Abstract
Absorption of UV light by nucleic acids can lead to damaging photoreactions, which may ultimately lead to mutations of the genetic code. The complexity of the photodynamical behavior of nucleobases in the DNA double-helix provides a great challenge to both experimental and computational chemists studying these processes. Starting from the initially excited states, the main question regards the understanding of the subsequent relaxation processes, which can either utilize monomer-like deactivation pathways or lead to excitonic or charge transfer species with new relaxation dynamics. After a review of photophysical processes in single nucleobases we outline the theoretical background relevant for interacting chromophores and assess a large variety of computational approaches relevant for the understanding of the nature and dynamics of excited states of DNA. The discussion continues with the analysis of calculations on excitonic and charge transfer states followed by the presentation of the dynamics of excited-state processes in DNA. The review is concluded by topics on proton transfer in DNA and photochemical dimer formation of nucleobases.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Ab initio calculations
- Charge transfer excited states
- Excitonic states
- Interaction of excited state nucleic acid bases
- Photodynamics
- UV absorption spectra
1 Introduction
Nucleic acids play a central role in biology as carriers of the genetic code. All nucleic acid bases are strongly absorbing species in the ultraviolet (UV) region and their excitation can result in production of harmful photoproducts leading to, e.g., mutations [1–3]. A well known example is the dimerization of pyrimidine type bases, i.e., formation of thymine (T<>T), cytosine (C<>C), and thymine–cytosine (C<>T) dimers [4–7]. This process is formally analogous to the dimerization of two ethylene molecules to cyclobutane, a textbook example of a reaction which is prohibited in the ground state but allowed in the excited state according to the Woodward–Hoffmann rules. Fortunately, the formation of such photoproducts occurs very rarely. The nucleic acid bases themselves show a high degree of photostability (see, e.g., [8–11]). The decay mechanisms of isolated nucleobases have been investigated by ultrafast time resolved spectroscopic techniques [11–15] and extended computational methods [16–28]. Most of the general features and many details are largely understood by now. It is well established [11, 29] that ultrafast (on the time scale of a few picoseconds) internal conversion is responsible for the relaxation of the system into the ground state without changing its chemical identity. This process occurs at structures near the crossing seam of excited-state and ground-state potential energy surfaces (PES) under conditions of a strong nonadiabatic coupling. Due to this mechanism nucleobases are photostable which in turn protects them against radiative damage.
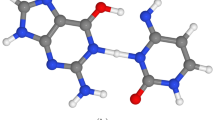
The situation becomes much more complex when a nucleobase is interacting with other bases within the nucleic acid polymer. The structure of nucleic acids allows for interaction within the same strand via stacking and, in the case of double-stranded DNA, also for interactions between two strands via hydrogen bonding as illustrated in Fig. 1. These interactions are further affected by the conformation of DNA (e.g., differences can be expected in B-DNA and RNA-like A-DNA conformations) and the flexibility of the sugar–phosphate backbone which changes the mutual orientation of adjacent nucleobases and thus their interaction. This complex picture of photodynamics of nucleic acids provides a great challenge to both experimental and computational chemists.
In the present contribution the current knowledge of the photodynamics of isolated nucleobases and the survey of experimental observations and their interpretations will be provided in Sect. 1. Only a brief discussion will be given since the scope of the paper is laid on the computational approaches to the UV absorption characteristics and photodynamics of nucleic acids. The theoretical concept of electronic coupling is introduced in Sect. 2. Computational methods and their reliability for the description of excited states in DNA are discussed in Sect. 3. Section 4 discusses the interpretation of the experimentally observed absorption spectra of DNA and its model systems. In Sect. 5 state-of-the-art computational studies on excited states processes, including the effect of DNA environment on the photophysics of nucleobases, excimer formation, excitation energy and proton transfer, and photochemical reactions are reported. The discussion on the latter is limited to formation of cyclobutane pyrimidine dimers as the most studied process.
1.1 Ultrafast Deactivation of Single Nucleobases
The understanding of the photodeactivation mechanism of individual nucleobases is important for analysis of the more complex deactivation patterns of nucleobases embedded in DNA or RNA. It is interesting to note that all five nucleobases (adenine, guanine, thymine, cytosine, and uracil; Scheme 1) show the property of photostability due to their ultrafast, radiationless deactivation to the ground state within a few picoseconds [8, 10]. This decay provides the possibility for the nucleobase to transfer quickly the harmful excess energy accumulated by photoabsorption of UV light into heat, which further on can be dissipated to the environment. These ultrafast processes observed for all five naturally occurring nucleobases contrast with the much longer lifetimes for nucleobase analogues not found in DNA or RNA [10, 30], indicating an evolutionary pressure in early biotic ages.
The photodeactivation of nucleobases in the gas phase has been studied in detail by means of ultrafast time-resolved spectroscopy [8, 10, 11, 17]. These investigations provide the main source for relaxation times which are obtained by fitting the measured deactivation curves in terms of pump-probe delay times. Information on the structural changes responsible for the photostability are not or barely available from these data. The ultrafast and radiationless character of these processes points to internal conversion governed by conical intersections located on the crossing seam between different energy surfaces. This hypothesis has been substantiated in great detail by many theoretical investigations. Starting from the Franck–Condon (FC) region, deactivation paths have been computed leading to characteristic crossing points which are usually chosen as the minima on the intersection seam. One of the nucleobases studied extensively is adenine for which a large variety of types of intersections has been found [16, 22, 23, 31–34]. The two energetically lowest S0–S1 intersections show puckering at the C2 atom and out-of-plane distortion of the NH2 group, respectively (Fig. 2). The former structure corresponds to an S0/π–π* intersection characterized as envelope 2E and the latter to an S0/n–π* intersection with a skew boat 6S1 puckering following the notation of ring puckering coordinates introduced by Cremer and Pople [35]. Starting at the Franck–Condon structure, pathways connecting the bright L a state with these intersections are supposed to have no or at most very small barriers so that both are good candidates for the explanation of the ultrafast decay of adenine. For a more detailed discussion of the outcomes of different dynamics simulations see below. In the case of the other purine base guanine, two ethylenic-type intersections (I and II) with strong out-of-plane distortion of the NH2 group and another intersection type (denoted oop-O) with puckering at C6 and concomitant out-of-plane motion of the oxygen atom are found [20, 28, 36–39]. The bright π–π* state is the lowest singlet excited state in the Franck–Condon region and is connected from there via a barrierless path with the ethylenic intersections [28].
In comparison to the purine bases, the pyrimidine bases are characterized by a more complex system of conical intersections and reaction paths. In the case of cytosine, three major conical intersections have been determined. The first [25] is characterized as semi-planar. It occurs in a region of mixing of the n O–π*, π–π* and closed shell states, which leads to a triple degeneracy as reported in [40, 41]. The second intersection leads to a puckering at N3 connected with an out-of-plane distortion of the amino group. The third intersection involves puckering at C6 (for more details see [42–44]). Reaction paths computed in [44] connecting the Franck–Condon region with the above-mentioned conical intersections show that these intersections are all energetically accessible and display similar qualitative features so that no a priori decision can be made concerning specific photodynamical reaction mechanisms.
Concerning theoretical investigations on the remaining nucleobases (see, e.g., [17, 21, 24, 27]), it should be mentioned at this point that not only the ring puckering motions are of relevance for the explanation of the ultrafast decay of the nucleobases but also that NH dissociation is of interest since it has been shown that this process leads to conical intersections [32] even though they are supposed to be accessible only by higher excitation energies beyond the first absorption band.
The above discussion concentrates on the singlet excited states. It should be noted that several studies discussing the triplet excited states appeared in the literature (see, e.g., [45–50]).
The large manifold of conical intersections and their reaction paths give a good picture of the different deactivation possibilities but makes conclusive predictions about the real deactivation mechanisms difficult if not impossible. Dynamics simulations have been performed to find the actual reaction paths and to obtain information about the characteristic times connected with the different processes. Owing to the strongly coupled motion involving many internal degrees of freedom, Tully surface hopping [51] and ab initio multiple spawning (AIMS) [52] appear to be methods of choice since all internal degrees of freedom are directly included in these dynamics simulations (see also [53] for a general discussion on the options and outlook of photodynamical simulations). The main bottleneck in these calculations is the necessity of performing a full quantum chemical calculation in each time step using extended methods able to describe several electronic states and their nonadiabatic coupling. Under these circumstances, mostly complete active space self-consistent field (CASSCF) or multireference configuration interaction with single excitations (MRCIS) approaches have been used [17, 19, 33, 38, 44, 54–56]. An overview of respective simulations of the photodynamics of nucleobases is available, e.g., in [18]. As an interesting alternative to the computationally expensive ab initio approaches, semi-empirical multireference methods based on the orthogonalization model 2 Hamiltonian (OM2) [34, 57, 58] and the fractional orbital occupation/AM1 model [59] have also been applied extensively in surface hopping dynamics. The performance of TDDFT using a larger variety of functionals has been investigated [60], and time-dependent tight binding density functional theory (TD-DFTB) [61] and DFTB mean field simulations [62] have been used as well. The restricted open shell Kohn–Sham (ROKS) method within the framework of Car–Parinello dynamics [63] and quantum dynamical simulations have also been performed [64].
From this multitude of widely differing approaches, we wanted to discuss the photodynamics of adenine and cytosine in more detail. Surface hopping dynamics simulations using an MRCIS approach [33] have been performed for adenine, starting the dynamics in the L a π–π*(S3) state. Already after 25 fs this state is completely depopulated and practically all trajectories have arrived in the S1 state after ~60 fs. By fitting an exponential function containing two decay constant, values of 22 and 538 fs were found, in good agreement with those of <50 and 750 fs measured by Ullrich et al. [9]. The analysis of the trajectories showed that almost exclusively the 2E with puckering at C2 was accessed. In contrast, OM2 photodynamics [34] shows a strong predominance of the NH2 out-of-plane conical intersection (1S6). In a recent study [60] the energy profiles along the reaction coordinates leading to both mentioned intersections were computed using several methods (OM2/MRCI, MRCIS, complete active space perturbation theory to second order (CASPT2), approximate coupled cluster singles- and doubles method (CC2), and a variety of different TDDFT methods). These profiles show a preference toward the 2E intersection in the case of MRCIS, CASPT2, and CC2 whereas the opposite trend is found for OM2. It was further shown that the MRCIS calculations using one lone pair (n) orbital in the active space were subject to a certain bias toward 2E which was partially lifted by including two n orbitals into the active space [65]. Thus, according to current understanding, the real dynamics should still be dominated by the 2E pathway but significant participation of the 1S6 should be expected as well. The investigations in [60] also show a severe failure of the TDDFT and TD-DFTB approaches since an insufficient number of hoppings to the ground state were observed. Inclusion of range-separated functionals did not improve the situation significantly. For more details see [60].
In the case of cytosine, a more complex dynamics than that described for adenine is observed [18]. All three intersections described above for cytosine participate in the dynamics. Initially, cytosine relaxes along the π–π* pathway to a region of strong mixture of π–π*, n–π*, and closed shell character, where in a first approach about 16% of trajectories switch to the ground state with a time constant of 13 fs. It should be noted that this semi-instantaneous deactivation of cytosine through the semi-planar intersection was also found in AIMS simulations [19]. In the remaining 84% of cases, cytosine quickly relaxes to the n–π* state from where it either can deactivate via the semi-planar intersection to the ground state or can overcome the barrier to the π–π* state and subsequently switch to the ground state. In contrast, CASSCF(2,2) AIMS [19] and OM2 [57] simulations do not show any significant deactivation in this region. The results of surface hopping dynamics at CASSCF(12,9) level [66] are similar to those found in [18] and show predominance of the semi-planar conical intersection. Due to this complex set of events it has to be expected that even rather subtle changes in the relative positions of the energy surfaces can lead to significant modifications of the photodynamical mechanisms and all discussed results have to be regarded with care. It can be expected that further quantitative changes will occur when more sophisticated quantum chemical methods will become available for photodynamical simulations.
Notable changes in the dynamics also have to be expected when it is performed in aqueous solution. Several computations were performed to elucidate the effect of solvent on the excited states of uracil [49, 67–78], thymine [68, 76, 77, 79, 80], guanine [81–84], cytosine [75, 77, 85, 86], and adenine and its model systems [87–89]. The aqueous solvent was shown to modify the photodynamics of nucleobases by changing relative positions of excited states of different characters in the FC region, changing the heights of reaction barriers on the paths leading to conical intersections and relative energies of excited state minima. The consequences such as a blue shift of the n–π* states of uracil (see, e.g., [68, 70, 71, 73, 76], stabilization of a polar S2(π–π*) [32] and π–σ* [87] states of adenine, and geometry changes of excited state minima of guanine [82] on photodynamics were discussed. Dynamics studies performed on adenine [89] have shown that the out-of-plane motion of the amino group is more pronounced in water, which can explain the faster decay of the S1 state compared to the gas phase. The overall relaxation mechanism was however the same as observed in the gas phase. This finding is in contrast to guanine [84] where the pathway via a conical intersection with out-of-plane distortion of the carbonyl oxygen becomes dominant in water solvent.
It should be noted that in extension of the above-described dynamics simulations restricted to the singlet manifold, first surface hopping dynamics simulations have been performed for cytosine combining nonadiabatic and spin-orbit effects [90, 91].
At first sight it seems that each of the nucleobases possesses its own characteristics. A general picture emerging from the dynamics simulations [18] can be given, however, by the observation that the purine bases adenine and guanine have quite a simple deactivation pattern following basically one excited state to the intersection with the ground state whereas, in the case of the pyrimidine bases cytosine, thymine, and uracil, a significantly more complex picture appears with the participation of several excited states and a significantly more complex deactivation pattern.
1.2 Survey of Experimental Studies of Interacting Nucleobases
The question of the nature of excited states of DNA was first addressed in the 1960s. Based on theoretical models, Tinoco et al. [92] invoked a delocalized character of excited states of nucleic acids in the FC region to explain the hypochromism observed in absorption spectra of polynucleotides. In contrast to this prediction, Eisinger et al. [93] proposed that the UV photon is absorbed by a single base. This prediction was made on the basis of comparison of absorption spectra of DNA with corresponding pyrimidine and purine bases. The DNA absorption spectra closely resembled the sum of the monomer spectra and showed theoretically predicted spectral shifts and splitting of the UV band around 260 nm. A discussion on this issue based on molecular dynamics simulations and quantum chemical calculations will be given later in this chapter.
Following the prediction of the locally excited states red-shifted and broadened fluorescence spectra of di- and polynucleotides were attributed to excimers formed from these bases [5]. More pronounced occurrences of excimers observed at lower temperatures are explained by a higher degree of stacking as compared to more disordered structures at higher temperatures. Increased charge transfer (CT) contributions in these excimers were also predicted by these authors [94, 95].
In contrast to the above-discussed local character of absorption, the excitonic character in absorption spectra of di- and polynucleotides of adenine and cytosine was predicted by Kononov et al. [7], with the exciton limited to two bases. The excitonic features were, however, not observed in absorption spectra of guanine and thymine.
The experiments performed during the last two decades utilizing femtosecond time-resolved spectroscopy stimulated lively discussion on the nature of excited states of DNA. In addition to the ultrafast components decaying on the order of picoseconds, transients with lifetimes of 10–100 ps and even nanosecond time scale were observed for single- and double-stranded oligonucleotides [14, 96–103]. The existence of large decay times occurring for single-stranded oligonucleotides [98] demonstrates that stacking interactions are of primary importance for the explanation of differences in the complex decay dynamics in DNA as opposed to that observed for individual nucleic acid bases.
The interpretation of the ultrafast component observed in the experiments mentioned in the previous paragraph was based on the existence of bases undergoing monomer-like photo decay in the disordered parts of the oligo- and polymers. Different hypotheses were used to explain the slow-decay component: (1) the excitation in the FC region is localized on a single base and the interaction with an adjacent base leads to the formation of excimers [98, 101] several picoseconds after the excitation and (2) the excitation is already delocalized during the initial FC excitation, resulting in the formation of excitonic states [14, 96, 104–106]. In the latter interpretation partial emission from localized bases was suggested as well [105, 107]. Beyond the above question of the delocalization, the importance of charge transfer states is being discussed as an important issue (see, e.g., [108–113]).
A special challenge lies in interpreting and reconciling the large number of different decay times reported in the different experiments (see, e.g., [97–99, 101, 102, 114–121]). The results depend not only on the general character of the experimental technique (i.e., time-dependent absorption or emission spectroscopy) but also on the details of the preparation of the DNA samples and the base sequence, both playing an important role [118, 122]. In light of all these challenges with respect to interpreting experimental results, computational investigations play an important role in providing complementary information, shedding new light on the complex photodynamical behavior of DNA.
The extent of exciton delocalization is another issue discussed intensively. For example, Kadhane et al. suggested that excitons are delocalized over no more than two bases [123] and Bucharov et al. [14] suggested delocalization extending over three bases. However, delocalization over six and more bases was also considered [124]. Extended work concerning this issue is based mainly on dynamics simulations in connection with excitonic models or quantum chemical calculations, and will be discussed later in the chapter.
According to current evidence, inter-strand hydrogen bonding may also play an important role and add to the complexity of the photodynamical behavior of nucleic acids. A unique excited state behavior of Watson–Crick (WC) base pairs was shown in a resonant multi-photon ionization experiment performed in the gas phase [125]. Faster fluorescence decay of base pairs compared to isolated bases was also observed by other authors as shown in [98, 126–128]. Furthermore, interplay between intra-strand CT and inter-strand proton transfer has been suggested [128].
2 Description of Electronic Coupling
Before discussing the character of excited state interactions between nucleobases, the underlying theory will be briefly discussed. Scheme 2 describes the simplest situation of the electronic interactions of two identical chromophores (A and B) in terms of locally and charge transfer excited states. At infinite separation the locally excited states (described by Ψ(A*B) and Ψ(AB*)) are degenerate. At finite separation, in an adiabatic representation the degeneracy is removed and the energy splitting may be directly used to measure the excited state interaction (electronic coupling). The situation is similar for the charge transfer states described by Ψ(A .− B .+) and Ψ(A .+ B −.) [129–131]. For non-symmetrical arrangements this approach is not so straightforward. Inside the polymer the environment of two bases is strongly non-symmetrical due to thermal fluctuations. In such a case the splitting will also reflect the difference in the environment of each chromophore, which causes a shift of the orbital and excitation energies that are not directly related to the electronic coupling. In the case of a stacked nucleobase pair with nucleobases mutually orientated as in a B-DNA structure, the changes in excitation energies observed during dimerization caused by non-symmetric effects are an order of magnitude larger than those caused by the ‘pure’ electronic interaction [132, 133].
Interaction between chromophores in electronically excited states can promote one of the following processes (Fig. 3):
-
1.
Delocalization of the excited states which results in exciton formation.
-
2.
Electronic excitation energy transfer.
-
3.
Formation of a strongly interacting complex – an excimer – which results from the orbital overlap.
-
4.
Formation of a new chemical species – a photochemical product.
-
5.
Inter- and/or intra-strand electron or proton transfer.
-
6.
Localization of the electronic excitation on a single base followed by a monomer-like relaxation.
The interactions during electronic excitations are governed by two main contributions: (1) Coulombic interactions which are often approximated by the interactions of transition dipole moments and higher multipole interaction terms and (2) short-range interactions, which depend on the orbital overlap between two chromophores [129–131].
While the former interactions operate over larger through-space separation R, and depend asymptotically on R −3, the latter attenuate exponentially with R being unimportant for the chromophore separation larger than approximately 6 Ǻ. Since these interactions depend on the orbital overlap they are greatly dependent on the mutual orientation of the chromophores. Thus, significantly larger short-range interactions were found for the nucleobases in the B-DNA-like orientation with almost parallel stacking as compared to A-DNA-like orientation with disordered stacking [132]. The character of the excited states also largely influences the resulting electronic interactions. For example, these interactions between the states of (n−π*) character with negligible transition dipole moments and small orbital overlaps are much smaller in relation to states of (π–π*) character which possess a larger transition dipole moment and a larger overlap of interacting orbitals [132].
2.1 Excimers/Exciplexes and Excitons
As already mentioned, the photoabsorption of two (or more) nucleobases within the nucleic acid structure can result in excimers or exciplexes being formed by two identical or different nucleobases, respectively. It should be emphasized that the excimers/exciplexes are not necessarily formed from one molecule in the ground state and the second from the excited state. They can be formed from different initial states, including excitons.
The wavefunction has the following general form [129, 134, 135]:
where A = B, c 1 = c 2, and c 3 = c 4 for excimers.
The first two terms correspond to locally excited states and their interaction results in exciton states (see below). At intermolecular separations below 5–6 Å, orbital interactions come into play [129, 134] mediating a mixing of the locally excited states with the charge transfer or radical-ion-pair states, described by the third and fourth terms [135]. Stabilization of the exciplex can not only occur through excitonic coupling and Coulombic interaction, which are directly derived from the excitonic and CT contributions. It has been pointed out that with decreasing separation distance a new type of strong interaction may come into play that can be identified as a quasi-bond in the geminate radical ion pair [136]. Such quasi-bonds may go along with strong structural distortions, and experimental evidence shows that electronic and steric effects play an important role [137].
In Fig. 4, excimer formation in the face-to-face stacked naphthalene dimer computed at the ab initio algebraic diagrammatic construction to second order (ADC(2)) level is illustrated as a representative example (see [138] for more details). While the ground state potential is only weakly attractive with a shallow minimum around 3.65 Å intermolecular separation, the excited state is strongly bound with a minimum at 3.2 Å and stabilization energy of 1.38 eV with respect to infinite separation. Aside from the energy, a measure for charge transfer, representing the weight of the c 3 and c 4 coefficients in the equation (1), is shown in Fig. 4 (bottom). At separations >5.0 Å the CT value is approximately zero, showing that the excited state corresponds to a pure Frenkel exciton. When the molecules are moved closer together, orbital interactions are important (cf. [129]) and the locally excited states mix with the CT states, giving a gradual increase of the CT character. At the excimer minimum the CT character reaches 50%, showing that the excited state is now an even mixture between Frenkel excitonic and charge resonance contributions. This state is accessed by a single transition between delocalized orbitals, which corresponds to the situation of a coherent homogeneous state extended over the whole complex (cf. [138, 139]).
Excited state energies of the S0 and S1 states (top) and charge transfer character of the S1 state (bottom) of a face-to-face stacked naphthalene dimer (see [138] for details)
In extended quasi-periodic systems excited states can be delocalized over a number of chromophores. Such states are usually called excitons. In this context it is important to distinguish whether the exciton can be described as a linear combination of locally excited states, forming a Frenkel exciton, or whether CT configurations also play a role.
The first case can be understood in terms of Frenkel exciton theory [140, 141]. In this framework a model Hamiltonian H is written as the sum of isolated chromophores H a and a coupling term V ab:
The singly excited states are described by the term
where Φ i a corresponds to the wavefunction of the state where chromophore a is in its excited state, while others are in their-ground state. The wavefunction of the excitonic state is then written as a linear combination of the wavefunction with locally excited states:
The diagonal and off-diagonal terms of the exciton matrix correspond to the excitation energies of the chromophore a and the exciton coupling terms, respectively. In this picture the excitation within some energy range populates a number of excited states delocalized over several nucleobases. Furthermore, if the initial electronic wavepacket is prepared as a non-stationary starting state, such a Hamiltonian automatically leads to excitation energy transfer. If in addition vibronic contributions are considered, the system can reach the lower part of the emission band via internal conversion (intra-band scattering) [118].
When there are strong orbital interactions between the different fragments, Frenkel exciton theory is no longer sufficient and it is necessary to include explicitly charge transfer configurations into the calculation. The theoretical treatment in such a case is more involved considering that a larger number of parameters are needed [142–144]. However, the advantage of such an approach is that charge separation processes can be easily modeled.
3 Assessment of Computational Methods
A wide range of computational methods have been applied to the description of electronically excited states of DNA fragments. In this section we review these approaches shortly. Unfortunately, no perfect method exists because of the difficulties in computing excited states and the relatively large sizes of the molecular systems involved. In the following we will characterize the main features of the individual methods, including their strong and weak points, in an attempt to aid in assessing the substantial amount of available computational literature and to explain some of the discrepancies obtained in the different studies. This section will be strongly focused on DNA fragments; for a more general overview we refer the reader to [53].
3.1 Electronic Structure Methods
In this section, the different electronic structure methods that were used to describe DNA fragments will be considered. The main focus will be laid on the computation of excited states, and additionally methods for a phenomenological analysis of excited states and the computation of interaction potentials will be discussed. The main challenge relates to the large system sizes that have to be considered. Therefore, exciton models are a logical choice and were applied by several groups [142, 145–147]. In such an approach the excited states on the different molecules and their interactions are treated by a model Hamiltonian. This Hamiltonian can be easily extended to large system sizes, allowing the study of long range effects and potentially extended delocalization. However, a major challenge in such an approach is a proper parameterization. In particular the electronic couplings are of concern and a number of different methods for estimating them were used. Usually only the Coulombic contribution is computed while the direct orbital term is assumed to be smaller. The simplest approach for the estimation of this interaction is to consider only dipole–dipole interactions. This approximation was used, e.g., by Georghiou et al. [95] and Bittner [142] to estimate the efficiency of energy transfer along the DNA helix. This approach is suitable for systems in which the separation of the two chromophores is significantly greater than the dimension of the molecule. Its validity is however questionable in the case of nucleobases, because their molecular dimension is of the same order as the intermolecular distance of a neighboring nucleobase pair. Several approximations can help to overcome this difficulty using, e.g., the atomic-transition charge distribution [148, 149] or transition-density-cube models [144]. Comparison of the electronic couplings calculated using dipole–dipole and transition-density-cube approaches show that the former significantly overestimate the interactions at shorter distances [144]. Alternatively, the hybrid-multipole model which represents a combination of a truncated-multipole and the extended-dipole model [148, 150] was used to estimate Coulombic interactions [132]. Within this hybrid model, the multipole interaction up to R −5 is covered exactly; all terms with higher orders are considered if they arise from the multipole expansion of the extended dipole. The Coulomb part is usually the major component of the interaction potential, but a complete description also requires consideration of the direct orbital contributions. While it was shown from explicit quantum chemical calculations that at the ground state equilibrium geometry these terms only made a small contribution (<100 cm) [132], the involvement of such terms could in fact be identified through a mixing of Frenkel excitonic and CT states [113]. An ad hoc addition of orbital overlap terms in a Frenkel exciton model is difficult as it has already been pointed out that a modest additional coupling of 100 cm had a dramatic effect by doubling the delocalization length [149]. A more extended treatment of orbital overlap and resulting charge separation is possible but requires a more involved formalism and a larger number of parameters [142, 151].
Considering the structural flexibility of DNA, an atomistic description is in many cases highly desirable. As one option semi-empirical methods were used [59, 152–155]. Due to the fact that they allow an efficient description of quite large systems they were used for extended dynamics simulations, providing interesting insights. However, similar to exciton models, semi-empirical methods rely on careful parameterization, a problem which can be avoided by the use of ab initio methods. Time-dependent density functional theory (TDDFT), which offers efficient excited state computations while relying on no or relatively few empirical parameters, has been used by a number of groups [109, 110, 156]. However, a major challenge for the application of TDDFT is the overstabilization of charge transfer states occurring when local functionals are used [157]. This problem can be overcome by using range separated hybrid functionals or functionals containing an overall high amount of Hartree–Fock exchange but in such cases the energy of CT states crucially depends on the parameters determining the admixture of Hartree–Fock exchange [110, 112]. For a comparison of the performance of different density functionals when applied to DNA stacks see, e.g., [158]. Finally, a large number of wavefunction-based ab initio calculations was performed using single- and multi-reference methods. In the first case, in particular efficient second order models like the approximate coupled cluster method CC2 [159] and the algebraic diagrammatic construction ADC(2) [160] in connection with the resolution of the identity approximation [161] were applied [112, 113, 132]. However, computationally more demanding models like the equation-of-motion coupled-cluster for excitation energies (EOM-EE-CC) with double excitations and even perturbative triple excitations have also been used [162–165]. In the case of strongly distorted structures and intersections between different states, multi-reference methods are needed to provide a reliable description. For this purpose the complete-active space self-consistent field (CASSCF) method, second order perturbation theory on this reference (CASPT2) [166], and multi-reference configuration interaction with single and double excitations (MR-CISD) [167] were performed [168]. While wavefunction-based methods offer the attractive property of systematic improvability toward the exact solution, they suffer from high computational demands. Unless massively parallel computer systems are available heavy truncations have to be carried out as far as the excitation level and/or the one electron basis set are concerned. For this purpose wavefunction-based ab initio methods also require careful testing and analysis before they can be successfully applied.
After the computation of excitation energies the next decisive task is to get a maximum of information for interpretive and phenomenological models. In quantum chemical calculations on DNA fragments this task may be quite challenging in many cases due to many interacting configurations, partially delocalized orbitals, and the presence of hidden charge resonance contributions. Several approaches have been taken to analyze the states in more detail. One useful indicator is Mulliken populations, which aside from simply quantifying charge transfer, can also be used to estimate the contribution of a given monomer to an electronic transition [169]. Furthermore, energetic criteria based on model Hamiltonians have been applied to estimate electronic couplings and to identify charge resonance states [132, 162]. A different route may be taken by considering that the transition density matrix (TDM) between the ground and excited states can be used to represent the structure of the exciton as an electron-hole pair [138, 139, 170]. By partitioning the TDM into blocks corresponding to the different fragments, locally excited, excitonic, and CT contributions can be readily identified [138]. This approach was applied to absorption spectra [113] and excimer formation [168].
Another critical problem of great importance in the description of excimers is the computation of interaction potentials. In particular, dispersion, which is the main force determining stacking interactions, is difficult to describe. In the case of non-correlated methods, dispersion is completely absent, while standard DFT functionals usually strongly underestimate it. By contrast, MP2 is known to overbind complexes [171] and from the pure ab initio methods usually only CCSD(T) with basis set extrapolation is considered to provide reliable results [172]. To obtain accurate interaction potentials a number of empirical corrections have been suggested, where in particular Grimme’s dispersion correction for DFT is widely used [173]. While parameterized methods can provide very good results for standard ground state geometries, it is not clear how they perform for excited states and in particular for strongly bound exciplex structures. It has been pointed out that standard force field parameters severely exaggerate repulsion at short intermolecular separations [172], which means that attempts to use such parameters in the description of excimers may result in an underestimation of binding energies and an overestimation of binding distances. Another problem arising from short intermolecular separations is increased basis set superposition error (BSSE). It was pointed out that the counterpoise (CP) correction for BSSE may strongly affect excimer structures and binding energies [23]. However, it was later also shown that the CP correction, when applied to smaller basis sets, may significantly overshoot, thereby incorrectly destabilizing the resulting exciplexes [168].
3.2 Environmental Models, Sampling, and Dynamics
Aside from a proper description of the electronic structure, new challenges arise with respect to the description of environmental effects, proper sampling to obtain statistically significant results, and the simulation of dynamical phenomena. For representing environmental effects, continuum models as well as atomistic descriptions have been applied. The former type of approach offers a simple well-defined way to include the main effects of environmental polarization where even a differentiation between slow and fast polarization effects is easily achieved. In particular the polarizable continuum model (PCM) [174] has been applied, e.g., in [109, 112, 175]. There are only a few adjustable parameters, most importantly the dielectric constant and the time-regime (equilibrium or non-equilibrium). However, there is some arbitrariness when choosing an effective dielectric constant to represent the effect of the heterogeneous and anisotropic surroundings (i.e., the other DNA bases, the backbone, the surrounding water molecules and counterions). To overcome this problem, quantum mechanics/molecular mechanics (QM/MM) coupling schemes for an atomistic description of the environment have been widely used [110, 113, 176, 177]. Usually the effects of the environment are considered at the level of electrostatic embedding only, and electronic polarization of the environment is neglected (see [178] for a definition of these terms). This should have an effect on vertical excitation energies, while in dynamics simulations the main solvent response can be included through orientational polarization. A critical observation, made from the PCM computations, is that the environment may have a strong impact on the stability of charge transfer states and that, aside from the time-regime (equilibrium or non-equilibrium), even the precise PCM implementation (linear-response [179] or state-specific [180]) makes a crucial difference [181]. A similar sensitivity to the environmental models is probably also present in a QM/MM framework. Thus, in summary, the importance of an accurate description of the environment should be highlighted.
The most straightforward way for sampling ground state DNA structures is by classical molecular dynamics (MD) using standard biomolecular force fields and such an approach has been taken by a number of groups [144, 149, 156, 176]. The situation becomes more complicated when effects of zero-point vibrations should also be included. For this purpose a hybrid approach based on mixed initial conditions was introduced: Large scale motions and solvent degrees of freedom are properly treated by MD while the Wigner distribution of the vibrational ground state is used to represent the central molecule of interest [182]. This approach was applied to produce initial conditions for QM/MM dynamics simulations of a DNA base [177], and a slight extension considering several active molecules was used for spectra simulations [113]. QM/MM geometry optimizations can be performed by using an averaged solvent electrostatic potential generated from MD sampling of the environmental motions [183]. This method was applied to excited state optimizations of the adenine dinucleotide [168].
To go beyond static calculations, a number of excited state dynamics studies have also been performed. A particular focus was placed on the description of non-adiabatic effects, which are essential for understanding, e.g., internal conversion, excitation energy transfer, and charge separation processes. Dynamics have been performed using exciton models [143] and wave packet propagation on parameterized surfaces [184] but in most cases on-the-fly surface hopping dynamics were applied [59, 154, 155, 176, 177]. In the latter case a QM/MM approach was often chosen to allow a real time polarization response of the environment. While the studies presented above considered only the singlet manifold, intersystem crossings to the triplet were also already considered in surface hopping dynamics [90].
4 UV Absorption
The nature of excited states in the Franck–Condon region significantly affects the excited state behavior of nucleic acids. Thus, understanding the character of absorption spectra is crucial for the further evaluation of the photodynamics of nucleic acids. As already mentioned in the Introduction, there is still controversy as to whether during photon absorption the nucleic acids form exciton or charge transfer states or whether they remain localized. In this chapter the survey of theoretical works suggesting both possibilities of delocalization is treated separately.
4.1 Excitonic Delocalization
The concept of excitons in nucleic acids was first introduced by Tinoco et al. [92, 185] and Rhodes [186]. However, a localized character of excited states then dominated the discussions for several decades. The question of exciton character was opened again by Bouvier et al. [148]. In this work the homogeneous (dA)20.(dT)20 and alternating (dAdT)10.(dAdT)10 oligonucleotides in idealized B-DNA geometry are investigated. The two lowest excited states of adenine and one excited state of thymine are considered. The interaction between the states is described by the atomic transition charges model [187] in which the transition dipoles are decomposed onto atomic orbitals. This procedure results in transition charges located on each atom. Using this approach, delocalization of the excited states was found for both oligomers. In (dA)20.(dT)20 the intra-strand coupling dominates with a strength of about 250 cm−1. In the alternating (dAdT)10.(dAdT)10 the inter-strand coupling is more important, the estimated value of the coupling being approximately 100 cm−1. In the following contribution [104] the influence of structure fluctuations on the character of the Frank–Condon excited states is analyzed for oligonucleotides (dA)10.(dT)10 and (dAdT)5.(dAdT)5. A ground-state molecular dynamics simulation to scan possible DNA structures resulting from the plasticity of the sugar-phosphate helix. Importantly, the diagonal energies of monomer excited states are not affected by structural fluctuations, with the change being smaller than 10 cm−1. Relatively large fluctuations of the off-diagonal terms for these structures were found, with the amplitude of the variations 35% and 45% for (dA)10.(dT)10 and (dAdT)5.(dAdT)5, respectively. The distribution of the oscillator strengths, on the other hand, is not significantly affected by the structural dynamics for the (dA)10.(dT)10 oligonucleotide. In fact, 90% of the oscillator strength remains concentrated on the same eigenstates. In the case of the alternating (dAdT)5.(dAdT)5 oligonucleotide the structural disorder results in a larger spreading of the oscillator strength among eigenstates. Studies performed on (dCdG)5.(dCdG)5 [149] resulted in similar conclusions as for the afore-mentioned (dAdT)5.(dAdT)5 case: small fluctuations of monomer excitation energies, less than 15 cm−1 , and a large perturbation of dipolar coupling due to the structural disorder. In all the cases mentioned above a delocalization of the excited states over at least two nucleobases was observed. Delocalization over three bases was found in the calculations of the absorption spectra using the TDDFT method in combination with ground-state MD simulations for single-stranded oligomers of adenine [156] and (adenine–thymine) oligomers [188]. Formation of delocalized exciton states was further investigated in studies which combine ground-state molecular dynamics and the evaluation of the exciton model for the poly(dA).poly(dT) [143] and (dA)12.(dT)12 [144]. The lattice model and the transition density cube model, based on the transition model calculated using the TDDFT method, were used in the former and later investigations, respectively. In agreement with the work of Bouvier et al. [104] and Emanuele et al. [149], changes in electronic coupling due to structural fluctuation were found. The results indicated that during the absorption process the electronic excitation is delocalized over at least six nucleobases and localizes upon relaxation on four nucleobases. Importantly, these authors predicted the presence of charge transfer excitons in their model [144]. Charge transfer character of excitons was also suggested by Starikov et al. [189]. The character of exciton states due to structural fluctuations of B-DNA conformations was investigated in short (dA) n .(dT) n and (dC) n .(dG) n oligomers (with n = 3,4) obtained by means of ground-state molecular dynamics [152]. The effect of different conformational modes was evaluated, among which the twist is predicted as the most powerful regulator of the exciton character. It was also shown that the effect of the twist angle on the localization of electronic states is stronger in poly(dG)-poly(dC) than in poly(dA)-poly(dT) [190]. A new study based on ab initio calculations of alternating duplexes after extended sampling [113] presented a picture of rather localized states, which were situated on one or at most two bases. Considering poly(dAdT)-poly(dAdT) and poly(dGdC)-poly(dGdC) duplexes with four stacked bases in the QM region and the remaining part of the system treated at the MM level, only about a third of the states are delocalized over 1.5 bases or more. These are, however, responsible for somewhat more than 50% of the intensity at the absorption maximum due to higher than average oscillator strengths. Intramolecular vibrations were identified as the main factor responsible for spectral broadening and for causing disorder resulting in localization. This strong coupling of intramolecular modes to the excited states suggests that these are also active in the early excited state dynamics. The picture of rather localized states was also drawn from another recent study that compared measured and computed circular dichroism spectra in (dA) n .(dT) n hairpins [147]. The TDDFT study on adenine stacks pointed out that the A5 spectrum is almost identical to that of A4 [191], i.e., there are no relevant effects either of excitonic or indirect nature which go beyond four bases. While most studies focused on singlet states, triplet excitations were studied in poly(dA)-poly(dT) sequences, and it was reported that they are confined to single nucleobases [192, 193].
4.2 Charge Transfer States
Compared to the studies performed within the framework of exciton models where the charge transfer configurations are usually not included, supermolecular quantum chemical calculations automatically provide a simultaneous treatment of excitonic and the charge transfer states. The interpretation of the absorption spectra in terms of delocalized, charge transfer and localized characters of excited states became a matter of discussion due to the performance of different methods. Suitability of various methods to describe long-range charge transfer states between nucleic acids has been tested in recent years (see, for example, [108, 110–112, 169, 194–196]).
In the text below the calculations of the character of excited states observed at the ground state geometries, i.e., upon UV absorption, as affected by base pairing and stacking interactions will be reviewed with the emphasis on a formation of charge transfer states. In initial ab initio calculations the effect of base pairing was studied [42, 197–199]. Although charge transfer states were detected at higher energies when using the CIS method [197], these were the lowest in calculations at the TDDFT level employing the LDA functional [198]. Stacking interactions were considered for the first time in calculations of Varsano et al. [200]. Other calculations studying the effects of stacking on excited states formed upon UV absorption in the gas phase have been reported, e.g., in [132, 133, 162, 201–203]. The difficulties of a correct description of charge transfer states in the absorption spectra is demonstrated in the case of homologous oligomers of adenine, both single-stranded and in double stranded A–T sequences, as well as alternating A–T sequences.
Santoro et al. [109] performed the first study on the excited states of stacked nucleobases in a water environment. In this contribution the absorption spectra of the adenine dimer are studied employing TDDFT with the PBE0 functional for the (9-Met-A)2 and (dA)2 models using a polarizable continuum model for solvent effects. The experimentally observed features when going from monomers to multimers [97, 105, 118], i.e., a blue-shift of the maximum, a red-shift of the lower-energy part, and hypochromic effect of the absorption spectra are already reproduced in the calculations of the former model. These effects reflect the coupling due to the orbital overlap in the stacking configuration of nucleobases. The symmetric combination of monomer bright transitions gives rise to the maximum of the absorption band. The character of the states responsible for low-lying energy transitions changes with the mutual orientation of the adjacent adenines, i.e., an increasing intra-strand charge transfer character of the first excited state was found in less symmetrical arrangements in (dA)2 in comparison to (9-Met-A)2 [109]. The same change of the maximum absorption peak, i.e., blue shift and decrease in the intensity with increasing number of stacked nucleobases, was already observed in the gas phase in calculations of Lange et al. [110] performed with a long-range-corrected PBE0 functional (LRC-ω-PBE0) on the homologous (A) n and (T) n , (n = 1–4). In contrast to the calculations performed with a non-corrected density functional [109], charge transfer character was not observed in the states responsible for the red tail of the absorption spectra. The effect of the solvent is studied by means of a QM/MM approach with water molecules in the first solvation shell (within 2.5 Ǻ) treated at the QM level. Depending on the solvent configuration the energies of charge transfer states span more than 1 eV and for some configurations they overlap with bright ππ* states. The average stabilization of the charge transfer states by the solvent is 0.1 eV.
A blue shift of the maximum absorption band is also observed for double-stranded (A)2.(T)2 with localization on the (A)2 dimer while excited states localized on the (T)2 dimer are responsible for the low energy part of the spectrum [169]. The calculations performed with the M052X functional place the intra-strand (A → A) and inter-strand (A → T) charge transfer states in the range of bright excited states of the adenine dimer. Note that in the PBE0 calculations the charge transfer states are the most stable. As in the case of single-stranded oligomers, the gas phase calculations performed with the LRC-ω-PBE0 functional [110] place the intra-strand charge transfer states above the bright states. The inclusion of solvent causes a stabilization of these states by about 0.1 eV on the average.
For a single-strand with an alternating sequence of adenine and thymine studied with the LRC-ω-PBE0 functional in the gas phase, Lange et al. [110] found that, due to a mismatch of ππ* state energies of these nucleobases, the intra-strand excitonic delocalization is missing and, consequently, the bright states are localized on a single base. In contrast to homologous oligomers, the CT states become resonant with the ππ* states of adenine and thymine. When the second strand is included, the intra-strand CT states are placed about 0.4 eV below the bright absorption peak of adenine, while the inter-strand CT states appear about 0.7 eV above this bright peak. Note that the ADC(2) calculations performed by Aquino et al. [112] placed the lowest intra-strand CT state 0.5 and 0.3 eV above the lowest ππ* states localized on adenine and thymine, respectively.
Recently Plasser et al. [113] reported the results of ADC(2) calculations of alternating oligomer duplexes in both gas phase and embedded in DNA environment employing a QM/MM scheme, together with a detailed analysis of the excited states. For both systems considered, poly(dAdT)-poly(dAdT) and poly(dGdC)-poly(dGdC), a similar picture was drawn: CT states were found energetically well above the bright states. Due to their low intensity they do not significantly contribute to the absorption spectra even when the DNA environment is involved. A statistical treatment of the QM/MM simulations shows that only about half of the states exhibit Frenkel exciton character (a CT contribution below 0.1 e) while the remaining states show a non-negligible admixture of CT contributions, which means that they may be misrepresented in a pure Frenkel exciton model. About 15% of the states considered show significant charge separation (>0.5 e).
5 Excited State Processes
A number of factors may play a role in DNA photoactivity. These may be divided into external factors like sterical hindrances and electrostatic interactions on the one hand, and electronic interactions on the other. From a computational point of view the significance is that for the former class it may be sufficient to treat only one base at the QM level and the environment at a lower, e.g., molecular mechanics, level (QM/MM method). By contrast, for the second class it is indispensable to consider several bases simultaneously at the QM level, which imposes restrictions on the available approaches. In this chapter, studies concerned with external interactions will be reviewed first. Then direct electronic interactions will be considered, which may occur between stacked bases (leading to excitons, CT states, and exciplexes) as well as hydrogen bonded bases (where proton coupled electron transfer processes are of special interest). Finally, further reactions determining the photochemistry of DNA will be discussed.
5.1 Sterical Hindrances and Electrostatic Interactions
The calculations of the excited state relaxation of adenine [204] performed at the CASPT2/CASSCF/AMBER level assumed that the main reaction channel involves the 2E conical intersection for the adenine molecule (see above, Fig. 2) in both vacuo and solvated (dA)10 · (dT)10 duplex. In the latter case the reaction path is flatter and features a small barrier of about 0.2 eV. These characteristics are suggested to be responsible for the slow decay component (>100 ps) observed in single and double-stranded systems with stacked adenine, thus questioning the importance of delocalized excited states. Nonadiabatic dynamics simulations employing the QM/MM method with semi-empirical treatment of the QM part (using OM2/MRCI) [58, 155] performed on an adenine molecule embedded in (dA)10 and (dA)10 · (dT)10 in water predict an elongation of the excited state lifetime of adenine in comparison with the gas phase but the decay still occurs within 10 ps. In particular, decay times of 5.7 and 4.1 ps for (dA)10 and (dA)10 · (dT)10, respectively, were predicted. In the single-stranded system relaxation proceeds mainly via the 6S1 conical intersection but the 2E pathway coexists, while in double-stranded DNA the 6S1 conical intersection is blocked due to inter-strand hydrogen bonding and only the 2E intersection is accessed.
Surface hopping dynamics simulations using the QM/MM approach with CASSCF (cytosine) and MR-CIS (guanine) wavefunction for the QM part were performed for these two nucleobases, each embedded individually in a DNA double strand helix [177]. The restraining influence of the inter-strand hydrogen bonding on the structural deformations necessary to reach conical intersections was studied. In the case of photoexcited cytosine the isolated molecule shows relatively small puckering of the structure at the conical intersections populated during the excited state relaxation. The geometrical restrictions exerted by the hydrogen bonds of the DNA environment thus do not inhibit the photodeactivation of cytosine and consequently its excited state lifetime. In contrast to this, the isolated guanine relaxes to the ground state with strong out-of-plane motions of the NH2 group. This motion is significantly restrained by inter-strand hydrogen bonds which results in a considerable elongation of the relaxation time.
The effect of the intra-strand interaction on the photodecay of adenine in a single-stranded DNA was studied in nonadiabatic dynamics simulations using 4-aminopyrimidine (4AP) [205] as a model for adenine. In these QM/MM calculations 4AP was treated at the CASSCF level. Comparison with the previously investigated dynamics of isolated 4AP [33] shows a very similar relaxation mechanism and only slight elongation of the excited state lifetime. During the dynamics of embedded 4AP the dynamical formation and breaking of intra-strand hydrogen bonds was observed. Interestingly, these bonds contribute to a faster decay component by enhancing the out-of-plane motion of the amino-group in relevant conical intersections.
5.2 Excimer Formation and Excitation Energy Transfer
An initial study used static calculations of stacked adenine dimers and trimers at the TDDFT/PBE0 level with PCM solvation to discuss the effect of stacking interactions on their excited state behavior [109]. This study describes a process starting from rather delocalized absorbing states, with a subsequent localization on one base on a subpicosecond time scale. After this, the system could either deactivate to the ground state or reach a low energy CT minimum on the S1 surface, which could act as a trapping site. A quantum dynamical study performed at the same computational level indicated that charge separation could be fast and effective [108]. Subsequent studies on a number of similar cases used, aside from PBE0, the range-separated CAM-B3LYP and the M05-2X functional containing a large percentage of non-local exchange. Consideration of the (dA)2 · (dT)2 system showed that the interaction between stacked bases was more important than between hydrogen bonded bases. Again, charge transfer excimers were found. A subsequent study on AG and GA stacks also emphasized the importance of charge transfer in these systems [181]. Furthermore, efficient coupling of the CT states to the locally excited states was reported. Later, excited state relaxation in (dA)4 was considered [206]. In this investigation several excited state minima were found pertaining to localized excited states, neutral excimer states, and a CT minimum. The CT minimum was identified as the most stable one. These studies used the newly implemented state-specific PCM formalism [180] to describe the solvent response to the excitation. In these calculations it was shown that this new PCM formalism led to results significantly different from the standard linear-response approach, in particular stabilizing the CT states. Similar results were also found in a more recent combined experimental and theoretical paper [191].
In contrast to the TDDFT studies, a CASPT2/CASSCF work described the formation of neutral excimers, which were especially stabilized in maximum-overlap face-to-face configurations, whereas charge transfer excimers played only a secondary role [175]. Similar to the above analysis, a bifurcation of the initial wavepacket was considered leading either to ultrafast monomer-like decay or toward the formation of excimer states. A dynamics survey using semi-empirical methods [154] described the formation of a long-lived excimer between two adenine molecules, which was structurally characterized by a short intermolecular separation of the C2 atoms (cf. Scheme 1) to an average distance of 2.2 Å. A new decay channel of this system was found, which occurred through an additional shortening of the C2–C2 distance to 1.8 Å and concurrent ring deformations. A subsequent study by this group, using dynamics at the DFTB level [207], even described the formation of a chemical bond between two adenine molecules after excitation, which was stable for about 2 ps. Whereas similar processes in pyrimidine bases can lead to photodimerization (see below), a non-reactive deactivation was observed in these dynamics investigations. Recent ab initio work using the ADC(2) and MR-CISD methods in a QM/MM framework found several minima on the S1 surface, which possessed different degrees of delocalization and charge transfer (see Fig. 5) [168]. The lowest energy minimum was of exciplex type stabilized neither by pure excitonic interactions nor charge transfer, but by direct orbital interactions, which were mediated by a close approach of the adenine C6 atoms (down to about 2.0 Å) and a concurrent strong distortion of the molecular structures. Indications for various decay channels were found, which were related to either a further approach of the two molecules (as in [154]) or a restoration of monomer-like excited states and decay through the 6S1 intersection. Partial charge transfer was observed in response to solvent polarization but it did not play a dominant role. A similar conclusion was also drawn from a recent study using the RI-CIS(D) method [208] presenting a bonded exciplex mediated by an approach of two C6 atoms. In that case no excited state minimum but only an intersection was found. However, it was pointed out that the solvent effects may destabilize this intersection. These newer results may be rationalized with the bonded exciplex model [136] (see also Sect. 2.1). Experimental evidence for this type of mechanism may be found in the unexpected fact of negative fluorescence anisotropy recorded for (dA)20, which was taken as evidence of an out-of-plane polarization of the emitting states [191], a phenomenon which is forbidden in planar molecules by symmetry selection rules. Strong orbital overlap, combining two bases to one effective chromophore, can lift this selection rule, and a transition dipole moment with a strong out-of-plane component was indeed found for the exciplex [168].
Excited state minima of ApA localized on the S1 surface: excitation localized on a single Ade (top), excitation from localized n orbital to delocalized π* orbital (middle), excitation from delocalized π to delocalized π* orbital (see [168] for details)
Aside from explicit quantum chemical calculations, exciton models have also been used to describe excited state dynamics in DNA. The poly(dA).poly(dT) duplex was studied using a model considering both Frenkel excitonic and charge-transfer interactions [143]. The dynamics in this study were dominated by base-stacking whereas inter-strand interactions only played a minor role. It was found that on the adenine side the exciton remained in a cohesive bound state (i.e., no charge transfer occurred between stacked adenine molecules). By contrast, electron-hole separation into mobile charge carriers was reported on the thymine side. It was hypothesized that such charge separation and subsequent recombination could lead to triplet formation.
While the above studies considered adenine and thymine molecules, stacking interactions between two cytosine molecules were also investigated at the CASPT2 level [209] showing an attractive potential between the two cytosine molecules identified in the excited state, which could lead to red-shifted fluorescence. The strength of this interaction experienced strong dependence on BSSE. Using the CP correction, an intermolecular separation of 3.08 Å and an excimer binding energy of 0.58 eV was obtained.
5.3 Proton Transfer Processes
The initial interest in the proton transfer process dates back to Watson and Crick pointing out that the occurrence of rare tautomeric forms of the nucleobases may lead to spontaneous mutations [210] and to Löwdin discussing that such tautomers may be formed through double proton tunneling in the DNA structure [211]. It has been pointed out by Guallar et al. [212] that such double proton transfer is not feasible in the ground state but that such a pathway could be accessed in the excited state. However, the computations suggested that it is unlikely that the rare tautomers would persist long enough to perturb the duplication of the genetic code [212]. Gorb et al. [213] reported that while the AT double proton transfer was completely inaccessible, the corresponding GC structure would show minimal population (with an equilibrium constant of about 10−6). The same group also reported excited state investigations on the AT and GC pairs [197], as well as the AU pair [197]. In both cases the presence of locally excited states and higher lying charge transfer states was reported.
In 2004 Sobolewski and Domcke put forward the hypothesis that a proton coupled electron transfer (PCET) along the central hydrogen bond in the Watson–Crick base pair of guanine and cytosine could serve as an efficient decay channel to the ground state [42]. This hypothesis was based on CASSCF/CASPT2 calculations on structures and reaction paths optimized at the CIS level. It was found that in the vicinity of the FC region a stable charge transfer state existed, and that starting in this state a transfer of the central hydrogen bonding proton (yielding in total a PCET) could neutralize the system and stabilize this state. Potential curves revealed a weakly avoided crossing between the CT and ground states, which might provide an efficient channel for the relaxation. The study also found a locally excited path for cytosine to the ethylenic intersection, which was only later described for the isolated molecule (see, e.g., [44, 214]). An experimental work on GC dimers in the gas phase later found that the Watson–Crick bound base pair did indeed possess a much broader UV spectrum as opposed to other structures of this system [125]. These results were rationalized by computations at the CC2 level, where the three lowest energy conformers of the GC system were considered [215]. In all three cases a proton transfer path could be identified. However, this path was only favored in the WC case, whereas for the other conformers significant barriers would have to be overcome due to the fact that the CT states were higher in energy for these structures. Subsequently, time-resolved fluorescence experiments in chloroform were carried out, which supported the existence of such a decay channel by showing a short lifetime of about 0.36 ps for the GC base pair [127]. However, transient absorption experiments in chloroform in connection with TDDFT calculations in PCM solvent contested this claim [126]. Regarding GC duplexes, it was pointed out from experiment that PCET may play a role in the case of alternating but not in homopolymeric sequences [128]. This idea was supported computationally by comparing TDDFT calculations of the GGG · CCC and GCG · CGC hexamer stacks [216].
Aside from static calculations, non-adiabatic dynamics simulations were also performed to get a deeper mechanistic insight into a potential proton transfer in GC and why it should or should not be effective. In particular, it is of interest to understand the coupling between the electronic and nuclear motion during the PCET process. In this regard the GC pair was studied by CASSCF using an approximate diabatic surface hopping method [176]. The simulations were performed in vacuo and embedded in a DNA double helix solvated in water using a QM/MM scheme. Dynamics were started in the S1 (CT) state and the processes leading to the ground state decay were monitored. After a quick initial proton transfer, the back transfer was briefly inhibited by a diabatic trapping situation leading to several recrossings before reaching the closed shell configuration on a 100 fs time scale. This phenomenon was explained by an unusual intersection topology, possessing effectively only a one-dimensional branching space. The authors showed that the inclusion of the environment could speed up the decay through a stabilization of the charge transfer state. At this point it should be mentioned that the intersection topology described in [176] may be interpreted as a signature of an electron transfer process, which necessarily occurs when switching from the charge transfer state to the neutral state. In fact, similar surface topologies and diabatic trapping dynamics were also obtained in electron transfer model systems [217], further supporting this interpretation. The PCET deactivation process in the WC paired GC system was also studied using non-adiabatic dynamics at the restricted open shell Kohn–Sham (ROKS) level, considering the system in vacuo as well as in water solvation [218]. A steering technique was used to accelerate the forward PCET. In this study a fast deactivation on the order of 100 fs was also reported. To date, no study has been performed attempting to describe the dynamics starting from the bright ππ* state to the CT state and to give a non-adiabatic description of the forward PCET process, considering that this would pose special challenges as a larger number of excited states would have to be described concurrently. However, a similar process was studied in the 2-pyridone dimer model system, using static calculations [219] and non-adiabatic dynamics, both performed at the TDDFT/BHLYP level [220] following time-resolved experiments [221]. The dynamics study highlighted that the PCET was not only governed by the energetics describing the proton transfer but also that non-adiabatic transition probabilities for the electron transfer (which were manifested by temporary hoppings into the adiabatic S2 state) played a decisive role. This observation may provide also an explanation for the discrepancies between the dynamics studies and the above-mentioned experiments on the GC base pair [101, 126]. An efficient deactivation channel may indeed exist once the proton is transferred from G to C. However, it is not clear whether the initial proton transfer occurs after excitation into one of the bright states.
While most studies focused on the GC pair, proton transfer in the AT pair was also studied at the CC2 level [222]. Similar to [215], it was also found for AT that the WC structure possessed an efficient decay channel related to proton transfer. While the CT state was 1.5 eV above the lowest bright transition at the FC region, this state was strongly stabilized by proton transfer. Barrierless deactivation could occur through the CT state by a sequence of conical intersections. Internal conversion to the nπ* state could serve as a competitive channel. As opposed to the WC structure no such efficient decay path was present in the most stable hydrogen bonded conformer of AT as the CT state was located at higher energy with respect to the minimum of the locally excited state. A TDDFT study of the WC bonded AT pair [223] highlighted the importance of solvent effects. In particular it was pointed out that a polar solvent could strongly stabilize the inter-strand CT state, thus disfavoring the subsequent proton transfer. The coupling via a πσ*state explains the time-resolved photoionization spectroscopy of adenine and the adenine dimer. In adenine this path is not dominating although it does play a role as indicated by the H-atom detection in nanosecond spectroscopic experiments. In the adenine dimer an increased population of πσ*transfer is observed. It is caused by stabilization of the relevant state in the dimer [87].
5.4 Photochemical Processes
The major photoproducts observed upon UV-irradiation of DNA include dimer or adduct formations of adjacent pyrimidine nucleobases, particularly cyclobutane pyrimidine dimers (CPDs) and the pyrimidine(6-4) pyrimidone dimers ((6-4)PP). The former products are observed more frequently (for review of experimental investigations see [224] and references therein). The dimerization proceeds via [2+2] cycloaddition of the C5–C6 double bonds of adjacent pyrimidine nucleobases and can be formed via both the singlet and triplet manifolds [224–227]. The thymine dimer (T<>T) is the major photoproduct occurring after DNA irradiation by UV light. The cytosine dimer (C<>C) on the other hand is more likely to induce mutations once it is formed [225, 228].
In the last decade, several results of experimental studies were reported, discussing the mechanism of CPD formation [226, 229–236]. For example, Schreier et al. showed that the T<>T is formed on the picosecond time scale which suggests that the formation proceeds in the singlet excited state [230]. The quantum yield of this reaction is, however, very low, of the order of 10−2. Based on time-resolved femtosecond transient absorption spectroscopy, Kwok et al. [232] suggested that both pathways might simultaneously contribute to the dimerization. In this contribution the role of other excited states for this process, in particular the role of the nπ* state, is discussed in the intersystem crossing to the triplet state.
The theoretical investigations focusing on the mechanism of the more frequently formed T<>T photoproduct considered both triplet and singlet pathways of the dimerization process. In the triplet state the stepwise mechanism was suggested based on the results of the TDDFT [237] and CASPT2 [238] calculations. The singlet pathway was proposed to proceed via a concerted mechanism [202, 203, 236, 237, 239–241] mediated via S1/S0 conical intersections leading to either the original ground state or to T<>T. Blancafort and Migani [240] suggested coexistence of the alternative singlet pathway where the excitation is first localized on a single thymine with a subsequent formation of quasi-minimum of the S1 state localized energetically above the relevant conical intersection. Based on the recently published joint experimental and theoretical investigations Banyasz et al. [236] put forward that the [2+2] cycloaddition occurs from the lowest excitonic state and follows a barrierless path. No CT states were identified in the reaction mechanism, in contrast to the formation of (6-4)PP adducts (for more detailed discussion on the mechanism of (6-4)PP formation see [242, 243]). The absence of charge transfer character in the former reaction might explain a decrease of the T<>T production in the systems with purine residues flanking the TT sequences by a formation of charge transfer purine-pyrimidine(T) exciplexes and subsequent decrease of the TT exciton population observed in the experiment [244].
Much effort has been made to explain the origin of very low quantum yield of the T<>T dimerization [229, 244–250]. It is now generally accepted that the ground state conformation of adjacent thymine controls the efficiency of this reaction with inter-molecular T…T distance and torsional angle between two double bonds (C5–C6 bond) on each thymine moiety being identified as the most important. While the agreement on the criteria for the former parameter was achieved [246–248, 250], the importance of the latter parameter is not clarified yet [246, 247]. The influence of the sugar conformation on the T<>T dimerization process was discussed as well [245, 250–252]. In a recently published contribution Improta suggested that a barrierless path from the delocalized exciton state towards T<>T is most effective when both sugars have C3-endo puckered conformation since the electronic coupling between thymines is larger compared to other sugar conformations [203].
The explanation of the experimentally observed lower yields of C<>C photoproduct as compared to T<>T [201, 239, 241, 253] has been the subject of several theoretical investigations. Results of these studies reveal a unified mechanism, i.e., concerted photocycloaddition of the BPD formation for both sequences. Possible competing reaction channels include monomer-like relaxation and excimer formation. The energy position of the conical intersection which mediates the cycloaddition (CIdimer (S0/S1)) with respect to stable excimer structures and CI corresponding to monomer-like relaxations determines the efficiency of the BPD formation. While CIdimer (S0/S1) is the lowest-energy structure in the case of thymine, it is comparable to the energy of the excimer in the case of cytosine and the monomer-like CI, opening other reaction funnels.
6 Conclusions
The area of light absorption by DNA and subsequent photophysical and photochemical processes is a fascinating research field with many important practical implications for everyday life. It is also an outstanding example for the interaction and influences between theory and experiment in an intensity which is very unique. In writing this chapter we have made an attempt to document the large variety of theoretical approaches and different, sometimes opposing, viewpoints to the photodynamics of DNA. Looking back on the history one can observe enormous effort and progress made in experiments and in theoretical simulations by collecting many facts and putting them in the proper perspective. However, the story is certainly not over yet and many interesting and fundamental insights are still waiting to be discovered.
References
Pfeifer GP, You YH, Besaratinia A (2005) Mutat Res Fund Mol M 571:19
Coulondre C, Miller JH (1977) J Mol Biol 117:577
Brash DE, Rudolph JA, Simon JA, Lin A, McKenna GJ, Baden HP, Halperin AJ, Ponten J (1991) Proc Natl Acad Sci U S A 88:10124
Becker MM, Wang Z (1989) J Mol Biol 210:429
Rochette PJ, Therrien JP, Drouin R, Perdiz D, Bastien N, Drobetsky EA, Sage E (2003) Nucleic Acids Res 31:2786
Beletskii A, Bhagwat AS (2001) J Bacteriol 183:6491
Douki T (2006) J Photoch Photobio B Biol 82:45
Ullrich S, Schultz T, Zgierski MZ, Stollow A (2004) Phys Chem Chem Phys 6:2796
Ullrich S, Schultz T, Zgierski MZ, Stolow A (2004) J Am Chem Soc 126:2262
Canuel C, Mons M, Piuzzi F, Tardivel B, Dimicoli I, Elhanine M (2005) J Chem Phys 122:074316
Crespo-Hernandez CE, Cohen B, Hare PM, Kohler B (2004) Chem Rev 104:1977
Doorley GW, McGovern DA, George MW, Towrie M, Parker AW, Kelly JM, Quinn SJ (2009) Angew Chem Int Ed 48:123
Peon J, Zewail AH (2001) Chem Phys Lett 348:255
Buchvarov I, Wang Q, Raytchev M, Trifonov A, Fiebig T (2007) Proc Natl Acad Sci U S A 104:4794
Markovitsi D, Sharonov A, Onidas D, Gustavsson T (2003) ChemPhysChem 4:303
Marian CM (2005) J Chem Phys 122:104314
Hudock HR, Levine BG, Thompson AL, Satzger H, Townsend D, Gador N, Ullrich S, Stolow A, Martinez TJ (2007) J Phys Chem A 111:8500
Barbatti M, Aquino AJA, Szymczak JJ, Nachtigallova D, Hobza P, Lischka H (2010) Proc Natl Acad Sci U S A 107:21453
Hudock HR, Martinez TJ (2008) ChemPhysChem 9:2486
Chen H, Li SH (2006) J Chem Phys 124:154315
Perun S, Sobolewski AL, Domcke W (2006) J Phys Chem A 110:13238
Perun S, Sobolewski AL, Domcke W (2005) J Am Chem Soc 127:6257
Serrano-Andres L, Merchan M, Borin AC (2006) Proc Natl Acad Sci U S A 103:8691
Matsika S (2004) J Phys Chem A 108:7584
Ismail N, Blancafort L, Olivucci M, Kohler B, Robb MA (2002) J Am Chem Soc 124:6818
Sobolewski AL, Domcke W (2002) Eur Phys J D 20:369
Epifanovsky E, Kowalski K, Fan PD, Valiev M, Matsika S, Krylov AI (2008) J Phys Chem A 112:9983
Serrano-Andres L, Merchan M, Borin AC (2008) J Am Chem Soc 130:2473
Hare PM, Crespo-Hernandez CE, Kohler B (2007) Proc Natl Acad Sci U S A 104:435
Blancafort L, Cohen B, Hare PM, Kohler B, Robb MA (2005) J Phys Chem A 109:4431
Chen H, Lis S (2005) J Phys Chem A 109:8443
Perun S, Sobolewski AL, Domcke W (2005) Chem Phys 313:107
Barbatti M, Lischka H (2008) J Am Chem Soc 130:6831
Fabiano E, Thiel W (2008) J Phys Chem A 112:6859
Cremer D, Pople JA (1975) J Am Chem Soc 97:1354
Marian CM (2007) J Phys Chem A 111:1545
Lan ZG, Fabiano E, Thiel W (2009) ChemPhysChem 10:1225
Barbatti M, Szymczak JJ, Aquino AJA, Nachtigallova D, Lischka H (2011) J Chem Phys 134:014304
Yamazaki S, Domcke W, Sobolewski AL (2008) J Phys Chem A 112:11965
Kistler KA, Matsika S (2008) J Chem Phys 128:215102
Blancafort L, Robb MA (2004) J Phys Chem A 108:10609
Sobolewski AL, Domcke W (2004) Phys Chem Chem Phys 6:2763
Zgierski MZ, Patchkovskii S, Fujiwara T, Lim EC (2005) J Phys Chem A 109:9384
Barbatti M, Aquino AJA, Szymczak JJ, Nachtigallova D, Lischka H (2011) Phys Chem Chem Phys 13:6145
Merchan M, Serrano-Andres L, Robb MA, Blancafort L (2005) J Am Chem Soc 127:1820
Gonzalez-Luque R, Climent T, Gonzalez-Ramirez I, Merchan M, Serrano-Andres L (2010) J Chem Theory Comput 6:2103
Etinski M, Fleig T, Marian CA (2009) J Phys Chem A 113:11809
Serrano-Perez JJ, Gonzalez-Luque R, Merchan M, Serrano-Andres L (2007) J Phys Chem B 111:11880
Marian CM, Schneider F, Kleinschmidt M, Tatchen J (2002) Eur Phys J D 20:357
Climent T, Gonzalez-Luque R, Merchan M, Serrano-Andres L (2007) Chem Phys Lett 441:327
Tully JC (1990) J Chem Phys 93:1061
Ben-Nun M, Martinez TJ (2002) Adv Chem Phys 121:439
Plasser F, Barbatti M, Aquino AJA, Lischka H (2012) Theor Chem Acc 131
Asturiol D, Lasorne B, Robb MA, Blancafort L (2009) J Phys Chem A 113:10211
Szymczak JJ, Barbatti M, Hoo JTS, Adkins JA, Windus TL, Nachtigallova D, Lischka H (2009) J Phys Chem A 113:12686
Nachtigallova D, Aquino AJA, Szymczak JJ, Barbatti M, Hobza P, Lischka H (2011) J Phys Chem A 115:5247
Lan ZG, Fabiano E, Thiel W (2009) J Phys Chem B 113:3548
Lu Y, Lan ZG, Thiel W (2012) J Comput Chem 33:1225
Alexandrova AN, Tully JC, Granucci G (2010) J Phys Chem B 114:12116
Barbatti M, Lan ZG, Crespo-Otero R, Szymczak JJ, Lischka H, Thiel W (2012) J Chem Phys 137:22A503
Mitric R, Werner U, Wohlgemuth M, Seifert G, Bonacic-Koutecky V (2009) J Phys Chem A 113:12700
Lei YB, Yuan SA, Dou YS, Wang YB, Wen ZY (2008) J Phys Chem A 112:8497
Langer H, Doltsinis NL, Marx D (2005) ChemPhysChem 6:1734
Picconi D, Barone V, Lami A, Santoro F, Improta R (2011) ChemPhysChem 12:1957
Szymczak JJ, Barbatti M, Lischka H (2011) Int J Quantum Chem 111:3307
Gonzalez-Vazquez J, Gonzalez L (2010) ChemPhysChem 11:3617
DeFusco A, Minezawa N, Slipchenko LV, Zahariev F, Gordon MS (2011) J Phys Chem Lett 2:2184
Gustavsson T, Banyasz A, Lazzarotto E, Markovitsi D, Scalmani G, Frisch MJ, Barone V, Improta R (2006) J Am Chem Soc 128:607
Gustavsson T, Sarkar N, Banyasz A, Markovitsi D, Improta R (2007) Photochem Photobiol 83:595
Improta R, Barone V (2004) J Am Chem Soc126:14320
Santoro F, Barone V, Gustavsson T, Improta R (2006) J Am Chem Soc 128:16312
Gustavsson T, Improta R, Markovitsi D (2010) J Phys Chem Lett 1:2025
Yoshikawa A, Matsika S (2008) Chem Phys 347:393
Mercier Y, Santoro F, Reguero M, Improta R (2008) J Phys Chem B 112:10769
Kistler KA, Matsika S (2009) J Phys Chem A 113:12396
Etinski M, Marian CM (2010) Phys Chem Chem Phys 12:4915
Rasmussen AM, Lind MC, Kim S, Schaefer HF (2010) J Chem Theory Comput 6:930
Olsen JM, Aidas K, Mikkelsen KV, Kongsted J (2010) J Chem Theory Comput 6:249
Nosenko Y, Kunitski M, Brutschy B (2011) J Phys Chem A 115:9429
Busker M, Nispel M, Haber T, Kleinermanns K, Etinski M, Fleig T (2008) ChemPhysChem 9:1570
Shukla MK, Leszczynski J (2008) J Phys Chem B 112:5139
Shukla MK, Leszczynski J (2009) Chem Phys Lett 478:254
Shukla MK, Leszczynski J (2010) Int J Quantum Chem 110:3027
Heggen B, Lan ZG, Thiel W (2012) Phys Chem Chem Phys 14:8137
Kistler KA, Matsika S (2010) Phys Chem Chem Phys 12:5024
Blancafort L, Migani A (2007) J Photochem Photobio A Chem 190:283
Ritze HH, Lippert H, Samoylova E, Smith VR, Hertel IV, Radloff W, Schultz T (2005) J Chem Phys 122:224320
Szymczak JJ, Muller T, Lischka H (2010) Chem Phys 375:110
Lan ZG, Lu Y, Fabiano E, Thiel W (2011) ChemPhysChem 12:1989
Richter M, Marquetand P, Gonzalez-Vazquez J, Sola I, Gonzalez L (2012) J Phys Chem Lett 3:3090
Mai S, Marquetand P, Richter M, Gonzales-Vazquez J, Gonzales L (2013) ChemPhysChem 14:1
Tinoco IJ (1960) J Am Chem Soc 82:4785
Eisinger J, Shulman RG (1968) Science 161:1311
Gueron M, Shulman RG, Eisinger J (1966) Proc Natl Acad Sci U S A 56:814
Georghiou S, Zhu S, Weidner R, Huang CR, Ge G (1990) J Biomol Struct Dyn 8:657
Markovitsi D, Gustavsson T, Sharonov A (2004) Photochem Photobiol 79:526
Crespo-Hernandez CE, Kohler B (2004) J Phys Chem B 108:11182
Crespo-Hernandez CE, Cohen B, Kohler B (2005) Nature 436:1141
Crespo-Hernandez CE, de La Harpe K, Kohler B (2008) J Am Chem Soc 130:10844
Kwok WM, Ma CS, Phillips DL (2006) J Am Chem Soc 128:11894
Middleton CT, de La Harpe K, Su C, Law YK, Crespo-Hernandez CE, Kohler B (2009) Annu Rev Phys Chem 60:217
Vaya I, Miannay FA, Gustavsson T, Markovitsi D (2010) ChemPhysChem 11:987
Plessow R, Brockhinke A, Eimer W, Kohse-Hoinghaus (2000) J Phys Chem B 104:3695
Bouvier B, Dognon JP, Lavery R, Markovitsi D, Millie P, Onidas D, Zakrzewska K (2003) J Phys Chem B 107:13512
Onidas D, Gustavsson T, Lazzarotto E, Markovitsi D (2007) J Phys Chem B 111:9644
Markovitsi D, Onidas D, Gustavsson T, Talbot F, Lazzarotto E (2005) J Am Chem Soc 127:17130
Onidas D, Gustavsson T, Lazzarotto E, Markovitsi D (2007) Phys Chem Chem Phys 9:5143
Santoro F, Barone V, Improta R (2009) J Am Chem Soc 131:15232
Santoro F, Barone V, Improta R (2007) Proc Natl Acad Sci U S A 104:9931
Lange AW, Herbert JM (2009) J Am Chem Soc 131:3913
Lange AW, Rohrdanz MA, Herbert JM (2008) J J Phys Chem B 112:6304
Aquino AJA, Nachtigallova D, Hobza P, Truhlar DG, Hättig C, Lischka H (2011) J Comp Chem 32:1217
Plasser F, Aquino AJA, Hase WL, Lischka H (2012) J Phys Chem A 116:11151
Crespo-Hernandez CE, Cohen B, Kohler B (2006) Nature 441:E8
Takaya T, Su C, de La Harpe K, Crespo-Hernandez CE, Kohler B (2008) Proc Natl Acad Sci U S A 105:10285
de La Harpe K, Crespo-Hernandez CE, Kohler B (2009) ChemPhysChem 10:1421
Vaya I, Gustavsson T, Miannay FA, Douki T, Markovitsi D (2010) J Am Chem Soc 132:11834
Markovitsi D, Gustavsson T, Talbot F (2007) Photochem Photobiol Sci 6:717
Markovitsi D, Gustavsson T, Vaya I (2010) J Phys Chem Lett 1:3271
Onidas D, Gustavsson T, Lazzarotto E, Markovitsi D (2007) Phys Chem Chem Phys 9:1
Vaya I, Changenet-Barret P, Gustavsson T, Zikich D, Kotlyar AB, Markovitsi D (2010) Photochem Photobiol Sci 9:1193
Schwalb NK, Temps F (2008) Science 322:243
Kadhane U, Holm AIS, Hoffmann SV, Nielsen SB (2008) Phys Rev E 77:021901
Markovitsi D, Gustavsson T, Banyasz A (2010) Mutat Res Rev Mutat 704:21
Abo-Riziq A, Grace L, Nir E, Kabelac M, Hobza P, de Vries MS (2005) Proc Natl Acad Sci U S A 102:20
Biemann L, Kovalenko SA, Kleinermanns K, Mahrwald R, Markert M, Improta R (2011) J Am Chem Soc 133:19664
Schwalb NK, Temps F (2007) J Am Chem Soc 129:9272
de La Harpe K, Crespo-Hernandez CE, Kohler B (2009) J Am Chem Soc 131:17557
Scholes GD, Ghiggino KP (1994) J Phys Chem 98:4580
Scholes GD (1996) J Phys Chem 100:18731
Scholes GD (1999) J Phys Chem B 103:2543
Nachtigallova D, Hobza P, Ritze HH (2008) Phys Chem Chem Phys 10:5689
Ritze HH, Hobza P, Nachtigallova D (2007) Phys Chem Chem Phys 9:1672
East ALL, Lim EC (2000) J Chem Phys 113:8981
Klessinger M, Michl J (1995) Excited states and photochemistry of organic molecules. Wiley-VCH, New York
Wang YS, Haze O, Dinnocenzo JP, Farid S, Farid RS, Gould IR (2007) J Org Chem 72:6970
Wang YS, Haze O, Dinnocenzo JP, Farid S, Farid RS, Gould IR (2008) J Phys Chem A 112:13088
Plasser F, Lischka H (2012) J Chem Theory Comput 8:2777
Tretiak S, Mukamel S (2002) Chem Rev 102:3171
Frenkel J (1931) Phys Rev 37:1276
Davydov AS (1971) Theory of molecular excitons. McGraw-Hill, New York
Bittner ER (2006) J Chem Phys 125:094909
Bittner ER (2007) J Photochem Photobio A Chem 190:328
Czader A, Bittner ER (2008) J Chem Phys 128:035101
Emanuele E, Markovitsi D, Millie P, Zakrzewska K (2005) ChemPhysChem 6:1387
Conwell EM, Bloch SM, McLaughlin PM, Basko DM (2007) J Am Chem Soc 129:9175
Patwardhan S, Tonzani S, Lewis FD, Siebbeles LDA, Schatz GC, Grozema FC (2012) J Phys Chem B 116:11447
Bouvier B, Gustavsson T, Markovitsi D, Millie P (2002) Chem Phys 275:75
Emanuele E, Zakrzewska K, Markovitsi D, Lavery R, Millie P (2005) J Phys Chem B 109:16109
Czikklely V, Forsterling HD, Kuhn H (1970) Chem Phys Lett 6:207
Conwell EM, McLaughlin PM, Bloch SM (2008) J Phys Chem B 112:2268
Starikov EB, Cuniberti G, Tanaka S (2009) J Phys Chem B 113:10428
Granucci G, Persico M (2007) J Chem Phys 126:134114
Zhang W, Yuan S, Whang Z, Qi Z, ZhaO J, Dou Y, Lo GV (2011) Chem Phys Lett 506:303
Lu Y, Lan ZG, Thiel W (2011) Angew Chem Int Ed 50:6864
Tonzani S, Schatz GC (2008) J Am Chem Soc 130:7607
Dreuw A, Weisman JL, Head-Gordon M (2003) J Chem Phys 119:2943
Improta R (2008) Phys Chem Chem Phys 10:2656
Christiansen O, Koch H, Jorgensen P (1995) Chem Phys Lett 243:409
Trofimov AB, Schirmer J (1995) J Phys B At Mol Opt 28:2299
Hattig C, Weigend F (2000) J Chem Phys 113:5154
Kozak CR, Kistler KA, Lu Z, Matsika S (2010) J Phys Chem B 114:1674
Szalay PG, Watson T, Perera A, Lotrich V, Fogarasi G, Bartlett RJ (2012) J Phys Chem A 116:8851
Szalay PG, Watson T, Perera A, Lotrich V, Bartlett RJ (2013) J Phys Chem A 117:3149
Szalay PG (2013) Int J Quantum Chem 113:1821
Andersson K, Malmqvist PA, Roos BO (1992) J Chem Phys 96:1218
Szalay PG, Muller T, Gidofalvi G, Lischka H, Shepard R (2012) Chem Rev 112:108
Plasser F, Lischka H (2013) Photochem Photobiol Sci 12:1440
Santoro F, Barone V, Improta R (2008) ChemPhysChem 9:2531
Luzanov AV, Zhikol OA (2010) Int J Quantum Chem 110:902
Riley KE, Platts JA, Rezac J, Hobza P, Hill JG (2012) J Phys Chem A 116:4159
Morgado CA, Jurecka P, Svozil D, Hobza P, Sponer J (2010) Phys Chem Chem Phys 12:3522
Grimme S (2004) J Comput Chem 25:1463
Cossi M, Barone V, Cammi R, Tomasi J (1996) Chem Phys Lett 255:327
Olaso-Gonzalez G, Merchan M, Serrano-Andres L (2009) J Am Chem Soc 131:4368
Groenhof G, Schafer LV, Boggio-Pasqua M, Goette M, Grubmuller H, Robb MA (2007) J Am Chem Soc 129:6812
Zeleny T, Ruckenbauer M, Aquino AJA, Muller T, Lankas F, Drsata T, Hase WL, Nachtigallova D, Lischka H (2012) J Am Chem Soc 134:13662
Bakowies D, Thiel W (1996) J Phys Chem 100:10580
Cossi M, Barone V (2001) J Chem Phys 115:4708
Improta R, Scalmani G, Frisch MJ, Barone V (2007) J Chem Phys 127
Santoro F, Barone V, Lami A, Improta R (2010) Phys Chem Chem Phys 12:4934
Ruckenbauer M, Barbatti M, Muller T, Lischka H (2010) J Phys Chem A 114:6757
Galvan IF, Sanchez ML, Martin ME, del Valle FJO, Aguilar MA (2003) J Chem Phys 118:255
Improta R, Santoro F, Barone V, Lami A (2009) J Phys Chem A 113:15346
Tinoco I, Bradley DF, Woody RW (1963) J Chem Phys 38:1317
Rhodes W (1961) J Am Chem Soc 83:3609
Claverie P (1978) In: Pullman B (ed) Intramolecular interactions – from diatomics to biopolymers. Wiley, New York, p 69
Voityuk AA (2013) Photochem Photobiol Sci 12:1303
Starikov EB, Lewis JP, Sankey OF (2005) Int J Mod Phys B 19:4331
Yamada H, Starikov EB, Hennig D, Archilla JFR (2005) Eur Phys J E 17:149
Banyasz A, Gustavsson T, Onidas D, Changenet-Barret P, Markovitsi D, Improta R (2013) Chem Eur J 19:3762
Curutchet C, Voityuk AA (2011) Chem Phys Lett 512:118
Curutchet C, Voityuk AA (2011) Angew Chem Int Ed 50:1820
Jensen L, Govind N (2009) J Phys Chem A 113:9761
Shukla MK, Leszczynski J (2010) Mol Phys 108:3131
Santoro F, Barone V, Improta R (2008) J Comput Chem 29:957
Shukla MK, Leszczynski J (2002) J Phys Chem A 106:1011
Tsokalidis A, Kaxiras E (2005) J Phys Chem A 109:2373
Wesolowski TA (2004) J Am Chem Soc 126:11444
Varsano D, Di Felice R, Marques MAL, Rubio A (2006) J Phys Chem B 110:7129
Roca-Sanjuan D, Olaso-Gonzalez G, Gonzalez-Ramirez I, Serrano-Andres L, Merchan M (2008) J Am Chem Soc 130:10768
Boggio-Pasqua M, Groenhof G, Schafer LV, Grubmuller H, Robb MA (2007) J Am Chem Soc 129:10996
Improta R (2012) J Phys Chem B 116:14261
Conti I, Altoe P, Stenta M, Garavelli M, Orlandi G (2010) Phys Chem Chem Phys 12:5016
Nachtigallova D, Zeleny T, Ruckenbauer M, Muller T, Barbatti M, Hobza P, Lischka H (2010) J Am Chem Soc 132:8261
Improta R, Barone V (2011) Angew Chem Int Ed 50:12016
Dou YS, Liu ZC, Yuan S, Zhang WY, Tang H, Zhao JS, Fang WH, Lo GV (2013) Int J Biol Macromol 52:358
Spata VA, Matsika S (2013) J Phys Chem A 117. doi:10.1021/jp4033194
Olaso-Gonzáles G, Roca-Sanjuán D, Serrano-Andres L, Merchan M (2006) J Chem Phys A 125:231102
Watson JD, Crick FHC (1953) Nature 171:737
Lowdin PO (1963) Rev Mod Phys 35:724
Guallar V, Douhal A, Moreno M, Lluch JM (1999) J Phys Chem A 103:6251
Gorb L, Podolyan Y, Dziekonski P, Sokalski WA, Leszczynski J (2004) J Am Chem Soc 126:10119
Blancafort L (2007) Photochem Photobiol 83:603
Sobolewski AL, Domcke W, Hattig C (2005) Proc Natl Acad Sci U S A 102:17903
Ko C, Hammes-Schiffer S (2013) J Phys Chem Lett 4:2540
Plasser F, Lischka H (2011) J Chem Phys 134:034309
Markwick PRL, Doltsinis NL (2007) J Chem Phys 126:175102
Sagvolden E, Furche F (2010) J Phys Chem A 114:6897
Plasser F, Granucci G, Pittner J, Barbatti M, Persico M, Lischka H (2012) J Chem Phys 137:22A514
Muller A, Talbot F, Leutwyler S (2002) J Chem Phys116:2836
Perun S, Sobolewski AL, Domcke W (2006) J Phys Chem A 110:9031
Dargiewicz M, Biczysko M, Improta R, Barone V (2012) Phys Chem Chem Phys 14:8981
Cadet J, Mouret S, Ravanat JL, Douki T (2012) Photochem Photobiol 88:1048
Ravanat JL, Douki T, Cadet J (2001) J Photochem Photobio B Biol 63:88
Cuquerella MC, Lhiaubet-Vallet V, Bosca F, Miranda MA (2011) Chem Sci 2:1219
Bosca F, Lhiaubet-Vallet V, Cuquerella MC, Castell JV, Miranda MA (2006) J Am Chem Soc 128:6318
Lee DH, Pfeifer GP (2003) J Biol Chem 278:10314
Schreier WJ, Schrader TE, Koller FO, Gilch P, Crespo-Hernandez CE, Swaminathan VN, Carell T, Zinth W, Kohler B (2007) Science 315:625
Schreier WJ, Kubon J, Regner N, Haiser K, Schrader TE, Zinth W, Clivio P, Gilch P (2009) J Am Chem Soc 131:5038
Marguet S, Markovitsi D (2005) J Am Chem Soc 127:5780
Kwok WM, Ma C, Phillips DL (2008) J Am Chem Soc 130:5131
Douki T, Cadet J (2003) Photochem Photobiol Sci 2:433
Mouret S, Philippe C, Gracia-Chantegrel J, Banyasz A, Karpati S, Markovitsi D, Douki T (2010) Org Biomol Chem 8:1706
Tommasi S, Denissenko MF, Pfeifer GP (1997) Cancer Res 57:4727
Banyasz A, Douki T, Improta R, Gustavsson T, Onidas D, Vaya I, Perron M, Markovitsi D (2012) J Am Chem Soc 134:14834
Zhang RB, Eriksson LA (2006) J Phys Chem B 110:7556
Climent T, Gonzalez-Ramirez I, Gonzalez-Luque R, Merchan M, Serrano-Andres L (2010) J Phys Chem Lett 1:2072
Gonzalez-Ramirez I, Roca-Sanjuan D, Climent T, Serrano-Perez JJ, Merchan M, Serrano-Andres L (2011) Theor Chem Acc 128:705
Blancafort L, Migani A (2007) J Am Chem Soc 129:14540
Serrano-Perez JJ, Gonzalez-Ramirez I, Coto PB, Merchan M, Serrano-Andres L (2008) J Phys Chem B 112:14096
Labet V, Jorge N, Morell C, Douki T, Grand A, Cadet J, Eriksson LA (2013) Photochem Photobiol Sci 12:1509
Giussani A, Serrano-Andres L, Merchan M, Roca-Sanjuan D, Garavelli M (2013) J Phys Chem B 117:1999
Pan ZZ, Hariharan M, Arkin JD, Jalilov AS, McCullagh M, Schatz GC, Lewis FD (2011) J Am Chem Soc 134:3611
Santini GPH, Pakleza C, Auffinger P, Moriou C, Favre A, Clivio P, Cognet JAH (2007) J Phys Chem B 111:9400
Law YK, Azadi J, Crespo-Hernandez CE, Olmon E, Kohler B (2008) Biophys J 94:3590
Johnson AT, Wiest O (2007) J Phys Chem B 111:14398
McCullagh M, Hariharan M, Lewis FD, Markovitsi D, Douki T, Schatz GC (2010) J Phys Chem B 114:5215
Pan Z, McCullagh M, Schatz GC, Lewis FD (2011) J Phys Chem Lett 2:1432
Hariharan M, McCullagh M, Schatz GC, Lewis FD (2010) J Am Chem Soc 132:12856
Desnous C, Babu BR, McIriou C, Mayo JUO, Favre A, Wengel J, Clivio P (2008) J Am Chem Soc 130:30
Ostrowski T, Maurizot JC, Adeline MT, Fourrey JL, Clivio P (2003) J Org Chem 68:6502
Yuan SA, Zhang WY, Liu LH, Dou YS, Fang WH, Lo GV (2011) J Phys Chem A 115:13291
Acknowledgments
This work was supported by the Austrian Science Fund within the framework of the Special Research Program F41, Vienna Computational Materials Laboratory (ViCoM). We also acknowledge technical support from and computer time at the Vienna Scientific Cluster (Projects 70019 and 70151). Support was also provided by the Robert A. Welch Foundation under Grant No. D-0005. FP is a recipient of a research fellowship by the Alexander von Humboldt Foundation. This work has been supported by the grants of the Grant Agency of the Czech Republic (P208/12/1318) and the grant of the Czech Ministry of Education, Youth and Sport (LH11021). The research at IOCB was part of the project RVO:61388963.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Plasser, F., Aquino, A.J.A., Lischka, H., Nachtigallová, D. (2014). Electronic Excitation Processes in Single-Strand and Double-Strand DNA: A Computational Approach. In: Barbatti, M., Borin, A., Ullrich, S. (eds) Photoinduced Phenomena in Nucleic Acids II. Topics in Current Chemistry, vol 356. Springer, Cham. https://doi.org/10.1007/128_2013_517
Download citation
DOI: https://doi.org/10.1007/128_2013_517
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-13271-6
Online ISBN: 978-3-319-13272-3
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)