Abstract
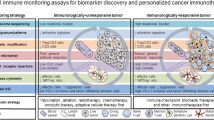
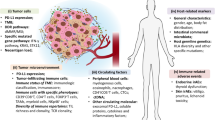
Cancer immunotherapy has revolutionized both the oncology research and treatment. Many of these treatment modalities have rapidly grown and gained approval over the last several years. Despite the durable and robust response to this outstanding class, the majority of patients remain resistant to such treatments. Hence, a crucial goal of on-going research is to broaden the scope of treatable tumors and patients and provide clinicians with tools that can predict patients’ responses to selected treatments. In this chapter, we shed the light on the importance of applying integrated molecular and computational approaches to address those research needs and potentially enhance the personalized medicine strategies. Numerous molecular high-throughput platforms are currently available and provide the means to perform a comprehensive immune profiling in the context of tumor immunotherapies. This allows the researchers to understand the reasons underlying the success or failure of these therapies and design strategies to mitigate any potential side effects. In addition, mathematical models have been used to predict the efficacy of therapies by recognizing and correlating specific immune features at the diagnosis time, which is crucial for selecting candidate patients who are more likely to benefit from specific immunotherapeutic treatments. Both experimental and computational tools complement each other and should be among the arsenal of tools in developing and optimizing immunotherapy approaches. The cutting-edge molecular and computational technologies exploited to predict the patients’ response to immunotherapy will be discussed throughout this chapter.
Similar content being viewed by others
References
Agur Z, Elishmereni M, Foryś U, Kogan Y (2020) Accelerating the development of personalized cancer immunotherapy by integrating molecular patients’ profiles with dynamic mathematical models. Clin Pharmacol Ther 108:515–527. https://doi.org/10.1002/cpt.1942
Alva A, Daniels GA, Wong MKK et al (2016) Contemporary experience with high-dose interleukin-2 therapy and impact on survival in patients with metastatic melanoma and metastatic renal cell carcinoma. Cancer Immunol Immunother 65:1533–1544. https://doi.org/10.1007/S00262-016-1910-X
Bendall SC, Nolan GP, Roederer M, Chattopadhyay PK (2012) A deep profiler’s guide to cytometry. Trends Immunol 33:323–332. https://doi.org/10.1016/j.it.2012.02.010
Berglund E, Maaskola J, Schultz N et al (2018) Spatial maps of prostate cancer transcriptomes reveal an unexplored landscape of heterogeneity. Nat Commun 91(9):1–13. https://doi.org/10.1038/s41467-018-04724-5
Blache U, Weiss R, Boldt A et al (2021) Advanced flow cytometry assays for immune monitoring of CAR-T cell applications. Front Immunol 12:1–17. https://doi.org/10.3389/fimmu.2021.658314
Borghaei H, Paz-Ares L, Horn L et al (2015) Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N Engl J Med 373:1627–1639. https://doi.org/10.1056/NEJMoa1507643
Bracci L, Fragale A, Gabriele L, Moschella F (2020) Towards a systems immunology approach to unravel responses to cancer immunotherapy. Front Immunol 11:2748
Brogden KA, Parashar D, Hallier AR et al (2018) Genomics of NSCLC patients both affirm PD-L1 expression and predict their clinical responses to anti-PD-1 immunotherapy. BMC Cancer 18:225. https://doi.org/10.1186/s12885-018-4134-y
Charoentong P, Finotello F, Angelova M et al (2017) Pan-cancer Immunogenomic analyses reveal genotype-Immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep 18:248–262. https://doi.org/10.1016/j.celrep.2016.12.019
Chen X (2020) Chapter 26: Single-cell epigenomics: methods and translation. In: Tollefsbol TBT-EM (ed) Translational epigenetics. Academic, pp 525–535
Chen JA, Ma W, Yuan J, Li T (2020a) Translational biomarkers and rationale strategies to overcome resistance to immune checkpoint inhibitors in solid tumors. Cancer Treat Res 180:251–279. https://doi.org/10.1007/978-3-030-38862-1_9
Chen L, Zhong F, Zhu J (2020b) Bridging targeted and untargeted mass spectrometry-based metabolomics via hybrid approaches. Meta 10:1–19. https://doi.org/10.3390/metabo10090348
Chen Y, Hu J, Bu F et al (2020c) Clinical characteristics of hyperprogressive disease in NSCLC after treatment with immune checkpoint inhibitor: a systematic review and meta-analysis. BMC Cancer 20:707. https://doi.org/10.1186/s12885-020-07206-4
Das S, Johnson DB (2019) Immune-related adverse events and anti-tumor efficacy of immune checkpoint inhibitors. J Immunother Cancer 7:306. https://doi.org/10.1186/s40425-019-0805-8
Davis MM, Tato CM, Furman D (2017) Systems immunology: just getting started. Nat Immunol 18:725–732. https://doi.org/10.1038/ni.3768
Depledge DP, Srinivas KP, Sadaoka T et al (2019) Direct RNA sequencing on nanopore arrays redefines the transcriptional complexity of a viral pathogen. Nat Commun 101(10):1–13. https://doi.org/10.1038/s41467-019-08734-9
Dine J, Gordon R, Shames Y et al (2017) Immune checkpoint inhibitors: an innovation in immunotherapy for the treatment and management of patients with cancer. Asia-Pacific J Oncol Nurs 4:127
Fancello L, Gandini S, Pelicci PG, Mazzarella L (2019) Tumor mutational burden quantification from targeted gene panels: major advancements and challenges. J Immunother Cancer 7:183. https://doi.org/10.1186/s40425-019-0647-4
Fister KR, Donnelly JH (2005) Immunotherapy: an optimal control theory approach. Math Biosci Eng 2:499–510. https://doi.org/10.3934/mbe.2005.2.499
Frei AP, Bava F-A, Zunder ER et al (2016) Highly multiplexed simultaneous detection of rnAs and proteins in single cells. https://doi.org/10.1038/nmeth.3742
Garon EB, Rizvi NA, Hui R et al (2015) Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med 372:2018–2028. https://doi.org/10.1056/NEJMoa1501824
Gebregiworgis T, Powers R (2012) Application of NMR metabolomics to search for human disease biomarkers. Comb Chem High Throughput Screen 15:595–610. https://doi.org/10.2174/138620712802650522
Heinrich S, Craig AJ, Ma L et al (2021) Understanding tumour cell heterogeneity and its implication for immunotherapy in liver cancer using single-cell analysis. J Hepatol 74:700–715. https://doi.org/10.1016/j.jhep.2020.11.036
Hellmann MD, Callahan MK, Awad MM et al (2018) Tumor mutational burden and efficacy of Nivolumab monotherapy and in combination with Ipilimumab in small-cell lung cancer. Cancer Cell 33:853–861.e4. https://doi.org/10.1016/j.ccell.2018.04.001
Hwang S, Kwon A-Y, Jeong J-Y et al (2020) Immune gene signatures for predicting durable clinical benefit of anti-PD-1 immunotherapy in patients with non-small cell lung cancer. Sci Rep 10:643. https://doi.org/10.1038/s41598-019-57218-9
Kargl J, Zhu X, Zhang H et al (2019) Neutrophil content predicts lymphocyte depletion and anti-PD1 treatment failure in NSCLC. JCI Insight 4. https://doi.org/10.1172/jci.insight.130850
Kirschner D, Panetta JC (1998) Modeling immunotherapy of the tumor-immune interaction. J Math Biol 37:235–252. https://doi.org/10.1007/s002850050127
Kogan Y, Halevi-Tobias K, Elishmereni M et al (2012) Reconsidering the paradigm of cancer immunotherapy by computationally aided real-time personalization. Cancer Res 72:2218–2227. https://doi.org/10.1158/0008-5472.CAN-11-4166
Krishnamurthy A, Jimeno A (2017) Atezolizumab: a novel PD-L1 inhibitor in cancer therapy with a focus in bladder and non-small cell lung cancers. Drugs Today 53:217–237. https://doi.org/10.1358/DOT.2017.53.4.2589163
Kronik N, Kogan Y, Vainstein V, Agur Z (2008) Improving alloreactive CTL immunotherapy for malignant gliomas using a simulation model of their interactive dynamics. Cancer Immunol Immunother 57:425–439. https://doi.org/10.1007/s00262-007-0387-z
Kronik N, Kogan Y, Elishmereni M et al (2010) Predicting outcomes of prostate cancer immunotherapy by personalized mathematical models. PLoS One 5:e15482. https://doi.org/10.1371/journal.pone.0015482
Kruger S, Ilmer M, Kobold S et al (2019) Advances in cancer immunotherapy 2019 – latest trends. J Exp Clin Cancer Res 38:268. https://doi.org/10.1186/s13046-019-1266-0
Kuai R, Yuan W, Son S et al (2018) Elimination of established tumors with nanodisc-based combination chemoimmunotherapy. Sci Adv, 4:eaao1736. https://doi.org/10.1126/sciadv.aao1736
Kuznetsov VA, Makalkin IA, Taylor MA, Perelson AS (1994) Nonlinear dynamics of immunogenic tumors: parameter estimation and global bifurcation analysis. Bull Math Biol 56:295–321. https://doi.org/10.1007/BF02460644
Lau D, Bobe AM, Khan AA (2019) RNA sequencing of the tumor microenvironment in precision cancer immunotherapy. Trends Cancer 5:149–156. https://doi.org/10.1016/j.trecan.2019.02.006
Le DT, Uram JN, Wang H et al (2015) PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med 372:2509–2520
Li H, Bullock K, Gurjao C et al (2019) Metabolomic adaptations and correlates of survival to immune checkpoint blockade. Nat Commun 10:1–6
Liao X, Makris M, Luo XM (2016) Fluorescence-activated cell sorting for purification of Plasmacytoid dendritic cells from the mouse bone marrow. J Vis Exp 2016:54641. https://doi.org/10.3791/54641
Longo SK, Guo MG, Ji AL, Khavari PA (2021) Integrating single-cell and spatial transcriptomics to elucidate intercellular tissue dynamics. Nat Rev Genet 22:627–644. https://doi.org/10.1038/s41576-021-00370-8
Ma S, Zhang B, LaFave LM et al (2020) Chromatin potential identified by shared single-cell profiling of RNA and chromatin. Cell 183:1103–1116.e20. https://doi.org/10.1016/j.cell.2020.09.056
Macklin A, Khan S, Kislinger T (2020) Recent advances in mass spectrometry based clinical proteomics: applications to cancer research. Clin Proteomics 17:1–25. https://doi.org/10.1186/s12014-020-09283-w
Masucci GV, Cesano A, Hawtin R et al (2016) Validation of biomarkers to predict response to immunotherapy in cancer: volume I — pre-analytical and analytical validation. J Immunother Cancer 4:76. https://doi.org/10.1186/s40425-016-0178-1
McCombie WR, McPherson JD, Mardis ER (2019) Next-generation sequencing technologies. Cold Spring Harb Perspect Med 9. https://doi.org/10.1101/CSHPERSPECT.A036798
Monsurrò V, Monsurrò M, Marincola FM (2011) Microarray analysis for a comprehensive immunological-status evaluation during cancer vaccine immune monitoring. J Biomed Biotechnol 2011. https://doi.org/10.1155/2011/307297
Muhammad II, Kong SL, Akmar Abdullah SN, Munusamy U (2019) RNA-seq and ChIP-seq as complementary approaches for comprehension of plant transcriptional regulatory mechanism. Int J Mol Sci 21. https://doi.org/10.3390/ijms21010167
Neu KE, Tang Q, Wilson PC, Khan AA (2017) Single-cell genomics: approaches and utility in immunology. Trends Immunol 38:140–149
Nie X, Xia L, Gao F et al (2021) Serum metabolite biomarkers predictive of response to PD-1 blockade therapy in non-small cell lung cancer. Front Mol Biosci 8:1–11. https://doi.org/10.3389/fmolb.2021.678753
Nishino M, Ramaiya NH, Hatabu H, Hodi FS (2017) Monitoring immune-checkpoint blockade: response evaluation and biomarker development. Nat Rev Clin Oncol 14:655–668. https://doi.org/10.1038/nrclinonc.2017.88
Patel SP, Kurzrock R (2015) PD-L1 expression as a predictive biomarker in cancer immunotherapy. Mol Cancer Ther 14:847–856. https://doi.org/10.1158/1535-7163.MCT-14-0983
Perlstein D, Shlagman O, Kogan Y et al (2019) Personal response to immune checkpoint inhibitors of patients with advanced melanoma explained by a computational model of cellular immunity, tumor growth, and drug. PLoS One 14:e0226869. https://doi.org/10.1371/journal.pone.0226869
Philip M, Fairchild L, Sun L et al (2017) Chromatin states define tumour-specific T cell dysfunction and reprogramming. Nature 545:452–456. https://doi.org/10.1038/nature22367
Picelli S, Björklund ÅK, Faridani OR et al (2013) Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat Methods 10:1096–1100. https://doi.org/10.1038/nmeth.2639
Rizvi NA, Hellmann MD, Snyder A et al (2015) Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 80(348):124–128. https://doi.org/10.1126/science.aaa1348
Rohaan MW, Wilgenhof S, Haanen JBAG (2019) Adoptive cellular therapies: the current landscape. Virchows Arch 474:449–461. https://doi.org/10.1007/s00428-018-2484-0
Russi AE, Brown MA (2016) Mass cytometry: single cells, many features. Cell 165:255–269. https://doi.org/10.1016/j.cell.2016.04.019.Mass
Samstein RM, Lee C-H, Shoushtari AN et al (2019) Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet 51:202–206. https://doi.org/10.1038/s41588-018-0312-8
Schiffman C, Lin C, Shi F et al (2017) SIDEseq: a cell similarity measure defined by shared identified differentially expressed genes for single-cell RNA sequencing data. Stat Biosci 9:200–216. https://doi.org/10.1007/s12561-017-9194-z
See P, Lum J, Chen J, Ginhoux F (2018) A single-cell sequencing guide for immunologists. Front Immunol 9:2425. https://doi.org/10.3389/FIMMU.2018.02425/BIBTEX
Shindo Y, Hazama S, Tsunedomi R et al (2019) Novel biomarkers for personalized cancer immunotherapy. Cancers (Basel) 11. https://doi.org/10.3390/cancers11091223
Shyr D, Liu Q (2014) Next generation sequencing in cancer research and clinical application. Omi Clin Pract Genom Pharmacogenom Proteomics Trans Clin Res:71–94. https://doi.org/10.1201/b17137
Siebert JC, Wagner BD, Juarez-Colunga E (2014) Integrating and mining diverse data in human immunological studies. Bioanalysis 6:209–223. https://doi.org/10.4155/bio.13.309
Snyder A, Makarov V, Merghoub T et al (2014) Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med 371:2189–2199. https://doi.org/10.1056/NEJMoa1406498
Ståhl PL, Salmén F, Vickovic S et al (2016) Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353:78–82. https://doi.org/10.1126/science.aaf2403
Stoeckius M, Hafemeister C, Stephenson W et al (2017) Simultaneous epitope and transcriptome measurement in single cells. Nat Methods 149(14):865–868. https://doi.org/10.1038/nmeth.4380
Sun L, Chen L, Li H (2019) Checkpoint-modulating immunotherapies in tumor treatment: targets, drugs, and mechanisms. Int Immunopharmacol 67:160–175. https://doi.org/10.1016/j.intimp.2018.12.006
Szymańska Z (2003) Analysis of immunotherapy models in the context of cancer dynamics. Int J Appl Math Comput Sci 13:407–418
Tang F, Barbacioru C, Nordman E et al (2010) RNA-Seq analysis to capture the transcriptome landscape of a single cell. Nat Protoc 5:516–535. https://doi.org/10.1038/nprot.2009.236
Trevino V, Falciani F, Barrera-Saldaña HA (2007) DNA microarrays: a powerful genomic tool for biomedical and clinical research. Mol Med 13:527–541. https://doi.org/10.2119/2006-00107.TREVINO/FIGURES/13
Tyanova S, Albrechtsen R, Kronqvist P et al (2016) Proteomic maps of breast cancer subtypes. Nat Commun 7:1–11
Van Allen EM, Miao D, Schilling B et al (2015) Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 80(350):207–211. https://doi.org/10.1126/science.aad0095
Waldman AD, Fritz JM, Lenardo MJ (2020) A guide to cancer immunotherapy: from T cell basic science to clinical practice. Nat Rev Immunol 20:651–668. https://doi.org/10.1038/s41577-020-0306-5
Wang X, He Y, Zhang Q et al (2021) Direct comparative analyses of 10X genomics chromium and Smart-seq2. Genomics Proteomics Bioinformatics 19:253. https://doi.org/10.1016/J.GPB.2020.02.005
Xie F, Zhang J, Wang J et al (2020) Multifactorial deep learning reveals pan-cancer genomic tumor clusters with distinct Immunogenomic landscape and response to immunotherapy. Clin Cancer Res 26:2908–2920. https://doi.org/10.1158/1078-0432.CCR-19-1744
Yang B, Zhou L, Zhong J et al (2021) Combination of computed tomography imaging-based radiomics and clinicopathological characteristics for predicting the clinical benefits of immune checkpoint inhibitors in lung cancer. Respir Res 22:189. https://doi.org/10.1186/s12931-021-01780-2
Yu J, Peng J, Chi H (2019) Systems immunology: integrating multi-omics data to infer regulatory networks and hidden drivers of immunity. Curr Opin Syst Biol 15:19–29. https://doi.org/10.1016/j.coisb.2019.03.003
Zachariadis V, Cheng H, Andrews N, Enge M (2020) A highly scalable method for joint whole-genome sequencing and gene-expression profiling of single cells. Mol Cell 80:541–553.e5
Zheng C, Zheng L, Yoo JK et al (2017) Landscape of infiltrating T cells in liver cancer revealed by single-cell sequencing. Cell 169:1342–1356.e16. https://doi.org/10.1016/j.cell.2017.05.035
Zheng P-P, Kros JM, Li J (2018) Approved CAR T cell therapies: ice bucket challenges on glaring safety risks and long-term impacts. Drug Discov Today 23:1175–1182. https://doi.org/10.1016/j.drudis.2018.02.012
Zhu C, Preissl S, Ren B (2020) Single-cell multimodal omics: the power of many. Nat Methods 17:11–14. https://doi.org/10.1038/s41592-019-0691-5
Ziegenhain C, Vieth B, Parekh S et al (2017) Comparative analysis of single-cell RNA sequencing methods. Mol Cell 65:631–643.e4. https://doi.org/10.1016/j.molcel.2017.01.023
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 Springer Nature Switzerland AG
About this entry
Cite this entry
Elghaish, R.A., Alaswad, Z., Abdelhafeez, S.H., Salem, O., Elserafy, M. (2023). Advancing Cancer Immunotherapy Through Integrating Molecular and Computational Approaches. In: Rezaei, N. (eds) Handbook of Cancer and Immunology. Springer, Cham. https://doi.org/10.1007/978-3-030-80962-1_316-1
Download citation
DOI: https://doi.org/10.1007/978-3-030-80962-1_316-1
Received:
Accepted:
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-80962-1
Online ISBN: 978-3-030-80962-1
eBook Packages: Springer Reference Biomedicine and Life SciencesReference Module Biomedical and Life Sciences