Abstract
Intrinsically disordered or unstructured regions in proteins are both common and biologically important, particularly in regulation, signaling, and modulating intermolecular recognition processes. From a practical point of view, however, such disordered regions often can pose significant challenges for crystallization. Disordered regions are also detrimental to NMR spectral quality, complicating the analysis of resonance assignments and three-dimensional protein structures by NMR methods. The DisMeta Server has been used by Northeastern Structural Genomics (NESG) consortium as a primary tool for construct design and optimization in preparing samples for both NMR and crystallization studies. It is a meta-server that generates a consensus analysis of eight different sequence-based disorder predictors to identify regions that are likely to be disordered. DisMeta also identifies predicted secretion signal peptides, transmembrane segments, and low-complexity regions. Identification of disordered regions, by either experimental or computational methods, is an important step in the NESG structure production pipeline, allowing the rational design of protein constructs that have improved expression and solubility, improved crystallization, and better quality NMR spectra.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Dyson HJ, Wright PE (2002) Coupling of folding and binding for unstructured proteins. Curr Opin Struct Biol 12:54–60
Iakoucheva LM, Brown CJ, Lawson JD et al (2002) Intrinsic disorder in cell-signaling and cancer-associated proteins. J Mol Biol 323:573–584
Radivojac P, Iakoucheva LM, Oldfield CJ et al (2007) Intrinsic disorder and functional proteomics. Biophys J 92:1439–1456
Kovacs D, Szabo B, Pancsa R et al (2012) Intrinsically disordered proteins undergo and assist folding transitions in the proteome. Arch Biochem Biophys 531:80–89
Liu J, Perumal NB, Oldfield CJ et al (2006) Intrinsic disorder in transcription factors. Biochemistry 45:6873–6888
Minezaki Y, Homma K, Kinjo AR et al (2006) Human transcription factors contain a high fraction of intrinsically disordered regions essential for transcriptional regulation. J Mol Biol 359:1137–1149
Balazs A, Csizmok V, Buday L et al (2009) High levels of structural disorder in scaffold proteins as exemplified by a novel neuronal protein, CASK-interactive protein1. FEBS J 276:3744–3756
Buday L, Tompa P (2010) Functional classification of scaffold proteins and related molecules. FEBS J 277:4348–4355
Tompa P, Csermely P (2004) The role of structural disorder in the function of RNA and protein chaperones. FASEB J 18:1169–1175
Dosztanyi Z, Chen J, Dunker AK et al (2006) Disorder and sequence repeats in hub proteins and their implications for network evolution. J Proteome Res 5:2985–2995
Haynes C, Oldfield CJ, Ji F et al (2006) Intrinsic disorder is a common feature of hub proteins from four eukaryotic interactomes. PLoS Comput Biol 2:e100
Pantazatos D, Kim JS, Klock HE et al (2004) Rapid refinement of crystallographic protein construct definition employing enhanced hydrogen/deuterium exchange MS. Proc Natl Acad Sci USA 101:751–756
Spraggon G, Pantazatos D, Klock HE et al (2004) On the use of DXMS to produce more crystallizable proteins: structures of the T. maritima proteins TM0160 and TM1171. Protein Sci 13:3187–3199
Sharma S, Zheng H, Huang YJ et al (2009) Construct optimization for protein NMR structure analysis using amide hydrogen/deuterium exchange mass spectrometry. Proteins 76:882–894
Netzer WJ, Hartl FU (1997) Recombination of protein domains facilitated by co-translational folding in eukaryotes. Nature 388:343–349
Emanuelsson O, Brunak S, von Heijne G et al (2007) Locating proteins in the cell using TargetP, SignalP and related tools. Nat Protoc 2:953–971
Krogh A, Larsson B, von Heijne G et al (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580
Wootton JC, Federhen S (1996) Analysis of compositionally biased regions in sequence databases. Methods Enzymol 266:554–571
Rost B, Yachdav G, Liu J (2004) The PredictProtein server. Nucleic Acids Res 32:W321–W326
McGuffin LJ, Bryson K, Jones DT (2000) The PSIPRED protein structure prediction server. Bioinformatics 16:404–405
Dosztanyi Z, Meszaros B, Simon I (2009) ANCHOR: web server for predicting protein binding regions in disordered proteins. Bioinformatics 25:2745–2746
Dessailly BH, Nair R, Jaroszewski L et al (2009) PSI-2: structural genomics to cover protein domain family space. Structure 17:869–881
Hunter S, Jones P, Mitchell A et al (2012) InterPro in 2011: new developments in the family and domain prediction database. Nucleic Acids Res 40:D306–D312
Graslund S, Sagemark J, Berglund H et al (2008) The use of systematic N- and C-terminal deletions to promote production and structural studies of recombinant proteins. Protein Expr Purif 58:210–221
Chikayama E, Kurotani A, Tanaka T et al (2010) Mathematical model for empirically optimizing large scale production of soluble protein domains. BMC Bioinformatics 11:113
Xiao R, Anderson S, Aramini J et al (2010) The high-throughput protein sample production platform of the Northeast Structural Genomics Consortium. J Struct Biol 172:21–33
Acton TB, Xiao R, Anderson S et al (2011) Preparation of protein samples for NMR structure, function, and small-molecule screening studies. Methods Enzymol 493:21–60
Aramini JM, Rossi P, Huang YJ et al (2008) Solution NMR structure of the NlpC/P60 domain of lipoprotein Spr from Escherichia coli: structural evidence for a novel cysteine peptidase catalytic triad. Biochemistry 47:9715–9717
Rossi P, Aramini JM, Xiao R et al (2009) Structural elucidation of the Cys-His-Glu-Asn proteolytic relay in the secreted CHAP domain enzyme from the human pathogen Staphylococcus saprophyticus. Proteins 74:515–519
Linding R, Jensen LJ, Diella F et al (2003) Protein disorder prediction: implications for structural proteomics. Structure 11:1453–1459
Ward JJ, Sodhi JS, McGuffin LJ et al (2004) Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J Mol Biol 337:635–645
Cheng JSM, Baldi P (2005) Accurate prediction of protein disordered regions by mining protein structure data. Data Min Knowl Discov 11:213–222
Prilusky J, Felder CE, Zeev-Ben-Mordehai T et al (2005) FoldIndex: a simple tool to predict whether a given protein sequence is intrinsically unfolded. Bioinformatics 21:3435–3438
Linding R, Russell RB, Neduva V et al (2003) GlobPlot: Exploring protein sequences for globularity and disorder. Nucleic Acids Res 31:3701–3708
Dosztanyi Z, Csizmok V, Tompa P et al (2005) The pairwise energy content estimated from amino acid composition discriminates between folded and intrinsically unstructured proteins. J Mol Biol 347:827–839
Yang ZR, Thomson R, McNeil P et al (2005) RONN: the bio-basis function neural network technique applied to the detection of natively disordered regions in proteins. Bioinformatics 21:3369–3376
Vucetic S, Brown CJ, Dunker AK et al (2003) Flavors of protein disorder. Proteins 52:573–584
Lupas A, Van Dyke M, Stock J (1991) Predicting coiled coils from protein sequences. Science 252:1162–1164
Romero P, Obradovic Z, Li X et al (2001) Sequence complexity of disordered protein. Proteins 42:38–48
Romero P, Obradovic Z, Dunker AK (1999) Folding minimal sequences: the lower bound for sequence complexity of globular proteins. FEBS Lett 462:363–367
Weathers EA, Paulaitis ME, Woolf TB et al (2007) Insights into protein structure and function from disorder-complexity space. Proteins 66:16–28
Finn RD, Mistry J, Tate J et al (2010) The Pfam protein families database. Nucleic Acids Res 38:D211–D222
Acknowledgements
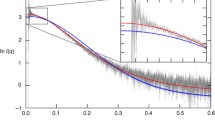
We thank H. Zheng, S. Sharma, A. Ertekin, and R. Xiao for providing the HDX-MS data illustrated in this chapter and J. Aramini for providing the NMR spectrum shown in Fig. 5. The NMR data shown in Fig. 3 were recorded by P. Rossi. This work was supported by a grant from the National Institute of General Medical Sciences Protein Structure Initiative U54-GM074958 (to G.T.M.).
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Huang, Y.J., Acton, T.B., Montelione, G.T. (2014). DisMeta: A Meta Server for Construct Design and Optimization. In: Chen, Y. (eds) Structural Genomics. Methods in Molecular Biology, vol 1091. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-691-7_1
Download citation
DOI: https://doi.org/10.1007/978-1-62703-691-7_1
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-690-0
Online ISBN: 978-1-62703-691-7
eBook Packages: Springer Protocols