Abstract
The red, iron containing tetrapyrrole heme is an essential cofactor of enzymes involved in the electron transport chain of energy generation and used for catalyzing chemically challenging reactions of the metabolism. It is also used for diatomic gas transport (O2, CO, CO2, NO, N2O), catalysis, and detection. Multiple transcriptional regulators and transporters bind heme. This chapter focuses on the highly unusual pathways for heme biosynthesis and the integration of protoheme into target proteins. Today, three different biosynthetic routes for heme formation are known. The general precursor molecule of all tetrapyrroles 5-aminolevulinic acid is formed by two different pathways starting either with glutamyl-tRNA or succinyl-CoA and glycine. The conversion of 5-aminolevulinic acid to uroporphyrinogen III is common to all biosynthetic paths. Then the pathway branches to a classical route via protoporphyrin and two currently known alternative routes via coproporpyhrin III and siroheme. Various steps are catalyzed by up to three structurally unrelated enzymes. Finally, formed protoheme (heme b) gets actively inserted into proteins by the “Radical SAM” protein HemW. A detailed description of involved intermediates, enzymes, and their mechanisms are depicted below.
Access provided by Autonomous University of Puebla. Download reference work entry PDF
Similar content being viewed by others
1 Introduction
Hemes are red colored, iron containing porphyrins which belong to the class of tetrapyrroles. The word heme is derived from the Greek αἷμα haima which stands for “blood.” These molecules are all composed of four pyrrole rings connected by methine bridges. The centrally coordinated iron can be used as electron source or sink and usually switches between the Fe(II), Fe (III), and Fe (IV) state. In humans, defects of the enzymes involved in heme biosynthesis lead to severe diseases called porphyrias (Kaufholz et al. 2013b).
Overall, multiple different closed circular and open chain tetrapyrroles are known. Cyclic tetrapyrroles reveal characteristic reduction states of the ring system and typical metal irons chelated in the center (Dailey et al. 2017; Heinemann et al. 2008; Jahn and Jahn 2012; Layer et al. 2010). The pyrrole moieties are substituted by propionate, vinyl, acetate, and methyl groups.
Prominent members of the tetrapyrroles are the magnesium coordinating, green bacteriochlorophylls and chlorophylls (Bröcker et al. 2012; Wang and Grimm 2015), the light absorbing pigments of oxygenic and anoxygenic photosynthesis. The yellow nickel containing coenzyme F430 plays an essential role in methanogenesis as part of the enzyme methyl coenzyme M (CoM) reductase (Moore et al. 2017; Zheng et al. 2016). Siroheme and heme d1 are technically no hemes (Boss et al. 2017). Siroheme is an iron containing isobacteriochlorin and serves as prosthetic group of assimilatory sulfite and nitrite reductases. Heme d1 is an iron containing dioxoisobacteriochlorin and the prosthetic group of the dissimilatory cd1 nitrite reductase. The cobalt containing corrins of the cobalamin or vitamin B12 class are part of methyltransferases, reductive dehalogenases, isomerases, and radical S-adensoyl-L-methionine (SAM) enzymes (Moore et al. 2017; Smith et al. 2018). However, the most ubiquitously found and versatile tetrapyrroles are hemes. They participate in various cellular processes including electron transfer, gas sensing and transport, signaling, catalysis, and transcriptional regulation.
2 Multiple Pathways for Heme Biosynthesis: An Overview
For many decades, a unique biosynthetic pathway for the biosynthesis of hemes in all organisms was proposed (Bogorad 1958; Bogorad and Granick 1953; Hoare and Heath 1958). First variations were observed in the late 1980s for the formation of the general precursor 5-aminolevulinc acid (ALA , (Jahn 1992). Originally, the presence of aminolevulinic acid synthase was assumed to be ubiquitous throughout all kingdoms of life (Figs. 1 and 2). But for plants and later on for most bacteria and archaea, a different so-called C5-pathway, originating from the C5 skeleton of glutamate and proceeding via a glutamyl-tRNA intermediate, was discovered (Czarnecki and Grimm 2013; Jahn et al. 1992). Later in the biosynthetic pathway, variation occurred at the enzymatic steps known to require molecular oxygen, the coproporphyrinogen III (HemF) and protoporphyrinogen IX oxidase (HemY) reactions (Fig. 2). For their anaerobic metabolism, bacteria had developed the “Radical SAM” enzyme coproporphyrinogen III dehydrogenase (HemN) and an electron chain coupled flavin protoporphyrinogen IX oxidase (HemG) (Boynton et al. 2009; Layer et al. 2004, 2005; Möbius et al. 2010). In the meantime, a third protoporphyrinogen IX oxidase (HemJ) was discovered in cyanobacteria (Kato et al. 2010; Skotnicova et al. 2018). With increasing access to multiple genomes, heme-synthesizing bacteria and archaea were discovered that lack genes for the enzymes of the late classical biosynthetic route to heme. Investigations from the late 1990s already proposed an alternative route via precorrin 2, an intermediate of the branch of tetrapyrrole biosynthesis toward cobalamin (B12), F430, siroheme, and heme d1 (Ishida et al. 1998). Novel enzyme activities were measured in Desulfovibrio vulgaris; however, due to the isolation of the reaction products as oxidized ester, their exact nature was not determined with absolute certainty. It was demonstrated in 2006 for the archaeon Methanosarcina barkeri and in 2009 for D. vulgaris that heme can be synthesized from precorrin 2 (Buchenau et al. 2006; Lobo et al. 2009). A novel pathway via siroheme was proposed and demonstrated (Bali et al. 2011; Kuhner et al. 2016; Storbeck et al. 2009). Additionally, in many Gram-positive bacteria, genes for coproporphyrinogen III oxidases/dehydrogenases were missing. Recently, for these organisms a novel pathway with coproporphyrin III as intermediate was described (Dailey et al. 2015; Lobo et al. 2015). Here, a coproporphyrinogen III oxidase produces this tetrapyrrole which in turn is subjected to iron insertion and final decarboxylation to yield heme (Dailey et al. 2015; Hansson et al. 1997a; Hobbs et al. 2016, 2017; Lobo et al. 2014). Interestingly, the siroheme and coproporphyrin pathways to heme share the final step, the decarboxylation of Fe-coproporphyrin III (coproheme) to form heme (Lobo et al. 2014). In summary, today we know three different pathways for the formation of heme with typical intermediates protoporphyrin, siroheme and coproporphyrin (Fig. 2).
3 Two Routes for 5-Aminolevulinic Acid Formation
The C5 compound ALA represents the general precursor of all known tetrapyrroles. Two different pathways for its formation are known. The most likely older pathway uses the C5 skeleton of glutamate as precursor (Beale and Castelfranco 1973). Glutamate is loaded onto tRNAGlu by glutamyl-tRNA synthetase, normally involved in protein biosynthesis (Schulze et al. 2006). An active site cysteine residue of glutamyl tRNA reductase (HemA, GtrR, GluTR) attacks the ester bond between the α-carbonyl of glutamate and tRNAGlu with the formation of an enzyme-bound thioester intermediate (Fig. 3) and the release of free tRNAGlu (Moser et al. 1999; Randau et al. 2004; Schauer et al. 2002). Hydride transfer from NADPH yields glutamate-1-semialdehyde (Lüer et al. 2007). Next, the pyridoxal-5′-phosphate/pyridoxamine-5′-phosphate-dependent glutamate-1-semialdehyde-2,1-aminomutase (HemL, GsaM) catalyzes the intramolecular transfer of an amino group using a modified aminotransferase mechanism (Grimm et al. 1992; Ilag and Jahn 1992). The product of both reactions is ALA. Currently, two crystal structures of GtrR and multiple structures of GsaM exist (Hennig et al. 1997; Li et al. 2018; Moser et al. 2001; Schulze et al. 2006; Zhao et al. 2014). Both enzymes form a stable channeling complex to protect the water-labile glutamate-1-semialdehyde intermediate (Lüer et al. 2005).
In a second, often termed Shemin pathway, 5-aminolevulinic acid synthase (HemA, AlaS) catalyzes the condensation of the C4 compound succinyl-CoA and the C2 amino acid glycine with elimination of CO2 to form ALA (Fig. 2 (Gibson et al. 1958; Kikuchi et al. 1958; Shemin and Rittenberg 1945)). The pyridoxal-5′-phosphate-dependent enzyme proceeds after lysine binding through an internal aldimine, after pro-R-hydrogen abstraction a quinonoid I intermediate is formed prior to succinyl-CoA binding of the 2-amino-3-ketoadipate and after the release of coenzyme A the quinonoid II intermediates (Fig. 3). Decarboxylation and protonation yields ALA (Kaufholz et al. 2013a; Stojanovski et al. 2014). Two crystal structures of the enzyme are known (Astner et al. 2005; Brown et al. 2018).
4 The Common Part of Heme Biosynthesis from 5-Aminolevulinic Acid to Uroporphyrinogen III
The monopyrrole porphobilinogen is formed via the asymmetric condensation of two ALA molecules (Dresel and Falk 1953; Granick 1954) by porphobilinogen synthase (Fig. 2, HemB, PbgS). For this purpose, the enzyme contains two ALA binding sites, termed A and P sites, referring to the contribution to the acetate or propionate moiety of porphobilinogen (Fig. 4). ALA 1 is bound via a Schiff base to a P site lysine, while ALA2 is also bound to a lysine residue of the sometimes metal containing A site (Spencer and Jordan 1995). This second Schiff base in the A site is converted to an enamine. In an aldol addition reaction, the C3 of ALA2 attacks the C4 of ALA1 with the formation of a C-C bond. Now the amino group of ALA 1 targets the Schiff base at the C4 atom of ALA2 yielding a C-N bond with the release from the lysine (Fig. 4). Subsequently, lysis of the remaining bond to the other lysine and aromatization of the formed ring system leads to porphobilinogen (Frere et al. 2002; Jaffe 2004). Overall three different binding sites for metal ions have been detected. These sites are filled in multiple combinations by zinc and magnesium (Jaffe 2016). Multiple crystal structures for the usually octameric enzymes have been elucidated (Erskine et al. 1997; Frankenberg et al. 1999; Frere et al. 2005; Jaffe et al. 2000). For the human enzyme, an equilibrium of functionally distinct octameric, hexameric, and two different dimers was described (Breinig et al. 2003; Jaffe and Lawrence 2012). Interestingly, the PbgS from Rhodobacter capsulatus is a hexamer without any metals (Bollivar et al. 2004).
During the next two enzymatic steps, the first circular closed tetrapyrrole uroporphyrinogen III is formed by hydroxymethylbilane synthase (previously porphobilinogen deaminase, HemC, HmbS) and uroporphyrinogen III synthase (HemD, UroS). Originally, these reactions were believed to be catalyzed by one enzyme. However, at end of the 1960s, both activities were separated (Stevens and Frydman 1968). During the 1970s, the new intermediate hydroxymethylbilane (pre-uroporphyrinogen) was identified via 13C NMR and shown to be the substrate for uroporphyrinogen III synthase (Burton et al. 1979; Jordan and Seehra 1979). Hydroxymethylbilane synthase catalyzes the polymerization of four porphobilinogen pyrroles starting with ring A followed by rings B,C, and D of the final tetrapyrrole (Battersby et al. 1979; Jordan and Seehra 1979). In 1987, the existence of covalently attached dipyrromethane cofactor, formed by the enzyme from two molecules of porphobilinogen, serving as primer for the polymerization reaction, was shown (Azim et al. 2014; Hart et al. 1987; Jordan et al. 1988; Warren and Jordan 1988). During catalysis first the dipyrromethane cofactor is assembled once after ribosomal enzyme formation. Incoming porphobilinogens are deaminated and polymerized at the cofactor. The roles of conserved aspartic acid and arginine amino acid residues have been described (Bung et al. 2018; Pluta et al. 2018; Woodcock and Jordan 1994). Finally, the bond between the cofactor and the tetrapyrrole is hydrolyzed and hydroxymethylbilane is released (Fig. 5). In water, it would cyclize to fully symmetric uroporphyrinogen I, an inhibitor of the next enzyme, uroporphyrinogen III synthase. Multiple crystal structures are available (Azim et al. 2014; Gill et al. 2009; Roberts et al. 2013). Recently, a regulation of hydroxymethylbilane synthase by heme was proposed (Uchida et al. 2018).
Uroporphyrinogen III synthase catalyzes the cyclization of linear hydroxymethylbilane with the inversion of ring D to form the cyclic but asymmetric uroporphyrinogen III. Uroporphyrinogen III as an intermediate of heme biosynthesis was proposed in the 1950s (Bogorad 1958). The enzyme was characterized and a mechanism proposed in the 1960s (Levin 1968; Mathewson and Corwin 1961). Catalysis starts with the loss of the hydroxyl group at ring A with the formation of the first azafulvene intermediate (Fig. 5). The reaction of the azafulvene with the substituted α-position of the D ring results in the formation of a spirocyclic pyrrolenine intermediate. A second azafulvene intermediate is formed on ring C by the breakage of the bond between rings C and D. This azafulvene reacts in the final steps with the free a-position before deprotonation, and rearrangement leads to the formation of uroporphyrinogen III (Hawker et al. 1998; Stark et al. 1985, 1986, 1993). Crystal structures for the enzyme have been published (Mathews et al. 2001; Peng et al. 2011; Schubert et al. 2008). Due to the low degree of amino acid conservation, the gene for the protein of a plant enzyme was discovered decades later (Tan et al. 2008).
5 The Classical Pathway via Protoporphyrin IX
Here, uroporphyrinogen III is first subject to a stepwise decarboxylation of each of the four pyrrole ring acyl side chains at the C2, C7, C12, and C18 to four methyl groups (Mauzerall and Granick 1958). The reaction is catalyzed by uroporphyrinogen III decarboxylase (HemE, UroD), which starts at ring D and proceeds clockwise via ring A and B to ring C (Jackson et al. 1976). The enzyme does not require any associated cofactor. The enzyme is a homodimer with juxtaposed and facing active sites. Two major different models were proposed for enzyme activity. In the first model, the homodimer shuttles a single substrate molecule forth and back between both active sites without solvent contact (Phillips et al. 2003; Whitby et al. 1998). In the second model, solely one active site of one subunit is used, the substrate is rotated by 90° following each decarboxylation. Interestingly, a dimer of an active and an inactive uroporphyrinogen III decarboxylase subunit is still able to perform the complete reaction pointing toward model 2 (Phillips et al. 2009). Several crystal structures have been reported (Fan et al. 2007; Martins et al. 2001; Phillips et al. 2003; Whitby et al. 1998). A uroporphyrinogen III decarboxylase structure with the product coproporphyrinogen III revealed that the substrate/product binds in a dome-shaped structure with the four NH groups facing a 2x hydrogen bond distance to a conserved aspartate residue. Furthermore, three conserved arginines, one conserved histidine, and one tyrosine residues were proposed to be involved in catalysis. One of the arginine residues was identified as the general acid catalyst. Upon its protonation the decarboxylation reaction becomes rate-limiting (Silva et al. 2010). Regardless of the exact mechanism, the enzyme was suggested as a “benchmark” for catalytic proficiency among enzymes without cofactors due to a calculated enzyme enhancement value of the various decarboxylation reaction of around 1017 (Lewis and Wolfenden 2008).
During the next biosynthetic step toward the formation of heme, the propionate side chain of coproporphyrinogen III ring A and B undergoes an oxidative decarboxylation to the corresponding vinyl groups by two different, structurally not related enzymes. The oxygen-dependent cofactor-free coproporphyrinogen III oxidase (HemF, CpgC) and the radical SAM enzyme coproporphyrinogen III dehydrogenase (HemN, CgdH) are catalyzing the conversion of coproporphyrinogen III into protoporphyrinogen IX. The general reaction was discovered in the 1950s (Granick and Mauzerall 1958) and the corresponding oxidase enzyme was described shortly thereafter (Sano and Granick 1961). First, it was shown that coproporphyrinogen III oxidase catalyzes the decarboxylation of ring A prior to that of ring B with the intermediate haderoporphyrinogen (Cavaleiro et al. 1974; Elder and Evans 1978). A mechanism for the oxygen-dependent reaction by the cofactor-free enzyme was proposed (Lash 2005; Silva and Ramos 2008). During the first step a base catalyzed deprotonation of the pyrrole NH-group yields an azacyclopentadienyl anion, which reacts with molecular oxygen at the α-position to form a pyrrole peroxide anion. Next, a proton at the β-positon of the substrate gets abstracted by the peroxide via a six-membered ring transition state with the formation of an exocyclic double bond (Fig. 6). Elimination of CO2 and H2O2 with the following bond rearrangements results in the formation of the vinyl group of the product protoporphyrinogen IX (Breckau et al. 2003). Solved crystal structures were without substrate (Lee et al. 2005; Phillips et al. 2004). Nevertheless, an aspartate and two conserved arginine residues were proposed to be in catalysis and substrate binding (Stephenson et al. 2007).
Many bacteria possess anaerobic heme containing respiratory chains for energy generation. Consequently, an oxygen-independent coproporphyrinogen III conversion was needed. At the end of the 1960s Tait described such enzyme activity for Rhodobacter sphaeroides (Tait 1969, 1972). Around 10 years later the identical stereochemistry of an initial pro-S-hydrogen abstraction at the β-carbon as observed for the oxygen-dependent catalysis was described for the coproporphyrinogen III dehydrogenase (Seehra et al. 1983). Similarly, haderoporphyrinogen was identified as reaction intermediate (Rand et al. 2010). Genetic approaches led to the isolation of the corresponding genes in the 1990s (Lieb et al. 1998; Troup et al. 1995; Xu and Elliott 1994). Intensive biochemical and structural analysis of recombinant coproporphyrinogen III dehydrogenase from Escherichia coli identified the protein as “Radical SAM” enzyme. It carries a [4Fe-4S] cluster coordinated by three cysteine residues and one S-adenosyl-L-methionine (SAM) molecule (Layer et al. 2002, 2005; Lieb et al. 1998; Troup et al. 1995; Xu and Elliott 1994). The reaction starts with the reduction of the [4Fe-4S] cluster by an unknown electron donor. The electron is subsequently transferred to SAM, which in turn undergoes a homolytic cleavage with the generation of methionine and a 5' deoxyadenosyl radical. The highly reactive radical abstracts stereo-specifically the pro-S-hydrogen at the β-carbon (Fig. 6). Finally, elimination of CO2, transfer of the remaining electron to a yet unknown electron acceptor, and structural rearrangements with formation of the vinyl group finalize the reaction which has to occur twice (Layer et al. 2004, 2006). The crystal structure revealed the presence of two SAM molecules, which would allow a complete reaction cycle without release of a reaction intermediate (Layer et al. 2003).
The six electron oxidation of protoporphyrinogen IX to the red colored protoporphyrin IX is catalyzed by three distinct enzymes all named protoporphyrinogen IX oxidase (HemY or PgoX, HemG or PgdH1, HemJ or PgdH2). Under aerobic conditions the reaction can occur auto-catalytically. Already in the early 1960s the corresponding enzyme was discovered (Porra and Falk 1961, 1964; Sano and Granick 1961). The FAD-containing, oxygen-dependent PgoX is found in all heme-synthesizing eukaryotes and a few Gram-negative bacteria. Currently, the crystal structure of three enzymes are known (Corradi et al. 2006; Koch et al. 2004; Qi et al. 2002), however, without bound substrate or product. A model for substrate binding was verified via kinetic studies (Heinemann et al. 2007). Due to the FAD cofactor one can assume that the reaction proceeds via three two-electron steps from porphyrinogen via tetrahydro and dihydro intermediates to the fully oxidized porphyrin. Kinetic studies revealed that three meso-carbon hydride ions are removed in a sequential fashion with the concomitant removal of the NH proton (Akhtar 2003). Alternatively, it was proposed that all hydride abstractions occur at the C-20 meso-carbon including total ring hydrogen rearrangement via enamine-imine tautomerizations (Koch et al. 2004). Based on the PgoX structure, a complex with the following enzyme ferrochelatase was proposed and finally demonstrated (Koch et al. 2004; Masoumi et al. 2008).
Again, PgoX is oxygen-dependent, and bacteria require an alternative system for anaerobic heme biosynthesis. First description of an oxygen-independent system channeling the six abstracted electrons into respiratory chains was reported in the 1970s from Jacobs and Jacobs (Ishihara et al. 1995; Jacobs and Jacobs 1978; Jacobs et al. 1970, 1971). In E. coli, the corresponding gene hemG was mapped and the corresponding mutant used for cloning of the gene (Nishimura et al. 1995; Sasarman et al. 1993). A detailed biochemical characterization of PgdH1 followed at the beginning of this decade (Boynton et al. 2009; Möbius et al. 2010). The FMN-containing protein belongs to the class of long chain flavodoxins. It was shown that the abstracted six electrons are transferred via ubiquinone to terminal oxidases (Cyo, Cyd) under aerobic conditions and via menaquinone to nitrate and fumarate reductase under anaerobic conditions (Möbius et al. 2010). A similar mechanism as proposed for PgoX is also possible for PgdH1 (Fig. 7). Interestingly, the only eukaryotic organism utilizing PgdH1 is the Leishmania major (Zwerschke et al. 2014).
For the third enzyme PgdH2 (HemJ), little is known. The corresponding hemJ gene was discovered in the cyanobacterium Synechocystis 6803 (Boynton et al. 2011; Kato et al. 2010). The corresponding membrane protein contained heme and was proposed to interact with coproporphyrinogen III oxidase (Skotnicova et al. 2018).
Protoporphyrin ferrochelatase (HemH, PpfC) catalyzes the final step of heme biosynthesis, the insertion of ferrous iron into protoporphyrin IX with the formation of protoheme (heme b). The first description of ferrochelatase from avian erythrocytes dates back to the 1950s (Ashenbrucker et al. 1956). Eukaryotic enzymes are usually membrane associated and contain with the exception of the plant enzymes one [2Fe-2S] cluster of unknown function per subunit (Shepherd et al. 2006). Bacterial enzymes are found with and without the cluster. The tetrapyrrole binds to the enzyme in an open conformation. Binding of the substrate triggers a rearrangement of a hydrogen bond network among conserved active site amino acid residues. Possibly, due to an abstraction of one pyrrole hydrogen a closed conformation is induced. Now the macrocycle is engulfed which causes an approximately 12 degree distortion of the bound tetrapyrrole. This distortion obviously facilitates metal chelation by the porphyrin with the simultaneous displacement of the second pyrrole hydrogen to a conserved histidine residue. The imidazole ring of the histidine moves, which causes structural rearrangements of the enzyme to adapt to the release conformation. Finally, heme b is released (Medlock et al. 2007, 2009; Sigfridsson and Ryde 2003; Wang et al. 2009, 2013).
6 The Alternative Pathway via Coproporphyrin III
Gram-positive bacteria utilize an alternative, only recently discovered pathway for heme biosynthesis. Coproporpyhrinogen III synthesized via the classical pathway gets oxidized to coproporphyrin III followed by metal insertion with the formation of Fe-coproheme III. Finally, the propionate side chains at ring A and B of Fe-coproheme III get decarboxylated to form heme b (Dailey et al. 2015; Hansson et al. 1997b; Hobbs et al. 2016, 2017; Lobo et al. 2015). Obviously, oxidation of the ring system of the porphyrinogen to form a porphyrin occurs at the level of coproporphyrinogen III instead of protoporphyrinogen IX as seen in the classical pathway. Consequently, iron insertion follows and the decarboxylation of ring A and B, as performed at the level of coproporphyrinogen by two different enzymes (CpgC, CgdH) in the classical pathway, utilizes the iron containing Fe-coproheme III in the novel pathway. The first committed step of the novel pathway, the oxidation of coproporpyhrinogen III to coproporphyrin III is performed by a coproporpyhrinogen III oxidase (HemY, CgoX). It was originally described for Bacillus subtilis as protoporphyrinogen oxidase (Corrigall et al. 1998; Hansson and Hederstedt 1994; Qin et al. 2010). However, already then it was reported that the enzyme catalyzes the oxidation of coproporpyhrinogen III to coproporphyrin III at a higher rate as the proposed protoporpyhrinogen oxidation (Han et al. 2013; Hansson et al. 1997b). With the discovery of the novel pathway this observation makes sense. The mechanism of the oxygen-dependent six electron oxidation performed by the FAD enzyme is highly similar to the oxygen-dependent protoporphyrinogen oxidase (FgoX) mechanism with the abstraction of six protons from the porphyrinogen and the formation of three molecules H2O2. However, the active site pocket of CgoXs is with 1173 Å3 is much larger than those of FgoX protoporphyrinogen IX oxidases with 527 to 440 Å3 and contains more positively charged surface areas (Qin et al. 2010). One explanation is the accommodation of the remaining propionate side chain containing coproporpyhrinogen III by CgoX. The oxygen-independent enzyme is currently unknown.
Next, coproporphyrinogen ferrochelatase (HemH, CpfC) catalyzes the insertion of ferrous iron into coproporpyhrin III to generate coproheme III. The best characterized enzyme is again from B. subtilis. It is water soluble and possesses a [2Fe-2S] cluster of unknown function. The mechanism is most likely highly similar to those of the ferrochelatase PpfC (Hansson et al. 2007; Karlberg et al. 2002; Lecerof et al. 2000, 2003; Olsson et al. 2002).
The last step of the novel pathway is the decarboxylation of coproheme III to heme b and was named coproheme decarboxylase or heme synthase (ChdC, HemQ) . An enzyme (AhbD) of identical catalysis is also part of the novel siroheme pathway for heme biosynthesis described below, but structurally not related to ChdC. While AhbD belongs to the family of “Radical SAM” enzymes, ChdC is a member of the chlorite dismutase family (Celis et al. 2015; Dailey et al. 2015; Pfanzagl et al. 2018). The structures of several incorrectly annotated as potential chlorite dismutase ChdCs were solved (PDB accession numbers 1T0T, 3DZT, 1VDH, 4WWS, SLOQ). The Listeria monocytogenes enzyme was co-crystallized with the product heme (Hofbauer et al. 2016). Two molecules of H2O2 are needed for catalysis (Celis et al. 2015; Hofbauer et al. 2014, 2016). During catalysis coproheme acts as substrate and cofactor. The coproheme ferric iron is coordinated by a conserved histidine residue. Under aerobic conditions O2 gets to the coproheme iron bound and oxidizes it to the ferric state. The subsequent second-order reaction between the ferric complex and H2O2 is slow and pH-dependent. First evidence for ferryl porphyrin cation radical was obtained (Streit et al. 2018). A tyrosine, hydrogen bonding to the propionate at ring A, is essential for decarboxylation. It is proposed that an oxidizing equivalent from the most likely radical allows for the formation of a tyrosine radical which abstracts the hydrogen from the propionate side chain. Migration of the unpaired propionyl electron back to the coproheme would yield ferric haderoheme and CO2. The propionate at ring B forms salt bridges to a lysine residue. Now, a similar pathway is proposed with this lysine as the essential proton shuttle for the second decarboxylation reaction (Celis et al. 2017). An alternative version of the ChdC protein called PitA, representing a fusion of ChdC with a monooxygenase domain, was described for the aerobic growth of Halferax volcanii (Kosugi et al. 2017).
7 The Second Alternative Pathway via Siroheme
The second alternative pathway starts directly at uroporphyrinogen III and uses the initial steps of cobalamin, F430, heme d1, and the complete siroheme biosynthesis (Bali et al. 2014; Kuhner et al. 2014). The pathway was originally proposed for archaea but subsequently found in multiple bacteria (Storbeck et al. 2010). The three reactions of siroheme formation are the SAM-dependent methylation of uroporphyrinogen II at position C-2 and C-7 to form precorrin-2 via precorrin-1, the following NAD-dependent ring dehydrogenation of precorrin-2 to form sirohydrochlorin and NADH, and the final iron insertion into sirohydrochlorin to yield siroheme. These three catalytic challenging reactions can be performed by one multifunctional enzyme called siroheme synthase (CysG) (Spencer et al. 1993; Warren et al. 1990). The enzyme contains two independent enzymatic modules, one for the methyltransferase and the other for the combined dehydrogenase-ferrochelatase function (Anderson et al. 2001; Lobo et al. 2009; Strey et al. 1999). Alternatively, the identical reactions are performed by three different enzymes, uroporphyrinogen-III C-methyltransferase (SUMT = NirE, CobA, SirA, Met1p) for the SAM-dependent methylation of uroporphyrinogen II at position C-2 and C-7 to form precorrin-2 via precorrin, the precorrin 2-dehydrogenase for the NAD-dependent ring dehydrogenation of precorrin-2 to form sirohydrochlorin, and the sirohydrochlorin ferrochelatase (SirB) for iron insertion. Interestingly, some bacteria carry a protein fusion of uroporphyrinogen III synthase and SUMT (Anderson et al. 2001; Lobo et al. 2009), most likely allowing direct channeling of the uroporphyrinogen III intermediate into the precorrin-2 pathway.
Uroporphyrinogen-III C-methyltransferase (SUMT = NirE, CobA, SirA, Met1p) was first described for cobalamin biosynthesis in Pseudomonas denitrificans (Blanche et al. 1989). The crystal structures of SUMT involved in cobalamin and heme d1 biosynthesis were solved (Rehse et al. 2005; Storbeck et al. 2011; Vevodova et al. 2004). The NirE structure revealed the coordination of the tetrapyrrole by three arginines, a histidine, and a methionine residue. A mechanism induced by the arginine-mediated proton abstraction from the C-20 position was proposed (Storbeck et al. 2011). The subsequent movement of electrons facilitates the nucleophilic attack of C-2 at the methyl group of SAM. Upon proton abstraction from C-20 a new double bond between C-20 and C-1 is formed. After methyl transfer, rearrangements of double bonds within the macrocycle occur to form presorrin-1. After the first round of methylation the intermediate and side product SAH are released from the active site. Precorrin-1 and new SAM have to bind again to initiate a novel round of methylation at C-7 (Storbeck et al. 2011).
Precorrin-2 dehydrogenase (SirC, Met8p) catalyzes the NAD+-dependent oxidation of precorrin-2 (dipyrrocorphin) to form sirohydrochlorin (Raux et al. 2003; Schubert et al. 2008). The identical reaction is catalyzed by domains of the multifunctional CysG and Met8p. The crystal structures of SirC and Met8p were solved. Both enzymes were found to bind metals including Co(II) and Cu(II). It was proposed that SirC evolved from a Met8p-type protein via loosing its chelatase domain (Raux et al. 2003; Schubert et al. 2008).
Sirohydrochlorin ferrochelatase (SirB, Met8p) catalyzes the insertion of iron into sirohydrochlorin to form siroheme (Schubert et al. 2002). SirB solely catalyzes the iron chelation reaction. The multifunctional Met8p carries one active site with a catalytically important aspartate residue for both reactions, the dehydrogenase and chelatase reaction (Schubert et al. 2002).
The heterodimeric Siroheme decarboxylase (NirDLGH, AhbAB) catalyzes the conversion of siroheme to didecarboxysiroheme (Palmer et al. 2014). The two acyl side chains attached to C-12 and C-18 are decarboxylated to the corresponding methyl groups. Surprisingly, the crystal structure of the Desulfovibrio desulfuricans protein was solved and revealed structural similarity to proteins of the Asn/Lrp transcriptional regulator family proteins. A enzyme mechanism was proposed (Palmer et al. 2014). The coproheme (Fe-coproporphyrin) synthase (AhbC) converts didecarboxysiroheme into coproheme (Fe-coproporphyrin). Recombinant AhbC protein from Methanosarcina barkeri was shown to transform in the presence of SAM and the reducing agent dithionite 12,18-didecarboxysiroheme into Fe-coproporphyrin III (coproheme). Thus, the enzyme catalyzes the loss of the acyl side chains at C2 and C7 (Bali et al. 2011). The exact enzymatic mechanism remains to be determined. Heme synthase (AhbD) catalyzes the oxidative decarboxylation of coproheme into heme b (Kuhner et al. 2016). The protein belongs to the “Radical SAM” family of enzymes. A close mechanistic relationship to the coproporphyrinogen III dehydrogenase reaction can be assumed. Interestingly, the protein contains two [4Fe-4S] clusters. Besides the “classical” [4Fe-4S] cluster I involved in coproporphyrinogen III dehydrogenase type catalysis, the second auxiliary [4Fe-4S] cluster was identified to be involved in the electron transfer to the final electron acceptor of the reaction (Kuhner et al. 2016).
8 Heme b Insertion by the Heme Chaperone HemW
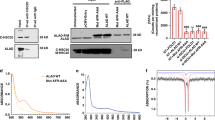
The “radical SAM” protein HemW revealed a high degree of amino acid sequence homology to coproporphyrinogen III dehydrogenases. However, the corresponding enzyme activity was never demonstrated for these proteins. A genetic investigation in Lactobacillus lactis indicated HemW’s participation in the generation of cytochromes. Subsequently, stable heme binding of the protein was demonstrated. Very recently the E. coli HemW was shown to be a heme chaperone involved in the active integration of heme b into the usually heme containing respiratory nitrate NarGHI which is involved in the anaerobic energy generation of the bacterium. The human counterpart RSAD1 was also shown to stably bind heme indicating its heme chaperone function (Abicht et al. 2012; Haskamp et al. 2018).
Like other radical SAM proteins, HemW contains three cysteines and one SAM coordinating a [4Fe-4S] cluster. The intact iron-sulfur cluster is required for HemW dimerization, which in turn causes membrane localization. The intact iron-sulfur cluster is not required for stable covalent heme binding. Bacterioferritins and the heme-containing subunit NarI of the respiratory nitrate reductase NarGHI were shown to interact directly with HemW. Bacterioferritins might serve as heme donors for HemW, while the cytochrome subunit NarI of nitrate reductase represents the target of HemW. During contact heme covalently bound to HemW gets actively transferred to a heme-depleted, catalytically inactive nitrate reductase, restoring its nitrate-reducing enzyme activity (Fig. 8). For the transfer process, an intact a [4Fe-4S] cluster is required. The exact mechanism of heme transfer remains to be determined (Abicht et al. 2012; Haskamp et al. 2018).
Model of E. coli HemW activity. Heme from heme biosynthesis is most likely transferred via bacterioferritin (Bfr, light green) to the heme chaperone HemW (blue). The [4Fe-4S] cluster-containing HemW dimerizes and localizes to the membrane, where it interacts with its target protein NarI (yellow), a subunit of the respiratory nitrate reductase NarGHI. After heme incorporation into apo-NarI, the holo-NarGHI catalyzes the reduction of nitrate to nitrite
9 Research Needs
Some of the enzyme activities of the various alternative pathways, including the oxygen-independent coproporphyrinogen oxidase for the formation of coproporphyrin III, are unknown. Moreover, the exact nature of various enzymatic mechanisms and enzyme structures require further investigations. Possibly, additional alternative pathways shall be discovered.
References
Abicht HK, Martinez J, Layer G, Jahn D, Solioz M (2012) Lactococcus lactis HemW (HemN) is a haem-binding protein with a putative role in haem trafficking. Biochem J 442(2):335–343
Akhtar M (2003) Coproporphyrinogen III and protoporphyrinogen IX oxidases. In: Kadish KM, Smith KM, Guilard R (eds) The porphyrin handbook, vol 12, the iron and cobalt pigments: biosynthesis, structure and degradation. Elsevier, New York, pp 75–92
Anderson PJ, Entsch B, McKay DB (2001) A gene, cobA + hemD, from Selenomonas ruminantium encodes a bifunctional enzyme involved in the synthesis of vitamin B12. Gene 281(1–2):63–70
Ashenbrucker H, Cartwright GE, Goldberg A, Wintrobe MM (1956) Studies on the biosynthesis of heme in vitro by avian erythrocytes. Blood 11(9):821–833
Astner I, Schulze JO, van den Heuvel J, Jahn D, Schubert WD, Heinz DW (2005) Crystal structure of 5-aminolevulinate synthase, the first enzyme of heme biosynthesis, and its link to XLSA in humans. EMBO J 24(18):3166–3177
Azim N, Deery E, Warren MJ, Wolfenden BA, Erskine P, Cooper JB, Coker A, Wood SP, Akhtar M (2014) Structural evidence for the partially oxidized dipyrromethene and dipyrromethanone forms of the cofactor of porphobilinogen deaminase: structures of the Bacillus megaterium enzyme at near-atomic resolution. Acta Crystallogr D Biol Crystallogr 70(Pt 3):744–751
Bali S, Lawrence AD, Lobo SA, Saraiva LM, Golding BT, Palmer DJ, Howard MJ, Ferguson SJ, Warren MJ (2011) Molecular hijacking of siroheme for the synthesis of heme and d1 heme. Proc Natl Acad Sci USA 108(45):18260–18265
Bali S, Palmer DJ, Schroeder S, Ferguson SJ, Warren MJ (2014) Recent advances in the biosynthesis of modified tetrapyrroles: the discovery of an alternative pathway for the formation of heme and heme d1. Cell Mol Life Sci 71(15):2837–2863
Battersby AR, Fookes CJR, Matcham GWJ, McDonald E (1979) Order of assembly of the four pyrrole rings during biosynthesis of the natural porphyrins. J Chem Soc Chem Commun 0:539–541
Beale SI, Castelfranco PA (1973) 14C incorporation from exogenous compounds into delta-aminolevulinic acid by greening cucumber cotyledons. Biochem Biophys Res Commun 52(1):143–149
Blanche F, Debussche L, Thibaut D, Crouzet J, Cameron B (1989) Purification and characterization of S-adenosyl-l-methionine: uroporphyrinogen III methyltransferase from Pseudomonas denitrificans. J Bacteriol 171(8):4222–4231
Bogorad L (1958) The enzymatic synthesis of porphyrins from porphobilinogen. II. Uroporphyrin III. J Biol Chem 233(2):510–515
Bogorad L, Granick S (1953) The enzymatic synthesis of porphyrins from porphobilinogen. Proc Natl Acad Sci USA 39(12):1176–1188
Bollivar DW, Clauson C, Lighthall R, Forbes S, Kokona B, Fairman R, Kundrat L, Jaffe EK (2004) Rhodobacter capsulatus porphobilinogen synthase, a high activity metal ion independent hexamer. BMC Biochem 5:17
Boss L, Oehme R, Billig S, Birkemeyer C, Layer G (2017) The radical SAM enzyme NirJ catalyzes the removal of two propionate side chains during heme d1 biosynthesis. FEBS J 284(24):4314–4327
Boynton TO, Daugherty LE, Dailey TA, Dailey HA (2009) Identification of Escherichia coli HemG as a novel, menadione-dependent flavodoxin with protoporphyrinogen oxidase activity. Biochemistry 48(29):6705–6711
Boynton TO, Gerdes S, Craven SH, Neidle EL, Phillips JD, Dailey HA (2011) Discovery of a gene involved in a third bacterial protoporphyrinogen oxidase activity through comparative genomic analysis and functional complementation. Appl Environ Microbiol 77(14):4795–4801
Breckau D, Mahlitz E, Sauerwald A, Layer G, Jahn D (2003) Oxygen-dependent coproporphyrinogen III oxidase (HemF) from Escherichia coli is stimulated by manganese. J Biol Chem 278(47):46625–46631
Breinig S, Kervinen J, Stith L, Wasson AS, Fairman R, Wlodawer A, Zdanov A, Jaffe EK (2003) Control of tetrapyrrole biosynthesis by alternate quaternary forms of porphobilinogen synthase. Nat Struct Biol 10(9):757–763
Bröcker M, Jahn D, Moser J (2012) Key enzymes of chlorophyll biosynthesis. In: Kadish K, Smith K, Guilard R (eds) World Scientific Publishing Co., Singapore Vol. 20, 1–43
Brown BL, Kardon JR, Sauer RT, Baker TA (2018) Structure of the mitochondrial aminolevulinic acid synthase, a key heme biosynthetic enzyme. Structure 26(4):580–589.e584
Buchenau B, Kahnt J, Heinemann IU, Jahn D, Thauer RK (2006) Heme biosynthesis in Methanosarcina barkeri via a pathway involving two methylation reactions. J Bacteriol 188(24):8666–8668
Bung N, Roy A, Chen B, Das D, Pradhan M, Yasuda M, New MI, Desnick RJ, Bulusu G (2018) Human hydroxymethylbilane synthase: molecular dynamics of the pyrrole chain elongation identifies step-specific residues that cause AIP. Proc Natl Acad Sci USA 115(17):E4071–E4080
Burton G, Fagerness PE, Hosozawa S, Jordan PM, Scott AI (1979) 13C NMR evidence for a new intermediate, pre-uroporphyrinogen, in the enzymic transformation of porphobilinogen into uroporphyrinogens I and III. J Chem Soc Chem Commun 1:202–204
Cavaleiro JA, Kenner GW, Smith KM (1974) Pyrroles and related compounds. XXXII. Biosynthesis of protoporphyrin-IX from coproporphyrinogen-3. J Chem Soc Perkin 1 10:1188–1194
Celis AI, Streit BR, Moraski GC, Kant R, Lash TD, Lukat-Rodgers GS, Rodgers KR, DuBois JL (2015) Unusual peroxide-dependent, heme-transforming reaction catalyzed by HemQ. Biochemistry 54(26):4022–4032
Celis AI, Gauss GH, Streit BR, Shisler K, Moraski GC, Rodgers KR, Lukat-Rodgers GS, Peters JW, DuBois JL (2017) Structure-based mechanism for oxidative decarboxylation reactions mediated by amino acids and heme propionates in coproheme decarboxylase (HemQ). J Am Chem Soc 139(5):1900–1911
Corradi HR, Corrigall AV, Boix E, Mohan CG, Sturrock ED, Meissner PN, Acharya KR (2006) Crystal structure of protoporphyrinogen oxidase from Myxococcus xanthus and its complex with the inhibitor acifluorfen. J Biol Chem 281(50):38625–38633
Corrigall AV, Siziba KB, Maneli MH, Shephard EG, Ziman M, Dailey TA, Dailey HA, Kirsch RE, Meissner PN (1998) Purification of and kinetic studies on a cloned protoporphyrinogen oxidase from the aerobic bacterium Bacillus subtilis. Arch Biochem Biophys 358(2):251–256
Czarnecki O, Grimm B (2013) New insights in the topology of the biosynthesis of 5-aminolevulinic acid. Plant Signal Behav 8(2):e23124
Dailey HA, Gerdes S, Dailey TA, Burch JS, Phillips JD (2015) Noncanonical coproporphyrin-dependent bacterial heme biosynthesis pathway that does not use protoporphyrin. Proc Natl Acad Sci USA 112(7):2210–2215
Dailey HA, Dailey TA, Gerdes S, Jahn D, Jahn M, O’Brian MR, Warren MJ (2017) Prokaryotic heme biosynthesis: multiple pathways to a common essential product. Microbiol Mol Biol Rev 81(1) pii: e00048-16
Dresel EI, Falk JE (1953) Conversion of alpha-aminolaevulinic acid to porphobilinogen in a tissue system. Nature 172(4391):1185
Elder GH, Evans JO (1978) Evidence that the coproporphyrinogen oxidase activity of rat liver is situated in the intermembrane space of mitochondria. Biochem J 172(2):345–347
Erskine PT, Senior N, Awan S, Lambert R, Lewis G, Tickle IJ, Sarwar M, Spencer P, Thomas P, Warren MJ et al (1997) X-ray structure of 5-aminolaevulinate dehydratase, a hybrid aldolase. Nat Struct Biol 4(12):1025–1031
Fan J, Liu Q, Hao Q, Teng M, Niu L (2007) Crystal structure of uroporphyrinogen decarboxylase from Bacillus subtilis. J Bacteriol 189(9):3573–3580
Frankenberg N, Erskine PT, Cooper JB, Shoolingin-Jordan PM, Jahn D, Heinz DW (1999) High resolution crystal structure of a Mg2+-dependent porphobilinogen synthase. J Mol Biol 289(3):591–602
Frere F, Schubert WD, Stauffer F, Frankenberg N, Neier R, Jahn D, Heinz DW (2002) Structure of porphobilinogen synthase from Pseudomonas aeruginosa in complex with 5-fluorolevulinic acid suggests a double Schiff base mechanism. J Mol Biol 320(2):237–247
Frere F, Reents H, Schubert WD, Heinz DW, Jahn D (2005) Tracking the evolution of porphobilinogen synthase metal dependence in vitro. J Mol Biol 345(5):1059–1070
Gibson KD, Laver WG, Neuberger A (1958) Initial stages in the biosynthesis of porphyrins. 2. The formation of delta-aminolaevulic acid from glycine and succinyl-coenzyme A by particles from chicken erythrocytes. Biochem J 70(1):71–81
Gill R, Kolstoe SE, Mohammed F, Al DBA, Mosely JE, Sarwar M, Cooper JB, Wood SP, Shoolingin-Jordan PM (2009) Structure of human porphobilinogen deaminase at 2.8 A: the molecular basis of acute intermittent porphyria. Biochem J 420(1):17–25
Granick S (1954) Enzymatic conversion of delta-amino levulinic acid to porphobilinogen. Science 120(3131):1105–1106
Granick S, Mauzerall D (1958) Pbrphyrin biosynthesis in erythrocytes. II. Enzymes converting gamma-aminolevulinic acid to coproporphyrinogen. J Biol Chem 232(2):1119–1140
Grimm B, Smith MA, von Wettstein D (1992) The role of Lys272 in the pyridoxal 5-phosphate active site of Synechococcus glutamate-1-semialdehyde aminotransferase. Eur J Biochem 206(2):579–585
Han J, Zhou Z, Bu X, Zhu S, Zhang H, Sun H, Yang B (2013) Employing aqueous CdTe quantum dots with diversified surface functionalities to discriminate between heme (Fe(II)) and hemin (Fe(III)). Analyst 138(12):3402–3408
Hansson M, Hederstedt L (1994) Bacillus subtilis HemY is a peripheral membrane protein essential for protoheme IX synthesis which can oxidize coproporphyrinogen III and protoporphyrinogen IX. J Bacteriol 176(19):5962–5970
Hansson M, Gustafsson MC, Kannangara CG, Hederstedt L (1997a) Isolated Bacillus subtilis HemY has coproporphyrinogen III to coproporphyrin III oxidase activity. Biochim Biophys Acta 1340(1):97–104
Hansson M, Gustafsson MC, Kannangara CG, Hederstedt L, Hansson M, Hederstedt L, Hansson M, Hederstedt L (1997b) Isolated Bacillus subtilis HemY has coproporphyrinogen III to coproporphyrin III oxidase activity. Bacillus subtilis HemY is a peripheral membrane protein essential for protoheme IX synthesis which can oxidize coproporphyrinogen III and protoporphyrinogen IX. Cloning and characterization of the Bacillus subtilis hemEHY gene cluster, which encodes protoheme IX biosynthetic enzymes. Biochim Biophys Acta 1340(1):97–104
Hansson MD, Karlberg T, Rahardja MA, Al-Karadaghi S, Hansson M (2007) Amino acid residues His183 and Glu264 in Bacillus subtilis ferrochelatase direct and facilitate the insertion of metal ion into protoporphyrin IX. Biochemistry 46(1):87–94
Hart GJ, Miller AD, Leeper FJ, Battersby AR (1987) Biosynthesis of the natural porphyrins: proof that hydroxymethylbilane synthase (porphobilinogen deaminase) uses a novel binding group in its catalytic action. J Chem Soc Chem Commun 0:1762–1765
Haskamp V, Karrie S, Mingers T, Barthels S, Alberge F, Magalon A, Muller K, Bill E, Lubitz W, Kleeberg K et al (2018) The radical SAM protein HemW is a heme chaperone. J Biol Chem 293(7):2558–2572
Hawker CJ, Spivey AC, Leeper FJ, Battersby AR (1998) The rearrangement of 2H-pyrroles (pyrrolenines) related to the proposed spiro-intermediate for porphyrin biosynthesis. J Chem Soc Perkin Trans 1:1509–1518. (Biosynthesis of porphyrins and related macrocycles. Part 48)
Heinemann, I. U., N. Diekmann, A. Masoumi, M. Koch, A. Messerschmidt, M. Jahn, and D. Jahn. 2007. Functional definition of the tobacco protoporphyrinogen IX oxidase substrate-binding site. Biochem J 402 (3):575–80
Heinemann IU, Jahn M, Jahn D (2008) The biochemistry of heme biosynthesis. Arch Biochem Biophys 474(2):238–251
Hennig M, Grimm B, Contestabile R, John RA, Jansonius JN (1997) Crystal structure of glutamate-1-semialdehyde aminomutase: an alpha2-dimeric vitamin B6-dependent enzyme with asymmetry in structure and active site reactivity. Proc Natl Acad Sci USA 94(10):4866–4871
Hoare DS, Heath H (1958) Intermediates in the biosynthesis of porphyrins from porphobilinogen by Rhodopseudomonas spheroides. Nature 181(4623):1592–1593
Hobbs C, Dailey HA, Shepherd M (2016) The HemQ coprohaem decarboxylase generates reactive oxygen species: implications for the evolution of classical haem biosynthesis. Biochem J 473(21):3997–4009
Hobbs C, Reid JD, Shepherd M (2017) The coproporphyrin ferrochelatase of Staphylococcus aureus: mechanistic insights into a regulatory iron-binding site. Biochem J 474(20):3513–3522
Hofbauer S, Gruber C, Pirker KF, Sundermann A, Schaffner I, Jakopitsch C, Oostenbrink C, Furtmuller PG, Obinger C (2014) Transiently produced hypochlorite is responsible for the irreversible inhibition of chlorite dismutase. Biochemistry 53(19):3145–3157
Hofbauer S, Dalla Sega M, Scheiblbrandner S, Jandova Z, Schaffner I, Mlynek G, Djinovic-Carugo K, Battistuzzi G, Furtmuller PG, Oostenbrink C et al (2016) Chemistry and molecular dynamics simulations of heme b-HemQ and coproheme-HemQ. Biochemistry 55(38):5398–5412
Ilag LL, Jahn D (1992) Activity and spectroscopic properties of the Escherichia coli glutamate 1-semialdehyde aminotransferase and the putative active site mutant K265R. Biochemistry 31(31):7143–7151
Ishida T, Yu L, Akutsu H, Ozawa K, Kawanishi S, Seto A, Inubushi T, Sano S (1998) A primitive pathway of porphyrin biosynthesis and enzymology in Desulfovibrio vulgaris. Proc Natl Acad Sci USA 95(9):4853–4858
Ishihara T, Tomita H, Hasegawa Y, Tsukagoshi N, Yamagata H, Udaka S (1995) Cloning and characterization of the gene for a protein thiol-disulfide oxidoreductase in Bacillus brevis. J Bacteriol 177(3):745–749
Jackson AH, Sancovich HA, Ferramola AM, Evans N, Games DE, Matlin SA, Elder GH, Smith SG (1976) Macrocyclic intermediates in the biosynthesis of porphyrins. Philos Trans R Soc Lond Ser B Biol Sci 273(924):191–206
Jacobs NJ, Jacobs JM (1978) Quinones as hydrogen carriers for a late step in anaerobic heme biosynthesis in Escherichia coli. Biochim Biophys Acta 544(3):540–546
Jacobs NJ, Jacobs JM, Brent P (1970) Formation of protoporphyrin from coproporphyrinogen in extracts of various bacteria. J Bacteriol 102(2):398–403
Jacobs NJ, Jacobs JM, Brent P (1971) Characterization of the late steps of microbial heme synthesis: conversion of coproporphyrinogen to protoporphyrin. J Bacteriol 107(1):203–209
Jaffe EK (2004) The porphobilinogen synthase catalyzed reaction mechanism. Bioorg Chem 32(5):316–325
Jaffe EK (2016) The remarkable character of porphobilinogen synthase. Acc Chem Res 49(11):2509–2517
Jaffe EK, Lawrence SH (2012) Allostery and the dynamic oligomerization of porphobilinogen synthase. Arch Biochem Biophys 519(2):144–153
Jaffe EK, Volin M, Bronson-Mullins CR, Dunbrack RL Jr, Kervinen J, Martins J, Quinlan JF Jr, Sazinsky MH, Steinhouse EM, Yeung AT (2000) An artificial gene for human porphobilinogen synthase allows comparison of an allelic variation implicated in susceptibility to lead poisoning. J Biol Chem 275(4):2619–2626
Jahn D (1992) Expression of the Chlamydomonas reinhardtii chloroplast tRNA(Glu) gene in a homologous in vitro transcription system is independent of upstream promoter elements. Arch Biochem Biophys 298(2):505–513
Jahn M, Jahn D (2012) Tetrapyrroles. In: Michal G, Schomburg D (eds) Biochemical pathways: an atlas of biochemistry and molecular biology. Wiley, pp 82–92
Jahn D, Verkamp E, Soll D (1992) Glutamyl-transfer RNA: a precursor of heme and chlorophyll biosynthesis. Trends Biochem Sci 17(6):215–218
Jordan PM, Seehra JS (1979) The biosynthesis of uroporphyrinogen III: order of assembly of the four porphobilinogen molecules in the formation of the tetrapyrrole ring. FEBS Lett 104(2):364–366
Jordan PM, Thomas SD, Warren MJ (1988) Purification, crystallization and properties of porphobilinogen deaminase from a recombinant strain of Escherichia coli K12. Biochem J 254(2):427–435
Karlberg T, Lecerof D, Gora M, Silvegren G, Labbe-Bois R, Hansson M, Al-Karadaghi S (2002) Metal binding to Saccharomyces cerevisiae ferrochelatase. Biochemistry 41(46):13499–13506
Kato K, Tanaka R, Sano S, Tanaka A, Hosaka H (2010) Identification of a gene essential for protoporphyrinogen IX oxidase activity in the cyanobacterium Synechocystis sp. PCC6803. Proc Natl Acad Sci USA 107:16649–16654
Kaufholz AL, Hunter GA, Ferreira GC, Lendrihas T, Hering V, Layer G, Jahn M, Jahn D (2013a) Aminolaevulinic acid synthase of Rhodobacter capsulatus: high-resolution kinetic investigation of the structural basis for substrate binding and catalysis. Biochem J 451(2):205–216
Kaufholz AL, Layer G, Heinz DW, Jahn M, Jahn D (2013b) The structural basis of porphyrias-defects of heme biosynthetic enzymes. In: Ferreira GC, Kadish KM, Smith KM (eds) Handbook of porphyrin science. World Scientific, Singapore, pp 1–42
Kikuchi G, Shemin D, Bachmann BJ (1958) The enzymic synthesis of delta-aminolevulinic acid. Biochim Biophys Acta 28(1):219–220
Koch M, Breithaupt C, Kiefersauer R, Freigang J, Huber R, Messerschmidt A (2004) Crystal structure of protoporphyrinogen IX oxidase: a key enzyme in haem and chlorophyll biosynthesis. EMBO J 23(8):1720–1728
Kosugi N, Araki T, Fujita J, Tanaka S, Fujiwara T (2017) Growth phenotype analysis of heme synthetic enzymes in a halophilic archaeon, Haloferax volcanii. PLoS One 12(12):e0189913
Kuhner M, Haufschildt K, Neumann A, Storbeck S, Streif J, Layer G (2014) The alternative route to heme in the methanogenic archaeon Methanosarcina barkeri. Archaea 2014:327637
Kuhner M, Schweyen P, Hoffmann M, Ramos JV, Reijerse EJ, Lubitz W, Broering M, Layer G (2016) The auxiliary [4Fe-4S] cluster of the radical SAM heme synthase from Methanosarcina barkeri is involved in electron transfer. Chem Sci 7:4633–4643
Lash TD (2005) The enigma of coproporphyrinogen oxidase: how does this unusual enzyme carry out oxidative decarboxylations to afford vinyl groups? Bioorg Med Chem Lett 15(20):4506–4509
Layer G, Verfurth K, Mahlitz E, Jahn D (2002) Oxygen-independent coproporphyrinogen-III oxidase HemN from Escherichia coli. J Biol Chem 277(37):34136–34142
Layer G, Moser J, Heinz DW, Jahn D, Schubert WD (2003) Crystal structure of coproporphyrinogen III oxidase reveals cofactor geometry of radical SAM enzymes. EMBO J 22(23):6214–6224
Layer G, Heinz DW, Jahn D, Schubert WD (2004) Structure and function of radical SAM enzymes. Curr Opin Chem Biol 8(5):468–476
Layer G, Kervio E, Morlock G, Heinz DW, Jahn D, Retey J, Schubert WD (2005) Structural and functional comparison of HemN to other radical SAM enzymes. Biol Chem 386(10):971–980
Layer G, Pierik AJ, Trost M, Rigby SE, Leech HK, Grage K, Breckau D, Astner I, Jansch L, Heathcote P et al (2006) The substrate radical of Escherichia coli oxygen-independent coproporphyrinogen III oxidase HemN. J Biol Chem 281(23):15727–15734
Layer G, Reichelt J, Jahn D, Heinz DW (2010) Structure and function of enzymes in heme biosynthesis. Protein Sci 19(6):1137–1161
Lecerof D, Fodje M, Hansson A, Hansson M, Al-Karadaghi S (2000) Structural and mechanistic basis of porphyrin metallation by ferrochelatase. J Mol Biol 297(1):221–232
Lecerof D, Fodje MN, Alvarez Leon R, Olsson U, Hansson A, Sigfridsson E, Ryde U, Hansson M, Al-Karadaghi S (2003) Metal binding to Bacillus subtilis ferrochelatase and interaction between metal sites. J Biol Inorg Chem 8(4):452–458
Lee DS, Flachsova E, Bodnarova M, Demeler B, Martasek P, Raman CS (2005) Structural basis of hereditary coproporphyria. Proc Natl Acad Sci USA 102(40):14232–14237
Levin EY (1968) Uroporphyrinogen 3 cosynthetase in bovine erythropoietic porphyria. Science 161(3844):907–908
Lewis CA Jr, Wolfenden R (2008) Uroporphyrinogen decarboxylation as a benchmark for the catalytic proficiency of enzymes. Proc Natl Acad Sci USA 105(45):17328–17333
Li S, Lou X, Xu Y, Teng X, Che S, Liu R, Bartlam M (2018) Crystal structure of a glutamate-1-semialdehyde-aminomutase from Pseudomonas aeruginosa PAO1. Biochem Biophys Res Commun 500(3):804–809
Lieb C, Siddiqui RA, Hippler B, Jahn D, Friedrich B (1998) The Alcaligenes eutrophus hemN gene encoding the oxygen-independent coproporphyrinogen III oxidase, is required for heme biosynthesis during anaerobic growth. Arch Microbiol 169(1):52–60
Lobo SA, Brindley A, Warren MJ, Saraiva LM (2009) Functional characterization of the early steps of tetrapyrrole biosynthesis and modification in Desulfovibrio vulgaris Hildenborough. Biochem J 420(2):317–325
Lobo SA, Lawrence AD, Romao CV, Warren MJ, Teixeira M, Saraiva LM (2014) Characterisation of Desulfovibrio vulgaris haem b synthase, a radical SAM family member. Biochim Biophys Acta 1844(7):1238–1247
Lobo SA, Scott A, Videira MA, Winpenny D, Gardner M, Palmer MJ, Schroeder S, Lawrence AD, Parkinson T, Warren MJ et al (2015) Staphylococcus aureus haem biosynthesis: characterisation of the enzymes involved in final steps of the pathway. Mol Microbiol 97(3):472–487
Lüer C, Schauer S, Möbius K, Schulze J, Schubert WD, Heinz DW, Jahn D, Moser J (2005) Complex formation between glutamyl-tRNA reductase and glutamate-1-semialdehyde 2,1-aminomutase in Escherichia coli during the initial reactions of porphyrin biosynthesis. J Biol Chem 280(19):18568–18572
Lüer C, Schauer S, Virus S, Schubert WD, Heinz DW, Moser J, Jahn D (2007) Glutamate recognition and hydride transfer by Escherichia coli glutamyl-tRNA reductase. FEBS J 274(17):4609–4614
Martins BM, Grimm B, Mock HP, Huber R, Messerschmidt A (2001) Crystal structure and substrate binding modeling of the uroporphyrinogen-III decarboxylase from Nicotiana tabacum. Implications for the catalytic mechanism. J Biol Chem 276(47):44108–44116
Masoumi A, Heinemann IU, Rohde M, Koch M, Jahn M, Jahn D (2008) Complex formation between protoporphyrinogen IX oxidase and ferrochelatase during haem biosynthesis in Thermosynechococcus elongatus. Microbiology 154(Pt 12):3707–3714
Mathews MA, Schubert HL, Whitby FG, Alexander KJ, Schadick K, Bergonia HA, Phillips JD, Hill CP (2001) Crystal structure of human uroporphyrinogen III synthase. EMBO J 20(21):5832–5839
Mathewson JH, Corwin AH (1961) Biosynthesis of pyrrole pigments: a mechanism for porphobilinogen polymerization. J Am Chem Soc 83:135–137
Mauzerall D, Granick S (1958) Porphyrin biosynthesis in erythrocytes. III. Uroporphyrinogen and its decarboxylase. J Biol Chem 232(2):1141–1162
Medlock AE, Dailey TA, Ross TA, Dailey HA, Lanzilotta WN (2007) A pi-helix switch selective for porphyrin deprotonation and product release in human ferrochelatase. J Mol Biol 373(4):1006–1016
Medlock AE, Carter M, Dailey TA, Dailey HA, Lanzilotta WN (2009) Product release rather than chelation determines metal specificity for ferrochelatase. J Mol Biol 393(2):308–319
Möbius K, Arias-Cartin R, Breckau D, Hännig AL, Riedmann K, Biedendieck R, Schroder S, Becher D, Magalon A, Moser J et al (2010) Heme biosynthesis is coupled to electron transport chains for energy generation. Proc Natl Acad Sci USA 107(23):10436–10441
Moore SJ, Sowa ST, Schuchardt C, Deery E, Lawrence AD, Ramos JV, Billig S, Birkemeyer C, Chivers PT, Howard MJ et al (2017) Elucidation of the biosynthesis of the methane catalyst coenzyme F430. Nature 543(7643):78–82
Moser J, Lorenz S, Hubschwerlen C, Rompf A, Jahn D (1999) Methanopyrus kandleri glutamyl-tRNA reductase. J Biol Chem 274(43):30679–30685
Moser J, Schubert WD, Beier V, Bringemeier I, Jahn D, Heinz DW (2001) V-shaped structure of glutamyl-tRNA reductase, the first enzyme of tRNA-dependent tetrapyrrole biosynthesis. EMBO J 20(23):6583–6590
Nishimura K, Taketani S, Inokuchi H (1995) Cloning of a human cDNA for protoporphyrinogen oxidase by complementation in vivo of a hemG mutant of Escherichia coli. J Biol Chem 270(14):8076–8080
Olsson U, Billberg A, Sjovall S, Al-Karadaghi S, Hansson M (2002) In vivo and in vitro studies of Bacillus subtilis ferrochelatase mutants suggest substrate channeling in the heme biosynthesis pathway. J Bacteriol 184(14):4018–4024
Palmer DJ, Schroeder S, Lawrence AD, Deery E, Lobo SA, Saraiva LM, McLean KJ, Munro AW, Ferguson SJ, Pickersgill RW et al (2014) The structure, function and properties of sirohaem decarboxylase–an enzyme with structural homology to a transcription factor family that is part of the alternative haem biosynthesis pathway. Mol Microbiol 93(2):247–261
Peng S, Zhang H, Gao Y, Pan X, Cao P, Li M, Chang W (2011) Crystal structure of uroporphyrinogen III synthase from Pseudomonas syringae pv. tomato DC3000. Biochem Biophys Res Commun 408(4):576–581
Pfanzagl V, Holcik L, Maresch D, Gorgone G, Michlits H, Furtmuller PG, Hofbauer S (2018) Coproheme decarboxylases – phylogenetic prediction versus biochemical experiments. Arch Biochem Biophys 640:27–36
Phillips JD, Whitby FG, Kushner JP, Hill CP (2003) Structural basis for tetrapyrrole coordination by uroporphyrinogen decarboxylase. EMBO J 22(23):6225–6233
Phillips JD, Whitby FG, Warby CA, Labbe P, Yang C, Pflugrath JW, Ferrara JD, Robinson H, Kushner JP, Hill CP (2004) Crystal structure of the oxygen-dependent coproporphyrinogen oxidase (Hem13p) of Saccharomyces cerevisiae. J Biol Chem 279(37):38960–38968
Phillips JD, Warby CA, Whitby FG, Kushner JP, Hill CP (2009) Substrate shuttling between active sites of uroporphyrinogen decarboxylase is not required to generate coproporphyrinogen. J Mol Biol 389(2):306–314
Pluta P, Roversi P, Bernardo-Seisdedos G, Rojas AL, Cooper JB, Gu S, Pickersgill RW, Millet O (2018) Structural basis of pyrrole polymerization in human porphobilinogen deaminase. Biochim Biophys Acta 1862:1948
Porra RJ, Falk JE (1961) Protein-bound porphyrins associated with protoporphyrin biosynthesis. Biochem Biophys Res Commun 5:179–184
Porra RJ, Falk JE (1964) The enzymic conversion of coproporphyrinogen 3 into protoporphyrin 9. Biochem J 90(1):69–75
Qi M, Lorenz M, Vogelpohl A (2002) Mathematical solution of the two-dimensional dispersion model. Chem Eng Technol 25(7):693–697
Qin X, Sun L, Wen X, Yang X, Tan Y, Jin H, Cao Q, Zhou W, Xi Z, Shen Y (2010) Structural insight into unique properties of protoporphyrinogen oxidase from Bacillus subtilis. J Struct Biol 170:76–82
Rand K, Noll C, Schiebel HM, Kemken D, Dulcks T, Kalesse M, Heinz DW, Layer G (2010) The oxygen-independent coproporphyrinogen III oxidase HemN utilizes harderoporphyrinogen as a reaction intermediate during conversion of coproporphyrinogen III to protoporphyrinogen IX. Biol Chem 391(1):55–63
Randau L, Schauer S, Ambrogelly A, Salazar JC, Moser J, Sekine S, Yokoyama S, Soll D, Jahn D (2004) tRNA recognition by glutamyl-tRNA reductase. J Biol Chem 279(33):34931–34937
Raux E, Leech HK, Beck R, Schubert HL, Santander PJ, Roessner CA, Scott AI, Martens JH, Jahn D, Thermes C et al (2003) Identification and functional analysis of enzymes required for precorrin-2 dehydrogenation and metal ion insertion in the biosynthesis of sirohaem and cobalamin in Bacillus megaterium. Biochem J 370(Pt 2):505–516
Rehse PH, Kitao T, Tahirov TH (2005) Structure of a closed-form uroporphyrinogen-III C-methyltransferase from Thermus thermophilus. Acta Crystallogr D Biol Crystallogr 61(Pt 7):913–919
Roberts A, Gill R, Hussey RJ, Mikolajek H, Erskine PT, Cooper JB, Wood SP, Chrystal EJ, Shoolingin-Jordan PM (2013) Insights into the mechanism of pyrrole polymerization catalysed by porphobilinogen deaminase: high-resolution X-ray studies of the Arabidopsis thaliana enzyme. Acta Crystallogr D Biol Crystallogr 69(Pt 3):471–485
Sano S, Granick S (1961) Mitochondrial coproporphyrinogen oxidase and protoporphyrin formation. J Biol Chem 236:1173–1180
Sasarman A, Letowski J, Czaika G, Ramirez V, Nead MA, Jacobs JM, Morais R (1993) Nucleotide sequence of the hemG gene involved in the protoporphyrinogen oxidase activity of Escherichia coli K12. Can J Microbiol 39(12):1155–1161
Schauer S, Chaturvedi S, Randau L, Moser J, Kitabatake M, Lorenz S, Verkamp E, Schubert WD, Nakayashiki T, Murai M et al (2002) Escherichia coli glutamyl-tRNA reductase. Trapping the thioester intermediate. J Biol Chem 277(50):48657–48663
Schubert HL, Raux E, Brindley AA, Leech HK, Wilson KS, Hill CP, Warren MJ (2002) The structure of Saccharomyces cerevisiae Met8p, a bifunctional dehydrogenase and ferrochelatase. EMBO J 21(9):2068–2075
Schubert HL, Phillips JD, Heroux A, Hill CP (2008) Structure and mechanistic implications of a uroporphyrinogen III synthase-product complex. Biochemistry 47(33):8648–8655
Schulze JO, Masoumi A, Nickel D, Jahn M, Jahn D, Schubert WD, Heinz DW (2006) Crystal structure of a non-discriminating glutamyl-tRNA synthetase. J Mol Biol 361(5):888–897
Seehra JS, Jordan PM, Akhtar M (1983) Anaerobic and aerobic coproporphyrinogen III oxidases of Rhodopseudomonas spheroides. Mechanism and stereochemistry of vinyl group formation. Biochem J 209(3):709–718
Shemin D, Rittenberg D (1945) The utilization of glycine for the synthesis of a porphyrin. J Biol Chem 159:567–568
Shepherd M, Dailey TA, Dailey HA (2006) A new class of [2Fe-2S]-cluster-containing protoporphyrin (IX) ferrochelatases. Biochem J 397(1):47–52
Sigfridsson E, Ryde U (2003) The importance of porphyrin distortions for the ferrochelatase reaction. J Biol Inorg Chem 8(3):273–282
Silva PJ, Ramos MJ (2008) A comparative density-functional study of the reaction mechanism of the O2-dependent coproporphyrinogen III oxidase. Bioorg Med Chem 16(6):2726–2733
Silva PJ, Schulz C, Jahn D, Jahn M, Ramos MJ (2010) A tale of two acids: when arginine is a more appropriate acid than H3O+. J Phys Chem B 114:8994–9001
Skotnicova P, Sobotka R, Shepherd M, Hajek J, Hrouzek P, Tichy M (2018) The cyanobacterial protoporphyrinogen oxidase HemJ is a new b-type heme protein functionally coupled with coproporphyrinogen III oxidase. J Biol Chem 293:12394
Smith AD, Warren MJ, Refsum H (2018) Vitamin B12. Adv Food Nutr Res 83:215–279
Spencer P, Jordan PM (1995) Characterization of the two 5-aminolaevulinic acid binding sites, the A- and P-sites, of 5-aminolaevulinic acid dehydratase from Escherichia coli. Biochem J 305(Pt 1):151–158
Spencer JB, Stolowich NJ, Roessner CA, Scott AI (1993) The Escherichia coli cysG gene encodes the multifunctional protein, siroheme synthase. FEBS Lett 335(1):57–60
Stark WM, Baker MG, Raithby PR, Leeper FJ, Battersby AR (1985) The spiro intermediate proposed for biosynthesis of the natural porphyrins: synthesis and properties of its macrocycle. J Chem Soc Chem Commun (19):1294–1296
Stark WM, Hart GJ, Battersby AR (1986) Synthetic studies on the proposed spiro intermediate for biosynthesis of the natural porphyrins: inhibition of cosynthetase. J Chem Soc Chem Commun (6):465–467
Stark MW, Hawker CJ, Hart GJ, Phillippides A, Petersen PM, Lewis DJ, Leeper FJ, Battersby AR (1993) Biosynthesis of porphyrins and related macrocycles. Part 40. Synthesis of a spiro-lactam related to the proposed spiro-intermediate for porphyrin biosynthesis: inhibition of cosynthetase. J Chem Soc Perkin Trans 1:2875–2892
Stephenson JR, Stacey JA, Morgenthaler JB, Friesen JA, Lash TD, Jones MA (2007) Role of aspartate 400, arginine 262, and arginine 401 in the catalytic mechanism of human coproporphyrinogen oxidase. Protein Sci 16(3):401–410
Stevens E, Frydman B (1968) Isolation and properties of wheat germ uroporphyrinogen 3 cosynthetase. Biochim Biophys Acta 151(2):429–437
Stojanovski BM, Hunter GA, Jahn M, Jahn D, Ferreira GC (2014) Unstable reaction intermediates and hysteresis during the catalytic cycle of 5-aminolevulinate synthase: implications from using pseudo and alternate substrates and a promiscuous enzyme variant. J Biol Chem 289(33):22915–22925
Storbeck S, Walther J, Muller J, Parmar V, Schiebel HM, Kemken D, Dulcks T, Warren MJ, Layer G (2009) The Pseudomonas aeruginosa nirE gene encodes the S-adenosyl-L-methionine-dependent uroporphyrinogen III methyltransferase required for heme d(1) biosynthesis. FEBS J 276(20):5973–5982
Storbeck S, Rolfes S, Raux-Deery E, Warren MJ, Jahn D, Layer G (2010) A novel pathway for the biosynthesis of heme in Archaea: genome-based bioinformatic predictions and experimental evidence. Archaea 2010:175050
Storbeck S, Saha S, Krausze J, Klink BU, Heinz DW, Layer G (2011) Crystal structure of the heme d1 biosynthesis enzyme NirE in complex with its substrate reveals new insights into the catalytic mechanism of S-adenosyl-l-methionine-dependent uroporphyrinogen III methyltransferases. J Biol Chem 286(30):26754–26767
Streit BR, Celis AI, Moraski GC, Shisler KA, Shepard EM, Rodgers KR, Lukat-Rodgers GS, DuBois JL (2018) Decarboxylation involving a ferryl, propionate, and a tyrosyl group in a radical relay yields heme b. J Biol Chem 293(11):3989–3999
Strey J, Wittchen KD, Meinhardt F (1999) Regulation of beta-galactosidase expression in Bacillus megaterium DSM319 by a XylS/AraC-type transcriptional activator. J Bacteriol 181(10):3288–3292
Tait GH (1969) Coproporphyrinogenase activity in extracts from Rhodopseudomonas spheroides. Biochem Biophys Res Commun 37(1):116–122
Tait GH (1972) Coproporphyrinogenase activities in extracts of Rhodopseudomonas spheroides and Chromatium strain D. Biochem J 128(5):1159–1169
Tan FC, Cheng Q, Saha K, Heinemann IU, Jahn M, Jahn D, Smith AG (2008) Identification and characterization of the Arabidopsis gene encoding the tetrapyrrole biosynthesis enzyme uroporphyrinogen III synthase. Biochem J 410(2):291–299
Troup B, Hungerer C, Jahn D (1995) Cloning and characterization of the Escherichia coli hemN gene encoding the oxygen-independent coproporphyrinogen III oxidase. J Bacteriol 177(11):3326–3331
Uchida T, Funamizu T, Chen M, Tanaka Y, Ishimori K (2018) Heme binding to porphobilinogen deaminase from Vibrio cholerae decelerates the formation of 1-hydroxymethylbilane. ACS Chem Biol 13(3):750–760
Vevodova J, Graham RM, Raux E, Schubert HL, Roper DI, Brindley AA, Ian Scott A, Roessner CA, Stamford NP, Elizabeth Stroupe M et al (2004) Structure/function studies on a S-adenosyl-l-methionine-dependent uroporphyrinogen III C methyltransferase (SUMT), a key regulatory enzyme of tetrapyrrole biosynthesis. J Mol Biol 344(2):419–433
Wang P, Grimm B (2015) Organization of chlorophyll biosynthesis and insertion of chlorophyll into the chlorophyll-binding proteins in chloroplasts. Photosynth Res 126(2–3):189–202
Wang Y, Shen Y, Ryde U (2009) QM/MM study of the insertion of metal ion into protoporphyrin IX by ferrochelatase. J Inorg Biochem 103(12):1680–1686
Wang B, Wen X, Qin X, Wang Z, Tan Y, Shen Y, Xi Z (2013) Quantitative structural insight into human variegate porphyria disease. J Biol Chem 288:11731
Warren MJ, Jordan PM (1988) Investigation into the nature of substrate binding to the dipyrromethane cofactor of Escherichia coli porphobilinogen deaminase. Biochemistry 27(25):9020–9030
Warren MJ, Roessner CA, Santander PJ, Scott AI (1990) The Escherichia coli cysG gene encodes S-adenosylmethionine-dependent uroporphyrinogen III methylase. Biochem J 265(3):725–729
Whitby FG, Phillips JD, Kushner JP, Hill CP (1998) Crystal structure of human uroporphyrinogen decarboxylase. EMBO J 17(9):2463–2471
Woodcock SC, Jordan PM (1994) Evidence for participation of aspartate-84 as a catalytic group at the active site of porphobilinogen deaminase obtained by site-directed mutagenesis of the hemC gene from Escherichia coli. Biochemistry 33(9):2688–2695
Xu K, Elliott T (1994) Cloning, DNA sequence, and complementation analysis of the Salmonella typhimurium hemN gene encoding a putative oxygen-independent coproporphyrinogen III oxidase. J Bacteriol 176(11):3196–3203
Zhao A, Fang Y, Chen X, Zhao S, Dong W, Lin Y, Gong W, Liu L (2014) Crystal structure of Arabidopsis glutamyl-tRNA reductase in complex with its stimulator protein. Proc Natl Acad Sci USA 111(18):6630–6635
Zheng K, Ngo PD, Owens VL, Yang XP, Mansoorabadi SO (2016) The biosynthetic pathway of coenzyme F430 in methanogenic and methanotrophic archaea. Science 354(6310):339–342
Zwerschke D, Karrie S, Jahn D, Jahn M (2014) Leishmania major possesses a unique HemG-type protoporphyrinogen IX oxidase. Biosci Rep 34(4):e00124dailey
Acknowledgments
We thank Stefan Barthels for his excellent technical assistance and are indebted to the Deutsche Forschungsgemeinschaft (GRK 2223, PROCOMPAS) for funding.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this entry
Cite this entry
Müller, K., Mingers, T., Haskamp, V., Jahn, D., Jahn, M. (2019). Biosynthesis and Insertion of Heme. In: Rojo, F. (eds) Aerobic Utilization of Hydrocarbons, Oils, and Lipids. Handbook of Hydrocarbon and Lipid Microbiology . Springer, Cham. https://doi.org/10.1007/978-3-319-50418-6_17
Download citation
DOI: https://doi.org/10.1007/978-3-319-50418-6_17
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-50417-9
Online ISBN: 978-3-319-50418-6
eBook Packages: Biomedical and Life SciencesReference Module Biomedical and Life Sciences