Abstract
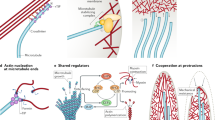
The dynamic nature of cellular cytoskeleton is vital to many cell functions. It was shown that all three filamentous components of the cytoskeleton (F-actin, microtubules, and intermediate filaments) share dynamic behavior. In this chapter we address briefly some basic features of cytoskeletal dynamics and then focus on the actin filament system. In this context we review the major classes of actin-interacting partners and their effects on actin cytoskeleton.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Chhabra ES, Higgs HN (2007) The many faces of actin: matching assembly factors with cellular structures. Nat Cell Biol 9:1110–1121
Kueh HY, Mitchison TJ (2009) Structural plasticity in actin and tubulin polymer dynamics. Science 325:960–963
Carlier MF, Gutfreund H, Bayley PM (1992) Nucleotide hydrolysis regulates the dynamics of actin filaments and microtubules [and discussion]. Philos Trans R Soc Lond B Biol Sci 336:93–97
Pollard TD, Borisy GG (2003) Cellular motility driven by assembly and disassembly of actin filaments. Cell 112:453–465
Mitchison T, Kirschner M (1984) Dynamic instability of microtubule growth. Nature 312:237–242
Schek HT III, Gardner MK, Cheng J et al (2007) Microtubule assembly dynamics at the nanoscale. Curr Biol 17:1445–1455
Pedigo S, Williams J (2002) Concentration dependence of variability in growth rates of microtubules. Biophys J 83:1809–1819
Chrétien D, Fuller SD, Karsenti E (1995) Structure of growing microtubule ends: two-dimensional sheets close into tubes at variable rates. J Cell Biol 129:1311–1328
Walker RA, O’Brien ET, Pryer NK et al (1988) Dynamic instability of individual microtubules analyzed by video light microscopy: rate constants and transition frequencies. J Cell Biol 107:1437–1448
Dent EW, Gupton SL, Gertler FB (2011) The growth cone cytoskeleton in axon outgrowth and guidance. Cold Spring Harb Perspect Biol 3(3):pii: a001800
Skau CT, Neidt EM, Kovar DR (2009) Role of tropomyosin in formin-mediated contractile ring assembly in fission yeast. Mol Biol Cell 20:2160–2173
Okabe S, Miyasaka H, Hirokawa N (1993) Dynamics of the neuronal intermediate filaments. J Cell Biol 121:375–386
Sheterline P, Clayton J, Sparrow JC (2002) Actin, 4th edn. Oxford University Press, Oxford
Orlova A, Shvetsov A, Galkin VE et al (2004) Actin-destabilizing factors disrupt filaments by means of a time reversal of polymerization. Proc Natl Acad Sci U S A 101:17664–17668
Kueh HY, Brieher WM, Mitchison TJ (2008) Dynamic stabilization of actin filaments. Proc Natl Acad Sci 105:16531–16536
Oztug Durer ZA, Diraviyam K, Sept D et al (2010) F-actin structure destabilization and DNase I binding loop fluctuations: mutational cross-linking and electron microscopy analysis of loop states and effects on F-actin. J Mol Biol 395:544–557
Scoville D, Stamm JD, Toledo-Warshaviak D et al (2006) Hydrophobic loop dynamics and actin filament stability. Biochemistry 45:13576–13584
Galkin VE, Orlova A, Schroder GF et al (2010) Structural polymorphism in F-actin. Nat Struct Mol Biol 17:1318–1323
Orlova A, Prochniewicz E, Egelman EH (1995) Structural dynamics of F-actin: II. Cooperativity in structural transitions. J Mol Biol 245:598–607
Fujii T, Iwane AH, Yanagida T et al (2010) Direct visualization of secondary structures of F-actin by electron cryomicroscopy. Nature 467:724–728
Dos Remedios CG, Chhabra D, Kekic M et al (2003) Actin binding proteins: regulation of cytoskeletal microfilaments. Physiol Rev 83:433–473
Fortin DA, Srivastava T, Soderling TR (2012) Structural modulation of dendritic spines during synaptic plasticity. Neuroscientist 18(4):326–341
Chesarone MA, Goode BL (2009) Actin nucleation and elongation factors: mechanisms and interplay. Curr Opin Cell Biol 21:28–37
Chesarone MA, DuPage AG, Goode BL (2010) Unleashing formins to remodel the actin and microtubule cytoskeletons. Nat Rev Mol Cell Biol 11:62–74
Xu XP, Rouiller I, Slaughter BD et al (2011) Three-dimensional reconstructions of Arp2/3 complex with bound nucleation promoting factors. EMBO J 31(1):236–247
Rouiller I, Xu XP, Amann KJ et al (2008) The structural basis of actin filament branching by the Arp2/3 complex. J Cell Biol 180:887–895
Le Clainche C, Pantaloni D, Carlier MF (2003) ATP hydrolysis on actin-related protein 2/3 complex causes debranching of dendritic actin arrays. Proc Natl Acad Sci 100:6337–6342
Cai L, Makhov AM, Schafer DA et al (2008) Coronin 1B antagonizes cortactin and remodels Arp2/3-containing actin branches in lamellipodia. Cell 134:828–842
Gandhi M, Smith BA, Bovellan M et al (2010) GMF is a cofilin homolog that binds Arp2/3 complex to stimulate filament debranching and inhibit actin nucleation. Curr Biol 20:861–867
Bugyi B, Didry D, Carlier MF (2010) How tropomyosin regulates lamellipodial actin-based motility: a combined biochemical and reconstituted motility approach. EMBO J 29:14–26
Weaver AM, Karginov AV, Kinley AW et al (2001) Cortactin promotes and stabilizes Arp2/3-induced actin filament network formation. Curr Biol 11:370–374
Urban E, Jacob S, Nemethova M et al (2010) Electron tomography reveals unbranched networks of actin filaments in lamellipodia. Nat Cell Biol 12:429–435
Mullins RD, Heuser JA, Pollard TD (1998) The interaction of Arp2/3 complex with actin: nucleation, high affinity pointed end capping, and formation of branching networks of filaments. Proc Natl Acad Sci 95:6181–6186
Svitkina TM, Borisy GG (1999) Arp2/3 complex and actin depolymerizing factor/cofilin in dendritic organization and treadmilling of actin filament array in lamellipodia. J Cell Biol 145:1009–1026
Yang C, Svitkina T (2011) Visualizing branched actin filaments in lamellipodia by electron tomography. Nat Cell Biol 13:1012–1013
Ydenberg CA, Smith BA, Breitsprecher D et al (2011) Cease-fire at the leading edge: new perspectives on actin filament branching, debranching, and cross-linking. Cytoskeleton 68:596–602
Quinlan ME, Heuser JE, Kerkhoff E et al (2005) Drosophila Spire is an actin nucleation factor. Nature 433:382–388
Bosch M, Le KHD, Bugyi B et al (2007) Analysis of the function of Spire in actin assembly and its synergy with formin and profilin. Mol Cell 28:555–568
Husson C, Renault L, Didry D et al (2011) Cordon-Bleu uses WH2 domains as multifunctional dynamizers of actin filament assembly. Mol Cell 43:464–477
Eisenmann KM, Harris ES, Kitchen SM et al (2007) Dia-interacting protein modulates formin-mediated actin assembly at the cell cortex. Curr Biol 17:579–591
Chesarone M, Gould CJ, Moseley JB et al (2009) Displacement of formins from growing barbed ends by bud14 is critical for actin cable architecture and function. Dev Cell 16:292–302
Quinlan ME, Hilgert S, Bedrossian A et al (2007) Regulatory interactions between two actin nucleators, Spire and cappuccino. J Cell Biol 179:117–128
Bear JE, Gertler FB (2009) Ena/VASP: towards resolving a pointed controversy at the barbed end. J Cell Sci 122:1947–1953
Hansen SD, Mullins RD (2010) VASP is a processive actin polymerase that requires monomeric actin for barbed end association. J Cell Biol 191:571–584
Ae M, Drubin D (2011) Building distinct actin filament networks in a common cytoplasm. Curr Biol 21:R560–R569
Chereau D, Boczkowska M, Skwarek-Maruszewska A et al (2008) Leiomodin is an actin filament nucleator in muscle cells. Science 320:239–243
Hesterkamp T, Weeds AG, Mannherz HG (1993) The actin monomers in the ternary gelsolin: 2 actin complex are in an antiparallel orientation. Eur J Biochem 218:507–513
Loisel TP, Boujemaa R, Pantaloni D et al (1999) Reconstitution of actin-based motility of Listeria and Shigella using pure proteins. Nature 401:613–616
Iwasa JH, Mullins RD (2007) Spatial and temporal relationships between actin-filament nucleation, capping, and disassembly. Curr Biol 17:395–406
Uruno T, Remmert K, Hammer JA (2006) CARMIL is a potent capping protein antagonist. J Biol Chem 281:10635–10650
Fischer RS, Fowler VM (2003) Tropomodulins: life at the slow end. Trends Cell Biol 13:593–601
Kostyukova AS (2008) Tropomodulin/tropomyosin interactions regulate actin pointed end dynamics. Adv Exp Med Biol 644:283–292, Landes Bioscience and Springer Science + Business Media
Fischer RS, Fritz-Six KL, Fowler VM (2003) Pointed-end capping by tropomodulin3 negatively regulates endothelial cell motility. J Cell Biol 161:371–380
Hotulainen P, Paunola E, Vartiainen MK et al (2005) Actin-depolymerizing factor and cofilin-1 play overlapping roles in promoting rapid F-actin depolymerization in mammalian nonmuscle cells. Mol Biol Cell 16:649–664
Hotulainen P, Llano O, Smirnov S et al (2009) Defining mechanisms of actin polymerization and depolymerization during dendritic spine morphogenesis. J Cell Biol 185:323–339
Andrianantoandro E, Pollard TD (2006) Mechanism of actin filament turnover by severing and nucleation at different concentrations of ADF/cofilin. Mol Cell 24:13–23
Pavlov D, Muhlrad A, Cooper J et al (2007) Actin filament severing by cofilin. J Mol Biol 365:1350–1358
Carlier MF, Laurent V, Santolini J et al (1997) Actin depolymerizing factor (ADF/cofilin) enhances the rate of filament turnover: implication in actin-based motility. J Cell Biol 136:1307–1322
Bobkov AA, Muhlrad A, Pavlov DA et al (2006) Cooperative effects of cofilin (ADF) on actin structure suggest allosteric mechanism of cofilin function. J Mol Biol 356:325–334
Moriyama K, Yahara I (1999) Two activities of cofilin, severing and accelerating directional depolymerization of actin filaments, are affected differentially by mutations around the actin-binding helix. EMBO J 18:6752–6761
Chen Q, Pollard TD (2011) Actin filament severing by cofilin is more important for assembly than constriction of the cytokinetic contractile ring. J Cell Biol 195:485–498
McGough A, Pope B, Chiu W et al (1997) Cofilin changes the twist of F-actin: implications for actin filament dynamics and cellular function. J Cell Biol 138:771–781
Suarez C, Roland J, Boujemaa-Paterski R et al (2011) Cofilin tunes the nucleotide state of actin filaments and severs at bare and decorated segment boundaries. Curr Biol 21:862–868
McCullough B, Grintsevich E, Chen C et al (2011) Cofilin-linked changes in actin filament flexibility promote severing. Biophys J 101:151–159
Kim E, Miller CJ, Reisler E (1996) Polymerization and in vitro motility properties of yeast actin: a comparison with rabbit skeletal α-actin. Biochemistry 35:16566–16572
Kueh HY, Charras GT, Mitchison TJ et al (2008) Actin disassembly by cofilin, coronin, and Aip1 occurs in bursts and is inhibited by barbed-end cappers. J Cell Biol 182:341–353
Gandhi M, Achard V, Blanchoin L et al (2009) Coronin switches roles in actin disassembly depending on the nucleotide state of actin. Mol Cell 34:364–374
Balcer HI, Goodman AL, Rodal AA et al (2003) Coordinated regulation of actin filament turnover by a high-molecular-weight Srv2/CAP complex, cofilin, profilin, and Aip1. Curr Biol 13:2159–2169
Dominguez R (2004) Actin-binding proteins—a unifying hypothesis. Trends Biochem Sci 29:572–578
Gutsche-Perelroizen I, Lepault J, Ott A et al (1999) Filament assembly from profilin-actin. J Biol Chem 274:6234–6243
Gunning P, O-Æneill G, Hardeman E (2008) Tropomyosin-based regulation of the actin cytoskeleton in time and space. Physiol Rev 88:1–35
Lindberg U, Schutt CE, Goldman RD et al (2008) Tropomyosin regulate the impact of actin binding proteins on actin filaments. Adv Exp Med Biol 644:223–231
Pruyne DW, Schott DH, Bretscher A (1998) Tropomyosin-containing actin cables direct the Myo2p-dependent polarized delivery of secretory vesicles in budding yeast. J Cell Biol 143:1931–1945
Nyåkern-Meazza M, Narayan K, Schutt CE et al (2002) Tropomyosin and gelsolin cooperate in controlling the microfilament system. J Biol Chem 277:28774–28779
Ishikawa R, Hayashi K, Shirao T et al (1994) Drebrin, a development-associated brain protein from rat embryo, causes the dissociation of tropomyosin from actin filaments. J Biol Chem 269:29928–29933
Grintsevich EE, Galkin VE, Orlova A et al (2010) Mapping of drebrin binding site on F-actin. J Mol Biol 398:542–554
Sharma S, Grintsevich EE, Phillips ML et al (2010) Atomic force microscopy reveals drebrin induced remodeling of F-actin with subnanometer resolution. Nano Lett 11:825–827
Claessens MMAE, Semmrich C, Ramos L et al (2008) Helical twist controls the thickness of F-actin bundles. Proc Natl Acad Sci 105:8819–8822
Hashimoto Y, Kim DJ, Adams JC (2011) The roles of fascins in health and disease. J Pathol 224:289–300
Lebart MC, Hubert F, Boiteau C et al (2004) Biochemical characterization of the L-plastin − actin interaction shows a resemblance with that of alpha-actinin and allows a distinction to be made between the two actin-binding domains of the molecule. Biochemistry 43:2428–2437
Schmoller KM, Semmrich C, Bausch AR (2011) Slow down of actin depolymerization by cross-linking molecules. J Struct Biol 173:350–357
Breitsprecher D, Koestler SA, Chizhov I et al (2011) Cofilin cooperates with fascin to disassemble filopodial actin filaments. J Cell Sci 124:3305–3318
Schmid MF, Sherman MB, Matsudaira P et al (2004) Structure of the acrosomal bundle. Nature 431:104–107
Nagy S, Rock RS (2010) Structured post-IQ domain governs selectivity of myosin X for fascin-actin bundles. J Biol Chem 285:26608–26617
Hirokawa N, Niwa S, Tanaka Y (2010) Molecular motors in neurons: transport mechanisms and roles in brain function, development, and disease. Neuron 68:610–638
Sweeney HL, Houdusse A (2010) Structural and functional insights into the myosin motor mechanism. Annu Rev Biophys 39:539–557
Acknowledgments
We recognize that due to space limitations many important and relevant studies were not cited directly in this review and we apologize for that to the authors of those publications. This work was supported by USPHS grant GM 077190.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Grintsevich, E.E., Reisler, E. (2013). Cytoskeleton Dynamics and Binding Factors. In: Dermietzel, R. (eds) The Cytoskeleton. Neuromethods, vol 79. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-266-7_4
Download citation
DOI: https://doi.org/10.1007/978-1-62703-266-7_4
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-265-0
Online ISBN: 978-1-62703-266-7
eBook Packages: Springer Protocols