Abstract
Microarray technology has made it possible to quantify gene expression of thousands of genes in a single experiment. With the technological advancement, it is now possible to quantify expression of all known genes using a single microarray chip. With this volume of data and the possibility of improper quantification of expression beyond our control, the challenge lies in appropriate experimental design and the data analysis.
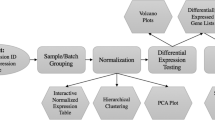
This chapter describes the different types of experimental design for experiments involving microarray analysis (with their specific advantages and disadvantages). It considers the optimum number of replicates for a particular type of experiment. Additionally, this chapter describes the fundamentals of data analysis and the data analysis pipeline to be followed in most common types of microarray experiment.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Derisi, J. (2001) Overview of nucleic acid arrays. Curr Protoc Mol Biol. Chapter 22, Unit 22.1.
Kerr, M.K, Churchill, G.A. (2007) Statistical design and the analysis of gene expression microarray data. Genet Res. 89, 509–14.
Vinciotti, V., Khanin, R., D’Alimonte, D., et al. An experimental evaluation of a loop versus a reference design for two-channel microarrays. Bioinformatics. 21, 492–501.
Lee, M.L., Kuo, F.C., Whitmore, G.A., Sklar, J. (2000) Importance of replication in microarray gene expression studies: statistical methods and evidence from repetitive cDNA hybridizations. Proc Natl Acad Sci USA. 97, 9834–9.
Pavlidis, P., Li, Q., Noble, W.S. (2003) The effect of replication on gene expression microarray experiments. Bioinformatics. 19, 1620–7.
Smyth, G.K., Speed, T. (2003) Normalization of cDNA microarray data. Methods. 31, 265–73.
Pepper, S.D., Saunders, E.K., Edwards, L.E., Wilson, C.L., Miller, C.J. (2007) The utility of MAS5 expression summary and detection call algorithms. BMC Bioinformatics. 8, 273.
Irizarry, R.A., Bolstad, B.M., Collin, F., Cope, L.M., Hobbs, B., Speed, T.P. (2003) Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 31, e15.
Li, C., Hung, Wong, W. (2001) Model-based analysis of oligonucleotide arrays: model validation, design issues and standard error application. Genome Biol. 2, RESEARCH0032.
Reiner, A., Yekutieli, D., Benjamini, Y. (2003) Identifying differentially expressed genes using false discovery rate controlling procedures. Bioinformatics. 19, 368–75.
Eisen, M.B., Spellman, P.T., Brown, P.O., Botstein, D. (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA. 95, 14863–8.
Alon, U., Barkai, N., Notterman, D.A., et al. (1999) Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proc Natl Acad Sci USA. 96, 6745–50.
Sorlie, T., Perou, C.M., Tibshirani, R., et al. (2001) Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA. 98, 10869–74.
Sorlie, T., Tibshirani, R., Parker, J., et al. (2003) Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA. 100, 8418–23.
Makretsov, N.A., Huntsman, D.G., Nielsen, T.O., et al. (2004) Hierarchical clustering analysis of tissue microarray immunostaining data identifies prognostically significant groups of breast carcinoma. Clin Cancer Res. 10, 6143–51.
Charafe-Jauffret, E., Ginestier, C., Monville, F., et al. (2006) Gene expression profiling of breast cell lines identifies potential new basal markers. Oncogene. 25, 2273–84.
Weigelt, B., Hu, Z., He, X., et al. (2005) Molecular portraits and 70-gene prognosis signature are preserved throughout the metastatic process of breast cancer. Cancer Res. 65, 9155–8.
Hu, Z., Fan, C., Oh, D.S., et al. (2006) The molecular portraits of breast tumors are conserved across microarray platforms. BMC Genomics. 7, 96.
Do, J.H., Choi, D.K. (2008) Clustering approaches to identifying gene expression patterns from DNA microarray data. Mol Cells. 25, 279–88.
Raychaudhuri, S., Stuart, J.M., Altman, R.B. (2000) Principal components analysis to summarize microarray experiments: application to sporulation time series. Pac Symp Biocomput. 455–66.
Ashburner, M., Ball, C.A., Blake, J.A., et al. (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 25, 25–9.
Khatri, P., Draghici, S. (2005) Ontological analysis of gene expression data: current tools, limitations, and open problems. Bioinformatics. 21, 3587–95.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Mehta, J.P. (2011). Microarray Analysis of mRNAs: Experimental Design and Data Analysis Fundamentals. In: O'Driscoll, L. (eds) Gene Expression Profiling. Methods in Molecular Biology, vol 784. Humana Press. https://doi.org/10.1007/978-1-61779-289-2_3
Download citation
DOI: https://doi.org/10.1007/978-1-61779-289-2_3
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-61779-288-5
Online ISBN: 978-1-61779-289-2
eBook Packages: Springer Protocols