Abstract
Förster resonance energy transfer (FRET) is a powerful spectroscopic technique to study conformational changes of nucleic acids and proteins and their molecular interactions. When combined with a single-molecule approach, FRET has the distinct advantage that it can monitor the conformational heterogeneity and dynamics of individual molecules and enable the observation of short-lived molecular intermediates usually hidden in ensemble experiments. This in turn makes single-molecule FRET an interesting tool for dynamic structural biology.
This chapter presents the principles of single-molecule FRET spectroscopy and the added information it gives compared to ensemble FRET spectroscopy. We describe different experimental implementations, primarily focusing on intensity-based approaches. Fluorescence from single molecules requires careful experimental procedures to maximize the inherently low signal intensity, and meticulous data analysis, which is introduced in this chapter, to quantify FRET detection. We comment on advantages and limitations of the technique, and its strength is illustrated by two application examples.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Conformational dynamics
- Fluorescence microscopy
- FRET spectroscopy
- Quantitative FRET
- Single-molecule FRET (smFRET)
1 Introduction
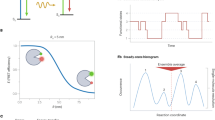
Life is dynamic and understanding its inner workings is a central scientific challenge. The cellular machinery is driven by interactions between DNA, RNA, proteins, and their environment. Molecular structure, binding affinity, and dynamics are each important regulators of these biochemical processes. A full view of this molecular dance, which allows resolving different molecular conformations, how they interconvert, and how molecules interact with each other, would greatly improve our mechanistic understanding of biology. It is unfortunately not currently possible to capture biological processes with atomistic spatial resolution and sub-millisecond time resolution. However, several techniques, alone or in combination, are striving to obtain high spatial and temporal resolutions toward dynamic structural biology [1]. They include, but are not limited to, X-ray crystallography, nuclear magnetic resonance (NMR), electron microscopy, molecular dynamics simulations, and Förster resonance energy transfer (FRET) approaches. The FRET mechanism describes non-radiative energy transfer through dipole–dipole coupling between a donor (D) and an acceptor (A) molecule [2, 3]. The energy transfer therefore strongly depends on the distance between the particular donor and acceptor pair typically in the range of 2–10 nm (Fig. 1). This approach is powerful in the context of dynamic structural biology as it allows distance measurements well below the optical diffraction limit, which can be used to identify and/or refine biomolecular structures. Furthermore, the coupling of FRET with single-molecule microscopy enables an exquisite quantification of dynamics.
(a) Illustration of the FRET mechanism. The donor (D) molecule is excited (blue arrow) and can relax from the excited state D∗ by several channels: radiationless relaxation (dashed gray arrow), fluorescence (photon emission with a radiative rate kD) (green wavy arrow), or energy transfer to the acceptor (A) molecule at rate kFRET. The acceptor molecule can return to the ground state by emitting a photon with a radiative rate kA (red wavy arrow) and/or by radiationless relaxation (dashed gray arrow). The FRET efficiency (E) is given by the rate of energy transfer between the donor and the acceptor, kFRET, divided by the sum of all radiative and non-radiative relaxation rates of the donor. Often both the donor and the acceptor are fluorescent molecules; however, only the donor needs to be fluorescent. (b) Distance dependence of FRET efficiency (E). R is the distance between the donor (colored green) and the acceptor (colored red). R0, the distance where the transfer efficiency is 50%, is the Förster radius specific for the donor–acceptor pair. The three different molecular conformations shown in the plot are illustrative of short, intermediate, and long donor–acceptor distances, respectively. The size of the glow around each D/A molecule is illustrative of the relative contribution of the green donor and the red acceptor for those conformations
In this chapter, we present an overview of smFRET principles and outputs and compare them to ensemble FRET experiments. We discuss experimental implementation basics focusing on intensity-based approaches and data analysis procedures so that the reader is introduced to the advantages and limitations of smFRET. Finally, we illustrate smFRET results by highlighting case studies in nucleic acid folding and protein–nucleic acid interactions.
2 Principles of Ensemble FRET
Most biomolecules are not intrinsically fluorescent and are typically modified by the addition of organic dyes to serve as FRET reporters on conformations. The dyes in a FRET pair are characterized by their chemical properties (molecular structure, size, charge distribution, hydrophobicity) and their electronic properties (spectral overlap of the emission spectrum of the donor and the absorption spectrum of the acceptor, extinction coefficients, quantum yields, and fluorescence lifetimes). The selection of suitable fluorophores for FRET depends on the nature of the biomolecule and experiment.
FRET spectroscopy at the ensemble level has been successfully used for many years to assess conformational changes, molecular interactions, and the average distance between fluorophores [3, 4]. A typical experimental approach based on measuring fluorescence intensities is illustrated in Fig. 2. The FRET efficiency, E, can be obtained by a number of experimental strategies, including quantifying the amount of donor fluorescence in the presence and absence of the acceptor, or by measuring the relative fluorescence of both the donor and the acceptor [5].
Illustration of the contrast between ensemble measurements and single-molecule FRET. (a) An illustration of the ensemble example where many molecules are averaged over both conformations and time. This approach gives information about donor and acceptor chromophore emission using fluorescence spectroscopy yielding an average FRET value for the entire population of molecules. (b) An illustration showing how temporal and conformational heterogeneity in a sample can be resolved using smFRET. FRET time traces (called single-molecule trajectories) yield information about the distribution of molecules in different conformational states as well as the kinetics of state-to-state transitions. Conformational distribution information for a population of molecules is illustrated in the smFRET efficiency histogram revealing three subpopulations. The black and dashed lines illustrate Gaussian fits to the underlying conformational distribution smFRET data. The dotted red line marks the average FRET efficiency 〈E〉 from the histogram, which can be compared to the value derived from ensemble experiments depicted in panel a
Ensemble FRET experiments can yield information on the rate of conformational changes by making use of fast-mixing techniques before equilibrium is established [6]. FRET efficiencies can also be determined by using lifetime measurements, and in certain cases, this can enable obtaining the distribution of molecular distances in solution [7, 8].
Ensemble measurements are inherently limited to averaging molecular properties because an ensemble measurement reports on many molecules at once. This limitation is overcome with single-molecule microscopy.
3 Principles of smFRET: Beyond the Ensemble Average
Since most biological reactions happen via the action of single enzymes, DNA molecules, or RNA molecules, the biomolecular dynamics are inherently stochastic and unsynchronized. The ability to directly follow single-molecule fluorescence enables the observation of biological reactions in a manner that is typically not accessible through the use of conventional ensemble techniques [9]. While an ensemble FRET measurement often yields an average FRET value for the entire population of molecules, smFRET probes each molecule individually over time, resulting in heterogeneous FRET distributions in both time and conformational space (Fig. 2b). There are a number of ways to determine FRET efficiencies at the single-molecule level [10, 11]; it is often advantageous to use both donor and acceptor fluorescence intensities. From the first demonstration on dehydrated samples in 1996 [12], smFRET was developed, to allow solution-based in vitro studies of biomolecular interactions and conformational dynamics of nucleic acids and proteins [13,14,15,16,17]. More recently, smFRET investigations in cells have become possible [18, 19].
3.1 Biomolecular Conformations and Dynamics
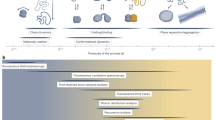
Biomolecular dynamics cover a wide timescale (Fig. 3). smFRET offers the ability to track conformational changes of an individual molecule or biomolecular complex as a function of time (Fig. 2b) covering a sub-microsecond to hours-long timescale. This approach enables the investigation of chain dynamics and domain motion taking place in the sub-microseconds and microseconds range, nucleic acid and protein folding in the microseconds to minutes range, and processes that can take hours or longer, such as protein aggregation [24,25,26]. Assigning conformational states to each of the experimentally identified FRET efficiency states enables the identification of important short-lived transient states, unraveling reaction and dynamic exchange pathways [20], and coupling dynamic and structural information with biochemical functions [27, 28]. Intermediate structures and the allowed pathway dynamics are crucial for understanding the overall mechanism of molecular folding and biomolecular interactions.
Range of biomolecular dynamics. Dynamic processes [14, 20,21,22] (top) and photophysical processes [23] (bottom) correlated with their respective time intervals. smFRET can report on processes occurring over timescales from sub-microseconds to several hours resolving a broad spectrum of dynamics from nucleic acid conformational changes to protein folding and molecular interactions
3.2 Quantitative FRET
The concept of using FRET quantitatively to determine intermolecular distance was first described in 1967 by Stryer and Haugland as the “spectroscopic ruler” and applied at the ensemble level [3]. For many applications, smFRET is often used to report on conformational changes and their dynamics without determining absolute FRET efficiencies. Quantitative smFRET distances are essential for FRET-based structural studies. They yield quantitative information on the conformations of large, heterogeneous, and dynamic biomolecules and their complexes [29].
Converting smFRET measurements to absolute distances requires a number of critical steps that include the determination of absolute FRET efficiencies that are not instrument-dependent (see Sect. 4.3) and modeling of fluorophore dynamics [11, 30,31,32]. A study involving 20 laboratories around the world investigated the precision and accuracy of smFRET measurements [11]. The quantitative assessment of smFRET intensities matched well with the expected structural distances using several double-stranded DNA constructs. Thus, smFRET is an interesting tool for structure determination and validation of flexible and dynamic large structures, which are difficult to solve with other approaches.
4 Experimental Considerations for smFRET
The choice of the smFRET modality, microscope, and experimental schemes, as well as data analysis, depends on several factors including the type of molecule being studied, the nature and scale of the intermolecular interaction, the timescale of the dynamics, and the available equipment. smFRET requires the molecules of interest to be labeled with fluorophores that are typically separated by distances in the range of 2–10 nm and thus relies on robust and site-specific labeling [33,34,35,36]. FRET pairs with high extinction coefficients and high quantum yields are a suitable choice for smFRET experiments. An understanding of the intrinsic photophysical properties of fluorophores is also important as they can affect experimental results [37]. If the interaction involves only two molecules, a single-FRET pair may be sufficient. If the interaction has three or more simultaneous interactions, a three- or four-color FRET scheme may be more appropriate [38].
4.1 Microscope Configurations: Confocal and Widefield Microscopy
There are two general types of single-molecule FRET experimental modalities: confocal microscopy and wide-field total internal reflection fluorescence (TIRF) microscopy (Fig. 4a, c). Confocal geometries allow studying molecules freely diffusing in solution, one at a time (Fig. 4a, b). They are fundamentally limited to an observation time window corresponding to the time it takes for molecules to diffuse through the femtoliter volume excited by a laser, generally in the order of 10 ms or less [39].
Single-molecule FRET experimental schematics. (a) Example of a confocal microscope. The laser enters the back-end of the objective and is focused to a femtoliter volume inside the sample. Freely diffusing molecules are excited as they pass through the laser focal spot. Fluorescence from the molecules is collected through the same objective, focused through a pinhole to reduce out-of-plane background, and sent to point detectors. (b) Sample data from confocal FRET microscopy. Bursts of photons are collected by the donor detector (colored green) and the acceptor detector (colored red). All events with fluorescence intensity above a user-defined threshold (dashed black lines, \( {I}_D^{\mathrm{min}} \) and \( {I}_A^{\mathrm{min}} \)) are interpreted as real events; from left to right: a high FRET, low FRET, acceptor-only, intermediate FRET, and donor-only signal. The yellow lines represent FRET values that make it into the final analysis after removing the low-intensity donor-only and acceptor-only subpopulations. (c) Example of a TIRF wide-field microscope: to perform prism-based TIRF, the incident light from a laser passes from the prism through index matching oil, into the quartz slide, and ultimately enters the sample at a super-critical angle. Alternatively, wide-field TIRF can be accomplished by exciting molecules bound to the bottom surface of a sample chamber in an objective-based geometry. The fluorescence from the sample is collected with an inverted microscope and sent to a sensitive camera. The microscope slide is often attached to a nanometer-precision XY-stage that can move the slide to new positions to collect more data as immobilized molecules photobleach. (d) Sample data from TIRF FRET microscopy. A split-screen image of immobilized donor and acceptor molecules and an example single-molecule trajectory. The green-colored spots are the molecules visible only in the donor channel, and the red-colored spots are the molecules visible only in the acceptor channel. The yellow spots are molecules where the signal is observed in both channels; these are used to determine the FRET efficiency. Donor and acceptor photobleaching is indicated in the single-molecule trajectories by a non-white background
If a chemical process or mechanism occurs over longer timescales, the molecules can be immobilized on a surface allowing for prolonged observation of many hours. Some commonly used methods to attach molecules to a microscope slide involve a biotin–streptavidin linkage [40]. If desired, the slide surface can be passivated to make it resistant to non-specific binding by molecules in the solution, for example, by the use of polyethylene glycol (PEG) [41, 42]. To image these surface-immobilized molecules, scanning confocal microscopy or wide-field microscopy can be used, the latter having the advantage to measure many molecules simultaneously. To selectively excite the molecules on the surface and not others in solution, and thus decrease background fluorescence, the phenomenon of total internal reflection (TIR) is utilized (Fig. 4c, d). TIRF can be implemented in a prism-based geometry and a microscope objective-based geometry.
4.2 Excitation Schemes and Multiplexed Detection
There are various ways to implement smFRET experiments that yield different information on conformational changes and dynamics. The techniques vary both in how the molecules are excited and in what signals are detected. The most common and simplest method for smFRET is single-pair (sp) FRET [12], which is compatible both with confocal and wide-field microscopy. spFRET uses single-color excitation of the donor molecule and simultaneous detection of the emission intensity of both donor and acceptor molecules. The absolute FRET efficiency, E, is given by Eq. 1.
where IFRET is the background-corrected fluorescence intensity of the acceptor dye resulting from energy transfer from the donor (see Sect. 4.3 for details of IFRET determination). \( {I}_{D_{\mathrm{em}}\mid {D}_{\mathrm{ex}}} \) and \( {I}_{A_{\mathrm{em}}\mid {D}_{\mathrm{ex}}} \) are detected to determine FRET efficiencies (Fig. 5a) and denote the intensity of the donor or acceptor emission, respectively, upon donor excitation. γ is a normalization constant that accounts for differences in chromophore quantum-yield (Φ) and detector-efficiency (η) and is given by Eq. 2.
where ηA and ηD are the detector efficiencies for the acceptor and the donor, and ΦA and ΦD are the quantum yields for the acceptor and the donor [11, 43,44,45].
(a) The four types of measurements in an ALEX experiment. Sequential donor and acceptor excitations give rise to the four detectable fluorescence signals, from top to bottom: Aem ∣ Dex (acceptor emission upon donor excitation), Dem ∣ Dex (donor emission upon donor excitation), Aem ∣ Aex (acceptor emission upon acceptor excitation), and Dem ∣ Aex (donor emission upon acceptor excitation), (b) E-S histogram. E (FRET efficiency) versus S (stoichiometry) histogram illustrating the division of measurements into three different categories: D-only (S = 1), distinct FRET populations (S = 0.5), and A-only (S = 0)
spFRET is often used to report on conformational changes through relative FRET efficiencies. Using two lasers to excite both the donor and acceptor molecules yields additional information. One of these approaches is alternating-laser excitation (ALEX) [46, 47], which can be implemented both with confocal and wide-field microscopy. This technique allows superior detection of the low FRET efficiency subpopulation, which can be difficult to distinguish from the donor-only population that is often a dominant contribution around E~0 using spFRET. Donor-only single molecules are created due to acceptor photobleaching. Furthermore, ALEX enables directly identifying eventual acceptor photophysical effects and separating those from conformational changes. With the direct excitation of the acceptor molecule made possible, one can measure two additional quantities compared to spFRET: the intensity of the donor after acceptor excitation, \( {I}_{D_{\mathrm{em}}\mid {A}_{\mathrm{ex}}} \), and the intensity of the acceptor after direct acceptor excitation, \( {I}_{A_{\mathrm{em}}\mid {A}_{\mathrm{ex}}} \) (Fig. 5a). These quantities allow the determination of the stoichiometry ratio, S, which quantifies the ratio of donor to acceptor molecules and is given by Eq. 3.
The stoichiometry ratio allows the set of single-molecule data to be categorized into subpopulations based on the amount of actively fluorescent donor and acceptor molecules present in a single-molecule measurement (Fig. 5b). A value of S = 1 indicates that the smFRET measurement has only a donor present and/or active, S = 0.5 means both donor and acceptor are present/active, and S = 0 means that only the acceptor is present/active. The subpopulation at S = 0.5 is the relevant distribution of FRET values in a sample.
Another way to implement excitation of both the acceptor and the donor is by using pulsed interleaved excitation (PIE), which is essentially a much faster version of ALEX using pulsed lasers. In PIE, laser pulses of different wavelengths are interleaved at a MHz rate such that the fluorescence emission caused by one wavelength laser is complete before the next pulse arrives [48]. This approach is used in combination with confocal microscopy and enables the mapping of each emitted photon to the laser which caused the excitation. PIE naturally enables obtaining fluorescence lifetimes of single molecules as the difference in time between the short laser pulse and the detected photon is measured. The simultaneous collection of fluorescence intensities and lifetimes enables increasing the information content of single-molecule experiments and even more parameters can be collected in an experiment including polarization anisotropy and fluorescence quantum yields [49, 50].
An additional way to increase the information content is to use additional fluorophores. smFRET with one donor and acceptor pair reports on one distance. Multi-color FRET is a way to monitor several distances simultaneously by incorporating more than two spectrally distinct fluorophores capable of transferring energy between them (Fig. 6). Three-color and four-color FRET is especially interesting for monitoring correlated changes in complex molecular dynamics or multi-component binding interactions. Single-molecule three-color FRET enabled the independent monitoring of the dynamics of two arms of a single DNA Holliday junction molecule [51], the conformation of a ligand and its binding to a target [52], and more. Four-color single molecule FRET has been demontrated [53]; at the time of this writing, there are only a few examples. Although information rich, the technical difficulty of executing a three- or four-color smFRET experiment is much higher than a two-color experiment. Multi-color FRET experiments typically make use of ALEX and require the spectral separation of the fluorescence from multiple chromophores.
Illustration of four-, three-, and two-color FRET. The four different colored spheres represent four spectrally distinct fluorophores with different excitation and emission spectra. The donor fluorophore is excited and can transfer energy to all other fluorophores either directly or via a chain of FRET events (colored lines in between spheres). The thickness of the line is proportional to the spectral overlap between fluorophores). Three- and four-color FRET experiments utilize multiple lasers consecutively to directly probe all the fluorophores and check for photobleaching
4.3 Data Analysis of Conformational Distributions and Dynamics
smFRET experiments yield data allowing the experimenter to follow conformational changes in a time-resolved manner. A qualitative view of conformational heterogeneity is obtained by determining the proximity ratio, directly using the measured signals \( {I}_{D_{\mathrm{em}}\mid {I}_{D_{\mathrm{ex}}}} \)and \( {I}_{A_{\mathrm{em}}\mid {D}_{\mathrm{ex}}} \) in Eq. 1, where IFRET is given by \( {I}_{A_{\mathrm{em}}\mid {D}_{\mathrm{ex}}} \) and γ = 1 (Fig. 7a).
Correction of single-molecule FRET data. Each panel shows the various correction parameters applied to single-molecule data sets following the procedure outlined in [11]. (a) First, background subtraction is performed yielding a proximity ratio distribution. (b) Next, donor leakage corrections (α) and indirect acceptor excitation corrections (δ) are performed yielding an apparent FRET distribution. (c) Lastly, detection normalization correction (γ) and excitation normalization correction (β) are applied to yield quantitative FRET distributions that can be turned into a distance distribution
Corrected smFRET data is necessary to obtain structural information about the different FRET populations. In order to transform the raw fluorescence-detected data into apparent and quantitative FRET measurements, various factors need to be accounted for (Fig. 7b, c). The fluorescence intensity of the FRET signal is given by:
as the data is corrected for crosstalk, i.e., leakage (spectral bleed-through) of the donor into the acceptor channel, with the parameter α, and contributions from direct acceptor excitation with the δ-factor [45]. The α- and δ-factors can be experimentally extracted and shift the donor-only population to E = 0 and S = 1 and the acceptor-only populations to E = 1 and S = 0 yielding apparent FRET efficiencies (Fig. 7b) [11]. Quantitative FRET efficiencies can be obtained, using Eq. 1, after determining the γ-factor. In multi-color experiments an additional correction factor, the β-factor, corrects the S ratio [11].
If these corrections are implemented, the FRET efficiency goes from describing qualitative information about relative distances to reporting on quantifiable distances between fluorophores. Quantitative FRET can be used to obtain novel structural information that can be difficult to obtain with other techniques.
Knowledge of the conformational dynamics of a molecular system and the network of state-to-state transition rates inform on the molecule’s free energy landscape. The outcome of such an analysis of smFRET data is a detailed model of relevant conformational transitions that a molecule can undergo in various conditions. The appropriate technique to analyze single-molecule conformational dwell times depends on the timescale of the measurements.
Fast timescale measurements performed on the sub-millisecond level are typically analyzed by calculating time-correlation functions of the signal. The rate and manner the time-correlation function decays from its initial value are related to molecular parameters such as the diffusion constant, the rate of conformational transitions, and the number of conformational macrostates [54, 55]. A maximum likelihood method used to analyze the photon-by-photon arrival times of smFRET experiments [56] has been shown to provide unique insights into conformational transitions of biomolecules, such as elucidating the time of protein folding transitions [24]. Additionally, different methods allow evaluating FRET distributions for the presence of dynamic heterogeneity and determining dynamics. They include the burst variance analysis (BVA) and photon distribution analysis (PDA) methods [57, 58]. For fast dynamics, it is important to consider photophysical phenomena such as photoblinking [10, 59] and fluorescence enhancement due to steric hindrances induced by other molecules, such as protein-induced fluorescence enhancement (PIFE) [60, 61].
Hidden Markov modeling (HMM) is typically used for relatively slower measurements occurring on the millisecond to second timescale regime [62, 63]. HMM analysis ultimately will give an estimate of the number of distinct FRET states in a sample, the connectivity of states, and the transition rate between the states. For example, in the case of an intermolecular binding event, HMM analysis will yield on/off rates and can be used to calculate a binding constant. In more complex systems, dwell-time distributions (a histogram of the length of time spent in a macrostate) constructed from the HMM are fit with single-exponential decays to indicate the approximate rate constant of a state-to-state transition [64, 65].
The appropriate type of smFRET analysis ultimately depends on the nature and timescale of the experiment.
5 smFRET Application Examples
smFRET is a powerful approach for studying the function of biomolecules in heterogeneous mixtures and distinguishing subpopulations of conformers and the dynamics of their inter-conversion. Its application to nucleic acids and proteins has given important mechanistic insights to a number of areas of biological research recently reviewed in [13, 15, 66] and also within nanotechnology [67]. Here, we give an example from nucleic acid folding and DNA–protein interaction to illustrate how smFRET can be used to obtain molecular information in heterogeneous mixtures on reaction pathways and free energy landscapes.
DNA can form a range of secondary structures [68, 69]. A study by Aznauryan et al. focused on the complex folding pathway of the G-rich DNA sequences found at the telomeres [70], which can form specific types of non-canonical DNA secondary structures called G-quadruplexes. These structures can be very stable and are believed to participate in a number of biological processes [71]. By studying the DNA’s conformational dynamics using TIRF-based smFRET, Aznauryan et al. identified the co-existence of several conformations with closely related stabilities and relatively slow conformational changes (Fig. 8a, b) and compiled a view of the folding energy landscape (Fig. 8c). The obtained thermodynamic and kinetic description of the folding process revealed parallel folding pathways through kinetic partitioning and trapping of the most stable conformation. Studies of this folding process using other biophysical techniques complement this view of the folding free energy landscape [72, 73]. DNA conformations, which are detectable by smFRET, can also be very transient [74].
DNA conformation dynamics with smFRET. (a) A smFRET trajectory illustrating conformational changes within a G-quadruplex-forming ssDNA strand. The trajectories were fit with a hidden Markov model using the variational Bayesian inference technique which selects the most likely N-state model. (b) A FRET efficiency histogram constructed from the smFRET trajectories of many molecules. The colored regions describe a four-Gaussian fit to the data, and the black line is the sum of the underlying fits. (c) One-dimensional free energy surface of G-quadruplex folding pathway constructed from smFRET data. The rate constants were extracted from single-exponential fits to the dwell-time distributions. Each structure represents a different G-quadraplex conformation. Adapted with permission from [70]
Proteins are capable of recognizing both specific nucleic acid sequences and structural conformations through direct and indirect readout mechanisms [75]. smFRET has had an enormous impact on the study of these interactions by uncovering key intermediates and important subpopulations not visible by ensemble measurements. A study by Hohlbein et al. used confocal single-molecule FRET to investigate protein/nucleic acid interactions involved with DNA replication in Escherichia coli [76]. The work showed that DNA polymerase I, the enzyme responsible for adding new nucleotides during DNA replication, has three distinct conformations: an open and closed conformation (corresponding to distinct FRET species observed before and after the correct insertion of a complementary nucleotide) and an intermediate conformation observed upon the incorporation of an incorrect nucleotide. These distinct conformations are important for the fidelity of DNA replication (Fig. 9). Later work by the same group used smFRET to obtain accurate distances for refining the structure of a DNA polymerase complex and in this way unveil a novel solution structure exhibiting a sharply bent DNA substrate [27]. The use of smFRET, and its combination with all-atomistic molecular dynamics simulations, is a very promising technique for quantifying structural dynamics. The bent DNA structure was also observed in live bacteria and a mechanism for substrate recognition was proposed [27]. In cellulo smFRET is complicated by the higher background from the crowded cellular environment and labeling difficulties, but is emerging as an exciting approach for studying molecular processes in cells [18, 19].
Polymerase conformations and interactions with DNA studied with smFRET. (a) The structure of the polymerase-I enzyme around double-stranded DNA. The open and closed conformations are observed before and after the correct addition of a nucleotide. The proposed structure of the partially closed conformation is adopted when an incorrect nucleotide is added to the nascent DNA strand. (b) The FRET efficiency histograms corresponding to the correct base pair and incorrect base pair. The dotted lines represent the open and closed states observed when the correct base pair is added, and the dashed line is the partially closed state only observed upon the addition of the incorrect nucleotide. (c) Proposed free energy landscapes which show that binding of a nucleotide stabilizes the polymerase complex with the partially closed conformation, either weakly (incorrect base pair A-dGTP) or strongly (correct base pair A-dTTP), and that equilibration to the fully closed conformation is favored when the correct nucleotide is incorporated. Panels a and c are adapted with permission, and panel b is redrawn using data from [76]
6 Concluding Remarks
Single-molecule fluorescence enables investigating molecular processes and biological events while minimizing both temporal and conformational averaging. Over two decades of smFRET applications have given rise to unprecedented views into interaction mechanisms and conformational heterogeneities and dynamics.
Combining smFRET with other techniques can be used to increase the information content of experiments. For example, smFRET can be used in tandem with PIFE and stacking-induced fluorescence enhancement (SIFI) to increase sensitivity at close distances [61, 77]. Molecular dynamics simulations can be used in conjunction with smFRET for novel structure determination. smFRET can also be combined with force manipulation techniques, such as optical and magnetic tweezers, to monitor local conformational rearrangements amid global molecular manipulation [78].
On its own or in combination with other techniques, smFRET is a very powerful approach. We have given here an introduction to this method for investigating biomolecular structural dynamics.
References
van den Bedem H, Fraser J (2015) Integrative, dynamic structural biology at atomic resolution–it’s about time. Nat Methods 12:307–318. https://doi.org/10.1038/NMETH.3324
Förster T (1948) Zwischenmolekulare Energiewanderung und Fluoreszenz. Ann Phys 437:55–75. https://doi.org/10.1002/andp.19484370105
Stryer L, Hauglandt RP (1967) Energy transfer: a spectroscopic ruler. Proc Natl Acad Sci U S A 58:719. https://doi.org/10.1073/pnas.58.2.719
Latt SA, Cheung T, Blout ER (1964) Energy transfer. A system with relatively fixed donor-acceptor separation. J Am Chem Soc 87:995–1003. https://doi.org/10.1021/ja01083a011
Clegg RM (1992) Fluorescence resonance energy transfer and nucleic acids. Methods Enzymol 211:353–388. https://doi.org/10.1016/0076-6879(92)11020-j
Haas E (2005) The study of protein folding and dynamics by determination of intramolecular distance distributions and their fluctuations using ensemble and single-molecule FRET measurements. ChemPhysChem 6:858–870. https://doi.org/10.1002/cphc.200400617
Haas E, Wilchek M, Katchalski-Katzir E, Steinberg IZ (1975) Distribution of end-to-end distances of oligopeptides in solution as estimated by energy transfer. Proc Natl Acad Sci U S A 72:1807–1811. https://doi.org/10.1073/pnas.72.5.1807
Grinvald A, Haas E, Steinberg IZ (1972) Evaluation of the distribution of distances between energy donors and acceptors by fluorescence decay. Proc Natl Acad Sci 69:2273–2277. https://doi.org/10.1073/pnas.69.8.2273
Tinoco I, Gonzalez RL (2011) Biological mechanisms, one molecule at a time. Genes Dev 25:1205–1231. https://doi.org/10.1101/GAD.2050011
Gopich IV, Szabo A (2012) Theory of the energy transfer efficiency and fluorescence lifetime distribution in single-molecule FRET. Proc Natl Acad Sci U S A 109:7747–7752. https://doi.org/10.1073/pnas.1205120109
Hellenkamp B, Schmid S, Doroshenko O et al (2018) Precision and accuracy of single-molecule FRET measurements – a multi-laboratory benchmark study. Nat Methods 15:669–676. https://doi.org/10.1038/s41592-018-0085-0
Ha T, Enderle T, Ogletree DF et al (1996) Probing the interaction between two single molecules: fluorescence resonance energy transfer between a single donor and a single acceptor. Proc Natl Acad Sci U S A 93:6264–6268. https://doi.org/10.1073/pnas.93.13.6264
Bacic L, Sabantsev A, Deindl S (2020) Recent advances in single-molecule fluorescence microscopy render structural biology dynamic. Curr Opin Struct Biol 65:61–68. https://doi.org/10.1016/J.SBI.2020.05.006
Lerner E, Barth A, Hendrix J et al (2021) FRET-based dynamic structural biology: challenges, perspectives and an appeal for open-science practices. elife 10:1–69. https://doi.org/10.7554/eLife.60416
Lerner E, Cordes T, Ingargiola A et al (2018) Toward dynamic structural biology: two decades of single-molecule Förster resonance energy transfer. Science 359(6373):eaan1133. https://doi.org/10.1126/science.aan1133
Sanders JC, Holmstrom ED (2021) Integrating single-molecule FRET and biomolecular simulations to study diverse interactions between nucleic acids and proteins. Essays Biochem 65:37–49. https://doi.org/10.1042/EBC20200022
Walter NG, Huang CY, Manzo AJ, Sobhy MA (2008) Do-it-yourself guide: how to use the modern single-molecule toolkit. Nat Methods 5:475–489. https://doi.org/10.1038/nmeth.1215
Sustarsic M, Kapanidis AN (2015) Taking the ruler to the jungle: single-molecule FRET for understanding biomolecular structure and dynamics in live cells. Curr Opin Struct Biol 34:52–59. https://doi.org/10.1016/j.sbi.2015.07.001
Asher WB, Geggier P, Holsey MD et al (2021) Single-molecule FRET imaging of GPCR dimers in living cells. Nat Methods 18:397–405. https://doi.org/10.1038/s41592-021-01081-y
Fenwick RB, Esteban-Martín S, Salvatella X (2011) Understanding biomolecular motion, recognition, and allostery by use of conformational ensembles. Eur Biophys J 40(12):1339–1355. https://doi.org/10.1007/S00249-011-0754-8
Fisette O, Lagüe P, Gagné S, Morin S (2012) Synergistic applications of MD and NMR for the study of biological systems. J Biomed Biotechnol. https://doi.org/10.1155/2012/254208
Zhong D (2007) Ultrafast catalytic processes in enzymes. Curr Opin Chem Biol 11:174–181. https://doi.org/10.1016/J.CBPA.2007.02.034
Jaffé HH, Miller AL (1966) The fates of electronic excitation energy. J Chem Educ 43:469–473. https://doi.org/10.1021/ED043P469
Chung HS, Eaton WA (2018) Direct protein folding transition path times from single molecule FRET. Curr Opin Struct Biol 48:30–39. https://doi.org/10.1016/j.sbi.2017.10.007
Kilic S, Felekyan S, Doroshenko O et al (2018) Single-molecule FRET reveals multiscale chromatin dynamics modulated by HP1α. Nat Commun 9. https://doi.org/10.1038/s41467-017-02619-5
Yang O, Ha T (2018) Single-molecule studies of ssDNA-binding proteins exchange. Methods Enzymol 600:463–477. https://doi.org/10.1016/bs.mie.2017.11.017
Craggs TD, Sustarsic M, Plochowietz A et al (2019) Substrate conformational dynamics facilitate structure-specific recognition of gapped DNA by DNA polymerase. Nucleic Acids Res 47:10788–10800. https://doi.org/10.1093/nar/gkz797
De Boer M, Gouridis G, Vietrov R et al (2019) Conformational and dynamic plasticity in substrate-binding proteins underlies selective transport in ABC importers. elife 8. https://doi.org/10.7554/ELIFE.44652
Kapanidis AN, Laurence TA, Nam KL et al (2005) Alternating-laser excitation of single molecules. Acc Chem Res 38:523–533. https://doi.org/10.1021/ar0401348
Beckers M, Drechsler F, Eilert T et al (2015) Quantitative structural information from single-molecule FRET. Faraday Discuss 184:117–129. https://doi.org/10.1039/C5FD00110B
Sindbert S, Kalinin S, Nguyen H et al (2011) Accurate distance determination of nucleic acids via Förster resonance energy transfer: implications of dye linker length and rigidity. J Am Chem Soc 133:2463–2480. https://doi.org/10.1021/ja105725e
Kalinin S, Peulen T, Sindbert S et al (2012) A toolkit and benchmark study for FRET-restrained high-precision structural modeling. Nat Methods 9:1218–1225. https://doi.org/10.1038/nmeth.2222
Kapanidis AN, Weiss S (2002) Fluorescent probes and bioconjugation chemistries for single-molecule fluorescence analysis of biomolecules. J Chem Phys 117:10953–10964. https://doi.org/10.1063/1.1521158
Zosel F, Holla A, Schuler B (2022) Labeling of proteins for single-molecule fluorescence spectroscopy. Methods Mol Biol 2376:207–233. https://doi.org/10.1007/978-1-0716-1716-8_12
Hanspach G, Trucks S, Hengesbach M (2019) Strategic labelling approaches for RNA single-molecule spectroscopy. RNA Biol 16:1119–1132. https://doi.org/10.1080/1547628620191593093
Gust A, Zander A, Gietl A et al (2014) A starting point for fluorescence-based single-molecule measurements in biomolecular research. Molecules 19:15824–15865. https://doi.org/10.3390/molecules191015824
Ha T, Tinnefeld P (2012) Photophysics of fluorescent probes for single-molecule biophysics and super-resolution imaging. Annu Rev-Physchem 63:595–617. https://doi.org/10.1146/ANNUREV-PHYSCHEM-032210-103340
Hohng S, Lee S, Lee J, Jo MH (2014) Maximizing information content of single-molecule FRET experiments: multi-color FRET and FRET combined with force or torque. Chem Soc Rev 43:1007–1013. https://doi.org/10.1039/c3cs60184f
Kim JY, Kim C, Lee NK (2015) Real-time submillisecond single-molecule FRET dynamics of freely diffusing molecules with liposome tethering. Nat Commun 6:1–9. https://doi.org/10.1038/ncomms7992
Roy R, Hohng S, Ha T (2008) A practical guide to single-molecule FRET. Nat Methods 5:507–516. https://doi.org/10.1038/nmeth.1208
Chandradoss SD, Haagsma AC, Lee YK et al (2014) Surface passivation for single-molecule protein studies. J Vis Exp:4–11. https://doi.org/10.3791/50549
Paul T, Ha T, Myong S (2021) Regeneration of PEG slide for multiple rounds of single-molecule measurements. Biophys J:1–12. https://doi.org/10.1016/j.bpj.2021.02.031
McCann JJ, Choi UB, Zheng L et al (2010) Optimizing methods to recover absolute FRET efficiency from immobilized single molecules. Biophys J 99:961–970. https://doi.org/10.1016/j.bpj.2010.04.063
Hildebrandt LL, Preus S, Birkedal V (2015) Quantitative single molecule FRET efficiencies using TIRF microscopy. Faraday Discuss 184:131–142. https://doi.org/10.1039/c5fd00100e
Lee NK, Kapanidis AN, Wang Y et al (2005) Accurate FRET measurements within single diffusing biomolecules using alternating-laser excitation. Biophys J 88:2939–2953. https://doi.org/10.1529/biophysj.104.054114
Kapanidis AN, Lee NK, Laurence TA et al (2004) Fluorescence-aided molecule sorting: analysis of structure and interactions by alternating-laser excitation of single molecules. Proc Natl Acad Sci U S A 101:8936–8941. https://doi.org/10.1073/pnas.0401690101
Hohlbein J, Craggs TD, Cordes T (2014) Alternating-laser excitation: single-molecule FRET and beyond. Chem Soc Rev 43:1156–1171. https://doi.org/10.1039/c3cs60233h
Müller BK, Zaychikov E, Bräuchle C, Lamb DC (2005) Pulsed interleaved excitation. Biophys J 89:3508–3522. https://doi.org/10.1529/biophysj.105.064766
Eggeling C, Berger S, Brand L et al (2001) Data registration and selective single-molecule analysis using multi-parameter fluorescence detection. J Biotechnol 86:163–180. https://doi.org/10.1016/S0168-1656(00)00412-0
Rothwell PJ, Berger S, Kensch O et al (2003) Multiparameter single-molecule fluorescence spectroscopy reveals heterogeneity of HIV-1 reverse transcriptase:primer/template complexes. Proc Natl Acad Sci U S A 100:1655–1660. https://doi.org/10.1073/pnas.0434003100
Hohng S, Joo C, Ha T (2004) Single-molecule three-color FRET. Biophys J 87:1328–1337. https://doi.org/10.1529/biophysj.104.043935
Kim E, Lee S, Jeon A et al (2013) A single-molecule dissection of ligand binding to a protein with intrinsic dynamics. Nat Chem Biol 9:313–318. https://doi.org/10.1038/nchembio.1213
Lee J, Lee S, Ragunathan K et al (2010) Single-molecule four-color FRET. Angew Chem Int Ed 49:9922–9925. https://doi.org/10.1002/ANIE.201005402
Phelps C, Israels B, Marsh MC et al (2016) Using multiorder time-correlation functions (TCFs) to elucidate biomolecular reaction pathways from microsecond single-molecule fluorescence experiments. J Phys Chem B 120:13003–13016. https://doi.org/10.1021/acs.jpcb.6b08449
Santoso Y, Joyce CM, Potapova O et al (2010) Conformational transitions in DNA polymerase I revealed by single-molecule FRET. Proc Natl Acad Sci U S A 107:715–720. https://doi.org/10.1073/pnas.0910909107
Chung HS, Gopich IV (2014) Fast single-molecule FRET spectroscopy: theory and experiment. Phys Chem Chem Phys 16:18644–18657. https://doi.org/10.1039/c4cp02489c
Torella JP, Holden SJ, Santoso Y et al (2011) Identifying molecular dynamics in single-molecule fret experiments with burst variance analysis. Biophys J 100:1568–1577. https://doi.org/10.1016/j.bpj.2011.01.066
Kalinin S, Valeri A, Antonik M et al (2010) Detection of structural dynamics by FRET: a photon distribution and fluorescence lifetime analysis of systems with multiple states. J Phys Chem B 114:7983–7995. https://doi.org/10.1021/jp102156t
Gopich I, Szabo A (2005) Theory of photon statistics in single-molecule Förster resonance energy transfer. J Chem Phys 122:14707. https://doi.org/10.1063/1.1812746
Lerner E, Ploetz E, Hohlbein J et al (2016) A quantitative theoretical framework for protein-induced fluorescence enhancement-Förster-type resonance energy transfer (PIFE-FRET). J Phys Chem B 120:6401–6410. https://doi.org/10.1021/acs.jpcb.6b03692
Hwang H, Myong S (2014) Protein induced fluorescence enhancement (PIFE) for probing protein-nucleic acid interactions. Chem Soc Rev 43:1221–1229. https://doi.org/10.1039/c3cs60201j
Bronson JE, Fei J, Hofman JM et al (2009) Learning rates and states from biophysical time series: a Bayesian approach to model selection and single-molecule FRET data. Biophys J 97:3196–3205. https://doi.org/10.1016/j.bpj.2009.09.031
McKinney SA, Joo C, Ha T (2006) Analysis of single-molecule FRET trajectories using hidden Markov modeling. Biophys J 91:1941–1951. https://doi.org/10.1529/biophysj.106.082487
Szoszkiewicz R, Ainavarapu SRK, Wiita AP et al (2008) Dwell time analysis of a single-molecule mechanochemical reaction. Langmuir 24:1356–1364. https://doi.org/10.1021/la702368b
Kinz-Thompson CD, Bailey NA, Gonzalez RL (2016) Precisely and accurately inferring single-molecule rate constants. Methods Enzymol 581:187–225. https://doi.org/10.1016/bs.mie.2016.08.021
Widom JR, Dhakal S, Heinicke LA, Walter NG (2014) Single-molecule tools for enzymology, structural biology, systems biology and nanotechnology: an update. Arch Toxicol 88:1965–1985. https://doi.org/10.1007/s00204-014-1357-9
Tsukanov R, Tomov TE, Liber M et al (2014) Developing DNA nanotechnology using single-molecule fluorescence. Acc Chem Res 47:1789–1798. https://doi.org/10.1021/ar500027d
Bandyopadhyay D, Mishra PP (2021) Decoding the structural dynamics and conformational alternations of DNA secondary structures by single-molecule FRET microspectroscopy. Front Mol Biosci 8:781. https://doi.org/10.3389/FMOLB.2021.725541/BIBTEX
Bochman ML, Paeschke K, Zakian VA (2012) DNA secondary structures: stability and function of G-quadruplex structures. Nat Rev Genet 13:770–780. https://doi.org/10.1038/nrg3296
Aznauryan M, Søndergaard S, Noer SL et al (2016) A direct view of the complex multi-pathway folding of telomeric G-quadruplexes. Nucleic Acids Res 44:11024–11032. https://doi.org/10.1093/nar/gkw1010
Varshney D, Spiegel J, Zyner K et al (2020) The regulation and functions of DNA and RNA G-quadruplexes. Nat Rev Mol Cell Biol 218(21):459–474. https://doi.org/10.1038/s41580-020-0236-x
Grün JT, Schwalbe H (2021) Folding dynamics of polymorphic G-quadruplex structures. Biopolymers:e23477. https://doi.org/10.1002/BIP.23477
Šponer J, Bussi G, Stadlbauer P et al (2017) Folding of guanine quadruplex molecules–funnel-like mechanism or kinetic partitioning? An overview from MD simulation studies. Biochim Biophys Acta, Gen Subj 1861:1246–1263. https://doi.org/10.1016/j.bbagen.2016.12.008
Israels B, Albrecht CS, Dang A et al (2021) Submillisecond conformational transitions of short single-stranded DNA lattices by photon correlation single-molecule Förster resonance energy transfer. J Phys Chem B. https://doi.org/10.1021/acs.jpcb.1c04119
Von Hippel PH, Berg OG (1986) On the specificity of DNA-protein interactions. Proc Natl Acad Sci USA 83:1608–1612. https://doi.org/10.1073/pnas.83.6.1608
Hohlbein J, Aigrain L, Craggs TD et al (2013) Conformational landscapes of DNA polymerase i and mutator derivatives establish fidelity checkpoints for nucleotide insertion. Nat Commun 4:1–11. https://doi.org/10.1038/ncomms3131
Morten MJ, Lopez SG, Emilie Steinmark I et al (2018) Stacking-induced fluorescence increase reveals allosteric interactions through DNA. Nucleic Acids Res 46:11618–11626. https://doi.org/10.1093/nar/gky887
Hohng S, Zhou R, Nahas MK et al (2007) Fluorescence-force spectroscopy maps two-dimensional reaction landscape of the Holliday junction. Science 318:279–283. https://doi.org/10.1126/science.1146113
Acknowledgments
We thank the reviewer for his/her feedback. We would like to acknowledge support from the Novo Nordisk Foundation (grant NNF20OC0061417) and from the Danish Council for Independent Research (grant DFF-9040-00323B).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Israels, B., Lund, L.M., Birkedal, V. (2022). Single-Molecule FRET: Principles and Analysis. In: Šachl, R., Amaro, M. (eds) Fluorescence Spectroscopy and Microscopy in Biology. Springer Series on Fluorescence, vol 20. Springer, Cham. https://doi.org/10.1007/4243_2022_32
Download citation
DOI: https://doi.org/10.1007/4243_2022_32
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-30361-6
Online ISBN: 978-3-031-30362-3
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)