Abstract
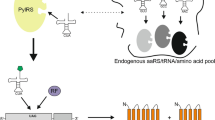
Cell-free protein synthesis (CFPS) offers a fast and inexpensive means to incorporate unnatural amino acids (UAAs) site specifically into proteins. This enables engineering of proteins and allows production of protein-based probes for analysis of their interactions with other molecules. Using dialysis Escherichia coli CFPS system in combination with aminoacyl-tRNA synthetase and suppressor tRNA evolved from Methanocaldococcus jannaschii high expression yield of proteins with site specifically incorporated UAAs can be achieved. Typically the target protein can be prepared at concentrations of about 1 mg/mL, which is generally sufficient for subsequent applications.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kigawa T, Yabuki T, Yoshida Y et al (1999) Cell-free production and stable-isotope labeling of milligram quantities of proteins. FEBS Lett 442:15–19
Renesto P, Raoult D (2003) In vitro expression of rickettsial proteins. Ann N Y Acad Sci 990:642–645
Kigawa T, Yamaguchi-Nunokawa E, Kodama K et al (2001) Selenomethionine incorporation into a protein by cell-free synthesis. J Struct Funct Genomics 2:29–35
Ozawa K, Headlam MJ, Mouradov D et al (2005) Translational incorporation of L-3,4-dihydroxyphenylalanine into proteins. FEBS J 272:3162–3171
Ozawa K, Jergic S, Crowther JA et al (2005) Cell-free protein synthesis in an autoinduction system for NMR studies of protein–protein interactions. J Biomol NMR 32:235–241
Ozawa K, Jergic S, Park AY et al (2008) The proofreading exonuclease subunit ε of Escherichia coli DNA polymerase III is tethered to the polymerase subunit α via a flexible linker. Nucleic Acids Res 36:5074–5082
Klammt C, Löhr F, Schäfer B et al (2004) High level cell-free expression and specific labeling of integral membrane proteins. Eur J Biochem 271:568–580
Klammt C, Schwarz D, Löhr F et al (2006) Cell-free expression as an emerging technique for the large scale production of integral membrane protein. FEBS J 273:4141–4153
Ozawa K, Dixon NE, Otting G (2005) Cell-free synthesis of 15N-labeled proteins for NMR studies. IUBMB Life 57:615–622
Ozawa K, Wu PSC, Dixon NE et al (2006) 15N-Labelled proteins by cell-free protein synthesis: strategies for high-throughput NMR studies of proteins and protein–ligand complexes. FEBS J 273:4154–4159
Shimizu Y, Kuruma Y, Ying BW et al (2006) Cell-free translation systems for protein engineering. FEBS J 273:4133–4140
Wang L, Schultz PG (2001) A general approach for the generation of orthogonal tRNAs. Chem Biol 8:883–890
Liu CC, Schultz PG (2010) Adding new chemistries to the genetic code. Annu Rev Biochem 79:413–444
Young TS, Ahmad I, Yin JA et al (2010) An enhanced system for unnatural amino acid mutagenesis in E. coli. J Mol Biol 395:361–374
Ozawa K, Loscha KV, Kuppan KV et al (2012) High-yield cell-free protein synthesis for site-specific incorporation of unnatural amino acids at two sites. Biochem Biophys Res Commun 418:652–656
Neylon C, Brown SE, Kralicek AV et al (2000) Interaction of the Escherichia coli replication terminator protein (Tus) with DNA: a model derived from DNA-binding studies of mutant proteins by surface plasmon resonance. Biochemistry 39:11989–11999
Wu PSC, Ozawa K, Lim SP et al (2007) Cell-free transcription/translation from PCR amplified DNA for high-throughput NMR studies. Angew Chem Int Ed 46:3356–3358
Fleissner MR, Brustad EM, Kálai T et al (2009) Site-directed spin labeling of a genetically encoded unnatural amino acid. Proc Natl Acad Sci U S A 106:21637–21642
Hong V, Presolski SI, Ma C et al (2009) Analysis and optimization of copper-catalyzed azide-alkyne cycloaddition for bioconjugation. Angew Chem Int Ed 48:9879–9883
Pratt JM (1984) Coupled transcription-translation in prokaryotic cell-free systems. In: Hames BD, Higgins SJ (eds) Transcription and translation. IRL Press, Oxford, pp 179–209
Apponyi M, Ozawa K, Dixon NE et al (2008) Cell-free protein synthesis for analysis by NMR spectroscopy. In: Kobe B, Guss M, Huber T (eds) Methods in molecular biology. Structural proteomics: high-throughput methods, vol 426. Humana, Totowa, NJ, pp 257–268
Jergic S, Ozawa K, Williams NK et al (2007) The unstructured C-terminus of the τ subunit of Escherichia coli DNA polymerase III holoenzyme is the site of interaction with the α subunit. Nucleic Acids Res 35:2813–2824
Loscha KV, Herlt AJ, Qi R, Huber T et al (2012) Multiple-site labeling of proteins with unnatural amino acids. Angew Chem Int Ed 51:2243–2246
Guo J, Melançon CE 3rd, Lee HS et al (2009) Evolution of amber suppressor tRNAs for efficient bacterial production of proteins containing nonnatural amino acids. Angew Chem Int Ed Engl 48(48):9148–9151
Chin JW, Santoro SW, Martin AB et al (2002) Addition of p-azido-l-phenylalanine to the genetic code of Escherichia coli. J Am Chem Soc 124(31):9026–9027
Acknowledgements
This work has been supported by Profs. Gottfried Otting (Australian National University, Canberra, Australia) and Nicholas E. Dixon (University of Wollongong, Wollongong, Australia) and the Australian Research Council, including a fellowship to K.O. K.O. thanks Dr. Andrew V. Kralicek (Plant and Food Research, Auckland, New Zealand) for critical reading of an earlier version of the manuscript.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Ozawa, K., Loh, C.T. (2014). Site-Specific Incorporation of Unnatural Amino Acids into Proteins by Cell-Free Protein Synthesis. In: Alexandrov, K., Johnston, W. (eds) Cell-Free Protein Synthesis. Methods in Molecular Biology, vol 1118. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-782-2_12
Download citation
DOI: https://doi.org/10.1007/978-1-62703-782-2_12
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-781-5
Online ISBN: 978-1-62703-782-2
eBook Packages: Springer Protocols