Abstract
The organisation of chromatin is first discussed to conclude that nucleosomes play both structural and transcription-regulatory roles. The presence of nucleosomes makes difficult the access of transcriptional factors to their target sequences and the action of RNA polymerases. The histone post-translational modifications and nucleosome remodelling are first discussed, from a historical point of view, as mechanisms to remove the obstacles imposed by chromatin structure to transcription. Instead of reviewing the state of the art of the whole field, this review is centred on some open questions. First, some “non-classical” histone modifications, such as short-chain acylations other than acetylation, are considered to conclude that their relationship with the concentration of metabolic intermediaries might make of them a sensor of the physiological state of the cells. Then attention is paid to the interest of studying chromatin organisation and epigenetic marks at a single nucleosome level as a complement to genome-wide approaches. Finally, as a consequence of the above questions, the review focuses on the presence of multiple histone post-translational modifications on a single nucleosome. The methods to detect them and their meaning, with special emphasis on bivalent marks, are discussed.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Chromatin
- Nucleosome
- Epigenetics
- Histone post-translational modifications
- Metabolism and histone modifications
1 Introduction
Eukaryotic nuclear DNA is not naked, but organized in chromatin, a complex in which histones and non-histone proteins, as well as some RNA molecules, are packed with DNA in nuclei. A somatic human cell, for instance, contains almost 2 m DNA within a nucleus of around 10 μm diameter at the most. It seems evident that this enormous compaction is an obstacle that the nuclear machinery has to surmount to play all the functions required for cell survival: replication, transcription, reparation and recombination of DNA. The present review deals with the mechanisms involved in allowing eukaryotic transcription to occur within the frame of a chromatin context. It will be centred in mammalian genes transcribed by RNA polymerase II and, rather than being a comprehensive review, particular attention will be paid to some open questions. It should be emphasized from the very beginning that, in spite of the above statement on the obstacles that chromatin structure raises to the action of RNA polymerases, chromatin plays a dualistic regulatory role. Its organisation may either repress or allow transcription often in a reversible way and, therefore, chromatin has to be considered as a highly dynamic regulatory complex. The present article starts by a brief account on how the chromatin structure interferes with the action of RNA pol II and how the difficulties are surmounted. These questions are presented from a historical perspective and then we focus on some issues that are attracting a deep attention.
2 The Landscape of Eukaryotic Transcription
The study of eukaryotic transcription would be senseless without considering the landscape in which it occurs, the chromatin. Histones are the main protein counterpart of DNA in chromatin, from both a quantitative and a functional point of view. A histone octamer made by two copies of each inner histones, namely H2A, H2B, H3 and H4, forms a tripartite structure (Arents et al. 1991), around which 147 bp of DNA were subsequently docked into 1.75 left-handed superhelical turns to yield the first atomic-level model of the nucleosome core, the structural unit of chromatin (Arents and Moudrianakis 1993). The co-crystal structure of the nucleosome core particle was subsequently solved (Luger et al. 1997). The results of this newer study contributed the following significant new specific information about nucleosome organisation. A variable length of linker DNA connects successive nucleosome cores. H1 (H5 in some cells), a fifth histone molecule, interacts with 20 bp of linker DNA at the entrance and exit of some nucleosome cores to form the chromatosome (Simpson 1978). While the structure of the nucleosome core crystals are known to a great detail, for the chromatosome, which has not been crystallized to date, only models for its structure have been described. A short but comprehensive review on nucleosome structure, including some chromatosome models, has been recently published and the interested reader is referred to it for details (Cutter and Hayes 2015).
The presence of nucleosomes gives to the chromatin fibres, when observed through the electron microscope at low ionic strength, the classical appearance of “beads on a string”. This aligned arrangement of nucleosomes constitutes the first level of chromatin structure. Increasing ionic strength results in the folding of chromatin, which acquires the appearance of a 30 nm fibre (van Holde 1989), often referred to as the secondary level of chromatin structure (Sajan and Hawkins 2012). Starting with the solenoidal model of Finch and Klug (Finch and Klug 1976), several possibilities have been proposed for the structure of this fibre, but the actual existence of the 30 nm in vivo remains a controversial issue. It has been suggested that, while 30 nm fibres may occur in the diluted chromatin solutions required for experimental analyses, the highly compacted chromatin within the nuclei adopts a disordered and interdigitated state (Eltsov et al. 2008). This and other experimental results have made the very existence of the 30 nm fibre in vivo a somewhat contested question (Grigoryev and Woodcock 2012; Li and Zhu 2015) and pose some doubts as to the actual interest of a debate on the structure of chromatin in diluted solutions, which would be mainly of academic interest. At any rate, the N-terminal tails of histones play a fundamental role in maintaining the internucleosomal interactions that stabilize the secondary structure of chromatin. When the nucleosome structure was described at 2.8 Å resolution, Richmond and his colleagues noticed that the 16–25 segment of the H4 tail, which contains a succession of basic amino acid residues, binds to the flat face of a neighbour nucleosome core in the crystal lattice. The binding is facilitated by the presence of a cluster of acidic amino acids in the H2A/H2B dimer surface, and the authors advanced that these interactions might take place in nuclei to stabilize chromatin superstructure (Luger et al. 1997). Some years later, an elegant piece of work from the same group showed that deletion of the H4 N-terminus dramatically destabilizes the folding of nucleosome arrays and they were able to identify residues 14–19 as responsible for acquisition of superstructure (Dorigo et al. 2003).
Due to the above reasons, the in situ structural organisation of chromatin within the nucleus is attracting more attention. It was early assumed that the 30 nm fibre –or their alternative secondary structures– is progressively folded to yield more compact structures (for a review, see (Maeshima et al. 2014)). It has been proposed that the different chromatin domains are organised in loops, often referred to as the tertiary level of compaction, but these loops do not represent a unique possibility for this structural level, because interdigitating layers of irregularly oriented nucleosomes have also been proposed to occur in interphase nuclei (Sajan and Hawkins 2012).
Curiously enough, the use of several electron microscopy techniques applied to the study of metaphasic chromosomes, which represent the state of highest compaction of chromatin, has prompted Daban and his colleagues to propose the “thin-plate” model. Layers of planar chromatin are organised perpendicular to the chromosome axis and nucleosomes would be irregularly disposed within these plates (Gállego et al. 2009; Daban 2015). If interdigitating layers of nucleosomes actually exist in interphase chromatin, the transition to metaphasic chromosomes would be easily explained.
It is within this complex landscape that eukaryotic transcription occurs. To do this, RNA polymerase has to bind DNA at the transcription start site (TSS) to form the pre-initiation complex together with the basal transcriptional factors. Then, to start transcription, RNA pol II has to unwind the DNA double helix and to process along the template chain. The presence of nucleosomes obstructs all these steps, and the binding of transcriptional factors to DNA, which often circularly embrace the double helix contour, also may be hindered by the presence of a nucleosome. Nevertheless, in many instances, nucleosomes are placed on the TSS or cover the cis regulatory sites, and the gene bodies always display a nucleosomal organisation. For transcription to occur it is, then, necessary that the nucleosomes be eliminated, displaced or structurally reorganised. This is achieved through two interconnected processes: epigenetic modifications and nucleosome remodelling.
3 Removing the Obstacles
The post-translational modification (PTM) of histones was first described more than 50 years ago in the seminal paper by Allfrey et al. (Allfrey et al. 1964). The authors described the acetylation and the methylation of the amino groups of the lysine side chains. They remarked that, as the acetamide group generated by lysine acetylation is not charged, this modification would loosen the interactions between histones and DNA. Histones were considered as repressors of transcription, and so it was assumed that their acetylation may result in a transcriptional activation. On the contrary, histone methylation does not modify the charge of lysyl residues and the relevance of this modification was passed over at the moment. The interest in histone acetylation as a possible activator of transcription led researchers to discover the fundamentals of histone acetylation. The acetyl group is transferred from acetyl-CoA to histones in a reaction catalysed by histone acetyltransferases (HATs) and hydrolytically removed by the histone deacetylases (HDACs). Nevertheless, research on histone acetylation experienced an impasse until 1988, when Peter Loidl proposed a signalling role for this PTM. The author emphasized that the specific acetylation of some residues may represent “a distinct signal for induction or maintenance of certain structural features of chromatin”. Of course, not all the combinations of acetylated lysines ought to be meaningful, but some examples were provided by the author in support of the model, assuming that site specificity of histone acetylation would drive chromatin to transcription, replication or differentiation (Loidl 1988).
The Loidl’s model implied that the enzymes involved in histone acetylation and deacetylation should be multiple, histone- and site-specific. The multiplicity of HATs was first described in our laboratory, working with the yeast Saccharomyces cerevisiae. We found at least three, chromatographically resolved, nuclear HATs, which displayed different histone specificity (López-Rodas et al. 1989, 1991a) and these results were soon extended to other organisms (López-Rodas et al. 1991b; Georgieva et al. 1991). HDACs also proved to display a limited multiplicity, associated with histone specificity (Sendra et al. 1988, 1991). It was also soon checked that HATs were not only histone-specific, but also site-specific. These and other findings aroused a great interest in cloning nuclear HAT genes, a goal which was first achieved in David Allis’ laboratory working with the ciliate Tetrahymena thermophila. The results were of exceptional interest, because the cloned gene was homologous to yeast Gcn5, which codes for a protein known as a transcriptional activator (Brownell et al. 1996), thus establishing a direct link between histone acetylation and transcriptional activation.
Almost contemporarily to the cloning of the Tetrahymena HAT, the cloning of a human HDAC was also reported (Taunton et al. 1996), and since this pioneering work many HAT and HDAC genes from a wide variety of eukaryotes were cloned and characterized. Nuclear HATs may be grouped into five families, based on sequence similarity (for a recent review, see (Wapenaar and Dekker 2016)), while the 18 HDACs described to date are grouped in 4 classes, one of them, class 2, comprising two subclasses (Haery et al. 2015). The seven members of class 3 are called sirtuins, after the first one characterized coded by the yeast Sir2 (silent mating-type information regulation 2) gene (Rine and Herskowitz 1987).
HDACs and HATs are involved in a complex network of interactions with many other cellular components. It is not the purpose of the present paper to analyze in detail the nature and properties of the HAT and HDAC complexes, and the reader may find useful data in some recent reviews (Haery et al. 2015; Wang and Dent 2014; Yang 2015; Su et al. 2016a). Both, HATs and HDACs, possess functional domains that enable them to interact with many other regulatory proteins, so the possibility exists that the components of the complexes change from cell to cell and from one physiological situation to another.
Histone methylation attracted at first less attention. This PTM, which occurs at the side chains of lysine and arginine, does not eliminate the positive charge of these residues and it was, therefore, supposed that it did not modify the histone-DNA interactions. Moreover, the enzymes involved in this histone modification, especially those catalysing removal of methyl groups, proved to be refractory to research. The landscape of histone methylation changed around the end of the last century, mainly due to the work in Allis’ laboratory (Strahl et al. 1999, 2001). The ε-amino group of lysines may be mono- di- or trimethylated, whereas the guanidinium group of arginines can be mono- and dimethylated in vivo. In the last instance, methyl groups may be added symmetrically (i.e., in each of the nitrogen atoms) or asymmetrically (both methyl groups in the same nitrogen). The reaction of methylation is catalysed by protein methyltransferases or histone methyltransferases and uses S-adenosylmethionine as methyl donor.
While histone acetylation was, and still is, associated with transcriptional activation, histone methylation may play activating or repressing roles depending upon the histone and residue modified as first shown by Noma et al. (2001), working with the pericentromeric loci in the yeast Saccharomyces cerevisiae. The subsequent discovery of histone demethylases (Shi et al. 2004) definitively raised the interest on this histone modification. There are two classes of lysine demethylases, the FAD-dependent enzymes, to which the first histone demethylase belongs, and the Jumonji-type family. The catalytic mechanism of the latter enzymes requires a 2-oxoglutarate dependent oxidation of methyl-lysines (for a recent review, see (Nowak et al. 2016)).
Some of these facts, together with knowledge that histones, like many other proteins, may be reversibly phosphorylated at serine or threonine residues, induced Strahl and Allis, extending the Loidl’s signalling hypothesis (Loidl 1988), to propose that “multiple histone modifications, acting in a combinatorial or sequential fashion on one or multiple histone tails, specify unique downstream functions” (Strahl and Allis 2000). This proposal was designated by the authors as the “histone code hypothesis” and, albeit it cannot be understand in a univocal way (Rando 2012) and PTMs of histones seem to operate in a much complex way than first envisaged (Rothbart and Strahl 2014), it had the undeniable merit of raising the interest on histone modifications to the level in which now stands. Some of the histone PTMs have been shown to be heritable, at least through mitotic division, and in this way they fulfil the criteria to be considered, together with DNA methylation at cytosine rings, as epigenetic modifications. In fact, nowadays the adjective epigenetic refers to those mitotically and/or meiotically heritable phenotypic traits that result from changes in gene expression without alterations in the DNA sequence (Berger et al. 2009; Rodríguez-Paredes and Esteller 2011).
Taking into account that there are two copies of each histone in a nucleosome, the PTMs may be symmetrical or asymmetrical. In the former case, both copies of the histone harbour the same modification; in the latter, either one copy is modified and the other is not, or each histone molecule has a different PTM. We will deal with this question later.
For PTMs of histones to play a regulatory role, they have to be reversible. To use a terminology that has gained relevance, mechanisms must exist not only to “write”, but also to “erase” them. Of course, the reversibility of histone PTMs may also result from histone turnover, although this process would be more time-consuming than an enzyme-catalysed removal of the epigenetic mark. But, obviously, writing and erasing do not suffice if histone PTMs have to play a signalling role. Strahl and Allis named one of the headings of their paper as “How is the histone code read?”, the subsequent one being “Who reads the code?” (Strahl and Allis 2000). A great progress has been made in identifying the “readers”, protein molecules carrying one or several domains capable of specifically interacting with a given epigenetic mark or with a combination of marks. Plant homeodomains (PHD) and ATRX-DNMT3-DNMT3L domains interact with unmodified lysine residues. Bromodomains (BRD) are the classical readers of acetylated lysines and the YNL107w, ENL, AF-9, and TFIIF small subunit (YEATS) domains also have recently proven to play this role. Ankyrin repeats, chromodomains, PHDs, bromo-adjacent homology domains, malignant brain tumour domains, ZF-CW, Pro-Trp-Trp-Pro (PWWP) domains, ATRX-DNMT3-DNMT3L domains and Tudor domains, recognize methylated lysines, the latter two being also able to interact with methylated arginine, together with triptophan-aspartic acid repeats, which also read both, methylated and unmodified lysine side chains (Rothbart and Strahl 2014; Su and Denu 2016). It has been recently proposed that some readers, for instance the heterochromatin proteins HP1-α, HP1-β and M-phase phosphoprotein 8, which contain a chromodomain, the histone lysine methyltransferase WHSC1L1, in which a PWWP domain is present, and the Tudor-containing proteins PHF19 and 53BP1 may be phosphorylated at a tyrosyl residue and that this modification alters or even suppress their capacity to recognise methylated histone residues (Irving-Hooper and Binda 2015). In this way, a possibility to regulate recognition of histone PTMs by the corresponding readers has been opened, and this offers more and more versatility to the histone code.
As mentioned above, histone acetylation is usually associated with transcriptionally active chromatin. When this PTM occurs in the N-terminal tails of histones, the neutralization of their positive charges results in loosening the internucleosomal interactions that stabilize higher order structures. Of course, this facilitates transcription, but the presence of nucleosomes may still represent an obstacle for the accession of the transcriptional machinery and transcriptional factors to DNA.
For many years this question was the most serious challenge the chromatin researchers have to face, but it began to be solved by the end of the last century, by a series of initially non-connected experiments, which ended in the discovery of the yeast SWI/SNF remodelling complex. One of the components of this complex, namely Swi2p/Snf2p is an ATPase stimulated by DNA (Laurent et al. 1993) and hydrolysis of ATP provides the energy required to alter the stable structure of nucleosomes. In the subsequent years, more remodelling complexes were described. All of them are formed by an ATPase molecule, accompanied by several regulatory proteins, and they are grouped into four families according to the nature of the ATPase molecule: SWI/SNF, ISWI (imitation of SWI), CHD (chromodomain helicase DNA-binding) and INO80, which have been found in almost all eukaryotes examined to date (for a recent review, see (Längst and Manelyte 2015)). Remodellers may promote the sliding of the histone octamer along the DNA, the loosening of the histone-DNA contacts, the eviction of H2A-H2B dimers or the eviction of the whole octamer. These events make accessible the DNA sequences to transcriptional factors, to RNA pol II and associated factors, to mediator complexes or to any other DNA-interacting molecule required for regulated transcription.
Either the ATPase molecules or their regulatory partners in remodelling complexes possess epigenetic reader domains, as well as domains capable of interacting with whole nucleosomes or naked DNA. In this way, the remodellers may act in combination with epigenetic modifiers to direct their action to specific nucleosomes. On these grounds, it can be understood how the position occupied by a given nucleosome contributes decisively to the regulation of transcription. A nucleosome may be positioned in such a way that hinders transcription, but after the coordinate action of epigenetic modifiers and remodellers the obstacle may be removed. Conversely, these coordinate actions may be used to repress transcription, so remodelling machines are often necessary for both, activation and repression of genes.
It is within this frame of chromatin structure, epigenetic modifications and nucleosome remodelling that eukaryotic transcription occurs. Much has been done in the last decades to unravel these complex relationships, but in spite of all those efforts, much more is still to solve. The following sections deal with some of these questions.
4 Are all the Histone Post-Translational Modifications Meaningful?
4.1 Histone PTMs: Much More Than Acetylation, Methylation and Phosphorylation
If any database is searched for histone PTMs, most of the results retrieved will deal with methylation and acetylation at their N-termini, followed by those describing phosphorylation of serine and threonine side chains. Nevertheless, many other histone PTMs have been described. For instance, histones can be ubiquitinated, sumoylated, ADP-ribosylated, citrullinated and glycosylated and a reasonably high number of articles may be found on these items. These PTMs have been dealt with, especially from the point of view of the complex relationships among the different epigenetic marks, in several recent and excellent reviews (see, for instance, (Rothbart and Strahl 2014; Su and Denu 2016; Lawrence et al. 2016; Noh et al. 2016)) and they will not be further commented here. Reports on other PTMs, such as proline isomerisation, formylation, butyrylation, malonylation, succinylation, glutathionylation, glutarylation, 2-hydroxyisobutyrylation, tyrosine hydroxylation and histidine phosphorylation are found in literature to a lesser degree. Novel PTMs or novel sites for classical modifications are being continuously described, especially through mass spectrometry (MS) methods.
Tan et al., by means of an integrated, mass spectrometry-based proteomics approach, reported in 2011 the occurrence of 67 previously non-described modifications. Most of them corresponded to novel sites for “old” modifications, several of them occurring within the inner residues of the histone octamer, but a novel modification, namely crotonylation of lysine residues (Tan et al. 2011) was described. A question immediately arises in view of this great wealth of histone PTMs: are all of them functionally meaningful? We will try to provide an answer for some of these PTMs in the following paragraphs. In so doing, it is necessary to keep in mind that for a PTM to be functionally significant, apart from writers of this modification, the existence of erasers and of readers would be convenient if that PTM plays a regulatory, reversible role. By saying “convenient” we mean that the existence of erasers may be circumvented by the passive removing of the modification by histone turnover and the existence of readers is not necessary if a given PTM interferes with the acquisition of another mark with a different function (Schmitges et al. 2011).
All five histones are formylated at some of their lysyl residues, but this modification results from a non-enzymatic reaction after damage of DNA (Jiang et al. 2007) and, although the possibility that an enzyme-driven formylation using formyl-tetrahydrofolate cannot be discarded (Lin et al. 2012), the hypothetical enzyme involved has not been identified. Due to the absence of definite writers, erasers and readers, and of a clear role for histone formylation, this PTM will not be further considered in the present review.
Aminylation also represents a controversial histone PTM. That neologism defines the covalent binding of primary amines to protein glutaminyl residues by the reaction catalysed by transglutaminases (TGs) (Lai et al. 2016), an acyl transfer reaction in which the γ-carboxamide group of a glutamine residue in a protein acts as acyl donor and several primary amines may act as acceptors. The side chain of glutamine results linked to the amine through a secondary amide bond, with the release of ammonia. Tissue transglutaminase (TG2), the most abundant animal TG, is present in various subcellular locations, including nuclei (for a review, see (Kuo et al. 2011)), and all four core histones are excellent glutaminyl substrates for TG2 in vitro and 9 out of their 18 glutamines incorporate polyamines. When nucleosomes were used as substrates, only two glutamines from the N-terminus of H3 and one of H2B are modified (Ballestar et al. 1996).
Several attempts to detect whether incorporation of polyamines to histones take place in vivo have been done and a model in which polyamination of H3 leads to chromatin compaction by increasing the positive charge of the histone tail (McConoughey et al. 2010; Basso and Ratan 2013) has been proposed. Nevertheless, the actual existence of polyaminated H3 in vivo has not yet been demonstrated, nor has the existence of a proper eraser, catalysing an ammoniolytic reaction to restore the glutaminyl residue of the histones. Probably, aminylation cannot be yet included among the bona fide histone functional PTMs.
4.2 Short-Chain Acylations Other Than Acetylation
Histone crotonylation affects lysines in all the four inner histones and linker histone H1. There are 21 crotonylation sites in human inner histones (Tan et al. 2011), most of them residing in the N- or C-termini of the histones that project out of the globular region of the histone octamer. Genome-wide studies also allowed Tan et al. to conclude that crotonylation mainly takes place at both sides of the TSS in active genes (Tan et al. 2011) and similar results were obtained by Sabari et al. when studying the localization of H3K18Cr (Sabari et al. 2015).
The p300 HAT catalyses the crotonylation in vitro of both H3 and H4 using crotonyl-CoA as acyl donor and genome-wide approaches show that peaks of H3K18Cr map together with p300, so it is probable that the enzyme be also responsible for in vivo crotonylation. p300, then, is able to transfer either acetyl or crotonyl groups to histones, the choice between them depending on the relative concentration of both acyl donors (Sabari et al. 2015). Therefore, this histone PTM is related to the metabolic state of the cell, a question that will be dealt with later.
Sirtuins 1, 2 and 3 are able to remove crotonyl marks from histone peptides in vitro, and there are some indications that Sirt3 may be able to do so in vivo, because knocking down the SIRT3 gene in HeLa cells results in the increase of global crotonylated histones and of H3K4Cr (Bao et al. 2014). Moreover, ChIP analyses with a pan anti-KCr antibody in some selected genes, which had been reported to be enriched in Sirt3 in U2OS cells (Iwahara et al. 2012), showed that the level of crotonylated histones increases near their TSSs (Bao et al. 2014).
Human Taf1, which contains two bromodomains, is able to interact with peptides containing crotonyllysine through its second bromodomain (Flynn et al. 2015). In a recent review, Su and Denu remarked that it will be interesting to examine whether the YEATS domain also has the structural plasticity to accommodate longer chain acyl groups (Su and Denu 2016). Recently, this question was positively answered by three independent groups of workers (Andrews et al. 2016; Li et al. 2016; Zhao et al. 2016). Kutateladze and her colleagues have shown that Taf14, one on the components of the yeast transcription factors TFIID and TFIIF that contains a YEATS domain, is the sole H3K9Cr reader in the yeast Saccharomyces cerevisiae and that their binding affinity (K d = 9.5 μM) is in the usual range of epigenetic readers (Andrews et al. 2016).
Similar results have been almost simultaneously reported for human AF9 (Li et al. 2016) and YEATS2 a subunit of the HAT ADA Two A-Containing (ATAC) complex (Zhao et al. 2016). AF9 recognizes crotonyllysine with high affinity. The dissociation constant for the crotonylated H3 peptides ranges between 2.1 and 5.7 μM, depending on the particular lysine modified (9 or 18) and the affinity towards the equivalent, acetylated peptides is 2–3 times lower. Other longer acyl derivatives, such as propionyl or butyryl peptides are also recognized by AF9, but the affinity is reduced due to steric constraints. YEATS2 shows a preference for crotonylation in H3K27. The crotonyl-reading activity of AF9 is not limited to crotonylated peptides, because it can also recognize this PTM in nucleosomes, in preference to acetylated nucleosomes. This may explain why AF9 co-localizes with H3K18Cr at actively transcribing loci. Moreover, a YEATS domain-dependent correlation exists between the presence of AF9 and transcription as the mutation F59A, which reduces the affinity of AF9 for crotonyllysine, does not result in gene activation (Li et al. 2016).
Lysine propionylation and butyrylation were first described in 2007 by means of a proteomic approach (Chen et al. 2007), whereas succinylation and malonylation were detected in 2012 by immunological methods and confirmed by MS analyses (Xie et al. 2012). All these four acylations use the corresponding acyl-CoA derivatives, which are important metabolites. Malonylation is the less frequent of these histone PTMs having only been reported to occur in human H2BK116 (Xie et al. 2012), whereas the other three acylations are more widespread among the lysyl residues of the four inner histones (Chen et al. 2007; Xie et al. 2012; Liu et al. 2009; Xu et al. 2014a, b; Goudarzi et al. 2016).
Although these acylations of histones have been studied in some detail in the last years, there are still many unanswered questions. First, only a few data, and often controversial, exist on the nature of the writers of these PTMs. It was first reported that, in contrast to other enzymes tested, such as Tip60, MOF and PCAF, acetyltransferases p300/CBP were able to transfer propionyl groups from its coenzyme A derivative to H4 (Chen et al. 2007) and H3 (Liu et al. 2009) histones in vitro. These acetyltransferases also catalyse the in vitro butyrylation of histones (Chen et al. 2007) and the reaction also takes place when histone octamers or chromatin templates are used as substrates (Goudarzi et al. 2016). p300/CBP have also been shown to catalyse propionylation of some non-histone proteins in vivo, but transfection of human cells with genes encoding MOF, PCAF, Tip60, and HBO1 deacetylases does not enhance the propionylation of proteins (Cheng et al. 2009), a result in accordance with the data of Liu et al. (2009), but conflicting with the results of Leemhuis et al., who found that PCAF may propionylate H3 peptides in vitro (Leemhuis et al. 2008). MYST, another HAT belonging to the same family as MOF and Tip60, is able to bind propionyl-CoA and Denu and his colleagues found after a detailed kinetic analysis that, although the K m is higher than for acetyl-CoA, the k cat is similar for both acyl donors (Berndsen et al. 2007).
Regarding the nature of the writers of succinylation and malonylation of histones, no conclusive data have been yet obtained and, therefore, more research is needed to identify the enzymes responsible for introducing all these histone PTMs.
On the contrary, many reports on the nature of acyl-histone mark erasers have been published. In 2009, Zhang et al. reported that propionylation and butyrylation of yeast histones was enhanced by trichostatin A and sodium butyrate (Zhang et al. 2009). Suberoylanilide hydroxamic acid also enhances butyrylation in neuroblastome (Xu et al. 2014b). These compounds are inhibitors of HDACs from class I and/or II, so these results suggest that these deacetylases may be involved in removing propionyl or butyryl groups from histones.
Nevertheless, sirtuins are the best characterized enzymes catalysing the removal of non-acetyl acyl groups in proteins. It is known since almost 10 years ago that these NAD+-dependent enzymes may depropionylate bacterial proteins (Garrity et al. 2007) and soon afterwards it was shown that they were also able to remove the propionyl group from an H3 peptide containing modified lysine 23 (Liu et al. 2009). The first evidence for the in vivo depropionylating activity of a specific mammalian sirtuin, namely SIRT1, was obtained with non-histone proteins (Cheng et al. 2009), but the most relevant data on the role of sirtuins were obtained in 2011, when it was, almost simultaneously, described that SIRT5 is a demalonylase of non-histone proteins (Peng et al. 2011) and that it has demalonylase and desuccinylase activity towards histone peptides in vitro (Du et al. 2011). These authors also suggested that the enzyme may desuccinylate proteins in vivo. Based on a 3D structural comparison of human SIRT5, which has a low deacetylating activity, and its orthologous Sir2Tm from Thermatoga maritima, which is a potent protein deacetylase, Du et al. reasoned that a peptide containing a lysine acyl derivative carrying an extra carboxyl group would be a substrate of SIRT5 better than an acetyl peptide. To test this hypothesis, they crystallised a complex of SIRT5 with NAD+ and a succinyl peptide and showed that, actually, the γ-carboxyl group of the succinyl moiety fits into the SIRT5 substrate pocket (Du et al. 2011).
SIRT5 is also an efficient desuccinylase of non-histone proteins (Sadhukhan et al. 2016), as well as a demalonylase (Du et al. 2011). When using N-terminal H3 peptides modified at the lysine 9 as substrates, the k cat/K m values for the demalonylation and desuccinylation reaction are, respectively, 6.1 and 4.3 × 103 M−1 s−1, a 1000-fold higher than for deacetylation (Du et al. 2011). Consistent with these in vitro data is the finding that the level of malonylation of both mitochondrial and cytosolic proteins is much higher in SIRT5 knocked-out mice than in wild-type animals (Nishida et al. 2015).
The above results raise an interesting question. SIRT5 was classically considered as a mitochondrial protein. How is this localization reconciled with the fact that cytosolic proteins are demalonylated by the enzyme? Have the data obtained in vitro with histone peptides any functional significance? The answer to these questions is provided taking into account that two SIRT5 isoforms exist. Both of them possess a cleavable mitochondrial signal in their common N-terminus, but the isoform 1-specific C-terminus also contains a cytoplasmic topogenic sequence, which may allow it to be localized in both, mitochondria and cytosol after the N-terminus is cleaved (Matsushita et al. 2011) and the nuclear translocation may occur through the nuclear pores. Actually, the nuclear localization of SIRT5 has recently be demonstrated by western blot analysis of nuclear extracts of livers from wild type and SIRT5 knocked-out mice (Park et al. 2013).
The first report of a reader for propionyllysine was published in 2010, when Vollmuth et al. found that the two bromodomains (BD1 and BD2) of BRD4, a protein from the BET (bromodomain and extraterminal domain) family, were able to bind propionylated peptides, while butyryllysine-containing peptides were much weakly bound. BD2 recognises both acetyllysine ans propionyllysine-containing peptides with comparable affinity, but BD1 is two to threefold more specific for acetyllysine (Vollmuth and Geyer 2010). More recently, Flynn et al. used peptide arrays from the N-terminal region of histone H3 to check the binding of 49 bromodomains (Flynn et al. 2015). Thirty-five out of them were able to bind both, acetyllysine and propionyllysine peptides and those of a subset of bromodomain-containing proteins, namely CECR2, BRD9 and TAF1 not only bind the short propionyl group, but also the larger butyryl chain. As mentioned above, the second bromodomain of TAF1 also recognises crotonyllysine. The analysis of the crystal structure of BRD9 has revealed that the ligand pocket of the bromodomain is flexible enough to accommodate the large butyryl chain with an affinity comparable to that of acetyllysine (Flynn et al. 2015).
There are only a few clues as to the possible functions of the short-chain acylations of histones studied in this section. The propionylation of H3K23 in the leukemia U937 cells decreases during monocytic differentiation, but the acetylation level does not change. This finding led Liu et al. to propose that propionylation level of H3K23 is associated with U937 cell differentiation (Liu et al. 2009). These authors also remarked that, in spite of the similarity in histone acetylation levels among different cell lines, their propionylation levels are markedly different. Taking into account that the writers, erasers and readers of acetylation and propionylation are common (Table 1), these differences among the levels of both histone PTMs might point to a similar and yet distinct role.
The role of histone butyrylation has been recently studied in detail by Goudarzi et al. First, these authors found that the chromatin distribution and dynamic changes of H4K5ac and H4K8ac is clearly different from those of their butyrylated counterparts in mouse spermatogenesis. The use of this biological model allowed them to detect, through genome-wide approaches, that butyrylated H4 maps around TSSs as acetylated H4 does. Although butyrylation of H4 is also associated with active transcription, it seems to play a role different from that of the acetylated histone, as butyrylation of H4K5, but not of H4K8, inhibits the binding of the bromodomain-containing protein Brdt to H4 acetylated peptides (Goudarzi et al. 2016).
4.3 Distinct Histone Acylation: A Sensor of the Metabolic State of the Cell?
The above results suggest that histone acetylation and other acylations may share similar functions, writers, erasers and readers, although some subtle and yet unresolved differences and competitiveness may be found among these PTMs. Obviously the histone code significance may be considerably enhanced in view of these novel modifications and a deep research will be needed to understand their meaning. Meanwhile, we wish to emphasize that the facts that all the histone PTMs mentioned in the present section use coenzyme A derivatives as acyl donors, that all of them are important metabolites, and that acyl-transfer enzymes may in many cases uses several of them, considerably expand our views on the relationships between metabolism and epigenetic modifications of histones. This issue has been the subject of several recent reviews (see, for instance, (Gut and Verdin 2013; Fan et al. 2015; Su et al. 2016b; Janke et al. 2015)).
As remarked by Lin et al. (2012 ), the differences in the ratio between histone acetylation and propionylation in several cell lines (Liu et al. 2009) may be simply due to the different concentrations of both acyl donors, acetyl-CoA and propionyl-CoA in the cells. Actually, in vitro experiments have shown that the ratio crotonyl-CoA/acetyl-CoA largely determines the level of crotonylation or acetylation of histones (Sabari et al. 2015). In our opinion, this is a crucial question. Of course, taking into account that many different histone acylations occur in nuclei it is obvious that the corresponding acyl-CoA substrates ought to be present in this cell compartment, but the diverse metabolic states of the cells may dramatically change the levels of the different nuclear acyl-CoA.
It is first pertinent to examine the metabolic origin of the different nuclear acyl-CoA molecules. First studies suggested that acetyl-CoA can diffuse across the nuclear pore in S. cerevisiae (Takahashi et al. 2006) and this idea has been peacefully extended to all eukaryotes, so it is frequent to speak about the “nucleocytosolic” compartment as opposed to the mitochondrial one when dealing with the distribution of acetyl-CoA. Some data seemed to sustain this assumption. For instance, the linking between the activity of ATP-citrate lyase and histone acetylation (Gut and Verdin 2013) was taken as a proof that the cytosol-generated acetyl-CoA was the substrate for histone acetylation. But, to the best of our belief, the free diffusion of CoA derivatives through the nuclear pore has not yet been unequivocally proven. Moreover, as recently remarked by Boukouris et al. (2016), the energy of hydrolysis of the acetyl-CoA thioester bond is high enough to make the molecule so unstable that it requires being formed in the location in which it has to be used.
The almost simultaneous discovery that ATP-citrate lyase and acetyl-CoA synthetase (Wellen et al. 2009) as well as carnitine acetyltransferase (Madiraju et al. 2009) are also present in nuclei opened a novel perspective. The first two enzymes are typically cytoplasmatic; ATP-citrate lyase cleaves cytosolic citrate to yield acetyl-CoA and oxalacetate and acetyl-CoA synthetase is able to catalyse the formation of acetyl-CoA from acetate. The reaction of carnitine acetyltransferase, a classical mitochondrial enzyme, also yields acetyl-CoA from acetyl-carnitine. Not only these enzymes are also present in nuclei but a link between them and histone acetylation was found (Wellen et al. 2009; Madiraju et al. 2009). More recently, it has been found that the pyruvate dehydrogenase complex is present in nuclei and that it is functional in catalysing the formation of the acetyl-CoA required for histone modification (Sutendra et al. 2014).
These results provided an answer to the dilemma of the origin of nuclear acetyl-CoA, as aptly discussed in a recent review (Boukouris et al. 2016), because the substrates of the above mentioned enzymes can freely diffuse into nuclei. Nevertheless, they leave many things to solve, for instance, the mechanisms by which the mitochondrial pyruvate dehydrogenase complex is exported to nucleus once the mitochondrial topogenic signal has been removed (Sutendra et al. 2014).
Much less is known about the origin of other nuclear short-chain acyl-CoA derivatives required for the non-acetyl histone PTMs mentioned above. It has been long recognized that acetyl-CoA synthetase accepts some other non-acetate molecules, propionate included, although butyrate is not a substrate for the enzyme (Patel and Walt 1987). It has been proposed, but not proven, that butyryl-CoA may be formed in nuclei at the expense of butyrate. Acyl-CoA synthetase 2, which is present in both, nuclei and cytosol of human HCT116 cells (Wellen et al. 2009) has been shown to catalyse the formation of crotonyl-CoA (Sabari et al. 2015).
The documented origin of nuclear acyl-CoA substrates is depicted in Fig. 1. It remains to be determined the origin of butyryl-, succinyl- and malonyl-CoA, including the possibility of their diffusion through the nuclear pores. At any rate, the metabolic origin of the different acyl-CoAs is different, so their relative concentrations may considerably change in response to the diverse metabolic states of the cells. We have already mentioned that in many instances a HAT enzyme may catalyse the transfer of several acyl groups to histone lysines and that the reaction rate may critically depend on the relative concentration of the donors, but no conclusive data on the possible functional role of the different acylations are available. In this regard, it should be mentioned that the level of both, histone acetylation and methylation depends on dietary factors (Gut and Verdin 2013) and that the epigenetic traits may be transgenerationally inherited (Vollmuth and Geyer 2010), although the mechanisms involved in this process are still controversial (Su et al. 2016b). The above considerations suggest that it would be worth studying in depth the dependence of the different histone acylations on the metabolic state of the cells.
Possible origin of the different nuclear short-chain acyl-CoA substrates used in histone acylation. The figure shows the relations between the nucleus (large oval) and the cytosol. The experimentally checked reactions are represented by red arrows (see the text for details on the corresponding enzymes). The putative reactions, including the translocation of coenzyme A derivatives, are denoted by dashed, gray arrows
5 From Local to Genome-Wide Approaches and Back Again
5.1 Early Local Studies
Initial studies on nucleosome remodelling and changes in histone PTMs were carried out at the level of selected loci. The indirect end-labelling technique, which was first described to map DNase I hypersensitive sites in Drosophila heat shock genes (Wu 1980), was soon adapted, in combination with micrococcal nuclease digestion, to map the accessible linker DNA, and so a picture of nucleosome positioning could be obtained. Although the resolution of the technique was low and it could only be applied to relatively small loci, its application allowed studying transcription-related changes in nucleosome positioning (see, for instance (Pérez-Ortín et al. 1987; Matallana et al. 1992)).
The locus-specific analysis of histone PTMs was possible once the chromatin immunoprecipitation (ChIP) technique started to be used (Crane-Robinson et al. 1997). Fragments of chromatin, obtained by micrococcal nuclease digestion or, more frequently, by sonication, were immunofractionated with antibodies raised against a given histone epigenetic mark. The recovery of immunoprecipitated DNA and its analysis allowed deciding which sequences are occupied by nucleosomes containing that epigenetic mark. In this way, the distribution of histone PTMs along a given gene or locus could be studied. Early methods, especially when chromatin fragments encompassing two or three nucleosomes were used, only gave medium resolution maps of the PTMs distribution, but the ChIP assays proved to be very useful to study transcription-associated changes in histone PTMs and to check the validity and limitations of the histone code hypothesis at the level of single genes or small loci. The ChIP assays may also be directed to locate a non-nucleosomal protein (a transcription factor, or a histone modifying enzyme, for instance) with the use of specific antibodies against that protein.
5.2 The Genomic Era
The advent of the omic approaches allowed analysing many aspects of chromatin structure over the entire genome. The analysis starts as in the single-gene approaches, but the identification of the DNA sequences is carried out by omic techniques, involving microarrays or high-throughput sequencing and the corresponding bioinformatic treatment of data. The use of these approaches to study nucleosome occupancy, the distribution of histone variants and PTMs, as well as the dynamic changes in them was dealt with in the excellent review of Rando and Chang (2009).
By means of these genome-wide approaches highly valuable data have been obtained. Regarding nucleosome positioning (Zhang and Pugh 2011), it is a common feature in active genes that a nucleosome-free region (NFR) spans the TSS, with nucleosomes positioned both upstream and downstream. These nucleosomes are correlatively designated with minus and plus signs respectively. A recent study has revealed that in mouse embryo stem cells there exist two types of promoters enriched in the epigenetic mark H3K4me3, which differ in the wideness of the NFR. The wide NFRs contain transcription-associated histone variants and PTMs, but they have a non-canonical chromatin organisation, sensitive to nuclease digestion. The histones are present in heminucleosomes or nucleosomes lacking one or two H2A-H2B dimers, maintained by the action of remodellers (de Dieuleveult et al. 2016). In other cases, the presence of NFRs is due to DNA sequence determinants.
The distribution of histone PTMs was also studied along the human genome (see, for instance, (Wang et al. 2008)) and these studies provided information as to the co-occurrence of epigenetic marks along the genome, as well as to the frequency of the association of these marks with the different regions of the genes in their transcriptionally active or inactive states (Karmodiya et al. 2012).
In spite of the great value of the genome-wide approaches, they have an obvious limitation. In a certain sense, it can be said that the information provided by genome-wide approaches is similar to that given by statistical data. It is possible to know, for instance, that roughly 55% of patients suffering from colorectal cancer will survive at least 5 years after diagnosis. But these data do not give information as to the outcome of the illness of a particular patient, and yet the oncologist has to treat particular patients and not anonymous groups. The doctor needs to know the personal profile of every patient if a personalized medicine has to be applied. In an analogous manner, the knowledge of the common patterns of distribution of nucleosomes and of epigenetic marks at the level of the entire genome is not warrant for assuring that this pattern is displayed by a given gene.
It is obvious that details on chromatin structure and histone PTMs at selected loci can be obtained through genomic-wide approaches, but these methods are too expensive and time-consuming if only information on a few genes is required. Moreover, this information is often needed at different developmental stages, metabolic situations, etc., a need that would do the use of omic methods prohibitive. In other words, the above considerations suggest that the studies at single loci should be developed in parallel with the genome-wide approaches.
5.3 Experimental Determination of Nucleosome Occupancy at Selected Loci
To study in detail the chromatin organisation of a genetic locus it is first necessary to know the positions occupied by the nucleosomes. Micrococcal nuclease protection (MNP, Fig. 2) offer a convenient and affordable way to do so. Formaldehyde cross-linked chromatin is first digested with the nuclease to give fragments of mononucleosomal size. The DNA isolated from these fragments is then used as template for RT-qPCR. To do this, it is essential to design primers defining tiled amplicons of about 100 bp. Obviously, only the amplicons in which both of the primers lie within the template DNA, i.e., those corresponding to micrococcal nuclease-protected regions, will be amplified, and the PCR signal will be proportional to the degree of protection, which, in turn, depends on the nucleosome occupancy. As the efficiency of the different sets of primers will ordinarily be different, a correction should be made by amplifying naked DNA of about 150–200 bp. This may be conveniently prepared by sonication of naked genomic DNA. If the corrected signal is plotted against the position of each amplicon (usually measured at its centre) a graphic depicting the position of the nucleosomes may be obtained (Fig. 2). It has to be noted that nucleosome positioning, especially when results from DNA sequence motifs, is not absolutely strict. In other words, rotational determinants for nucleosome positioning usually define a series of positions differing by 10 bp, rather than a unique position. This is the reason by which more or less bell-shaped curves result in the plotting. Of course, the size of the amplicons influences the accuracy of the results and the maximum of the curve does not necessarily coincide with the position of the dyad axis of the nucleosomes.
The micrococcal nuclease protection (MNP) assay. Cross-linked chromatin is digested with micrococcal nuclease to give mononucleosomal fragments. The most common mechanism of sequence-directed nucleosome positioning involves rotational determinants and, as shown in the top row, each nucleosome actually is a family of 10 bp-spaced nucleosomes. Therefore, the protected DNA extends more than 147 bp, with the protection level diminishing at both sides. A PCR analysis with primers defining a series of tiled amplicons of about 100 bp will give the results depicted in the lower part of the figure. When RT-qPCR is used, and the DNA concentration is plotted against the position of the centre of each amplicons, the resulting graphic roughly depicts the position occupied by the nucleosome or family of nucleosomes
We first used this method in 2011, also known as nucleosome-scanning (Infante et al. 2012), in combination with semi-quantitative PCR, to detect the position of nucleosomes in the promoter of murine Gas1 gene (Sacilotto et al. 2011). An advantage of MNP is that it allows obtaining fine details in the transcription-related changes in nucleosome organisation. For instance, the murine Egr1 gene has resulted to be a good example of the use of this method. This immediate-early gene is expressed to a very low level in the non-stimulated mouse progenitor hepatocyte MLP29cell line, but it is rapidly induced after treating with 12-O-tetradecanoylphorbol-13-acetate (TPA). The RNAPol-ChIP technique, which measures real-time transcriptional rate (Sandoval and Rodríguez 2004), allows detecting a clear increase in transcription as soon as 5 min after adding TPA. Transcription reaches a maximum at 30 min and by 180 min it returns to the basal situation (Fig. 3), because the own product of the gene, once bound to a specific site in the promoter, recruits the NAB1/NAB2 repressors (Tur et al. 2010). By using a MNP assay, we found that the EGR1 site in the promoter of the gene is covered by nucleosome N − 2. Nevertheless, as shown in Fig. 4, soon after TPA induction a remodelling event results in the downstream sliding of N − 2, which allows the accessibility of the product of the gene to its target sequence, thus initiating the events that eventually led to the repression of the gene (Riffo-Campos et al. 2015). Nucleosome N − 1 is also remodelled upon transcription, and results partially evicted (Fig. 4), probably to facilitate the assembly of the mediator complex, that binds the Egr1 promoter between the TSS and −400 (Wang et al. 2005).
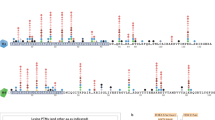
Time-course of changes in nucleosome occupancy in the promoter and proximal coding region of the murine Egr1 gene after TPA induction. The protection against nuclease digestion was determined and plotted against the distance to TSS. To facilitate comparisons, the nuclease protection at t = 0 min was shown in all the panels. The plotted experimental points correspond to the mean ± standard error of three determinations. This research was originally published in Riffo-Campos et al. (2015) © the American Society for Biochemistry and Molecular Biology
5.4 The Identification of Histone PTMs and/or Chromatin-Interacting Factors at Single-Nucleosome Level
The location of epigenetic histone marks and/or protein factors at the level of a given, single nucleosome may be easily obtained through the Nuc-ChIP method (Sacilotto et al. 2011), which has been used by many authors. Essentially, Nuc-ChIP is a classical ChIP assay in which the immunoprecipitation step is carried out with mononucleosomes, isolated as described above for MNP assays (see Fig. 2). The DNA recovered from the immunoprecipitate is then amplified with the appropriate set of primers, corresponding to the amplicons covered by the nucleosomes of choice. In this way the presence of a given histone mark in a specific nucleosome can be detected.
This offers several interesting possibilities. First of all, the method detects a PTM only when it is present in a nucleosome-assembled histone, and not when a free, modified histone is bound to DNA, as it occurs in the above-mentioned wide NFRs (de Dieuleveult et al. 2016). A second example is given by the distribution of the double mark H3S10phK14ac. It has been known for some time that the promoter of immediate-early genes acquire this double mark as a prerequisite to their induction, as shown by classical ChIP analysis of sonicated chromatin in mouse Fos (Cheung et al. 2000; Thomson et al. 2001), Jun (Thomson et al. 2001) and Egr1 (Tur et al. 2010), among other genes. A Nuc-ChIP assay allowed us to specify that H3 phosphoacetylation mainly affects nucleosome N + 1 in murine Egr1 (Riffo-Campos et al. 2015). Working with the same biological model, it was found that the histone activating marks H3K9ac and H4K16ac are characteristic of nucleosome N − 1, while H3K4me3, another activating mark, is mainly found in N + 1 (Fig. 5). The latter finding of Nuc-ChIP agrees with the data obtained from genome-wide analyses, which detected this histone PTM at the start of actively transcribed genes (Kimura 2013), but in other instances, the results obtained by Nuc-ChIP do not coincide with those of genomic approaches. For instance, Nuc-ChIP analysis showed that H3K9me3 is present in N + 1 of murine Egr1 (Fig. 5D), while Wang et al. found that this mark is virtually absent from the environment of the TSS of 1000 active genes (Wang et al. 2008).
Time-dependent changes in histone PTMs in nucleosomes −2, −1 and +1 of Egr1 gene after adding TPA to MLP29 cells. Panels a–h show the levels of different modifications in a representative experiment, plotted as percent of input, for two amplicons in every nucleosome. Error bars represent the standard deviation of three determinations. The times after adding TPA are given in the inset at the right of the lower row. This research was originally published in Riffo-Campos et al. (2015) © the American Society for Biochemistry and Molecular Biology
To further explore this apparent contradiction, we took advantage of another of the possibilities of Nuc-ChIP, namely, the analysis of non-histone proteins bound to a given nucleosome. Following this methodology, we explored the presence of the heterochromatin protein HP1-γ around the Egr1 TSS, because HP1 proteins are known to recognise methylated lysine 9 of H3 through their chromodomain. HP1 proteins are characteristic of heterochromatin and they are typically involved in gene silencing. Recent genome-wide studies describe a mechanism for the silencing caused by HP1 proteins, which co-localize with H3K9me3 (Hiragami-Hamada et al. 2016). Curiously enough, HP1-γ is preferentially found in Egr1 N + 1 (Fig. 6) and it co-maps with H3K9me3 (compare Figs. 5D and 6). Our results agree with those of Vakoc et al., also working with HP1-γ (Vakoc et al. 2005). Therefore, studies at single-nucleosome level, carried out by Nuc-ChIP, may aid to find exceptions to the general rules derived from genome-wide approaches. In the present instance, they suggest a distinct role for the different isoforms of HP1. This question deserves further study; it has been already mentioned that the interaction of HP1-α and HP1-β with H3K9me3 can be modulated by phosphorylation (Irving-Hooper and Binda 2015), and the possibility of a similar modulation in HP1-γ should be explored.
Time-dependent binding of HP1-γ to nucleosomes −1 and +1 of Egr1 gene after adding TPA to MLP29 cells. The experiment was carried out by Nuc-ChIP and the plot was done exactly as those in Fig. 5. The original experiment was carried out in our laboratory by A. L. Riffo-Campos, J. Castillo, G. López-Rodas and L. Franco (unpublished results)
The Nuc-ChIP analysis can be carried out in a time-dependent manner after activation of a gene. In this way, the sequence of histone PTMs can be ascertained at single-nucleosome level. The order in which the different epigenetic events take place during gene activation or repression has represented a classical challenge in chromatin research (see (Mellor 2006) for a review), but early data were obtained by classical ChIP. Kim and Kim (Kim and Kim 2010), studying the β-globin gene and its locus control region, obtained valuable data with ChIP analysis of chromatin digested with micrococcal nuclease to a mononucleosomal level. Nevertheless, as the nucleosome positioning was not determined, the results cannot be explained at single-nucleosome resolution.
The TPA-induced transient induction of Egr1 gene offers a convenient model to carry out these time-dependent, Nuc-ChIP studies (Fig. 5). A general observation is that, with the exception mentioned below, the modification state of the three nucleosomes studies returns to the basal level when the transient expression of the gene ceases. Apart from this general question, several conclusions can be drawn from these studies. First, H3 phosphoacetylation, which is an initial event in the induction of immediate-early genes (Crosio et al. 2003) is not required for sustained transcription, as it clearly diminishes prior to the peak of transcriptional rate of the gene (see Fig. 3). Second, H3K14ac and H4K16ac in the promoter precede other activating modifications, such as H3K9ac. Third, H3K4me3, which, as previously mentioned, fundamentally occurs in N + 1, does not return to the basal level at 180 min after stimulation. Probably, the permanence of this activating PTM, which co-occurs with the repressing mark H3K9me3 may assure an immediate activation of the gene in an eventual second round of transcription (see below for the meaning of the co-occurrence of bivalent marks).
6 The Presence of Multiple Histone PTMs
6.1 How Can Combinatorial Marks in a Single Nucleosome Be Detected?
It has been just shown how the nucleosomes of Egr1 promoter are labelled by several different histone PTMs. Of course, this is not an isolated example and the number of similar cases reported in the literature is too huge to be listed here. Strahl and Davis, when proposing the histone code hypothesis (Strahl and Allis 2000), advanced that a combination of different histone PTMs may specify a given function, so the simultaneous presence of some epigenetic marks in nucleosomes may be functionally significant. It may also be possible that two or more histone distinct PTMs drive a given function when they are sequentially introduced in chromatin. Most of the published work on multiple histone PTMs deal with the combinatorial significance of several epigenetic marks. In the next sections, we will describe the methods used to detect the simultaneous presence of different PTMs, to analyse afterwards some functional properties of combinatorial marks.
Nuc-ChIP, just as classical ChIP, has a limitation. Let us suppose that a single nucleosome (N1 in Fig. 7A) contains two different epigenetic marks in its histones. If a Nuc-ChIP assay was carried out with the antibodies against these marks, and the recovered nucleosomal DNA is PCR-amplified with the primers defining the corresponding amplicons, the results will indicate that these two marks are present in N1. But this result does not necessarily imply a co-occurrence of the two marks in the same nucleosome. It might occur that one of the marks is present in N1 in a given cell while the other mark is present, also in N1 but in other cell of the sample used. The possibility also exists that being both PTMs present in the same cell, one of them occurs in one of the alleles and the second one in the other. In both instances, as shown in Fig. 7B, the results of the Nuc-ChIP analysis will be the same as in the case of co-occurrence of the PTMs (Sacilotto et al. 2011).
A limitation of Nuc-ChIP exemplified by the case of co-occurrence of two different histone PTMs. They may actually co-occur in the same nucleosome N1, as in (A), but one of them may be present in N1 from a subset of cells of the analysed population and the second one in N1 from a different subset of cells (B). A similar situation would result if both marks are distributed between the two alleles in every cell. The finding of marks in N2 and N3, being present in every copy of the corresponding nucleosome, does not involve any ambiguity
To decide between both possibilities, reChIP (Furlan-Magaril et al. 2009), which can be adapted to mononucleosomes and then known as sequential Nuc-ChIP (Sacilotto et al. 2011) may be carried out. The nucleosomes immunoprecipitated with a first antibody were recovered and subjected to a second Nuc-ChIP assay with an antibody against a different protein or epigenetic mark. The final immunoprecipitate was then PCR-analysed so that the amplified sequences correspond to nucleosomes carrying simultaneously both epitopes. Of course, the above method can be scaled up to obtain information at a genome-wide level and several methods have been recently described to improve these approaches (Sadeh et al. 2016; Shema et al. 2016).
6.2 Combinatorial Epigenetic Marks in Nucleosomes
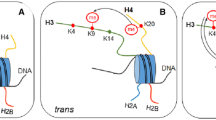
In the last years, evidence in favour of the ubiquitous presence of combined marks is continuously increasing. Combination of PTMs may occur in a trans-histone or in a cis-histone mode (Rothbart and Strahl 2014; Su and Denu 2016). In the former case, two marks are present in different histone molecule (either in the same nucleosome or in neighbouring nucleosomes). In the latter, the marks belong to the same histone molecule. Some histone PTMs occur in a “paired” manner, i. e., when one of the modifications is present in a nucleosome the second one is also frequently present. For instance, the pairs H3K27ac-H3K4me3 and H3K36me3-H3K27ac are characteristic, respectively, of active promoters and enhancers (Xiao et al. 2012) and the pair H3K9ac-H3K14ac is a hallmark of gene activation (Karmodiya et al. 2012).
The presence of a combination of epigenetic marks in a single nucleosome, either on the same histone molecule or in different histone copies within the octamer, requires an appropriate combination of reader domains, which may reside either in the same or in different protein molecules.
There are proteins harbouring two different reader domains that may bind their specific marks. This is, for instance, the case of 14-3-3 proteins, which have been typically known as phospho-binding proteins. Both, mammalian 14-3-3 and their yeast homologues Bmh1 and Bmh2 are able to bind H3S10phK14ac, under stringent conditions (500 mM NaCl), but non-acetylated H3S10ph is bound with much less affinity, as revealed by isothermal titration calorimetry (Walter et al. 2008). Human BPTF (bromodomain PHD finger transcription factor) specifically recognises the pair H3K4me3-H4K16ac in a single nucleosome and the structural basis for this specificity has been established (Ruthenburg et al. 2011). The above examples illustrate the existence of readers for cis- and trans-histone combinations of marks.
Another recent example of a protein that recognises two marks is zinc finger MYND (Myeloid, Nervy and DEAF-1)-type containing 8 (ZMYND8), which specifically binds the combination of activating marks H3.1K36me2/H4K16ac. A structural basis for the simultaneous recognition of both marks has been postulated, and it is worth noting that ZMYND8 shows a strong preference for the canonical histone H3.1, when compared with the variant H3.3 (Adhikary et al. 2016). The interested reader may find more examples in the recent authoritative review of Su and Denu (Su and Denu 2016).
In other instances, recognition of combinatorial epigenetic marks is carried out by protein complexes in which there are at least two components, each of them harbouring a different reader domain. Many often, these complexes also contain components with catalytic domains that may either “write” or “erase” other histone PTMs in the same nucleosome or in a neighbour one. For instance, the HBO1 HAT complex possesses three PHD domains in two different subunits, namely ING4/5 and JADE1/2/3. The pull-down experiments of Saksouk et al. (2009) confirmed that the PDH finger domains of ING4/5 specifically bind H3K4me3. The structural bases for this specificity were cleared up when the crystal structure of the ING4PHD-H3K4me3 complex was solved (Hung et al. 2009). The PHD domains of JADE are essential for the association of HBO1 to chromatin (Saksouk et al. 2009). In this way, the HBO1 complex may be recruited to H3K4me3-containing nucleosomes and catalyse the acetylation of H3 (Hung et al. 2009).
The MOZ/MORF HAT complex provides another excellent example of multiple readers distributed among several subunits. The KAT6A/KAT6B subunit, which contains the catalytic HAT activity, also contains two PHD domains capable of recognising acetylated H3. A third PHD domain is present in the BRPF adaptor subunit, which also contains a bromodomain and an H3K36me3-interacting PWWP domain. Finally, an additional PHD domain, which binds H3K4me3, is present in the ING5 subunit (Klein et al. 2014). In this way, a complex network of epigenetic regulation is established, in which the combination of pre-existing histone PTMs determine the further activity of a “writer”.
6.3 The Meaning of Bivalent Marks
Sequential Nuc-ChIP has been of invaluable aid to report the existence of bivalent epigenetic marks, i. e., histone PTMs of opposing meaning (activating and repressing, for instance), that co-occur in the same nucleosome. The presence of these apparently contradictory epigenetic modifications was first reported in some stem cell genes. In an early review, this combination of activating and repressive marks was designed as “bivalent” and the authors reasoned that the coexistence of both marks may play the role of priming the genes both to expression or silencing as demanded by differentiation to specific cell lineages (Gan et al. 2007). Since then, the presence of bivalent marks and their association with differentiation and development have been widely documented (for a recent review, see (Harikumar and Meshorer 2015)). Recent examples include the PLZF gene, which encodes a transcription factor that drives T cell differentiation into natural killer T cell lineage. It contains nucleosomes in their promoter harbouring the bivalent marks H3K27me3 and H3K4me3, as determined by sequential ChIP (Dobenecker et al. 2015). More recently, by using an original sequential ChIP protocol at a genomic scale, have identified the widespread occurrence of the same bivalent marks in CD4+ memory T cells (Kinkley et al. 2016).
The co-occurrence of bivalent marks is not a property exclusive to stem cells or to cells involved in developmental commitment, and some examples have been recently found in tumour suppressor genes. By means of classical ChIP analysis, the presence of the activating marks H3K9ac, H3K14ac, H3K4me3 and the silencing mark H3K27me3 has been found in the promoter of the tumour suppressor TXNIP gene, although the results do not allow to know which of these marks occur at a single nucleosome level (Baldan et al. 2015). A genome-wide analysis, conducted in 8 glioma stem cell lines, allowed Lin et al. to conclude that H3K4me3 and H3K27me3 co-map in a set of genes. The suppressor gene SLC17A7, whose overexpression reduces the oncogenic phenotype of glioblastoma cells, is an interesting member of that set of genes (Lin et al. 2015). Sequential ChIP analyses, carried out in our laboratory, detected that the repressing mark H3K9me2 co-occurs in the N-1 nucleosome of the mouse suppressor gene Gas1 with the activating mark H3K27ac and, to a lesser extent, with H3K4me3 and H4R3me2 (Sacilotto et al. 2011).
Although sequential ChIP has not been done to assess the co-occurrence of bivalent marks at single-nucleosome level, Fig. 5 shows that H3K4me3 and H3K27me3 are simultaneously present in the region of nucleosome +1 of murine Egr1. The functional meaning of these bivalent marks in suppressor genes and, perhaps, in immediate-early genes may be explained by the analogy proposed in Fig. 1 of the review by Harikumar and Meshorer (2015). A bivalent promoter is comparable to an anchored boat with the sails up. Should the sails be lowered, the boat will remain still, but if the anchor is removed, she will immediately sail. In a bivalent gene, if the repressing mark is erased, transcription may immediately start, but if the activating mark is removed, the gene will be fixed in a repressed state.
It should be mentioned that the recently described method for the traceless synthesis of asymmetrically bivalent nucleosomes (Lechner et al. 2016) may allow obtaining valuable data on the mechanisms by which histone PTMs cross-talk.
7 Concluding Remarks
The field of histone PTMs is a rapidly changing area of research and every novel finding poses new questions to solve. Some of them have been put forward in the present review. The expanding views on “non-classical” histone PTMs require a deeper analysis of their enzymology as well as of the possible “readers” of these modifications.
The relationships between metabolism and epigenetics, which has attracted a deep attention in the last years, are far to be solved and many questions challenge the researchers in this field.
On the other hand, both the novel PTMs and some contradictory results on the distribution and meaning of the PTMs, which result from comparing genome-wide analyses with local studies, may imply a novel formulation of the histone code.
Abbreviations
- BRD:
-
bromodomain
- ChIP:
-
chromatin immunoprecipitation
- HAT:
-
histone acetyltransferase
- HDAC:
-
histone deacetylase
- HP1:
-
heterochromatin protein 1
- MNP:
-
micrococcal nuclease protection
- NFR:
-
nucleosome-free region
- PCR:
-
polymerase chain reaction
- PHD:
-
plant homeodomain
- PTM:
-
post-translational modification
- RT-qPCR:
-
real time quantitative PCR
- TG:
-
transglutaminase
- TPA:
-
12-O-tetradecanoylphorbol-13-acetate
- TSS:
-
Transcription start site
- YEATS:
-
YNL107w, ENL, AF-9, and TFIIF small subunit
- ZMYND8:
-
zinc finger MYND (Myeloid, Nervy and DEAF-1)-type containing 8
References
Adhikary S, Sanyal S, Basu M, Sengupta I, Sen S, Srivastava DK, Roy S, Das C (2016) Selective recognition of H3.1K36 dimethylation/H4K16 acetylation facilitates the regulation of all-trans-retinoic acid (ATRA)-responsive genes by putative chromatin reader ZMYND8. J Biol Chem 291:2664–2681
Allfrey VG, Faulkner R, Mirsky AE (1964) Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci U S A 315:786–794
Andrews FH, Shinsky SA, Shanle EK, Bridgers JB, Gest A, Tsun IK, Krajewski K, Shi X, Strahl BD, Kutateladze TG (2016) The Taf14 YEATS domain is a reader of histone crotonylation. Nat Chem Biol 12:396–398
Arents G, Moudrianakis EN (1993) Topography of the histone octamer surface: repeating structural motifs utilized in the docking of nucleosomal DNA. Proc Natl Acad Sci U S A 90:10489–10493
Arents G, Burlingame RW, Wang BC, Love WE, Moudrianakis EN (1991) The nucleosomal core histone octamer at 3.1 Å resolution: a tripartite protein assembly and a left-handed superhelix. Proc Natl Acad Sci U S A 88:10148–10152
Baldan F, Mio C, Lavarone E, Di Loreto C, Puglisi F, Damante G, Puppin C (2015) Epigenetic bivalent marking is permissive to the synergy of HDAC and PARP inhibitors on TXNIP expression in breast cancer cells. Oncol Rep 33:2199–2206
Ballestar E, Abad C, Franco L (1996) Core histones are glutaminyl substrates for tissue transglutaminase. J Biol Chem 271:18817–18824
Bao X, Wang Y, Li X, Li X, Liu Z, Yang T (2014) Identification of “erasers” for lysine crotonylated histone marks using a chemical proteomics approach. eLife 3:e02999
Basso M, Ratan RR (2013) Transglutaminase is a therapeutic target for oxidative stress, excitotoxicity and stroke: a new epigenetic kid on the CNS block. J Cereb Blood Flow Metab 33:809–818
Berger SL, Kouzarides T, Shiekhattar R, Shilatifard A (2009) An operational definition of epigenetics. Genes Dev 23:781–783
Berndsen CE, Albaugh BN, Tan S, Denu JM (2007) Catalytic mechanism of a MYST family histone acetyltransferase. Biochemistry 46:623–629
Boukouris AE, Zervopoulos SD, Michelakis ED (2016) Metabolic enzymes moonlighting in the nucleus: metabolic regulation of gene transcription. Trends Biochem Sci 41:712–730
Brownell JE, Zhou J, Ranalli T, Kobayashi R, Edmondson DG, Roth SY, Allis CD (1996) Tetrahymena histone acetyltransferase A: a homolog to yeast Gcn5p linking histone acetylation to gene activation. Cell 84:843–851
Chen Y, Sprung R, Tang Y, Ball H, Sangras B, Kim SC, Falck JR, Peng J, Gu W, Zhao Y (2007) Lysine propionylation and butyrylation are novel post-translational modifications in histones. Mol Cell Proteomics 6:812–819
Cheng Z, Tang Y, Chen Y, Kim S, Liu H, Li SSC, Gu W, Zhao Y (2009) Molecular characterization of propionyllysines in non-histone proteins. Mol Cell Proteomics 8:45–52
Cheung P, Tanner KG, Cheung WL, Sassone-Corsi P, Denu JM, Allis CD (2000) Synergistic coupling of histone H3 phosphorylation and acetylation in response to epidermal growth factor stimulation. Mol Cell 5:905–915
Crane-Robinson C, Hebbes TR, Clayton AL, Thorne AW (1997) Chromosomal mapping of core histone acetylation by immunoselection. Methods 12:48–56
Crosio C, Heitz E, Allis CD, Borrelli E, Sassone-Corsi P (2003) Chromatin remodeling and neuronal response: multiple signaling pathways induce specific histone H3 modifications and early gene expression in hippocampal neurons. J Cell Sci 116:4905–4914
Cutter AR, Hayes JJ (2015) A brief review of nucleosome structure. FEBS Lett 589:2914–2922
Daban J-R (2015) Stacked thin layers of metaphase chromatin explain the geometry of chromosome rearrangements and banding. Sci Rep 5:14891
de Dieuleveult M, Yen K, Hmitou I, Depaux A, Boussouar F, Dargham DB, Jounier S, Humbertclaude H, Ribierre F, Baulard C, Farrell NP, Park B, Keime C, Carrière L, Berlivet S, Gut M, Gut I, Werner M, Deleuze J-F, Olaso R, Aude J-C, Chantalat S, Pugh BF, Gérard M (2016) Genome-wide nucleosome specificity and function of chromatin remodellers in ES cells. Nature 530:113–116
Dobenecker M-W, Kim JK, Marcello J, Fang TC, Prinjha R, Bosselut R, Tarakhovsky A (2015) Coupling of T cell receptor specificity to natural killer T cell development by bivalent histone H3 methylation. J Exp Med 212:297–306
Dorigo B, Schalch T, Bystricky K, Richmond TJ (2003) Chromatin fiber folding: requirement for the histone H4 N-terminal tail. J Mol Biol 327:85–96
Du J, Zhou Y, Su X, Yu JJ, Khan S, Jiang H, Kim JH, Woo J, Kim JH, Choi BH, He B, Chen W, Zhang S, Cerione RA, Auwerx J, Hao Q, Lin H (2011) Sirt5 is a NAD-dependent protein lysine demalonylase and desuccinylase. Science 334:806–809
Eltsov M, Maclellan KM, Maeshima K, Frangakis AS, Dubochet J (2008) Analysis of cryo-electron microscopy images does not support the existence of 30-nm chromatin fibers in mitotic chromosomes in situ. Proc Natl Acad Sci U S A 105:19732–19737
Fan J, Krautkramer KA, Feldman JL, Denu JM (2015) Metabolic regulation of histone post-translational modifications. ACS Chem Biol 10:95–108
Finch JT, Klug A (1976) Solenoidal model for superstructure in chromatin. Proc Natl Acad Sci U S A 73:1897–1901
Flynn EM, Huang OW, Poy F, Oppikofer M, Bellon SF, Tang Y, Cochran AG (2015) A subset of human bromodomains recognizes butyryllysine and crotonyllysine histone peptide modifications. Structure 23:1801–1814
Furlan-Magaril M, Rincón-Arano H, Recillas-Targa F (2009) Sequential chromatin immunoprecipitation protocol: ChIP-reChIP. Methods Mol Biol 543:253–266
Gállego I, Castro-Hartmann P, Caravaca JM, Caño S, Daban JR (2009) Dense chromatin plates in metaphase chromosomes. Eur Biophys J 38:503–522
Gan Q, Yoshida T, McDonald OG, Owens GK (2007) Concise review: epigenetic mechanisms contribute to pluripotency and cell lineage determination of embryonic stem cells. Stem Cells 25:2–9
Garrity J, Gardner JG, Hawse W, Wolberger C, Escalante-Semerena JC (2007) N-lysine propionylation controls the activity of propionyl-CoA synthetase. J Biol Chem 282:30239–30245
Georgieva EI, López-Rodas G, Sendra R, Grobner P, Loidl P (1991) Histone acetylation in Zea mays. II biological significance of post- translational histone acetylation during embryo germination. J Biol Chem 266:18751–18760
Goudarzi A, Zhang D, Huang H, Barral S, Kwon OK, Qi S, Tang Z, Buchou T, Vitte AL, He T, Cheng Z, Montellier E, Gaucher J, Curtet S, Debernardi A, Charbonnier G, Puthier D, Petosa C, Panne D, Rousseaux S, Roeder RG, Zhao Y, Khochbin S (2016) Dynamic competing histone H4 K5K8 acetylation and butyrylation are hallmarks of highly active gene promoters. Mol Cell 62:169–180
Grigoryev SA, Woodcock CL (2012) Chromatin organization – the 30 nm fiber. Exp Cell Res 318:1448–1455
Gut P, Verdin E (2013) The nexus of chromatin regulation and intermediary metabolism. Nature 502:489–498
Haery L, Thompson RC, Gilmore TD (2015) Histone acetyltransferases and histone deacetylases in B- and T-cell development, physiology and malignancy. Genes Cancer 6:184–213
Harikumar A, Meshorer E (2015) Chromatin remodeling and bivalent histone modifications in embryonic stem cells. EMBO Rep 16:1609–1619
Hiragami-Hamada K, Soeroes S, Nikolov M, Wilkins B, Kreuz S, Chen C, De La Rosa-Velázquez IA, Zenn HM, Kost N, Pohl W, Chernev A, Schwarzer D, Jenuwein T, Lorincz M, Zimmermann B, Walla PJ, Neumann H, Baubec T, Urlaub H, Fischle W (2016) Dynamic and flexible H3K9me3 bridging via HP1β dimerization establishes a plastic state of condensed chromatin. Nat Commun 7:11310
Hung T, Binda O, Champagne KS, Kuo AJ, Johnson K, Chang HY, Simon MD, Kutateladze TG, Gozani O (2009) ING4 mediates crosstalk between histone H3 K4 trimethylation and H3 acetylation to attenuate cellular transformation. Mol Cell 33:248–256
Infante JJ, Law GL, Young ET (2012) Analysis of nucleosome positioning using a nucleosome-scanning assay. Methods Mol Biol 833:63–87
Irving-Hooper BK, Binda O (2015) A phosphotyrosine switch controls the association of histone mark readers with methylated proteins. Biochemistry 55:1631–1634
Iwahara T, Bonasio R, Narendra V, Reinberg D (2012) SIRT3 functions in the nucleus in the control of stress-related gene expression. Mol Cell Biol 32:5022–5034
Janke R, Dodson AE, Rine J (2015) Metabolism and epigenetics. Annu Rev Cell Dev Biol 31:473–496
Jiang T, Zhou X, Taghizadeh K, Dong M, Dedon PC (2007) N-formylation of lysine in histone proteins as a secondary modification arising from oxidative DNA damage. Proc Natl Acad Sci U S A 104:60–65
Karmodiya K, Krebs AR, Oulad-Abdelghani M, Kimura H, Tora L (2012) H3K9 and H3K14 acetylation co-occur at many gene regulatory elements, while H3K14ac marks a subset of inactive inducible promoters in mouse embryonic stem cells. BMC Genomics 13:424
Kim K, Kim A (2010) Sequential changes in chromatin structure during transcriptional activation in the β globin LCR and its target gene. Int J Biochem Cell Biol 42:1517–1524
Kimura H (2013) Histone modifications for human epigenome analysis. J Hum Genet 58:439–445
Kinkley S, Helmuth J, Polansky JK, Dunkel I, Gasparoni G, Fröhler S, Chen W, Walter J, Hamann A, Chung HR (2016) reChIP-seq reveals widespread bivalency of H3K4me3 and H3K27me3 in CD4+ memory T cells. Nat Commun 7:12514
Klein BJ, Lalonde M-E, Côté J, Yang XJ, Kutateladze TG (2014) Crosstalk between epigenetic readers regulates the MOZ/MORF HAT complexes. Epigenetics 9:186–193
Kuo TF, Tatsukawa H, Kojima S (2011) New insights into the functions and localization of nuclear transglutaminase 2. FEBS J 278:4756–4767
Lai T-S, Lin C-J, Greenberg CS (2016) Role of tissue transglutaminase-2 (TG2)-mediated aminylation in biological processes. Amino Acids. doi:10.1007/s00726-016-2270-8
Längst G, Manelyte L (2015) Chromatin remodelers: from function to dysfunction. Genes (Basel) 6:299–324
Laurent BC, Treich I, Carlson M (1993) The yeast SNF2/SWI2 protein has DNA-stimulated ATPase activity required for transcriptional activation. Genes Dev 7:583–591
Lawrence M, Daujat S, Schneider R (2016) Lateral thinking: how histone modifications regulate gene expression. Trends Genet 32:42–56
Lechner CC, Agashe ND, Fierz B (2016) Traceless synthesis of asymmetrically modified bivalent nucleosomes. Angew Chem Int Ed 55:2903–2906
Leemhuis H, Packman LC, Nightingale KP, Hollfelder F (2008) The human histone acetyltransferase P/CAF is a promiscuous histone propionyltransferase. Chembiochem 9:499–503
Li G, Zhu P (2015) Structure and organization of chromatin fiber in the nucleus. FEBS Lett 589:2893–2904
Li Y, Sabari BR, Panchenko T, Wen H, Zhao D, Guan H, Wan L, Huang H, Tang Z, Zhao Y, Roeder RG, Shi X, Allis CD, Li H (2016) Molecular coupling of histone crotonylation and active transcription by AF9 YEATS domain. Mol Cell 62:181–193
Lin H, Su X, He B (2012) Protein lysine acylation and cysteine succination by intermediates of energy metabolism. ACS Chem Biol 7:947–960
Lin B, Lee H, Yoon J-G, Madan A, Wayner E, Tonning S, Hothi P, Schroeder B, Ulasov I, Foltz G, Hood L, Cobbs C (2015) Global analysis of H3K4me3 and H3K27me3 profiles in glioblastoma stem cells and identification of SLC17A7 as a bivalent tumor suppressor gene. Oncotarget 6:5369–5381
Liu B, Lin Y, Darwanto A, Song X, Xu G, Zhang K (2009) Identification and characterization of propionylation at histone H3 lysine 23 in mammalian cells. J Biol Chem 284:32288–32295
Loidl P (1988) Towards an understanding of the biological function of histone acetylation. FEBS Lett 227:91–95
López-Rodas G, Tordera V, Sánchez del Pino MM, Franco L (1989) Yeast contains multiple forms of histone acetyltransferase. J Biol Chem 264:19028–19033
López-Rodas G, Tordera V, Sánchez del Pino MM, Franco L (1991a) Subcellular localization and nucleosome specificity of yeast histone acetyltransferases. Biochemistry 30:3728–3732
López-Rodas G, Georgieva EI, Sendra R, Loidl P (1991b) Histone acetylation in Zea mays I: activities of histone acetyltransferases and histone deacetylases. J Biol Chem 266:18745–18750
Luger K, Mäder AW, Richmond RK, Sargent DF, Richmond TJ (1997) Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389:251–260
Madiraju P, Pande SV, Prentki M, Madiraju SRM (2009) Mitochondrial acetylcarnitine provides acetyl groups for nuclear histone acetylation. Epigenetics 4:399–403
Maeshima K, Imai R, Tamura S, Nozaki T (2014) Chromatin as dynamic 10-nm fibers. Chromosoma 123:225–237
Matallana E, Franco L, Pérez-Ortín JE (1992) Chromatin structure of the yeast SUC2 promoter in regulatory mutants. Mol Gen Genet 231:395–400
Matsushita N, Yonashiro R, Ogata Y, Sugiura A, Nagashima S, Fukuda T, Inatome R, Yanagi S (2011) Distinct regulation of mitochondrial localization and stability of two human Sirt5 isoforms. Genes Cells 16:190–202
McConoughey SJ, Basso M, Niatsetskaya ZV, Sleiman SF, Smirnova NA, Langley BC, Mahishi L, Cooper AJL, Antonyak MA, Cerione RA, Li B, Starkov A, Chaturvedi RK, Bea MF, Coppola G, Geschwind DH, Ryu H, Xia L, Iismaa SE, Pallos J, Pasternack R, Hils M, Fan J, Raymond LA, Marsh JL, Thompson LM, Ratan RR (2010) Inhibition of transglutaminase 2 mitigates transcriptional dysregulation in models of Huntington’s disease. EMBO Mol Med 2:349–370
Mellor J (2006) Dynamic nucleosomes and gene transcription. Trends Genet 22:320–329
Nishida Y, Rardin MJ, Carrico C, He W, Sahu AK, Gut P, Najjar R, Fitch M, Hellerstein M, Gibson BW, Verdin E (2015) SIRT5 regulates both cytosolic and mitochondrial protein malonylation with glycolysis as a major target. Mol Cell 59:321–332
Noh KM, Allis CD, Li H (2016) Reading between the lines: “ADD”-ing histone and DNA methylation marks toward a new epigenetic “Sum.” ACS Chem Biol 11:554–563
Noma K, Allis CD, Grewal SIS (2001) Transitions in distinct histone H3 methylation patterns at the heterochromatin domain boundaries. Science 293:1150–1155
Nowak RP, Tumber A, Johansson C, Che KH, Brennan P, Owen D, Oppermann U (2016) Advances and challenges in understanding histone demethylase biology. Curr Opin Chem Biol 33:151–159
Park J, Chen Y, Tishkoff DX, Peng C, Tan M, Dai L, Xie Z, Zhang Y, Zwaans BMM, Skinner ME, Lombard DB, Zhao Y (2013) SIRT5-mediated lysine desuccinylation impacts diverse metabolic pathways. Mol Cell 50:919–930
Patel SS, Walt DR (1987) Substrate specificity of acetyl coenzyme A synthetase. J Biol Chem 262:7132–7134
Peng C, Lu Z, Xie Z, Cheng Z, Chen Y, Tan M, Luo H, Zhang Y, He W, Yang K, Zwaans BMM, Tishkoff D, Ho L, Lombard D, He T-C, Dai J, Verdin E, Ye Y, Zhao Y (2011) The first identification of lysine malonylation substrates and its regulatory enzyme. Mol Cell Proteomics 10:M111.012658
Pérez-Ortín JE, Estruch F, Matallana E, Franco L (1987) Fine analysis of the chromatin structure of the yeast SUC2 gene and of its changes upon derepression. Comparison between the chromosomal and plasmid-inserted genes. Nucleic Acids Res 15:6937–6956
Rando OJ (2012) Combinatorial complexity in chromatin structure and function: revisiting the histone code. Curr Opin Genet Dev 22:148–155
Rando OJ, Chang HY (2009) Genome-wide views of chromatin structure. Annu Rev Biochem 78:245–271
Riffo-Campos ÁL, Castillo J, Tur G, González-Figueroa P, Georgieva EI, Rodríguez JL, López-Rodas G, Rodrigo MI, Franco L (2015) Nucleosome-specific, time-dependent changes in histone modifications during activation of the early growth response 1 (Egr1) gene. J Biol Chem 290:197–208
Rine J, Herskowitz I (1987) Four genes responsible for a position effect on expression from HML and HMR in Saccharomyces cerevisiae. Genetics 116:9–22
Rodríguez-Paredes M, Esteller M (2011) Cancer epigenetics reaches mainstream oncology. Nat Med 17:330–339
Rothbart SB, Strahl BD (2014) Interpreting the language of histone and DNA modifications. Biochim Biophys Acta 1839:627–643
Ruthenburg AJ, Li H, Milne TA, Dewell S, McGinty RK, Yuen M, Ueberheide B, Dou Y, Muir TW, Patel DJ, Allis CD (2011) Recognition of a mononucleosomal histone modification pattern by BPTF via multivalent interactions. Cell 145:692–706
Sabari BR, Tang Z, Huang H, Yong-Gonzalez V, Molina H, Kong HE, Dai L, Shimada M, Cross JR, Zhao Y, Roeder RG, Allis CD (2015) Intracellular crotonyl-CoA stimulates transcription through p300-catalyzed histone crotonylation. Mol Cell 58:203–215
Sacilotto N, Espert A, Castillo J, Franco L, López-Rodas G (2011) Epigenetic transcriptional regulation of the growth arrest-specific gene 1 (Gas1) in hepatic cell proliferation at mononucleosomal resolution. PLoS One 6:e23318
Sadeh R, Launer-Wachs R, Wandel H, Rahat A, Friedman N (2016) Elucidating combinatorial chromatin states at single-nucleosome resolution. Mol Cell 63:1–9
Sadhukhan S, Liu X, Ryu D, Nelson OD, Stupinski JA, Li Z, Chen W, Zhang S, Weiss RS, Locasale JW, Auwerx J, Lin H (2016) Metabolomics-assisted proteomics identifies succinylation and SIRT5 as important regulators of cardiac function. Proc Natl Acad Sci U S A 113:4320–4325
Sajan SA, Hawkins RD (2012) Methods for identifying higher-order chromatin structure. Annu Rev Genomics Hum Genet 13:59–82
Saksouk N, Avvakumov N, Champagne KS, Hung T, Doyon Y, Cayrou C, Paquet E, Ullah M, Landry AJ, Côté V, Yang XJ, Gozani O, Kutateladze TG, Côté J (2009) HBO1 HAT complexes target chromatin throughout gene coding regions via multiple PHD finger interactions with histone H3 tail. Mol Cell 33:257–265
Sandoval J, Rodríguez J (2004) RNAPol-ChIP: a novel application of chromatin immunoprecipitation to the analysis of real-time gene transcription. Nucleic Acids Res 32:e88
Schmitges FW, Prusty AB, Faty M, Stützer A, Lingaraju GM, Aiwazian J, Sack R, Hess D, Li L, Zhou S, Bunker RD, Wirth U, Bouwmeester T, Bauer A, Ly-Hartig N, Zhao K, Chan H, Gu J, Gut H, Fischle W, Müller J, Thomä NH (2011) Histone methylation by PRC2 is inhibited by active chromatin marks. Mol Cell 42:330–341
Sendra R, Rodrigo I, Salvador ML, Franco L (1988) Characterization of pea histone deacetylases. Plant Mol Biol 11:857–866
Sendra R, Salvador ML, Franco L (1991) A chromatin-associated histone deacetylase from pea (Pisum sativum). Plant Sci 78:43–51
Shema E, Jones D, Shoresh N, Donohue L, Ram O, Bernstein BE (2016) Single-molecule decoding of combinatorially modified nucleosomes. Science 352:717–721
Shi Y, Lan F, Matson C, Mulligan P, Whetstine JR, Cole PA, Casero RA, Shi Y (2004) Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119:941–953
Simpson RT (1978) Structure of the chromatosome, a chromatin particle containing 160 base pairs of DNA and all the histones. Biochemistry 17:5524–5531
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403:41–45
Strahl BD, Ohba R, Cook RG, Allis CD (1999) Methylation of histone H3 at lysine 4 is highly conserved and correlates with transcriptionally active nuclei in Tetrahymena. Proc Natl Acad Sci U S A 96:14967–14972
Strahl BD, Briggs SD, Brame CJ, Caldwell JA, Koh SS, Ma H, Cook RG, Shabanowitz J, Hunt DF, Stallcup MR, Allis CD (2001) Methylation of histone H4 at arginine 3 occurs in vivo and is mediated by the nuclear receptor coactivator PRMT1. Curr Biol 11:996–1000
Su Z, Denu JM (2016) Reading the combinatorial histone language. ACS Chem Biol 11:564–574
Su J, Wang F, Cai Y, Jin J (2016a) The functional analysis of histone acetyltransferase MOF in tumorigenesis. Int J Mol Sci 17:99
Su X, Wellen KE, Rabinowitz JD (2016b) Metabolic control of methylation and acetylation. Curr Opin Chem Biol 30:52–60. doi:10.1016/j.cbpa.2015.10.030
Sutendra G, Kinnaird A, Dromparis P, Paulin R, Stenson TH, Haromy A, Hashimoto K, Zhang N, Flaim E, Michelakis ED (2014) A nuclear pyruvate dehydrogenase complex is important for the generation of Acetyl-CoA and histone acetylation. Cell 158:84–97
Takahashi H, McCaffery JM, Irizarry RA, Boeke JD (2006) Nucleocytosolic acetyl-coenzyme A synthetase is required for histone acetylation and global transcription. Mol Cell 23:207–217
Tan M, Luo H, Lee S, Jin F, Yang JS, Montellier E, Buchou T, Cheng Z, Rousseaux S, Rajagopal N, Lu Z, Ye Z, Zhu Q, Wysocka J, Ye Y, Khochbin S, Ren B, Zhao Y (2011) Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell 146:1016–1028
Taunton J, Hassig CA, Schreiber SL (1996) A mammalian histone deacetylase related to the yeast transcriptional regulator Rpd3p. Science 272:408–411
Thomson S, Clayton AL, Mahadevan LC (2001) Independent dynamic regulation of histone phosphorylation and acetylation during immediate-early gene induction. Mol Cell 8:1231–1241
Tur G, Georgieva E, Gagete A, López-Rodas G, Rodríguez J, Franco L (2010) Factor binding and chromatin modification in the promoter of murine Egr1 gene upon induction. Cell Mol Life Sci:4065–4077
Vakoc CR, Mandat SA, Olenchock BA, Blobel GA (2005) Histone H3 lysine 9 methylation and HP1γ are associated with transcription elongation through mammalian chromatin. Mol Cell 19:381–391
van Holde KE (1989) Chromatin. Springer, New York
Vollmuth F, Geyer M (2010) Interaction of propionylated and butyrylated histone H3 lysine marks with Brd4 bromodomains. Angew Chem Int Ed Engl 49:6768–6772
Walter W, Clynes D, Tang Y, Marmorstein R, Mellor J, Berger SL (2008) 14-3-3 interaction with histone H3 involves a dual modification pattern of phosphoacetylation. Mol Cell Biol 28:2840–2849
Wang L, Dent SYR (2014) Functions of SAGA in development and disease. Epigenomics 6:329–339
Wang G, Balamotis MA, Stevens JL, Yamaguchi Y, Handa H, Berk AJ (2005) Mediator requirement for both recruitment and postrecruitment steps in transcription initiation. Mol Cell 17:683–694
Wang Z, Zang C, Rosenfeld JA, Schones DE, Barski A, Cuddapah S, Cui K, Roh T-Y, Peng W, Zhang MQ, Zhao K (2008) Combinatorial patterns of histone acetylations and methylations in the human genome. Nat Genet 40:897–903
Wapenaar H, Dekker FJ (2016) Histone acetyltransferases: challenges in targeting bi-substrate enzymes. Clin Epigenetics 8:59
Wellen KE, Hatzivassiliou G, Sachdeva UM, Bui TV, Cross JR, Thompson CB (2009) ATP-citrate lyase links cellular metabolism to histone acetylation. Science 324:1076–1080
Wu C (1980) The 5′ ends of Drosophila heat shock genes in chromatin are hypersensitive to DNase I. Nature 286:854–860
Xiao S, Xie D, Cao X, Yu P, Xing X, Chen C-C, Musselman M, Xie M, West FD, Lewin HA, Wang T, Zhong S (2012) Comparative epigenomic annotation of regulatory DNA. Cell 149:1381–1392
Xie Z, Dai J, Dai L, Tan M, Cheng Z, Wu Y, Boeke JD, Zhao Y (2012) Lysine succinylation and lysine malonylation in histones. Mol Cell Proteomics 11:100–107
Xu YM, Du JY, Lau ATY (2014a) Posttranslational modifications of human histone H3: an update. Proteomics 14:2047–2060
Xu G, Wang J, Wu Z, Qian L, Dai L, Wan X, Tan M, Zhao Y, Wu Y (2014b) SAHA regulates histone acetylation, butyrylation, and protein expression in neuroblastoma. J Proteome Res 13:4211–4219
Yang XJ (2015) MOZ and MORF acetyltransferases: Molecular interaction, animal development and human disease. Biochim Biophys Acta 1853:1818–1826
Zhang Z, Pugh BF (2011) High-resolution genome-wide mapping of the primary structure of chromatin. Cell 144:175–186
Zhang K, Chen Y, Zhang Z, Zhao Y (2009) Identification and verification of lysine propionylation and butyrylation in yeast core histones using PTMap software. J Proteome Res 8:900–906
Zhao D, Guan H, Zhao Z, Mi W, Wen H, Li Y, Zhao Y, Allis CD, Shi X, Li H (2016) YEATS2 is a selective histone crotonylation reader. Cell Res 26:629–632
Acknowledgments
This work was supported by grants from the Spanish Ministerio de Economía y Competitividad (FIS PI12/02110) and from Generalitat Valenciana (PROMETEO 2013–005). We are very indebted to Prof. Juan R. Viña for attracting our attention to the metabolic enzymes that “moonlight” in the nucleus.
Conflict of Interest
The authors declare no conflict of interests.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Castillo, J., López-Rodas, G., Franco, L. (2017). Histone Post-Translational Modifications and Nucleosome Organisation in Transcriptional Regulation: Some Open Questions. In: Atassi, M. (eds) Protein Reviews. Advances in Experimental Medicine and Biology(), vol 966. Springer, Singapore. https://doi.org/10.1007/5584_2017_58
Download citation
DOI: https://doi.org/10.1007/5584_2017_58
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-6921-5
Online ISBN: 978-981-10-6922-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)