Abstract
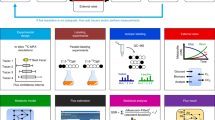
Mass spectrometry (MS) offers a sensitive, reliable, and highly accurate method for measurement of isotopic labeling, which is required for generating comprehensive flux maps using metabolic flux analysis (MFA). We present protocols for assessing isotope labeling in a wide range of biochemical species, including proteinogenic amino acids, free organic and amino acids, sugar phosphates, lipids, starch-glucose, and RNA-ribose. We describe the steps of sample preparation, MS analysis, and data handling required to obtain high-quality isotope labeling measurements that are applicable to MFA. By selecting target analytes that maximize identifiability of the key fluxes of interest, MS measurements of isotope labeling can provide a powerful platform for assessing metabolic fluxes in complex biochemical networks.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Sauer U (2006) Metabolic networks in motion: 13C-based flux analysis. Mol Syst Biol 2:62
Zamboni N, Fendt SM, Ruhl M, Sauer U (2009) (13)C-based metabolic flux analysis. Nat Protoc 4:878–892
Paula Alonso A, Dale VL, Shachar-Hill Y (2010) Understanding fatty acid synthesis in developing maize embryos using metabolic flux analysis. Metab Eng 12:488–497
Yang C, Hua Q, Shimizu K (2002) Metabolic flux analysis in Synechocystis using isotope distribution from 13C-labeled glucose. Metab Eng 4:202–216
Allen D, Ratcliffe R (2009) Quantification of isotope label. In: Schwender J (ed) Plant Metabolic Networks. Springer, New York, pp 105–149
Kitson FG, Larsen BS, McEwen CN (1996) Gas chromatography and mass spectrometry : a practical guide. Academic, San Diego
Wolfe RR, Chinkes DL (2005) Isotope Tracers in Metabolic Research: Principles and Practice of Kinetic Analysis. Wiley, Hoboken, NJ
Lu W, Bennett BD, Rabinowitz JD (2008) Analytical strategies for LC-MS-based targeted metabolomics. J Chromatogr B Analyt Technol Biomed Life Sci 871:236–242
Dauner M, Sauer U (2000) GC-MS analysis of amino acids rapidly provides rich information for isotopomer balancing. Biotechnol Prog 16:642–649
Young JD, Walther JL, Antoniewicz MR, Yoo H, Stephanopoulos G (2008) An elementary metabolite unit (EMU) based method of isotopically nonstationary flux analysis. Biotechnol Bioeng 99:686–699
Allen DK, Shachar-Hill Y, Ohlrogge JB (2007) Compartment-specific labeling information in 13C metabolic flux analysis of plants. Phytochemistry 68:2197–2210
Zamboni N, Sauer U (2009) Novel biological insights through metabolomics and 13C-flux analysis. Curr Opin Microbiol 12:553–558
Luo B, Groenke K, Takors R, Wandrey C, Oldiges M (2007) Simultaneous determination of multiple intracellular metabolites in glycolysis, pentose phosphate pathway and tricarboxylic acid cycle by liquid chromatography-mass spectrometry. J Chromatogr A 1147:153–164
Ausloos P, Clifton CL, Lias SG, Mikaya AI, Stein SE, Tchekhovskoi DV, Sparkman OD, Zaikin V, Zhu D (1999) The critical evaluation of a comprehensive mass spectral library. J Am Soc Mass Spectrom 10:287–299
Kopka J, Schauer N, Krueger S, Birkemeyer C, Usadel B, Bergmuller E, Dormann P, Weckwerth W, Gibon Y, Stitt M, Willmitzer L, Fernie AR, Steinhauser D (2005) GMD@CSB.DB: the Golm Metabolome Database. Bioinformatics 21:1635–1638
Kind T, Wohlgemuth G, Lee do Y, Lu Y, Palazoglu M, Shahbaz S, Fiehn O (2009) FiehnLib: mass spectral and retention index libraries for metabolomics based on quadrupole and time-of-flight gas chromatography/mass spectrometry. Anal Chem 81:10038–10048
Smith CA, O’Maille G, Want EJ, Qin C, Trauger SA, Brandon TR, Custodio DE, Abagyan R, Siuzdak G (2005) METLIN: a metabolite mass spectral database. Ther Drug Monit 27:747–751
Wishart DS, Tzur D, Knox C, Eisner R, Guo AC, Young N, Cheng D, Jewell K, Arndt D, Sawhney S, Fung C, Nikolai L, Lewis M, Coutouly MA, Forsythe I, Tang P, Shrivastava S, Jeroncic K, Stothard P, Amegbey G, Block D, Hau DD, Wagner J, Miniaci J, Clements M, Gebremedhin M, Guo N, Zhang Y, Duggan GE, Macinnis GD, Weljie AM, Dowlatabadi R, Bamforth F, Clive D, Greiner R, Li L, Marrie T, Sykes BD, Vogel HJ, Querengesser L (2007) HMDB: the Human Metabolome Database. Nucleic Acids Res 35:D521–D526
Horai H, Arita M, Kanaya S, Nihei Y, Ikeda T, Suwa K, Ojima Y, Tanaka K, Tanaka S, Aoshima K, Oda Y, Kakazu Y, Kusano M, Tohge T, Matsuda F, Sawada Y, Hirai MY, Nakanishi H, Ikeda K, Akimoto N, Maoka T, Takahashi H, Ara T, Sakurai N, Suzuki H, Shibata D, Neumann S, Iida T, Funatsu K, Matsuura F, Soga T, Taguchi R, Saito K, Nishioka T (2010) MassBank: a public repository for sharing mass spectral data for life sciences. J Mass Spectrom 45:703–714
Antoniewicz MR, Kelleher JK, Stephanopoulos G (2007) Accurate assessment of amino acid mass isotopomer distributions for metabolic flux analysis. Anal Chem 79:7554–7559
Price NP (2004) Acylic sugar derivatives for GC/MS analysis of 13C-enrichment during carbohydrate metabolism. Anal Chem 76:6566–6574
Antoniewicz MR (2006) Comprehensive Analysis of Metabolic Pathways Through the Combined Use of Multiple Isotopic Tracers, In Chemical Engineering, p 370. Massachusetts Institute of Technology, Cambridge, MA
Christie WW (1993) Preparation of ester derivatives of fatty acids for chromatographic analysis. Advances in lipid methodology 2:69–111
Yang L, Kasumov T, Yu L, Jobbins KA, David F, Previs SF, Kelleher JK, Brunengraber H (2006) Metabolomic assays of the concentration and mass isotopomer distribution of gluconeogenic and citric acid cycle intermediates. Metabolomics 2:85–94
Shastri AA (2008) Metabolic flux analysis of photosynthetic systems, In School of Chemical Engineering. Purdue University, West Lafayette, IN
Fernandez CA, Des Rosiers C, Previs SF, David F, Brunengraber H (1996) Correction of 13C mass isotopomer distributions for natural stable isotope abundance. J Mass Spectrom 31:255–262
Coplen T, Böhlke J, De Bievre P, Ding T, Holden N, Hopple J, Krouse H, Lamberty A, Peiser H, Revesz K (2002) Isotope-abundance variations of selected elements:(IUPAC technical report). Pure Applied Chem 74:1987–2017
Kiefer P, Nicolas C, Letisse F, Portais JC (2007) Determination of carbon labeling distribution of intracellular metabolites from single fragment ions by ion chromatography tandem mass spectrometry. Anal Biochem 360:182–188
Berg IA, Kockelkorn D, Ramos-Vera WH, Say RF, Zarzycki J, Hugler M, Alber BE, Fuchs G (2010) Autotrophic carbon fixation in archaea. Nat Rev Microbiol 8:447–460
Lonien J, Schwender J (2009) Analysis of metabolic flux phenotypes for two Arabidopsis mutants with severe impairment in seed storage lipid synthesis. Plant Physiol 151:1617–1634
Acknowledgments
This work has been supported by NSF CAREER Award CBET-0955251 (to J.D.Y).
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media, New York
About this protocol
Cite this protocol
Young, J.D., Allen, D.K., Morgan, J.A. (2014). Isotopomer Measurement Techniques in Metabolic Flux Analysis II: Mass Spectrometry. In: Sriram, G. (eds) Plant Metabolism. Methods in Molecular Biology, vol 1083. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-661-0_7
Download citation
DOI: https://doi.org/10.1007/978-1-62703-661-0_7
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-660-3
Online ISBN: 978-1-62703-661-0
eBook Packages: Springer Protocols