Abstract
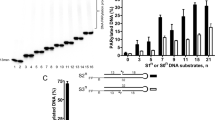
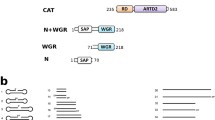
The enzymatic function of poly(adenosine diphosphate (ADP)-ribose) polymerase (PARP) is central to many of its function as a component of DNA repair machinery, modulator of gene transcription, and cell differentiation. While the auto-modification domain of PARP has been shown to be a primary acceptor site of poly-ADP ribose (pADPr), other DNA binding nuclear proteins are also modified by pADPr. It is generally accepted that pADPr polymer is built upon the carboxyl side chain of specific Glu, Asp, and/or Lys residues within the target protein. Identification of the unique amino acid acceptors of pADPr in these nuclear proteins is an active area of study. Because of the heterogeneity of pADPr chain on modified protein targets, the resulting modified proteins have unpredictable final masses, making it difficult to identify acceptor amino acids. Using recombinant proteins, in vitro pADP ribosylation assay and mass spectrometry, we have been able to identify conserved Glu residue in transcription factor NFAT that is enzymatically modified in vitro with pADPr by PARP-1. We discuss this protocol here as a model approach for identifying pADPr acceptors in other nuclear proteins.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Hassa PO, Hottiger MO (2002) The functional role of poly(ADP-ribose)polymerase 1 as novel coactivator of NF-kappaB in inflammatory disorders. Cell Mol Life Sci 59(9):1534–1553
Schreiber V, Dantzer F, Ame JC, de Murcia G (2006) Poly(ADP-ribose): novel functions for an old molecule. Nat Rev Mol Cell Biol 7(7):517–528
Kraus WL (2008) Transcriptional control by PARP-1: chromatin modulation, enhancer-binding, coregulation, and insulation. Curr Opin Cell Biol 20(3):294–302
Altmeyer M, Messner S, Hassa PO, Fey M, Hottiger MO (2009) Molecular mechanism of poly(ADP-ribosyl)ation by PARP1 and identification of lysine residues as ADP-ribose acceptor sites. Nucleic Acids Res 37(11):3723–3738
Suzuki H, Quesada P, Farina B, Leone EJ (1986) In vitro poly(ADP-ribosyl)ation of seminal ribonuclease. J Biol Chem 261(13):6048–6055
Riquelme PT, Burzio LO, Koide SS (1979) ADP ribosylation of rat liver lysine-rich histone in vitro. J Biol Chem 254(8):3018–3028
Olabisi OA, Soto-Nieves N, Nieves E, Yang TT, Yang X, Yu RY, Suk HY, Macian F, Chow CW (2008) Regulation of transcription factor NFAT by ADP-ribosylation. Mol Cell Biol 28(9):2860–2871
Mendoza-Alvarez H, Alvarez-Gonzalez R (2001) Regulation of p53 sequence-specific DNA-binding by covalent poly(ADP-ribosyl)ation. J Biol Chem 276(39):36425–36430
Walker J, Pearson CK (1981) NAD+, ADP-ribosylation and transcription in permeabilized mammalian cells. Biochem J 199(3):813–817
D’Amours D, Desnoyers S, D’Silva I, Poirier GG (1999) Poly(ADP-ribosyl)ation reactions in the regulation of nuclear functions. Biochem J 342(2):249–268
Acknowledgments
The authors would like to acknowledge the past and current members of the Chow laboratory for their input, discussion, and trouble shooting of the protocol. We are also grateful for the assistance from the Laboratory of Macromolecular Analysis and Proteomics (Einstein) with mass spectrometry analysis. O.A.O. was supported by 1F31-GM66607.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Olabisi, O.A., Chow, CW. (2011). Assay for Protein Modification by Poly-ADP-Ribose In Vitro. In: Tulin, A. (eds) Poly(ADP-ribose) Polymerase. Methods in Molecular Biology, vol 780. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-61779-270-0_3
Download citation
DOI: https://doi.org/10.1007/978-1-61779-270-0_3
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-61779-269-4
Online ISBN: 978-1-61779-270-0
eBook Packages: Springer Protocols