Abstract
Oral cavity cancers are the seventh tumor by diffusion worldwide with more than 90 % being diagnosed as oral squamous cell carcinomas (OSCCs). According to the latest WHO statistics, OSCC accounts for 5 % of the cancer deaths worldwide, being the eighth more lethal cancer entity. Early identification of cancer relapses would have the potentiality to improve the disease control and the patient survival.
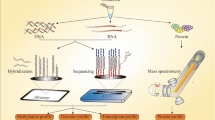
NeoMark is a European co-funded research project (Seventh Framework Program, Information and Communication Technologies: EU-FP7-ICT-2007-2-22483-NeoMark) that has the objective to identify relevant biomarkers of OSCC recurrence. It integrates high-throughput gene expression analysis in tumor cells and IT-assisted imaging with traditional staging and follow-up protocols to improve the recurrence risk stratification and to obtain the earlier identification of locoregional relapses.
The architecture of the project is based on the following key points:
-
Creation of a web application tool: a unified interface that helps the storage and management of all information
-
NeoMark database: the heterogeneous NeoMark data (demographics and risk factors; clinical, pathological, and immunohistochemical parameters; filtered and cleaned genomic and imaging data) are stored in a single database – the Integrated Health Record Repository (IHRR) – on a central NeoMark server. The server contains the marker definition functional environment (MDFE), a data analysis module. Based on the heterogeneous input data, it estimates the likelihood of a relapse and identifies OSCC risk factors.
-
Imaging biomarker extraction: several biomarkers are obtained from medical images such as CT and MRI scans (size, amount of necrosis from tumor and lymph nodes, etc.). To extract those features, a custom software tool – called the NeoMark Image Processing Tool – has specifically been developed.
-
Genomic data cleaning and filtering: extraction of genomic data and filtering of genes with low data quality and of those with high number of missing values.
The NeoMark system was trained and initially validated in a multicenter pilot study (three European clinical centers involved: two in Italy and one in Spain) basing on 86 patients affected by OSCC with a minimum follow-up of 12 months.
The clinicians recognized the usefulness of the disease bioprofile (or disease-specific profile) identified by NeoMark to evaluate the risk of disease reoccurrence of a patient at diagnosis, to stratify patients affected by OSCC at baseline according to the risk of recurrence, and to reserve a “tailored therapy” to each case.
Similar content being viewed by others
Abbreviations
- DBN:
-
Dynamic Bayesian Network
- FE:
-
Feature Extraction
- ICT:
-
Information and Communication Technologies
- IHRR:
-
Integrated Health Record Repository
- MDFE:
-
Marker Definition Functional Environment
- OSCC:
-
Oral Squamous Cell Carcinomas
- PCR:
-
Polymerase Chain Reaction
- RECIST:
-
Response Evaluation Criteria in Solid Tumors
- ROI:
-
Region(s) of Interest
- RT-PCR:
-
Real-Time PCR
- SW:
-
Software
- WHO:
-
World Health Organization
References
Bisdas S, et al. Outcome prediction after surgery and chemioradiation of squamous cell carcinoma in the oral cavity, oropharynx and hypopharynx: use of baseline perfusion CT microcirculatory parameters vs tumor volume. Int J Radiat Oncol Biol Phys. 2009;73(5):1313–8. Epub 2008 Oct 27.
Chen C, et al. Gene expression profiling identifies genes predictive of oral squamous cell carcinoma. Cancer Epidemiol Biomarkers Prev. 2008;17(8):2152–62.
Eden E, Navon R, Steinfeld I, Lipson D, Yakhin Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinf. 2009;10:48. doi:10.1186/1471-2105-10-48.
Exarchos KP, Goletsis Y, Fotiadis DI. A multiscale and multiparametric approach for modeling the progression of oral cancer. BMC Med Inform Decis Mak. 2012;12:136.
Fascina PA, et al. Evaluation of mathematical models for breast cancer risk assessment in routine clinical use. Eur J Cancer Prev. 2007;16(3):216–24.
Gene Ontology enRIchment anaLysis and visuaLizAtion tool. http://cbl-gorilla.cs.technion.ac.il.
Gerds TA, et al. The performance of risk prediction models. Biom J. 2008;50(4):457–79. Review.
Gil Z, et al. Lymph node density is a significant predictor of outcome in patients with oral cancer. Cancer. 2009;115(24):5700–10.
Giles CW, et al. Molecular classification of oral cancer by cDNA microarrays identifies overexpressed genes correlated with nodal metastasis. Int J Cancer. 2004;110:857–68.
Guarino N, Oberle D, Staab S. What is an ontology? In: Handbook on ontologies, International handbooks on information systems. Berlin/Heidelberg: Springer; 2009. p. 1–17.
Liao CT, Lee LY, Huang SF, Chen IH, Kang CJ, Lin CY, Fan KH, Wang HM, Ng SH, Yen TC. Outcome analysis of patients with oral cavity cancer and extracapsular spread in neck lymph nodes. Int J Radiat Oncol Biol Phys. 2011 Nov 15;81(4):930–7.
Liu X, Yu J, Jiang L, Wang A, Shi F, Ye H, Zhou X. MicroRNA-222 regulates cell invasion by targeting matrix metalloproteinase 1 (MMP1) and manganese superoxide dismutase 2 (SOD2) in tongue squamous cell carcinoma cell lines. Cancer Genomics Proteomics. 2009;6(3):131–9.
Liu X, Wang A, Lo Muzio L, Kolokythas A, Sheng S, Rubini C, Ye H, Shi F, Yu T, Crowe DL, Zhou X. Deregulation of manganese superoxide dismutase (SOD2) expression and lymph node metastasis in tongue squamous cell carcinoma. BMC Cancer. 2010;10:365.
Murphy KP. Dynamic Bayesian networks: representation, inference and learning. In: Computer science division. Berkeley: University of California; 2002.
Peng C-H, Liao C-T, Peng S-C, Chen Y-J, Cheng A-J, et al. A novel molecular signature identified by systems genetics approach predicts prognosis in oral squamous cell carcinoma. PLoS One. 2011;6(8):e23452. doi:10.1371/journal.pone.0023452.
Pillai R, et al. Do standardised prognostic algorithms reflect local practice? Application of EORTC risk tables for non-muscle invasive (pTa/pT1) bladder cancer recurrence and progression in a local cohort. Sci World J. 2011;11:751–9.
Pusapati RV, Weaks RL, Rounbehler RJ, McArthur MJ, Johnson DG. E2F2 suppresses Myc-induced proliferation and tumorigenesis. Mol Carcinog. 2010;49(2):152–6.
Reis PP, Waldron L, Perez-Ordonex B, Pintilie M, Naranjo Galloni N, Xuan Y, Cervigne NK, Warner GC, Makitie AA, Simpson C, et al. A gene signature in histologically normal surgical margins is predictive of oral carcinoma recurrence. BMC Cancer. 2011;11:437.doi:10.1186/1471-2407-11-437, Epub http://www.biomedcentral.com/1471-2407/11/437.
Rogers SN, et al. Survival following primary surgery for oral cancer. Oral Oncol. 2009;45(3):201–11. Epub 2008 July 31.
Saintigny P, et al. Gene expression profiling predicts the development of oral cancer. Cancer Prev Res (Phila). 2011;4(2):218–29.
Schmidt M, et al. Long-term outcome prediction by clinicopathological risk classification algorithms in node-negative breast cancer--comparison between Adjuvant!, St Gallen, and a novel risk algorithm used in the prospective randomized Node-Negative-Breast Cancer-3 (NNBC-3) trial. Ann Oncol. 2009;20(2):258–64. Epub 2008 Sept 29.
Smith B, Ashburner M, Rosse C, Bard J, Bug W, Ceusters W, Goldberg LJ, Eilbeck K, Ireland A, Mungall CJ, the OBI Consortium, Leontis N, Rocca-Serra P, Ruttenberg A, Sansone S-A, Scheuermann RH, Shah N, Whetzel PL, Lewis S. The OBO foundry: coordinated evolution of ontologies to support biomedical data integration. Nat Biotechnol. 2007;25(11):1251–5.
Steger S. Local rigid registration for multimodal texture feature extraction from medical images. In: SPIE medical imaging, Orlando; 2011.
Steger S, Keil M. Automated initialization and region of interest detection for successful head registration of truncated CT/MR head&neck images. In: Information technology and applications in biomedicine (ITAB), 2010 10th IEEE international conference on, Corfu; 2010.
Steger S, Wesarg S. Automated skeleton based multi-modal deformable registration of head&neck datasets. In: Ayache N, Delingette H, Golland P, Mori K, editors. Medical image computing and computer-assisted intervention – MICCAI 2012, Nice, vol. 7511. Berlin/Heidelberg: Springer; 2012. p. 66–73.
Steger S, Erdt M, Chiari G, Sakas G. Feature extraction from medical images for an oral cancer reoccurrence prediction. In: World congress on medical physics and biomedical engineering. Munich; 2009.
Steger S, Franco F, Sverzellati N, Chiari G, Colomer R. 3D assessment of lymph nodes vs. RECIST 1.1. Acad Radiol. 2011;18(3):391–4.
Steger S, Sakas G. FIST: fast interactive segmentation of tumors. In: Abdominal imaging. Computational and clinical applications, Toronto; 2012.
Steger S, Kirschner M, Wesarg S. Articulated atlas for segmentation of the skeleton from head&neck CT datasets. In: Biomedical imaging (ISBI), 2012 9th IEEE international symposium on, Barcelona; 2012.
Steger S, Bozoglu N, Kuijper A, Wesarg S. On the segmentation of cervical lymph nodes from CT images using radial rays. IEEE Trans Med Imaging. 2013;32(5):888–900.
Wang Z, Zhang B, Jiang L, Zeng X, Chen Y, Feng X, Guo Y, Chen Q. RACK1, an excellent predictor for poor clinical outcome in oral squamous carcinoma, similar to Ki67. Eur J Cancer. 2009;45(3):490–6.
Warner GC, et al. Molecular classification of oral cancer by cDNA microarrays identifies overexpressed genes correlated with nodal metastasis. Int J Cancer. 2004;110(6):857–68.
Woolgar JA. Histopathological prognostic factors in oral and oropharyngeal squamous cell carcinoma. Oral Oncol. 2006;42(3):229–39. Epub 2005 Sept 16. Review.
Woolgar JA, et al. Determinants of outcome following surgery for oral squamous cell carcinoma. Future Oncol. 2009;5(1):51–61. Review.
Ye H, Wang A, Lee BS, Yu T, Sheng S, Peng T, Hu S, Crowe DL, Zhou X. Proteomic based identification of manganese superoxide dismutase 2 (SOD2) as a metastasis marker for oral squamous cell carcinoma. Cancer Genomics Proteomics. 2008;5(2):85–94.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media Dordrecht
About this entry
Cite this entry
Poli, T. et al. (2014). Biomarkers in NeoMark European Project for Oral Cancers. In: Preedy, V., Patel, V. (eds) Biomarkers in Cancer. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-7744-6_12-1
Download citation
DOI: https://doi.org/10.1007/978-94-007-7744-6_12-1
Received:
Accepted:
Published:
Publisher Name: Springer, Dordrecht
Online ISBN: 978-94-007-7744-6
eBook Packages: Springer Reference Biomedicine and Life SciencesReference Module Biomedical and Life Sciences