doi.org/10.1101/gr.4759706
Zomer A, Otsuka N, Hiramatsu Y, Kamachi K, Nishimura N, Ozaki T, Poolman J, Geurtsen J (2018) Bordetella pertussis population dynamics and phylogeny in Japan after adoption of acellular pertussis vaccines. Microb Genom. https://doi.org/10.1099/mgen.0.000180
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Ring, N., Abrahams, J.S., Bagby, S., Preston, A., MacArthur, I. (2019). How Genomics Is Changing What We Know About the Evolution and Genome of Bordetella pertussis . In: Fedele, G., Ausiello, C. (eds) Pertussis Infection and Vaccines. Advances in Experimental Medicine and Biology(), vol 1183. Springer, Cham. https://doi.org/10.1007/5584_2019_401
Download citation
DOI: https://doi.org/10.1007/5584_2019_401
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-33248-8
Online ISBN: 978-3-030-33249-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)