Abstract
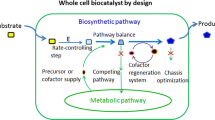
Although cell-free biosystems have been used as a tool for investigating fundamental aspects of biological systems for more than 100 years, they are becoming an emerging biomanufacturing platform in the production of low-value biocommodities (e.g., H2, ethanol, and isobutanol), fine chemicals, and high-value protein and carbohydrate drugs and their precursors. Here we would like to define the cell-free biosystems containing more than three catalytic components in a single reaction vessel, which although different from one-, two-, or three-enzyme biocatalysis can be regarded as a straightforward extension of multienzymatic biocatalysis. In this chapter, we compare the advantages and disadvantages of cell-free biosystems versus living organisms, briefly review the history of cell-free biosystems, highlight a few examples, analyze any remaining obstacles to the scale-up of cell-free biosystems, and suggest potential solutions. Cell-free biosystems could become a disruptive technology to microbial fermentation, especially in the production of high-impact low-value biocommodities mainly due to the very high product yields and potentially low production costs.
Graphical Abstract

Revised book chapter for Advances in Biochemical Engineering/Biotechnology.
Volume: Future Trends in Biotechnology.
Volume editor: Jian-Jiang Zhong.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Biocommodity engineering
- Bioeconomy
- Biofuels
- Biomanufacturing
- Cascade enzyme biocatalyst
- Cell-free synthetic biology
- Synthetic pathway biotransformation
1 Introduction
Biomanufacturing is defined as manufacturing the desired products by using living biological organisms (e.g., bacteria, yeasts, plants) or some components from one or several biological organisms. Biomanufacturing has great potential to become a defining technology, especially in the sustainability revolution [1, 2]. Research and development in the fields of biotechnology, bioengineering, and nanomaterials have dramatic impacts on both the products that we are able to create and the ways in which we create them. Potential products that could be produced through biomanufacturing can be listed in an increasing order of selling prices: biocommodities ($0.3–several US dollars per kg), specialties and biomaterials (tens of dollars per kg), fine chemicals (hundreds of dollars per kg), pharmaceuticals (thousands of dollars/kg), to protein drugs (more than tens of thousands of dollars per kg) (Fig. 1).
Biomanufacturing can be classified into two platforms: living organisms and cell-free biosystems [3, 4]. Living organisms, especially microorganisms, have been utilized for several thousand years to produce a number of products that meet mankind’s needs. As a result, living entities dominate as whole-cell biocatalysts in current biomanufacturing systems. With the development of genetic engineering, protein engineering, systems biology, and synthetic biology, we have gained the ability to modify living organisms to produce natural products at high yields or produce non-natural products. However, the potential of cell-free biosystems in biomanufacturing is often ignored. Herein we define cell-free biosystems composed of more than three catalytic components in one reactor. Thus, we intentionally exclude the use of one to three catalytic components as has been widely used to produce fructose from glucose, produce glucose from starch, and produce chiral alcohols, on large scales [5–9]. Therefore, we do not review traditional enzyme-mediated biocatalysis in this chapter.
Cell-free biosystems have many advantages over living-organism biomanufacturing, such as high product yield, fast reaction rate, high product titer, unprecedented level of control and freedom of design, and broad reaction conditions [3, 4, 10–12] (Table 1). Cell-free biomanufacturing usually gives a yield of the desired product that is often close to a theoretical value when the reactions are irreversible or when the products can be removed in situ. Living systems suffer from low product yields [13, 14] because a significant fraction of carbohydrate and/or energy is used for duplication of cells and formation of side-products when using living organism for manufacturing [15]. For example, all hydrogen-producing microorganisms regardless of whether natural or genetically modified cannot produce hydrogen at yields of more than four moles per mole of glucose, called the Thauer limit [16], because complete oxidation of glucose with water as an oxidant cannot generate any ATP supporting living systems. Conversely, cell-free enzyme cocktails are able to produce nearly 12 moles of hydrogen per mole of glucose [17]. Much higher reaction rates are believed to be possible for cell-free systems than living organisms because (i) neither cell membrane nor wall is present to slow down substrate/product transport, (ii) no energy is needed for transport of substrate/product across the membrane, and (iii) much higher concentrations of biocatalysts can be present in the reactors and no side reactions “slow” the production of the desired product [11]. For example, the highest power density of microbial fuel cells is expected to be around 0.79 mW/cm2 [18], while enzymatic fuel cells can generate power densities of up to 8.5–24 mW/cm2 [19, 20]. Sometimes too slow productivities mean low potential in industrial biomanufacturing [14, 21]. A high concentration of fermentation products can greatly decrease product separation costs [15] while often inhibiting the growth of living organisms, resulting in low product titers. Cell-free biosystems can be controlled much more easily than living organism because the latter has complicated feedback control for gene regulation, protein transcription, translation, and metabolite fluxes. Therefore, cell-free protein synthesis (CFPS) has become an alternative choice for fast synthesis of recombinant proteins in academic laboratories. Furthermore, cell-free protein synthesis has reached a 100-L milestone [22]. In principal, enzymes, especially thermostable enzymes, can tolerate higher levels of organic solvents or toxic chemical compounds than living organisms, resulting in high product titers. Recently, Sieber and co-workers demonstrated that the enzyme cocktails can produce isobutanol even at a concentration of more than 4 % [23]. Additionally, cell-free biosystems can be conducted under much broader reaction conditions than living organisms, for example in organic solvents, ionic liquids, or in the presence of compounds that are toxic to microorganisms [24]. Most building blocks of cell-free systems are highly exchangeable [25]. For example, high-yield enzymatic hydrogen can be produced by using enzymes isolated or originated from bacterium, yeast, plant, rabbit, and archaebacterium [26]. Given the advantages of cell-free systems over living organisms, cell-free biosystems are emerging as a powerful biomanufacturing platform to expand the capabilities of natural biological systems without using whole-cell cells.
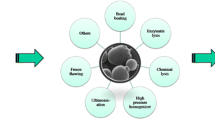
Cell-free biosystems may be classified into two distinctive platforms according to their preparation method. The first is based on whole cell extract by breaking the cell membrane (Fig. 2a). The second is based on purified enzymes from different sources that are then mixed together (Fig. 2b). The cell extract can be prepared easily but some cellular components may influence the whole system’s performance and the whole cell extract has a short half-life time [27, 28]. Systems made of purified components require labor-intensive preparation and seem costly but they can be stabilized after each component is carefully optimized and their performance is more precisely controlled [14, 29]. Specific attributes of two cell-free biosystems enable their applicability to different products.
2 History of Cell-Free Biosystems
The power of cell-free biosystems has been appreciated as a fundamental research tool for more than 100 years. In 1897, Eduard Buchner discovered a yeast extract (not living yeast) that can convert glucose to ethanol [30], leading to his Nobel Prize in Chemistry in 1907. Later, many scientists used cell-free systems to study the basic biological mechanisms and some have received the Nobel Prize for their efforts. An incomplete winners list includes Arthur Harden and Hans von Euler-Chelpin for their investigation of the fermentation of sugar and fermentative enzymes [31], Otto Warburg for his discovery of the nature and mode of action of respiratory enzymes [32], Carl and Gerty Cori for their discovery of the course of the catalytic conversion of glycogen [33, 34], Hans Krebs for his elucidation of the citric acid cycle [35], Melvin Calvin for his investigation of carbon dioxide assimilation in plants [36], and Nirenberg for his elucidation of the genetic codon by using a cell-free system to translate a poly-uracil RNA sequence [37].
Enzyme-based biotransformation became a manufacturing tool approximately 50 years after the discovery of enzymes. The developments of enzyme-based biotransformation can be divided roughly into three phases:
Phase 1 (1960s)—one-enzyme biotransformation [5, 38]. To solve enzyme stability and recycling issues, enzyme immobilization technology was developed. Invertase may be the first immobilized enzyme used commercially for the production of Golden Syrup by Tate and Lyle during World War II. Tanabe Seiyaku Co. (Japan) started the industrial production of l-methionine by using immobilized aminoacylase in a packed bed reactor in 1969. The Clinton Corn Processing Company (USA) produced fructose syrup using glucose isomerase in 1967. Current annual fructose production exceeds 9 million tons [38] and the longest published working lifetime of immobilized glucose isomerase is 687 days at 55 °C and pH 7.5 (Kato Kagaku, Japan) [5]. Semisynthetic beta-lactamase-resistant beta-lactam antibiotics (e.g., cloxacillin, flucloxacillin) are produced by using amidases [5, 39]. Enzymatic acrylamide production was initiated in 1985 and more than 100,000 metric tons of acrylamide per year is produced by using immobilized nitrile hydratases [38]. Progress is accelerating due to fast developments in protein engineering tools including directed evolution, rational design, and their combination [40], high cell-density fermentation for low-cost recombinant protein production [41], the discovery and utilization of thermoenzymes [42], and enzyme immobilization in nanomaterials [3]. As a result, enzyme-mediated biocatalysis is an alternative choice for numerous transformations from commodities to pharmaceuticals [43, 44].
Phase 2 (1990s)—multienzyme one pot for relatively complicated biotransformation. Multienzyme one pot has numerous benefits compared to single-enzyme reactors in cascade: fewer unit operations, smaller reactor volume, higher volumetric and space–time yields, shorter cycle times, and less waste generated. Also, by coupling steps together, unfavorable equilibria can be driven towards the formation of desired products [4, 45, 46]. For instance, enzymatic hydrolysis of crystalline cellulose requires a synergetic action of endoglucanase, cellobiohydrolase, and beta-glucosidase because a single enzyme cannot hydrolyze cellulose efficiently [47, 48]. For cofactor-dependent enzyme reactions that consume reduced NAD(P)H, in situ NAD(P)H-regenerated by another enzyme is becoming more and more accepted, especially for the synthesis of high-value chiral compounds in the pharmaceutical industry [3, 49, 50]. Similarly, NAD(P)H is usually generated by using a pair of a hydrogen-donor substrate and a single enzyme, including formate/formate dehydrogenase [51], glucose/glucose dehydrogenase [52], glucose-6-phosphate/glucose-6-phosphate dehydrogenase [42], dihydrogen/hydrogenase [53], and phosphite/phosphite dehydrogenase [54]. In the organic chemistry field, the synthesis of monosaccharides, activated monosaccharides, oligosaccharides, and glycopeptides by using multienzyme one pot has been intensively investigated [55–58].
Phase 3 (2000s)—the utilization of numerous enzymes (i.e., more than three) for implementing very complicated biotransformations. This cell-free biotransformation scheme has three representative directions: (i) cell-free protein synthesis, which utilizes natural protein synthesis systems in cell lysates for fast synthesis of proteins for research purposes and the production of high-value antibodies or other proteins [59, 60], (ii) in vitro synthetic biology for the production of high-value fine chemicals and pharmaceuticals [27, 61–64], and (iii) synthetic pathway biotransformation (SyPaB) for the production of low-value biocommodities [3, 4, 25]. Herein, we focus on the use of more than three catalytic units in one reactor because of its ability to implement biochemical reactions that living organisms cannot.
3 Examples of Cell-Free Biosystems for Biomanufacturing
We highlight several examples of cell-free biosystems so that readers can easily understand the advantages and applications of cell-free biosystems.
3.1 Cell-Free Biosystems for Biocommodity Engineering
Biocommodities are low-value and large-volume products, including biofuels, biochemicals, bioplatics, feed and food [4, 65]. Among them, transportation biofuels have received wide attention due to their nearly zero net greenhouse gas emissions, enhanced energy security, and new biomanufacturing job creation. Because feedstock costs usually account for more than half of biocommodity selling prices [65, 66], economically viable production of biocommodities requires high product yields and low production costs. Although living microorganisms can duplicate themselves easily and have seemingly low biocatalyst preparation costs, we argue that cell-free biosystems could be an alternative choice due to the unique advantages mentioned earlier: high product yields, fast reaction rates, and potentially very low biocatalyst costs as bulk enzymes become stable and are produced at low cost in the future [3].
There is no question that hydrogen would be the best energy carrier for the transport sector in the future due to the high-energy utilization efficiency through fuel cells and nearly zero pollutants for end-users. Hydrogen can be produced through a number of approaches based on a variety of feedstocks [67]. The production of biohydrogen from low-cost biomass is an excellent solution for producing low-cost hydrogen without net carbon emissions [16, 68, 69]. However, natural and genetically modified hydrogen-producing microbes cannot produce hydrogen in yields of more than four moles hydrogen per mole glucose, called the Thauer limit [14, 16, 70]. Chemical catalysis, such as gasification and aqueous phase reforming, suffers from low product yields due to low chemical selectivity. To solve these problems, a non-natural enzymatic pathway composed of enzymes from bacteria, yeasts, animals, plants, and archae was constructed for generating high-yield hydrogen from starch in 2007 [26] (Fig. 3). The Royal Society (the UK’s National Academy of Science) has praised this breakthrough as “the beginning stage of a cheap, green and high-yield hydrogen production” and as a good example of synthetic biology [71]. This synthetic pathway contains (i) a chain-shortening phosphorylation reaction on glucan for producing glucose-1-phosphate (G1P) catalyzed by glucan phosphorylase (Eq. 1); (ii) the conversion of G1P to glucose-6-phosphate (G6P) catalyzed by phosphoglucomutase (Eq. 2); (iii) a pentose phosphate pathway and gluconeogenesis pathway containing ten enzymes for producing 12 NADPH and 6 CO2 per G6P (Eq. 3); and (iv) the generation of hydrogen from NADPH catalyzed by hydrogenase (Eq. 4).
The synthetic pathway for complete conversion of glucan and water to hydrogen and carbon dioxide. PPP pentose phosphate pathway. The enzymes are: GNP glucan phosphorylase; PGM phosphoglucomutase; G6PDH G-6-P dehydrogenase; 6PGDH 6-phosphogluconate dehydrogenase; R5PI phosphoribose isomerase; Ru5PE ribulose 5-phosphate epimerase; TKL transketolase; TAL transaldolase; TIM triose phosphate isomerase; ALD aldolase; FBP fructose-1,6-bisphosphatase; PGI phosphoglucose isomerase; and H2ase, hydrogenase. The metabolites and chemicals are: g1p glucose-1-phosphate; g6p glucose-6-phosphate; 6pg 6-phosphogluconate; ru5p ribulose-5-phosphate; x5p xylulose-5-phosphate; r5p ribose-5-phosphate; s7p sedoheptulose-7-phosphate; g3p glyceraldehyde-3-phosphate; e4p erythrose-4-phosphate; dhap dihydroxyacetone phosphate; fdp fructose-1,6-diphosphate; f6p fructose-6-phosphate; and Pi inorganic phosphate. Modified from Ref. [26]
The combination of Eqs. (1)–(4) results in Eq. 5:
Thermodynamic analysis of Eq. (5) suggests that the overall reactions from starch or cellulosic materials and water are spontaneous and endothermic (∆G = –50 kJ/mol and ∆Ho = +598 kJ/mol) [17, 26]. These reactions are driven by entropy gains rather than enthalpy losses. The removal of gaseous products, H2 and CO2, from the aqueous phase favors the unidirectional reaction for hydrogen formation. When cellobiose is used as the substrate with a reaction time in the range of one week for a complete reaction, the overall yields of H2 and CO2 are 11.2 moles of H2 and 5.64 moles of CO2 per mole of anhydroglucose unit of cellobiose, corresponding to 93.1 and 94 % of the theoretical yields (Eq. 5). In principle, the theoretical yield of hydrogen (i.e., 12 H2 per glucose equivalent) can be obtained when a continuous reaction is conducted. Over the past few years, intensive efforts have been made pertaining to (i) decreasing enzyme costs and prolonging enzyme lifetime [17, 42, 72–75], (ii) producing all recombinant cytoplasmic enzymes at low costs [42, 76], and (iii) purifying recombinant proteins at low cost using heat precipitation [42, 75], ammonia sulfate precipitation [77, 78], cellulose-binding module-based adsorption and immobilization [79–82].
Ethanol is the most important gasoline additive because it can decrease air pollutants generated by internal combustion (Otto) engines. It can be produced through microbial anaerobic fermentation by the yeast Saccharomyces cerevisiae or by other microorganisms such as Escherichia coli and Zymomonas mobilis [83]. As early as 1897, Eduard Buchner discovered a yeast extract that converted glucose to ethanol [30]. Much later, Welch and Scopes (1985) started investigating the feasibility of high-yield production of ethanol by a reconstituted yeast glycolytic enzyme system—12 enzymes in total including ten enzymes required for the conversion of glucose to pyruvate and two enzymes required for the conversion of pyruvate to ethanol [84]. In this system, ATP accumulation (i.e., two ATP produced per glucose) prevents complete conversion of glucose to ethanol so that costly ATPase or highly toxic arsenate had to be carefully supplemented in Welch’s systems to achieve the high yield of ethanol. To solve this problem, Volker and co-workers designed a non-natural synthetic ethanol-producing pathway comprising six enzyme-catalyzed reactions only [23] (Fig. 4). In it, only four enzymes are required for the conversion of glucose to pyruvate. This synthetic pathway requires neither ATP nor CoA and has a balanced NAD cofactor. Since this pathway has a much lower Gibbs energy than those in yeast and Z. mobilis, it is anticipated that this cell-free pathway could have very fast production rates after optimization.
Schematic representation of a novel cell-free reaction pathway from glucose to ethanol and isobutanol. In the first part of the reaction (top box), glucose is converted into two molecules of pyruvate by four enzymes, then pyruvate can be either directed to ethanol (lower right box) or isobutanol synthesis (lower left box) in the second part of the reaction cascade. The molecules of CO2 and H2O are not shown for clarity. Enzymes are GDH glucose dehydrogenase; DHAD (gluconate/glycerate) dihydroxy acid dehydratase; ALDH glyceraldehyde dehydrogenase; KDGA 2-keto-3-desoxygluconate aldolase; ALS acetolactate synthase; KARI ketolacid reductoisomerase; KDC 2-ketoacid decarboxylase; PDC pyruvate decarboxylase complex; and ADH alcohol dehydrogenase. Modified from Ref. [23]
Isobutanol is a four-carbon liquid alcohol that has several advantages over ethanol, such as, lower water absorption, better blending ability, higher energy density, and compatibility with current internal combustion engines [85]. Volker and co-workers also designed a synthetic pathway that can produce isobutanol from glucose by using a cell-free enzyme mixture (Fig. 4). This pathway is composed of two parts: the production of pyruvate from glucose and the generation of isobutanol through the enzymes in valine biosynthesis pathways along with 2-keto-acid decarboxylases (KDC) and alcohol dehydrogenase (ADH). One of the most beautiful aspects of this cell-free system is that this enzyme system can work well even in the presence of 4 % (v/v) isobutanol, whereas low levels of isobutanol (e.g., 1–2 % v/v) stops microbial isobutanol production [86].
Fructose is the sweetest monosaccharide. High-fructose corn syrup (HFCS) is a mixture of fructose and glucose produced from starch, which has simply replaced sucrose as a sweetener. On average, one person in the USA consumes approximately 20 kg of HFCS per year. HFCS is made from starch through a series of enzymatic conversions: (i) starch liquefaction mediated by amylose, (ii) starch saccharification to glucose mediated by glucoamylase, and (iii) conversion of glucose to fructose mediated by glucose (xylose) isomerase (Fig. 5). In this typical process, fructose yield cannot be very high because the last step is reversible and the process runs at equilibrium. To solve this problem, Benner and his coworker designed a novel enzymatic pathway in one pot containing (i) starch phosphorylase, (ii) phosphoglucomutase, (iii) phosphoglucose isomerase, and (iv) fructose-6-phosphatase (Fig. 5). In the last step, the net hydrolysis of fructose-6-phosphate yields fructose via a transaldolase-catalyzed reaction between fructose-6-phosphate and glyceraldehyde to yield fructose and glyceraldehyde-3-phosphate, which is then hydrolyzed to regenerate glyceraldehyde and inorganic phosphate by using a 3-phosphoglycerate phosphatase. Because the final step is exerogenic and irreversible, this process pulls all of the equilibrium intermediates to the desired product. This pathway design illustrates a general idea that the energetics of a pathway should be considered when designing multistep biocatalytic transformations [87].
Schematic representation of enzymatic fructose production from starch. GNP alpha-glucan phosphorylase; PGM phosphoglucomutase; PGI phosphoglucose isomerase; F6P fructose-6-phosphatase. Arrows in red are reversible reactions. Modified from Ref. [87]
3.2 Cell-Free Biosystems for High-Value Product Synthesis
Cell-free systems can be used to produce high-value pharmaceuticals. Organic chemical synthesis plays a dominant role in producing chemicals in the modern pharmaceutical industry, although it is being challenged by enzyme-mediated biocatalysis [44]. For example, fondaparinux (trade name Arixtra) is a synthetic antithrombin III-binding pentasaccharide. Its chemical synthesis involves a large number of chemical synthesis reactions, resulting in very low yields of products (e.g., less than 0.5 %) and the generation of toxic intermediates [88]. As a result, Arixtra is a costly drug. In 2011, Xu et al. reported 10- and 12-step chemoenzymatic synthesis of two structurally homogeneous ultralow molecular weight heparins in 45 and 37 % overall yield, respectively, starting from simple disaccharides [89]. These ultra-low molecular weight heparins display excellent in vitro anticoagulant activity and comparable pharmacokinetic properties to Arixtra in a rabbit model. This enzymatic approach may be scaled up easily and shows great potential for a more cost-efficient way to synthesize this important drug class.
Sheldon and his coworker developed a one-pot procedure including four enzymes for the synthesis of carbohydrates from glycerol and an aldehyde (i.e., butanal) (Fig. 6). The first step is the conversion of glycerol to glycerol-3-phosphate at a cost of pyrophosphate mediated by phytase. The second enzyme (glycerol phosphate oxidase) converts glycerol-3-phosphate to DHAP and generates H2O2 as a by-product, where a third enzyme catalase is responsible for degrading H2O2 to water and oxygen. The fourth enzyme fructose-1,6-bisphosphate aldolase links DHAP and butanal to butanal-DHAP. At the last step, the first enzyme phytase releases phosphate from butanal-DHAP and generates 5-deoxy-5-ethyl-d-xylulose. Although the four enzymes have different optimal pHs, the four enzymes are put in one pot during the whole process. The pH of the reaction vessel is changed from pH 4.0 to pH 7.0 and then to pH 4.0, ensuring the cascading reactions occur in the desired order. The phytase on/off-switch at its respective pH 4.0 and 7.0 was the key to controlling phosphorylation and dephosphorylation [55].
Schematic representation of enzymatic transformation from glycerol to carbohydrates in one pot with pH shifts. GPO glycerol phosphate oxidase; ALD fructose-1,6-bisphosphate aldolase. Modified from Ref. [158]
Cell-free biosystems have been developed for the biosynthesis of radio-labeled purines (ATP and GTP) and pyrimidines (UTP and CTP) from glucose and ammonia, which requires 28 and 18 enzymes, respectively [63, 64]. Compared to the chemical synthesis, both purine and pyrimidine biosynthesis pathways can be reconstructed de novo for the incorporation of isotopes into specific sites. This work enables NMR detection for probing structural and dynamic characteristics of nucleic acids.
Enzymatic fuel cells (EFCs) represent a new type of fuel cell devices that can convert chemical energy stored in fuels into electricity mediated by redox enzymes [4, 90]. EFCs are an appealing micro-power source suitable for powering portable electronics because they have a number of features such as high-energy storage density; no flammability or explosion risk; biodegradability; low fuel costs; no costly, rare or heavy metals needed; and rapid “recharge” by injecting a sugar solution [91]. Nearly all EFCs extract only a small fraction of chemical energy in fuels by utilizing only one or a few oxidoreductase enzymes. The complete oxidation of fuels into electricity through engineered cascade pathways would have four benefits: (i) high energy utilization efficiency, (ii) high energy storage density, (iii) low product inhibition, and (iv) high power density [4, 92–94]. EFCs can utilize a large range of chemical compounds as fuels, including methanol, ethanol, glycerol, pyruvate, and glucose (these fuels are listed in increasing order of carbon number in the compound). To increase the fuel utilization efficiency, cascade enzymes can be employed. For example, three cascading redox enzymes have been used in an anode for the complete oxidization of one-carbon methanol to CO2 [95]. Similarly, two-carbon ethanol has been oxidized using an 11-enzyme pathway to generate more electrons [96]. Three-carbon glycerol and pyruvate have been oxidized using two cascading dehydrogenases [97] and enzymes in the Krebs cycle [98, 99], respectively. Furthermore, Minteer et al. have proposed the complete oxidation of glucose using glycolysis and the TCA cycle [92]. Their pathway design is more complicated than the one proposed here and involves ATP/GTP and acetyl-CoA. However, acetyl-CoA and ATP are labile and cannot be utilized for an extended period of time. Recently, Xu and Minteer [100] published another paper on glucose oxidation to CO2 through a new pathway that does not involve ATP and acetyl-CoA (Fig. 7). However, this design suffers from a very low power density because aldolase that breaks the C–C bond has a very low activity on non-phosphorylated carbohydrates [101, 102].
Schematic representation of complete oxidation of glucose for electricity generation through six enzymes. Modified from Ref. [100]
Zhang and coworkers utilized four thermophilic enzymes in cascade for deep oxidation of glucose without the use of ATP [94]. Polyphosphate glucokinase converts glucose to glucose-6-phosphate using low-cost, stable polyphosphate rather than costly ATP [81]. Two NAD-dependent dehydrogenases (glucose-6-phosphate dehydrogenase and 6-phosphogluconate dehydrogenase) that were immobilized on the bioanode were responsible for generating two NADH per glucose-6-phosphate (i.e., four electrons were generated per glucose via a diaphorase-vitamin K(3) electron shuttle system at the anode). When the temperature was increased to 50 °C, the maximum power density increased to 0.322 mW/cm−2, which was approximately eight times higher than that based on mesophilic enzymes at the same temperature. These results suggest that the deep oxidation of glucose could be achieved by using multiple dehydrogenases in synthetic cascade pathways and that high power output could be achieved by using thermostable enzymes at elevated temperatures.
3.3 Cell-Free Protein Synthesis
While cell-free protein synthesis (CFPS) has been used for decades as a foundational research tool for understanding transcription and translation, recent advances have made possible cost-effective micro-scale to manufacturing scale synthesis of complex proteins [22]. CFPS is becoming a more acceptable tool for the fast synthesis of recombinant proteins, especially when the proteins are in vivo cytotoxic, regulatory, or unstable proteins that are difficult to express in living cells and/or the proteins contain unnatural amino acids [10, 59, 103]. CFPS is usually conducted by using a crude cell lysate from any given organism (e.g., bacterial, plant, or animal cells) supplemented with the DNA template encoding the desired protein, NTPs, a highly processive RNA polymerase, amino acids, and an energy supply while the cell lysate provides the translational machinery, accessory enzymes, tRNA, and cofactors [10]. The use of cell lysate greatly simplifies CFPS but it may have some negative impacts due to other cellular components in cell extracts, for example, a rapid depletion of energy charge [59] and the degradation of protein products or template nucleic acids by proteases or nucleases [29]. In 2001, Shimizu et al. reported a protein-synthesizing system reconstituted from recombinant tagged protein factors purified to homogeneity [29]. The system – termed the “protein synthesis using recombinant elements” (PURE) system—contains all necessary translation factors, purified with high specific activity, and allows efficient protein production. This system has more than 100 well-defined molecules for implementing this complicated protein biosynthesis. The PURE system exhibits high translational efficiency with the added advantage of simple manipulation of reaction conditions and easy purification of untagged protein product. As a result, New England Biolabs Inc. sells the PURE fast protein synthesis kit.
Currently CFPS can produce protein in yields exceeding grams of target protein per liter of reaction volume, in batch reactions lasting multiple hours, with cost decreases to several orders of magnitude, and at scales reaching the 100-L milestone by SutroBio [11]. These advances have inspired new applications in the synthesis of protein libraries for functional genomics and structural biology, the production of protein therapeutics [104], personalized medicines [105], vaccines [106], short antimicrobial peptides [107], and the expression of virus-like particles, among others.
4 Challenges
The potential of cell-free biosystems for biomanufacturing is often ignored or underappreciated by many bioengineers and scientists because this paradigm shift is an out-of-the-box solution so that most do not realize that technological breakthroughs in other fields may be game changing in their field. A good example is no-till farming replacing tilling or discing farmland. Mankind has tilled land for thousands of years so it was easy to forget that the primary goal of tilling was to kill weeds. Beveridge described it as conditional thinking in his famous book entitled “The art of Scientific Investigation” [108]. Even long after the invention of selective herbicides and complementary herbicide-resistant seeds, farmers continued to till their land due to the ruling paradigm. The situation changed when several agricultural engineering pioneers in the nonpoint water pollution field realized that tilling was not necessary with proper herbicide and improved seed use. In reality, these no-tilling pioneers did not invent any new technologies but proposed a new concept to solve a key problem in their field. Now no-till farming is widely adopted because of the environmental and economic benefits [109]. Similarly, doubts regarding cell-free biosystems may include (i) enzymes cannot be produced and purified at low cost, (ii) enzymes are not stable enough, (iii) coenzymes are expensive and labile, and (iv) optimal conditions for numerous enzymes are different (Table 2). To address the above challenges, the respective solutions are listed in Table 2.
4.1 Low-Cost Enzyme Production and Purification
Cell-free protein synthesis usually uses cell extracts to decrease enzyme costs, while most cell-free biosystems prefer using partially purified enzymes that avoid unnecessary side-reactions and may prolong reaction time up to weeks, months, or even years. Low-cost production of bulk enzymes has been achieved but most academic researchers do not know this because most purchase costly enzymes from Sigma or other enzyme vendors. For example, bulk enzymes, such as protease and amylase produced by Bacillus sp., cellulase produced by Trichoderma and Aspergillus sp., have selling prices of ca. 5–10 US dollars per kg of dry protein [4, 110]. The cost of protein production is highly related to its expression level. The higher the expression level, the lower the cost of the purified protein. Codon usage optimization is a common way to enhance recombinant protein expression levels. For example, more than 500-fold improvement in the expression of soluble Thermotoga maritima 6-phosphogluconate dehydrogenase has been achieved in E. coli by codon usage optimization, accounting for >30 % of the total cellular protein [42]. Some recombinant formate dehydrogenase expression levels in E. coli are as high as 50 % cellular proteins [111]. It is estimated that current costs of recombinant proteins produced by E. coli BL 21 are approximately 100 US dollars per kg of dry protein including materials, labor, and capital depreciation [43]. As enzyme production is scaled up, production costs would decrease further. Dr. Tao at EnzymeWorks (China) pointed out that current enzyme costs in his company were approximately 70 US dollars per kg of enzyme because they can grow the E. coli cell densities of more than 100 g dry cell weight per liter without the use of costly pure oxygen (personal communication). It is anticipated that the cost of bulk recombinant enzymes will decrease greatly when their markets are ready.
The impression of costly enzyme purification is often gained from high-purity enzymes produced by academic labs and for pharmaceutical protein drugs, which are purified through a series of chromatographic methods. However, cell-free biosystems can utilize relatively low-quality enzymes as building blocks. Please bear in mind that cell extracts without purification could work well. In addition, several low-cost scalable protein purifications are available and have been developed recently. For example, ammonia sulfate precipitation can be used as the first step of most protein purifications [77]. Instead of using commercial costly protein purification resins like Ni–NTA resin, and glutathione Sepharose 4B, low-cost cellulosic materials can be used for protein adsorption/desorption, purification, and immobilization [79–82]. For highly thermophilic enzymes expressed by E. coli, heat precipitation could be the simplest way to obtain purified proteins from E. coli cell extracts [42, 75]. In our laboratory, we have systematically investigated heat precipitation for the purification of recombinant enzymes produced from E. coli. We found that six recombinant enzymes cloned from T. maritima and produced in E. coli, for example, 6-phosphogluconate dehydrogenase [42], ribose-5-phosphate isomerase [75], aldolase [112], transaldolase [113], transketolase, and xylulokinase can be purified to more than 80–90 % homogeneity by simple heat treatment at 80 °C for 20 min.
Inspired by integrated circuits and natural metabolons [114], we developed a new protein purification method which can enrich, purify, and immobilize three cascade enzymes in one step (Fig. 8a). This method was based on the highly species-specific interaction between cohesins and dockerins from natural cellulosomes [112, 115–117]. A synthetic protein scaffold called scaffoldin was constructed containing a CBM3 module and three cohesins. The CBM3 module can tightly bind on cellulosic materials and cohesins are responsible for binding with the specific dockerin-containing enzymes. Dockerins can be located in either the N- or C-terminal of the target protein [118]. After mixing four cell extracts containing the synthetic scaffoldin and dockerin-containing target proteins with cellulosic materials (e.g., regenerated amorphous cellulose, RAC) and centrifugation, the synthetic three-enzyme metabolon can be purified and immobilized on RAC (Fig. 8a). Triosephosphate isomerase (TIM), aldolase (ALD), and fructose 1,6-bisphosphatase (FBP) were chosen for demonstration purposes. The protein expression levels of dockerin-containing TIM, ALD, and FBP were low (Fig. 8b, Lane 1, 2, 3), but they can be easily enriched and purified by mixing with the synthetic scaffoldin (Lane 5, 6, and 7). The self-assembled three-enzyme metabolon can be obtained in one step (Fig. 8b, Lane 9 and Fig. 8c). Such an enzyme complex also has the unique feature of substrate channeling because of the proximity of the cascade enzymes [114]. This enzyme complex showed more than one order of magnitude enhancements on reaction rates compared to the noncomplexed TIM, ALD, and FBP mixture [112] (Fig. 9). In conclusion, numerous methods can decrease enzyme purification costs.
a Schematic representation for the purification and co-immobilization of the synthetic three-enzyme complex, where the mini-scaffoldin contained three different types of cohesins and one family 3 carbohydrate-binding module, and three enzymes contained respective dockerin. b SDS-PAGE analysis of the E. coli cell extracts containing the recombinant proteins and RAC pull-down proteins (a). Lane M, protein marker; Lane 1–4, cell extract containing TIM-Doc1, ALD-Doc2, FBP-Doc3, and CBM-Scaf3, respectively; Lane 5, RAC adsorbed mini-scaffoldin, Lane 6–8, RAC adsorbed CBM-Scaf3 and TIM-Doc1, ALD-Doc2, and FBP-Doc3, respectively; and Lane 9, RAC adsorbed CBM-Scaf3, TIM-Doc1, ALD-DoC2, and FBP-Doc3. c Schematic representation of the self-assembled three-enzyme complex containing TIM-Doc1, ALD-Doc2, FBP-Doc3, and CBM-Scaf3 containing three different types of cohesins and one family 3 carbohydrate-binding module. Modified from Ref. [159]
Profiles of fructose-6-phosphate production catalyzed by 2-μM synthetic enzyme complex (■) and 2-μM noncomplexed enzyme mixture (●) in 2.5-mM glycerealdehyde-3-phosphate at 60 °C, where the slope of the dashed lines was defined as the initial reaction rate. Modified from Ref. [112]
4.2 Prolonging Enzyme Lifetime
Most enzymes in academic laboratories deactivate rapidly, giving researchers an impression that enzymes are not stable. In reality, a few enzymes are very stable with a shelf lifetime of years and have been widely used in daily life, for example, protease used in detergents, or enzymes used in diabetic test strips. Another famous example is immobilized glucose isomerase lasting for more than 2 years [5].
Enzyme deactivation can be addressed by using thermoenzymes, enzyme immobilization, protein engineering through directed evolution and rational design, and their combination. Our economic analyses pertaining to enzymatic hydrogen production suggest that enzyme costs would be minimal when total turnover numbers (TTN) of all enzymes are larger than 107–108 mol of product per mol of enzyme [4, 77]. When cell-free biosystems are used to produce high value products, acceptable TTN values could be lower. On the basis of our experiences, it is very easy to obtain enzymes with such high TTN values by using natural thermostable enzymes (Table 3). Also, Bommarius and his coworker suggest a very simple way to calculate the TTN value of an enzyme: TTN = k cat/k d, where k cat and k d are the turn-over number and degradation constant of the enzyme, respectively [119].
The discovery and utilization of thermoenzymes may be the simplest strategy. During the past few years, we have produced a number of thermophilic enzymes in E. coli, such as Clostridium thermocellum cellodextrin phosphorylase [17], C. thermocellum cellobiose phosphorylase [17], C. thermocellum glucan phosphorylase [72], C. thermocellum phosphoglucomutase [73], T. maritima 6-phophogluconate dehydrogenase [42], T. maritima fructose bisphosphatase [74], and T. maritima pentose phosphate isomerase [75]. Most of them have TTN values of more than 107 as shown in Table 3.
Enzyme immobilization technology has been used to prolong the lifetime of enzymes for a long time [3, 120–122]. For instance, a one-step protein purification and immobilization method has been developed by using low-cost, ultra-high adsorption capacity RAC to adsorb CBM-tagged thermophilic C. thermocellum phosphoglucose isomerase (PGI) [82]. The resulting immobilized PGI is highly active and ultra-stable compared to the nonimmobilized PGI, with a TTN of more than 109 mol of product per mol of enzyme at 60 °C (Table 3). In the food industry, immobilized thermophilic glucose isomerase exhibits TTN values of ~5 × 108 mol of product per mol of enzyme. As a result, no company has a motivation to further prolong the lifetime of this enzyme.
Directed evolution and rational design are powerful approaches to enhancing the thermostability of enzymes [123, 124]. Among different desired properties of engineered enzymes, improving enzyme stability is the easiest. A number of companies and academic laboratories have developed tools for enhancing enzyme lifetime. For example, some enzyme companies and start-ups, such as Codexis, Biomethodes, EnzymeWorks, and Arzeda have demonstrated a number of successful examples for enhancing enzyme stability meeting their customers’ needs. For example, Codexis is developing a stable carbonic andrase used for capturing high-temperature and low-pH CO2 from the waste gas of power stations. Also, a combination of rational and random design also have a significant effect on enzyme stability [40].
4.3 Redox Enzyme Engineering
Since both NADP and NAD are not stable in vitro and are costly, it is important to replace them with low-cost biomimetic cofactors especially when cell-free systems are used to produce low-value biofuels and biochemicals (Fig. 10). The bulk prices of NADP, NAD, and nicotinate mononucleotide (NMN) are $4,500, $1,500, and $250 per kg, respectively (personal communication from Alex Tao). Biomimetic cofactors, such as NMN and 1-benzyl-1,4-dihydronicotinamide (BDN), not only have much lower selling prices but also have much better stability.
Redox enzyme engineering was initiated by Perham’s group over 20 years ago [125]. By using molecular modeling and comparing amino acid sequences responsible for cofactor binding sites, they changed NADP-preferred glutathione dihydrogen to NAD-preferred by site directed mutagenesis [125]. After this, by using rational design, a number of studies were conducted by swapping cofactor preferences from NADP to NAD [126–132], from NAD to NADP [133–137], and relaxing or broadening cofactor specificity [138–142].
In the 1990s, Lowe and coworkers developed a series of biomimetic analogues of NAD(P) based on triazine dyes [143–147]. Some natural dehydrogenases, such as horse liver alcohol dehydrogenase, can utilize such biomimetic cofactors for implementing redox reactions [146]. Later, Fish et al. determined that the pyrophosphate and adenosine groups associated with NAD are not essential in the hydride transfer and proposed the use of BDN or its analogues to replace natural cofactors [148]. Later, they also showed that wild-type horse liver alcohol dehydrogenase [149] and monooxygenase [150] can work on such biomimics. However, most wild-type redox enzymes cannot work on such biomimics. Clark and Fish demonstrated that a P450 mutant with two amino acid changes can work on these biomimics [151]. Also, another group demonstrated that engineered P450 can utilize Zn dust as an electron source rather than natural cofactors [152, 153]. In 2011, Zhao and coworkers presented a bio-orthogonal system that catalyzed the oxidative decarboxylation of l-malate with a dedicated biomimetic cofactor, nicotinamide flucytosine dinucleotide, where the redox enzymes were engineered by using saturation mutagenesis of the key amino acid sites [154].
NMN is a precursor of NAD(P) with a much smaller size as compared to NAD(P) (Fig. 6). A few wild-type redox enzymes function using NMN, including liver alcohol dehydrogenase [155] and glutamic dehydrogenase [156]. Recently, Scott et al. demonstrated that engineered Pyrococcus furiosus alcohol dehydrogenase has an ability to work on NMN [157].
Although the importance of redox enzyme engineering is gaining recognition for future biomanufacturing [1, 4, 14, 28], redox enzyme engineering remains at an early stage because no framework or general rules exist for engineering redox enzymes on non-natural cofactors [157]. This direction may become one of the top R&D priorities of cell-free biosystems, especially for the production of biocommodities but not for the production of pharmaceuticals that can use more costly natural cofactors.
4.4 Compromised Reaction Conditions
Although numerous enzymes used for the proof-of-concept cell-free biotransformation experiments have different optimal conditions [17, 26], they can work together under compromised conditions (e.g., 30–32 °C) where hyperthermophilic hydrogenase exhibited a very low activity. According to our experiences and literature data, most enzymes from different sources are highly exchangeable in cell-free biosystems [25]. In future applications, we will discover and utilize nearly all enzymes from one source and/or engineer some unmatched enzymes to make their optimal condition match most of other enzymes.
5 Conclusion
The significant advantages provided by cell-free biosystems (Table 1) are motivating their transition from basic research tools to a future biomanufacturing platform. Although a few obstacles to cell-free biosystems remain, all of them can be addressed by using well-known technologies (Table 2). The development of cell-free biosystems may have a similar trend to modern computers. The first prototypes were extremely costly, with low performance, and few applications. After the performance of each part (e.g., CPU, RAM) was improved greatly and standard parts were produced on a large scale, it was simple and less costly to assemble a customized high-performance computer at low prices by using the available standardized parts. Now it is time to discover and develop more stable enzymes as standardized building blocks and engineer redox enzymes that can work with less costly and more stable biomimic cofactors. Cell-free biosystems will eventually become a new biotechnology platform for biomanufacturing numerous products, especially for biocommodities because they are highly cost-sensitive to both product yields (or energy efficiencies) and production costs.
References
Zhang Y-HP, Huang W-D (2012) Constructing the electricity-carbohydrate-hydrogen cycle for a sustainability revolution. Trends Biotechnol 30:301–306
Thiel KA (2004) Biomanufacturing, from bust to boom…to bubble? Nat Biotechnol 22:1365–1372
Zhang Y-HP, Myung S, You C, Zhu ZG, Rollin J (2011) Toward low-cost biomanufacturing through cell-free synthetic biology: bottom-up design. J Mater Chem 21:18877–18886
Zhang Y-HP (2010) Production of biocommodities and bioelectricity by cell-free synthetic enzymatic pathway biotransformations: challenges and opportunities. Biotechnol Bioeng 105:663–677
Vasic-Racki D (2006) History of industrial biotransformations—Dreams and realities. In: Liese A, Seebald S, Wandrey C (eds) Industrial biotransformations. Wiley-VCH, Weinheim, pp 1–37
Lopez-Gallego F, Schmidt-Dannert C (2010) Multi-enzymatic synthesis. Curr Opin Chem Biol 14:174–183
Ricca E, Brucher B, Schrittwieser JH (2011) Multi-enzymatic cascade reactions: overview and perspectives. Adv Synth Catal 353:2239–2262
Santacoloma PA, Sin Gr, Gernaey KV, Woodley JM (2010) Multienzyme-catalyzed processes: next-generation biocatalysis. Org Proc Res Dev 15:203–212
Schoffelen S, van Hest JCM (2012) Multi-enzyme systems: bringing enzymes together in vitro. Soft Matter 8:1736–1746
Katzen F, Chang G, Kudlicki W (2005) The past, present and future of cell-free protein synthesis. Trends Biotechnol 23:150–156
Hodgman CE, Jewett MC (2012) Cell-free synthetic biology: thinking outside the cell. Metab Eng 14:261–269
Bujara M, Schümperli M, Billerbeck S, Heinemann M, Panke S (2010) Exploiting cell-free systems: Implementation and debugging of a system of biotransformations. Biotechnol Bioeng 106:376–389
Zhang YHP, You C, Chen H, Feng R (2012) Surpassing photosynthesis: High-efficiency and scalable CO2 utilization through artificial photosynthesis. In ACS Symposium Series . Recent Advances in Post-Combustion CO2 Capture Chemistry. American Chemical Society, pp275–292
Zhang Y-HP (2011) Simpler is better: high-yield and potential low-cost biofuels production through cell-free synthetic pathway biotransformation (SyPaB). ACS Catal 1:998–1009
Huang WD, Zhang Y-HP (2011) Analysis of biofuels production from sugar based on three criteria: Thermodynamics, bioenergetics, and product separation. Energy Environ Sci 4:784–792
Maeda T, Sanchez-Torres V, Wood TK (2012) Hydrogen production by recombinant Escherichia coli strains. Microb Biotechnol. doi: 10.1111/j.1751-7915.2011.00282.x
Ye X, Wang Y, Hopkins RC, Adams MWW, Evans BR, Mielenz JR, Zhang Y-HP (2009) Spontaneous high-yield production of hydrogen from cellulosic materials and water catalyzed by enzyme cocktails. ChemSusChem 2:149–152
Logan BE (2009) Exoelectrogenic bacteria that power microbial fuel cells. Nat Rev Microbiol 7:375–381
Gellett W, Schumacher J, Kesmez M, Le D, Minteer SD (2010) High current density air-breathing laccase biocathode. J Electrochem Soc 157:B557–B562
Zebda A, Gondran C, Le Goff A, Holzinger M, Cinquin P, Cosnier S (2011) Mediatorless high-power glucose biofuel cells based on compressed carbon nanotube-enzyme electrodes. Nat Commun 2:370
Zhang Y-HP (2010) Renewable carbohydrates are a potential high density hydrogen carrier. Int J Hydrogen Energy 35:10334–10342
Carlson ED, Gan R, Hodgman CE, Jewett MC (2012) Cell-free protein synthesis: applications come of age. Biotechnol Adv 30:1185–1194
Guterl J-K, Garbe D, Carsten J, Steffler F, Sommer B, Reiße S, Philipp A, Haack M, Rühmann B, Kettling U, et al (2012) Cell-free metabolic engineering—production of chemicals via minimized reaction cascades. ChemSusChem. doi: 10.1002/cssc.201200365
Wang Y, Huang W, Sathitsuksanoh N, Zhu Z, Zhang Y-HP (2011) Biohydrogenation from biomass sugar mediated by in vitro synthetic enzymatic pathways. Chem Biol 18:372–380
Zhang Y-HP, Sun J-B, Zhong J–J (2010) Biofuel production by in vitro synthetic pathway transformation. Curr Opin Biotechnol 21:663–669
Zhang Y-HP, Evans BR, Mielenz JR, Hopkins RC, Adams MWW (2007) High-yield hydrogen production from starch and water by a synthetic enzymatic pathway. PLoS One 2:e456
Bujara M, Schümperli M, Pellaux R, Heinemann M, Panke S (2011) Optimization of a blueprint for in vitro glycolysis by metabolic real-time analysis. Nat Chem Biol 7:271–277
Swartz JR (2011) Transforming biochemical engineering with cell-free biology. AIChE J 58:5–13
Shimizu Y, Inoue A, Tomari Y, Suzuki T, Yokogawa T, Nishikawa K, Ueda T (2001) Cell-free translation reconstituted with purified components. Nat Biotechnol 19:751–755
Buchner E (1897) Alkoholische Gärung ohne Hefezellen (Vorläufige Mitteilung). Berichte der Deutschen Chemischen Gesellschaft 30:117–124
Harden A, Young WJ (1907) The alcoholic ferment of yeast-juice. Proc Roy Soc London 77B:405–422
Warburg OH (1926) Über den Stoffwechsel der Tumoren. Springer, Berlin
Cori CF (1931) Mammalian carbohydrate metabolism. Physiol Rev 11:143–275
Cori GT, Cori CF (1936) The formation of hexosephosphate esters in frog muscle. J Biol Chem 116:119–128
Krebs HA, Eggleston LV (1944) Metabolism of acetoacetic acid in animal tissues. Nature 154:209–210
Calvin M, Benson AA (1948) The path of carbon in photosynthesis. Science 107:476–480
Nirenberg MW, Matthaei JH (1961) The dependence of cell-free protein synthesis in E. coli upon naturally occurring or synthetic polyribonucleotides. Proc Natl Acad Sci USA 47:1588–1602
Michels P, Rosazza J (2009) The evolution of microbial transformations for industrial applications. SIM News 2009:36–52
Demain AL (2004) Pickles, pectin, and penicillin. Annu Rev Microbiol 58:1–42
Ye X, Zhang C, Zhang YHP (2012) Engineering a large protein by combined rational and random approaches: stabilizing the Clostridium thermocellum cellobiose phosphorylase. Mol BioSyst 8:1815–1823
Shiloach J, Fass R (2005) Growing E. coli to high cell density—a historical perspective on method development. Biotechnol Adv 23:345–357
Wang Y, Zhang Y-HP (2009) Overexpression and simple purification of the Thermotoga maritima 6-phosphogluconate dehydrogenase in Escherichia coli and its application for NADPH regeneration. Microb Cell Fact 8:30
Tufvesson Pr, Lima-Ramos J, Nordblad M, Woodley JM (2011) Guidelines and cost analysis for catalyst production in biocatalytic processes. Org Proc Res Dev 15:266–274
Bornscheuer UT, Huisman GW, Kazlauskas RJ, Lutz S, Moore JC, Robins K (2012) Engineering the third wave of biocatalysis. Nature 485:185–194
Daines AM, Maltman BA, Flitsch SL (2004) Synthesis and modifications of carbohydrates, using biotransformations. Curr Opin Chem Biol 8:106–113
Chi Y, Scroggins ST, Frechet JMJ (2008) One-pot multi-component asymmetric cascade reactions catalyzed by soluble star polymers with highly branched non-interpenetrating catalytic cores. J Am Chem Soc 130:6322–6323
Lynd LR, Weimer PJ, van Zyl WH, Pretorius IS (2002) Microbial cellulose utilization: fundamentals and biotechnology. Microbiol Mol Biol Rev 66:506–577
Liao HH, Zhang XZ, Rollin JA, Zhang Y-HP (2011) A minimal set of bacterial cellulases for consolidated bioprocessing of lignocellulose. Biotechnol J 6:1409–1418
Wildeman SMAD, Sonke T, Schoemaker HE, May O (2007) Biocatalytic reductions: from lab curiosity to “first choice”. Acc Chem Res 40:1260–1266
Wichmann R, Vasic-Racki D (2005) Cofactor regeneration at the lab scale. Adv Biochem Eng Biotechnol 92:225–260
Bozic M, Pricelius S, Guebitz GM, Kokol V (2010) Enzymatic reduction of complex redox dyes using NADH-dependent reductase from Bacillus subtilis coupled with cofactor regeneration. Appl Microbiol Biotechnol 85:563–571
Xu Z, Jing K, Liu Y, Cen P (2007) High-level expression of recombinant glucose dehydrogenase and its application in NADPH regeneration. J Ind Microbiol Biotechnol 34:83–90
Mertens R, Liese A (2004) Biotechnological applications of hydrogenases. Curr Opin Biotechnol 15:343–348
Johannes TW, Woodyer RD, Zhao H (2007) Efficient regeneration of NADPH using an engineered phosphite dehydrogenase. Biotechnol Bioeng 96:18–26
Schoevaart R, van Rantwijk F, Sheldon RA (2000) A four-step enzymatic cascade for the one-pot synthesis of non-natural carbohydrates from glycerol. J Org Chem 65:6940–6943
Zhang J, Shao J, Kowal P, Wang PG (2005) Enzymatic Synthesis of Oligosaccharides. Wiley-VCH Verlag GmbH & Co.KGaA, Weinheim
Fessner W-D, Helaine V (2001) Biocatalytic synthesis of hydroxylated natural products using aldolases and related enzymes. Curr Opin Biotechnol 12:574–586
Endo T, Koizumi S (2000) Large-scale production of oligosaccharides using engineered bacteria. Curr Opin Struct Biol 10:536–541
Wang Y, Zhang Y-HP (2009) Cell-free protein synthesis energized by slowly-metabolized maltodextrin. BMC Biotechnol 9:58
Calhoun KA, Swartz JR (2005) An economical method for cell-free protein synthesis using glucose and nucleoside monophosphates. Biotechnol Prog 21:1146–1153
Hold C, Panke S (2009) Towards the engineering of in vitro systems. J Royal Soc Interface 6:S507–S521
Panke S, Held M, Wubbolts M (2004) Trends and innovations in industrial biocatalysis for the production of fine chemicals. Curr Opin Biotechnol 15:272–279
Schultheisz HL, Szymczyna BR, Scott LG, Williamson JR (2008) Pathway engineered enzymatic de Novo purine nucleotide synthesis. ACS Chem Biol 3:499–511
Schultheisz HL, Szymczyna BR, Williamson JR (2009) Enzymatic synthesis and structural characterization of 13C, 15 N-poly(ADP-ribose). J Am Chem Soc 131:14571–14578
Lynd LR, Wyman CE, Gerngross TU (1999) Biocommodity engineering. Biotechnol Prog 15:777–793
Zhang Y-HP (2011) What is vital (and not vital) to advance economically-competitive biofuels production. Proc Biochem 46:2091–2110
Zhang Y-HP (2011) Hydrogen production from carbohydrates: a mini-review. ACS Symp Ser 1067:203–216
Adams MWW, Stiefel EI (1998) Biological hydrogen production: not so elementary. Science 282:1842–1843
Cortright RD, Davda RR, Dumesic JA (2002) Hydrogen from catalytic reforming of biomass-derived hydrocarbons in liquid water. Nature 418:964–967
Thauer K, Jungermann K, Decker K (1977) Energy conservation in chemotrophic anaerobic bacteria. Bacteriol Rev 41:100–180
The Royal Society of the UK (2007) Synthetic biology: call for views. http://royalsociety.org/page.asp?changes=0&latest=1&id=6731
Ye X, Rollin J, Zhang Y-HP (2010) Thermophilic α-glucan phosphorylase from Clostridium thermocellum: cloning, Characterization and enhanced thermostability. J Mol Cat B Enzym 65:110–116
Wang Y, Zhang Y-HP (2010) A highly active phosphoglucomutase from Clostridium thermocellum: Cloning, purification, characterization, and enhanced thermostability. J Appl Microbiol 108:39–46
Myung S, Wang YR, Zhang Y-HP (2010) Fructose-1,6-bisphosphatase from a hyper-thermophilic bacterium Thermotoga maritima: Characterization, metabolite stability and its implications. Proc Biochem 45:1882–1887
Sun FF, Zhang XZ, Myung S, Zhang Y-HP (2012) Thermophilic Thermotoga maritima ribose-5-phosphate isomerase RpiB: Optimized heat treatment purification and basic characterization. Protein Expr Purif 82:302–307
Sun J, Hopkins RC, Jenney FE, McTernan PM, Adams MWW (2010) Heterologous expression and maturation of an NADP-dependent [NiFe]-hydrogenase: a key enzyme in biofuel production. PLoS One 5:e10526
Zhang Y-HP, Mielenz JR (2011) Renewable hydrogen carrier—carbohydrate: constructing the carbon-neutral carbohydrate economy. Energies 4:254–275
Scopes RK (1993) Protein purification: principles and practice, 3rd edn. Springer, New York
Hong J, Wang Y, Ye X, Zhang Y-HP (2008) Simple protein purification through affinity adsorption on regenerated amorphous cellulose followed by intein self-cleavage. J Chromatogr A 1194:150–154
Hong J, Ye X, Wang Y, Zhang Y-HP (2008) Bioseparation of recombinant cellulose binding module-protein by affinity adsorption on an ultra-high-capacity cellulosic adsorbent. Anal Chim Acta 621:193–199
Liao HH, Myung S, Zhang Y-HP (2012) One-step purification and immobilization of thermophilic polyphosphate glucokinase from Thermobifida fusca YX: glucose-6-phosphate generation without ATP. Appl. Microbiol Biotechnol 93:1109–1117
Myung S, Zhang X-Z, Zhang Y-HP (2011) Ultra-stable phosphoglucose isomerase through immobilization of cellulose-binding module-tagged thermophilic enzyme on low-cost high-capacity cellulosic adsorbent. Biotechnol Prog 27:969–975
Bai FW, Anderson WA, Moo-Young M (2008) Ethanol fermentation technologies from sugar and starch feedstocks. Biotechnol Adv 26:89–105
Welch P, Scopes RK (1985) Studies on cell-free metabolism: Ethanol production by a yeast glycolytic system reconstituted from purified enzymes. J Biotechnol 2:257–273
Li S, Wen J, Jia X (2011) Engineering Bacillus subtilis for isobutanol production by heterologous Ehrlich pathway construction and the biosynthetic 2-ketoisovalerate precursor pathway overexpression. Appl Microbiol Biotechnol 91:577–589
Atsumi S, Hanai T, Liao JC (2008) Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 451:86–89
Moradian A, Benner SA (1992) A biomimetic biotechnological process for converting starch to fructose: thermodynamic and evolutionary considerations in applied enzymology. J Am Chem Soc 114:6980–6987
Petitou M, van Boeckel CAA (2004) A synthetic antithrombin iii binding pentasaccharide is now a drug! What comes next? Angew Chem Int Ed 43:3118–3133
Xu Y, Masuko S, Takieddin M, Xu H, Liu R, Jing J, Mousa SA, Linhardt RJ, Liu J (2011) Chemoenzymatic synthesis of homogeneous ultralow molecular weight heparins. Science 334:498–501
Moehlenbrock M, Minteer S (2008) Extended lifetime biofuel cells. Chem Soc Rev 37:1188–1196
Zhang Y-HP, Xu J-H, Zhong JJ (2012) A new high-energy density hydrogen carrier - carbohydrate - might be better than methanol. Int. J. Energy Res. Epub, doi: 10.1002/er.2897
Minteer SD, Liaw BY, Cooney MJ (2007) Enzyme-based biofuel cells. Curr. Opin. Biotechnol. 18:228–234
Cooney MJ, Svoboda V, Lau C, Martin G, Minteer SD (2008) Enzyme catalysed biofuel cells. Energy Environ Sci 1:320–337
Zhu ZG, Sun F, Zhang X, Zhang Y-HP (2012) Deep oxidation of glucose in enzymatic fuel cells through a synthetic enzymatic pathway containing a cascade of two thermostable dehydrogenases. Biosens Bioelectron 36:110–115
Palmore GTR, Bertschy H, Bergens SH, Whitesides GM (1998) A methanol/dioxygen biofuel cell that uses NAD+-dependent dehydrogenases as catalysts: application of an electro-enzymatic method to regenerate nicotinamide adenine dinucleotide at low overpotentials. J Electroanal Chem 443:155–161
Sokic-Lazic D, Minteer SD (2008) Citric acid cycle biomimic on a carbon electrode. Biosens Bioelectron 24:939–944
Arechederra RL, Treu BL, Minteer SD (2007) Development of glycerol/O-2 biofuel cell. J Power Sources 173:156–161
Sokic-Lazic D, Minteer SD (2009) Pyruvate/air enzymatic biofuel cell capable of complete oxidation. Electrochem Solid-State Lett 12:F26–F28
Moehlenbrock MJ, Toby TK, Waheed A, Minteer SD (2010) Metabolon catalyzed pyruvate/air biofuel cell. J Am Chem Soc 132:6288–6289
Xu S, Minteer SD (2011) Enzymatic biofuel cell for oxidation of glucose to CO2. ACS Catal 1:91–94
Marsh JJ, Lebherz HG (1992) Fructose-bisphosphate aldolases: an evolutionary history. Trends Biochem Sci 17:110–113
Hibbert EG, Senussi T, Costelloe SJ, Lei W, Smith MEB, Ward JM, Hailes HC, Dalby PA (2007) Directed evolution of transketolase activity on non-phosphorylated substrates. J Biotechnol 131:425–432
Boyer ME, Stapleton JA, Kuchenreuther JM, Wang C-w, Swartz JR (2008) Cell-free synthesis and maturation of [FeFe] hydrogenases. Biotechnol Bioeng 99:59–67
Kim D-M, Swartz JR (2004) Efficient production of a bioactive, multiple disulfide-bonded protein using modified extracts of Escherichia coli. Biotechnol Bioeng 85:122–129
Kanter G, Yang J, Voloshin A, Levy S, Swartz JR, Levy R (2007) Cell-free production of scFv fusion proteins: an efficient approach for personalized lymphoma vaccines. Blood 109:3393–3399
Bundy BC, Franciszkowicz MJ, Swartz JR (2008) Escherichia coli-based cell-free synthesis of virus-like particles. Biotechnol Bioeng 100:28–37
Lee K-H, Kwon Y-C, Yoo SJ, Kim D-M (2010) Ribosomal synthesis and in situ isolation of peptide molecules in a cell-free translation system. Protein Expr Purif 71:16–20
Beveridge WIB (1960) The art of scientific investigation. Vintage, New York
Smith P, Powlson DS, Glendining MJ, Smith JU (1998) Preliminary estimates of the potential for carbon mitigation in European soils through no-till farming. Glob Change Biol 4:679–685
Klein-Marcuschamer D, Oleskowicz-Popiel P, Simmons BA, Blanch HW (2012) The challenge of enzyme cost in the production of lignocellulosic biofuels. Biotechnol Bioeng 109:1083–1087
Tishkov VI, Popov VO (2006) Protein engineering of formate dehydrogenase. Biomol Eng 23:89–110
You C, Myung S, Zhang Y-HP (2012) Facilitated substrate channeling in a self-assembled trifunctional enzyme complex. Angew Chem Int Ed 51:8787–8790
Huang SY, Zhang Y-HP, Zhong JJ (2012) A thermostable recombinant transaldolase with high activity over a broad pH range. Appl Microbiol Biotechnol 93:2403–2410
Zhang Y-HP (2011) Substrate channeling and enzyme complexes for biotechnological applications. Biotechnol Adv 29:715–725
Bayer EA, Morag E, Lamed R (1994) The cellulosome–a treasure-trove for biotechnology. Trends Biotechnol 12:379–386
You C, Zhang X-Z, Sathitsuksanoh N, Lynd LR, Zhang Y-HP (2012) Enhanced microbial cellulose utilization of recalcitrant cellulose by an ex vivo cellulosome-microbe complex. Appl Environ Microbiol 78:1437–1444
You C, Zhang X-Z, Zhang YHP (2012) Mini-scaffoldin enhanced mini-cellulosome hydrolysis performance on low-accessibility cellulose (Avicel) more than on high-accessibility amorphous cellulose. Biochem Eng J 63:57–65
Moraïs S, Barak Y, Hadar Y, Wilson DB, Shoham Y, Lamed R, Bayer EA (2011) Assembly of xylanases into designer cellulosomes promotes efficient hydrolysis of the xylan component of a natural recalcitrant cellulosic substrate. MBio 2 e00233-11
Rogers TA, Bommarius AS (2010) Utilizing simple biochemical measurements to predict lifetime output of biocatalysts in continuous isothermal processes. Chem Eng Sci 65:2118–2124
Cao L, Langen Lv, Sheldon RA (2003) Immobilised enzymes: carrier-bound or carrier-free? Curr Opin Biotechnol 14:387–394
Cao L (2005) Immobilised enzymes: science or art? Curr Opin Chem Biol 9:217–226
Hartmann M, Jung D (2010) Biocatalysis with enzymes immobilized on mesoporous hosts: the status quo and future trends. J Mater Chem 20:844–857
Arnold FH, Volkov AA (1999) Directed evolution of biocatalysts. Curr Opin Chem Biol 3:54–59
Eijsink VG, Bjork A, Gaseidnes S, Sirevag R, Synstad B, van den Burg B, Vriend G (2004) Rational engineering of enzyme stability. J Biotechnol 113:105–120
Scrutton NS, Berry A, Perham RN (1990) Redesign of the coenzyme specificity of a dehydrogenase by protein engineering. Nature 343:38–43
Zhang L, Ahvazi B, Szittner R, Vrielink A, Meighen E (1999) Change of nucleotide specificity and enhancement of catalytic efficiency in single point mutants of Vibrio harveyi aldehyde dehydrogenase. Biochemistry 38:11440–11447
Yaoi T, Miyazaki K, Oshima T, Komukai Y, Go M (1996) Conversion of the coenzyme specificity of isocitrate dehydrogenase by module replacement. J Biochem 119:1014–1018
Bastian S, Liu X, Meyerowitz JT, Snow CD, Chen MMY, Arnold FH (2011) Engineered ketol-acid reductoisomerase and alcohol dehydrogenase enable anaerobic 2-methylpropan-1-ol production at theoretical yield in Escherichia coli. Metab Eng 13:345–352
Rosell A, Valencia E, Ochoa WF, Fita I, Pares X, Farres J (2003) Complete reversal of coenzyme specificity by concerted mutation of three consecutive residues in alcohol dehydrogenase. J Biol Chem 278:40573–40580
Döhr O, Paine MJI, Friedberg T, Roberts GCK, Wolf CR (2001) Engineering of a functional human NADH-dependent cytochrome P450 system. Proc Natl Acad Sci USA 98:81–86
Banta S, Swanson BA, Wu S, Jarnagin A, Anderson S (2002) Alteration of the specificity of the cofactor-binding pocket of Corynebacterium 2,5-diketo-D-gluconic acid reductase A. Protein Eng Des Sel 15:131–140
Banta S, Swanson BA, Wu S, Jarnagin A, Anderson S (2002) Optimizing an artificial metabolic pathway: Engineering the cofactor specificity of Corynebacterium 2,5-Diketo-D-gluconic acid reductase for use in vitamin C biosynthesis. Biochemistry 41:6226–6236
Bocanegra JA, Scrutton NS, Perham RN (1993) Creation of an NADP-dependent pyruvate dehydrogenase multienzyme complex by protein engineering. Biochemistry 32:2737–2740
Mittl PRE, Berry A, Scrutton NS, Perham RN, Schulz GE (1993) Structural differences between wild-type NADP-dependent glutathione reductase from Escherichia coli and a redesigned NAD-dependent mutant. J Mol Biol 231:191–195
Steen IH, Lien T, Madsen MS, Birkeland N-K (2002) Identification of cofactor discrimination sites in NAD-isocitrate dehydrogenase from Pyrococcus furiosus. Arch Microbiol 178:297–300
Watanabe S, Kodaki T, Makino K (2005) Complete reversal of coenzyme specificity of xylitol dehydrogenase and increase of thermostability by the introduction of structural zinc. J Biol Chem 280:10340–10349
Glykys DJ, Banta S (2009) Metabolic control analysis of an enzymatic biofuel cell. Biotechnol Bioeng 102:1624–1635
Woodyer RD, van der Donk WA, Zhao H (2003) Relaxing the nicotinamide cofactor specificity of phosphite dehydrogenase by rational design. Biochemistry 42:11604–11614
Wiegert T, Sahm H, Sprenger GA (1997) The substitution of a single amino acid residue (Ser-116 → Asp) alters NADP-containing glucose-fructose oxidoreductase of Zymomonas mobilis into a glucose dehydrogenase with dual coenzyme specificity. J Biol Chem 272:13126–13133
Katzberg M, Skorupa-Parachin N, Gorwa-Grauslund M-F, Bertau M (2010) Engineering cofactor preference of ketone reducing biocatalysts: A mutagenesis study on a γ-Diketone reductase from the yeast Saccharomyces cerevisiae serving as an example. Int J Mol Sci 11:1735–1758
Sanli G, Banta S, Anderson S, Blaber M (2004) Structural alteration of cofactor specificity in Corynebacterium 2,5-diketo-D-gluconic acid reductase. Protein Eng 13:504–512
Campbell E, Wheeldon IR, Banta S (2010) Broadening the cofactor specificity of a thermostable alcohol dehydrogenase using rational protein design introduces novel kinetic transient behavior. Biotechnol Bioeng 107:763–774
Burton SJ, Vivian Stead C, Ansell RJ, Lowe CR (1996) An artificial redox coenzyme based on a triazine dye template. Enzym Microb Technol 18:570–580
Ansell RJ, Dilmaghanian S, Stead CV, Lowe CR (1997) Synthesis and properties of new coenzyme mimics based on the artificial coenzyme Blue N-3. Enzym Microb Technol 21:327–334
Ansell RJ, Small DAP, Lowe CR (1997) Characterisation of the artificial coenzyme CL4. J Mol Catal B Enzym 3:239–252
Ansell RJ, Lowe CR (1999) Artificial redox coenzymes: biomimetic analogues of NAD+. Appl Microbiol Biotechnol 51:703–710
Ansell RJ, Small DAP, Lowe CR (1999) Synthesis and properties of new coenzyme mimics based on the artificial coenzyme CL4. J Mol Recognit 12:45–56
Lo HC, Leiva C, Buriez O, Kerr JB, Olmstead MM, Fish RH (2001) Bioorganometallic chemistry. 13. regioselective reduction of NAD+ models, 1-benzylnicotinamde triflate and beta-nicotinamide ribose-5′-methyl phosphate, with in situ generated [Cp*Rh(Bpy)H]+: structure–activity relationships, kinetics, and mechanistic aspects in the formation of the 1,4-NADH derivatives. Inorg Chem 40:6705–6716
Lo HC, Fish RH (2002) Biomimetic NAD+ models for tandem cofactor regeneration, horse liver alcohol dehydrogenase recognition of 1,4-NADH derivatives, and chiral synthesis. Angew Chem Int Ed 41:478–481
Lutz J, Hollmann F, Ho TV, Schnyder A, Fish RH, Schmid A (2004) Bioorganometallic chemistry: biocatalytic oxidation reactions with biomimetic NAD+/NADH co-factors and [Cp*Rh(bpy)H]+ for selective organic synthesis. J Organomet Chem 689:4783–4790
Ryan JD, Fish RH, Clark DS (2008) Engineering cytochrome P450 enzymes for improved activity towards biomimetic 1,4-NADH cofactors. ChemBioChem 9:2579–2582
Nazor J, Schwaneberg U (2006) Laboratory evolution of P450 BM-3 for mediated electron transfer. ChemBioChem 7:638–644
Nazor J, Dannenmann S, Adjei RO, Fordjour YB, Ghampson IT, Blanusa M, Roccatano D, Schwaneberg U (2008) Laboratory evolution of P450 BM3 for mediated electron transfer yielding an activity-improved and reductase-independent variant. Protein Eng Des Sel 21:29–35
Ji D, Wang L, Hou S, Liu W, Wang J, Wang Q, Zhao ZK (2011) Creation of bioorthogonal redox systems depending on nicotinamide flucytosine dinucleotide. J Am Chem Soc 133:20857–20862
Plapp BV, Sogin DC, Dworschack RT, Bohlken DP, Woenckhaus C, Jeck R (1986) Kinetics and native and modified liver alcohol dehydrogenase with coenzyme analogs: isomerization of enzyme-nicotinamide adenine dinucleotide complex. Biochemistry 25:5396–5402
Fisher HF, McGregor LL (1969) The ability of reduced nicotinamide mononucleotide to function as a hydrogen donor in the glutamic dehydrogenase reaction. Biochem Biophys Res Commun 34:627–632
Campbell E, Meredith M, Minteer SD, Banta S (2012) Enzymatic biofuel cells utilizing a biomimetic cofactor. Chem Commun 48:1898–1900
Schoevaart R, van Rantwijk F, Sheldon RA (1999) Carbohydrates from glycerol: an enzymatic four-step, one-pot synthesis. Chem Commun 31:2465–2466
You C, Zhang Y-HP (2012) Self-assembly of synthetic metabolons through synthetic protein scaffolds: one-step purification, co-immobilization, and substrate channeling. ACS Syn. Biol. doi: 10.1021/sb300068g
Studier FW (2005) Protein production by auto-induction in high density shaking cultures. Protein Expr Purif 41:207–234
Banki MR, Feng L, Wood DW (2005) Simple bioseparations using self-cleaving elastin-like polypeptide tags. Nat Methods 2:659–662
Iturrate L, Sanchez-Moreno I, Doyaguez EG, Garcia-Junceda E (2009) Substrate channelling in an engineered bifunctional aldolase/kinase enzyme confers catalytic advantage for C–C bond formation. Chem Commun 2009:1721–1723
Bulow L, Ljungcrantz P, Mosbach K (1985) Preparation of a soluble bifunctional enzyme by gene fusion. Nat Biotechnol 3:821–823
Chen X, Liu Z, Zhang J, Zhang W, Kowal P, Wang P (2002) Reassembled biosynthetic pathway for large-scale carbohydrate synthesis: α-gal epitope producing “superbug”. ChemBioChem 4:47–53
Nahalka J, Liu Z, Chen X, Wang PG (2003) Superbeads: Immobilization in “sweet” chemistry. Chem Eur J 9:372–377
Demain AL, Vaishnav P (2009) Production of recombinant proteins by microbes and higher organisms. Biotechnol Adv 27:297–306
Kirk O, Borchert TV, Fuglsang CC (2002) Industrial enzyme applications. Curr Opin Biotechnol 13:345–351
Liu W, Wang P (2007) Cofactor regeneration for sustainable enzymatic biosynthesis. Biotechnol Adv 25:369–384
Chen K, Arnold FH (1993) Turning the activity of an enzyme for unusual environments: sequential random mutagenesis of subtilisin E for catalysis in dimethylformamide. Proc Natl Acad Sci USA 90:5618–5622
Acknowledgments
This work was supported partially by the Shell Game Charger Program, DOE BioEnergy Science Center (BESC), DOE ARPA-E Petro project, the College of Agriculture and Life Sciences Bioprocessing and Biodesign Research Center at Virginia Tech, and NSF SBIR.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
You, C., Zhang, YH.P. (2012). Cell-Free Biosystems for Biomanufacturing. In: Zhong, JJ. (eds) Future Trends in Biotechnology. Advances in Biochemical Engineering/Biotechnology, vol 131. Springer, Berlin, Heidelberg. https://doi.org/10.1007/10_2012_159
Download citation
DOI: https://doi.org/10.1007/10_2012_159
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-36507-2
Online ISBN: 978-3-642-36508-9
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)