Abstract
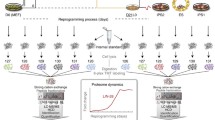
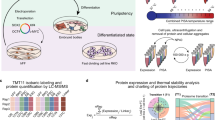
Despite advances in understanding pluripotency through traditional cell biology and gene expression profiling, the signaling networks responsible for maintenance of pluripotency and lineage-specific differentiation are poorly defined. To aid in an improved understanding of these networks at the systems level, we present procedures for the combined analysis of the total proteome and total phosphoproteome (termed (phospho)proteome) from human embryonic stem cells (hESCs), human induced pluripotent stem cells (hiPSCs), and their differentiated derivatives. Because there has been considerable heterogeneity in the literature on the culture of pluripotent cells, we first briefly describe our feeder-free cell culture protocol. The focus, however, is on procedures necessary to generate large-scale (phospho)proteomic data from the cells. Human cells are described here, but the (phospho)proteomic procedures are broadly applicable. Detailed procedures are given for lysis of the cells, protein sample preparation and digestion, multidimensional liquid chromatography, analysis by tandem mass spectrometry, and database searches for peptide/protein identification (ID). We summarize additional data analysis procedures, the subject of ongoing efforts.
Authors Junjie Hou and Brian T.D. Tope contributed equally to this chapter.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, Yamanaka S (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131:861–872
Thomson JA, Itskovitz-Eldor J, Shapiro SS, Waknitz MA, Swiergiel JJ, Marshall VS, Jones JM (1998) Embryonic stem cell lines derived from human blastocysts. Science 282:1145–1147
Singec I, Jandial R, Crain A, Nikkhah G, Snyder EY (2007) The leading edge of stem cell therapeutics. Annu Rev Med 58:313–328
Brandenberger R, Wei H, Zhang S, Lei S, Murage J, Fisk GJ, Li Y, Xu C, Fang R, Guegler K, Rao MS, Mandalam R, Lebkowski J, Stanton LW (2004) Transcriptome characterization elucidates signaling networks that control human ES cell growth and differentiation. Nat Biotechnol 22:707–716
Ramalho-Santos M, Yoon S, Matsuzaki Y, Mulligan RC, Melton DA (2002) “Stemness”: transcriptional profiling of embryonic and adult stem cells. Science 298:597–600
Ideker T, Thorsson V, Ranish JA, Christmas R, Buhler J, Eng JK, Bumgarner R, Goodlett DR, Aebersold R, Hood L (2001) Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 292:929–934
Blume-Jensen P, Hunter T (2001) Oncogenic kinase signalling. Nature 411:355–365
Hunter T (2009) Tyrosine phosphorylation: thirty years and counting. Curr Opin Cell Biol 21:140–146
Schlessinger J (2000) Cell signaling by receptor tyrosine kinases. Cell 103:211–225
Aebersold R, Mann M (2003) Mass spectrometry-based proteomics. Nature 422:198–207
Brill LM, Xiong W, Lee KB, Ficarro SB, Crain A, Xu Y, Terskikh A, Snyder EY, Ding S (2009) Phosphoproteomic analysis of human embryonic stem cells. Cell Stem Cell 5:204–213
Brill LM, Salomon AR, Ficarro SB, Mukherji M, Stettler-Gill M, Peters EC (2004) Robust phosphoproteomic profiling of tyrosine phosphorylation sites from human T cells using immobilized metal affinity chromatography and tandem mass spectrometry. Anal Chem 76:2763–2772
Brill LM, Motamedchaboki K, Wu S, Wolf DA (2009) Comprehensive proteomic analysis of Schizosaccharomyces pombe by two-dimensional HPLC-tandem mass spectrometry. Methods 48:311–319
Hou J, Cui Z, Xie Z, Xue P, Wu P, Chen X, Li J, Cai T, Yang F (2010) Phosphoproteome analysis of rat L6 myotubes using reversed-phase C18 prefractionation and titanium dioxide enrichment. J Proteome Res 9:777–788
Mitulovic G, Stingl C, Steinmacher I, Hudecz O, Hutchins JR, Peters JM, Mechtler K (2009) Preventing carryover of peptides and proteins in nano LC-MS separations. Anal Chem 81:5955–5960
Swaney DL, McAlister GC, Coon JJ (2008) Decision tree-driven tandem mass spectrometry for shotgun proteomics. Nat Methods 5:959–964
McAlister GC, Phanstiel D, Wenger CD, Lee MV, Coon JJ (2010) Analysis of tandem mass spectra by FTMS for improved large-scale proteomics with superior protein quantification. Anal Chem 82:316–322
Olsen JV, Schwartz JC, Griep-Raming J, Nielsen ML, Damoc E, Denisov E, Lange O, Remes P, Taylor D, Splendore M, Wouters ER, Senko M, Makarov A, Mann M, Horning S (2009) A dual pressure linear ion trap Orbitrap instrument with very high sequencing speed. Mol Cell Proteomics 8:2759–2769
Good DM, Wenger CD, McAlister GC, Bai DL, Hunt DF, Coon JJ (2009) Post-acquisition ETD spectral processing for increased peptide identifications. J Am Soc Mass Spectrom 20:1435–1440
Chalkley RJ, Medzihradszky KF, Lynn AJ, Baker PR, Burlingame AL (2010) Statistical analysis of peptide electron transfer dissociation fragmentation mass spectrometry. Anal Chem 82:579–584
Syka JE, Coon JJ, Schroeder MJ, Shabanowitz J, Hunt DF (2004) Peptide and protein sequence analysis by electron transfer dissociation mass spectrometry. Proc Natl Acad Sci U S A 101:9528–9533
Zhou JY, Schepmoes AA, Zhang X, Moore RJ, Monroe ME, Lee JH, Camp DG, Smith RD, Qian WJ (2010) Improved LC-MS/MS spectral counting statistics by recovering low-scoring spectra matched to confidently identified peptide sequences. J Proteome Res 9:5698–5704
Singec I, Hoo J, Crain AM, Tobe BTD, Talantova M, Doctor KS et al. Deep Phosphoproteomic Profiling of Human Pluripotency and neural lineage commitment (In Preparation)
Zhang Y, Wen Z, Washburn MP, Florens L (2010) Refinements to label free proteome quantitation: how to deal with peptides shared by multiple proteins. Anal Chem 82:2272–2281
Li QR, Ning ZB, Tang JS, Nie S, Zeng R (2009) Effect of peptide-to-TiO2 beads ratio on phosphopeptide enrichment selectivity. J Proteome Res 8:5375–5381
Acknowledgments
We thank Anjum Khan and Dante Bencivengo of Thermo Fisher Scientific; Peter Kent, Lori Ann Upton, David Mintline, and Kerry Nugent of Michrom Bioresources for support with instrumentation; David Chiang and Patrick Chu from SageN and Kutbuddin Doctor from the Sanford-Burnham Medical Research Institute (SBMRI) for bioinformatics support; as well as Khatereh Motamedchaboki and Wenhong Zhu from SBMRI for help with some of the experiments. Support was provided by the SBMRI, NCI Cancer Center Support Grant 5 P30 CA30199-28, The La Jolla Interdisciplinary Neuroscience Center Cores Grant 5 P30 NS057096 from NINDS, and RC2 MH090011 from NIMH.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this protocol
Cite this protocol
Hou, J. et al. (2013). Combined Total Proteomic and Phosphoproteomic Analysis of Human Pluripotent Stem Cells. In: Zavazava, N. (eds) Embryonic Stem Cell Immunobiology. Methods in Molecular Biology, vol 1029. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-478-4_12
Download citation
DOI: https://doi.org/10.1007/978-1-62703-478-4_12
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-477-7
Online ISBN: 978-1-62703-478-4
eBook Packages: Springer Protocols